Abstract
Objective
Artificial intelligence (AI)-driven precision medicine is moving beyond standalone prediction toward individualized treatment strategies built on multimodal evidence. Yet the field's rapid, interdisciplinary growth has produced a fragmented knowledge structure, while most bibliometric syntheses remain single-database and therefore vulnerable to coverage bias and unstable thematic inference. A cross-validated, implementation-oriented knowledge map is needed to clarify how AI innovations are converging on tailored treatment pathways.
Methods
We established a dual-database cross-validation framework using the Web of Science Core Collection (WoSCC) and Scopus (2015–2025). After parallel screening, preprocessing, and standardized metadata/keyword normalization, 810 WoSCC records and 999 Scopus records were analyzed independently. Science-mapping was performed across publication dynamics, collaboration networks, journal/reference co-citation structures, keyword clustering, burst detection, and longitudinal thematic evolution, followed by explicit cross-database consistency checks.
Results
Across 1809 publications, both databases showed concordant structural patterns: the United States and China formed the central global hubs, while institution-level networks were more stable and coherent than author-level clusters, led by Harvard Medical School, Stanford University, and major Chinese universities/academies. The field evolved in stages from feasibility and data-source expansion to multimodal/multi-omics integration, then toward mechanism-informed interpretability and clinically oriented stratification. Recent frontiers converge on three axes: biologically grounded multimodal inference, explainable AI constrained by biological priors, and deployable decision systems integrating imaging AI, hybrid decision support, and emerging LLM-agent paradigms.
Conclusion
This dual-database, cross-validated atlas delineates a robust trajectory toward clinically executable, AI-enabled personalized medicine. By linking thematic evolution to deployability and interpretability, it provides a transferable framework to prioritize patient stratification, therapy optimization, and workflow-integrated decision support in real-world settings.
Keywords
Introduction
Artificial intelligence (AI) has rapidly evolved from a supporting analytical tool into a foundational driver of modern precision medicine, reshaping how diseases are characterized, stratified, and managed at the individual level. 1 Advances in machine learning and data-driven modeling have enabled the integration of complex biomedical information across molecular, cellular, and clinical dimensions, thereby challenging traditional one-size-fits-all paradigms in healthcare. 2 Precision medicine increasingly relies on the capacity of AI systems to synthesize heterogeneous data sources—ranging from genomics and proteomics to imaging and physiological signals—into actionable insights that inform diagnosis, prognosis, and therapeutic decision-making. 3 This transformation is particularly evident in domains where data complexity and biological heterogeneity are pronounced. Oncology, radiology, and systems medicine have emerged as early adopters, demonstrating how AI-assisted frameworks can enhance risk stratification, treatment planning, and outcome prediction beyond conventional statistical approaches. 4 Parallel developments in computational infrastructure and analytical methodologies have further expanded the scope of precision medicine toward real-time, adaptive, and multiscale clinical applications. 5 Collectively, these advances underscore a broader paradigm shift: precision medicine is no longer defined solely by biomarker discovery, but by the coordinated integration of multi-omics, imaging, and clinical data within AI-enabled analytical ecosystems. 6 Understanding this structural evolution is essential for contextualizing current capabilities and informing translational trajectories, particularly in the context of digital health and clinically embedded AI systems that extend beyond algorithm development toward real-world execution.
Early applications of AI in precision medicine were largely built on single-modality data, such as genomics, imaging, or electronic health records, analyzed in isolation. However, accumulating evidence indicates that such reductionist approaches are insufficient to capture the intrinsic complexity of human disease. Converging studies emphasize that clinically actionable stratification and treatment optimization require coordinated integration of heterogeneous data sources across molecular, cellular, tissue, and clinical scales. AI-driven frameworks spanning drug discovery, biomarker identification, and clinical stratification rely fundamentally on multi-omics and multimodal integration rather than isolated predictive models. 7 Multimodal models integrating imaging, pathology, and clinical variables substantially outperform single-source approaches in gastric cancer management, underscoring the real-world complexity faced in clinical decision-making. 8 Systems-level perspectives incorporating host-microenvironment interactions further reinforce the necessity of integrative paradigms, 9 while advances in spatial and single-cell technologies introduce new analytical and computational challenges for precision medicine. 10 Methodologically, multi-omics and multimodal integration are increasingly recognized not as an optional enhancement, but as a technical prerequisite for next-generation precision medicine.11,12
Despite rapid advances in AI-driven precision medicine, translation from computational models to routine clinical practice remains uneven, placing digital health and implementation science at the center of contemporary precision-medicine discourse. Evidence from digital health research indicates that algorithmic performance alone does not guarantee clinical impact. AI-enabled mHealth systems demonstrate value in personalized monitoring and intervention delivery, yet their effectiveness depends on patient engagement and integration within care pathways. Similar patterns are observed in chronic disease management, where AI increasingly interacts with patient self-management rather than acting as isolated decision tools. 13 From a healthcare operations perspective, successful implementation relies on compatibility with nursing practice, interdisciplinary coordination, and clinical workflows. 11 Trust, transparency, and ethical governance further shape real-world adoption, particularly for large language models influencing clinical decisions. 14 Persistent barriers—including cognitive burden and organizational resistance—limit clinical decision support deployment. 15 Medical imaging remains a comparatively mature entry point, benefiting from standardized data and validation pathways.
Despite the rapid expansion of bibliometric studies examining AI in medicine, existing analyses remain largely constrained by single-database dependency and subfield-specific perspectives, resulting in a fragmented understanding of the broader research landscape. Recent bibliometric investigations have explored AI applications within digital health ecosystems, 16 disease-oriented modeling frameworks, 17 surgical intelligence, 18 and pharmacology-AI intersections.19,20 However, these studies predominantly rely on a single bibliographic source, limiting their capacity to assess structural robustness and cross-disciplinary consistency. Although a limited number of studies have attempted dual-database approaches, their scope has largely been confined to narrowly defined clinical domains, such as specific oncological contexts,21,22 without establishing a scalable or reusable methodological paradigm. In an era where multimodal AI, multi-omics integration, and clinically embedded decision pathways are increasingly convergent, there is a critical need for a comprehensive bibliometric framework capable of cross-database validation. To address this gap, the present study implements a dual-database cross-validated bibliometric synthesis to systematically map the intellectual structure, thematic evolution, and emerging frontiers of AI-driven precision medicine, with explicit emphasis on digital-health contexts and clinically executable trajectories, thereby enhancing methodological rigor and providing a more generalizable evidence base for the field.
Materials and methods
Data sources, study design, and search strategy
This study adopted a dual-database cross-validation bibliometric design to systematically map the research landscape of AI-driven precision medicine. Web of Science Core Collection (WoSCC) and Scopus were selected as data sources due to their complementary strengths in journal coverage, indexing strategies, and disciplinary emphasis. WoSCC is widely recognized for its comprehensive citation tracking and long-standing coverage of high-impact core journals, particularly in biomedical and life sciences, whereas Scopus offers broader journal inclusion, faster indexing updates, and enhanced representation of interdisciplinary and emerging research. In this study, dual-database cross-validation does not refer to a simple merger of records from different databases, but rather to the independent implementation of conceptually aligned yet database-adapted search strategies, followed by parallel bibliometric analyses and subsequent cross-database consistency checks of key trends and knowledge structures. This approach aims to minimize database-specific indexing bias and enhance the robustness and generalizability of the findings. Literature searches in WoSCC and Scopus covered the period from January 1, 2015, to December 25, 2025. All records were downloaded on December 26, 2025, and no additional data retrieval or modification was performed thereafter. In WoSCC, retrieval was conducted using the topic search (TS) field, which covers titles, abstracts, author keywords, and keywords plus, whereas in Scopus the TITLE-ABS-KEY field was used to accommodate its indexing system. Conceptually comparable but database-adapted Boolean search strategies were developed to capture AI-related methodologies, precision/personalized medicine terminology, and multimodal or integrative analytical frameworks. In the present study, “AI-enabled precision care” was operationalized primarily through integrative, multimodal, or systems-oriented precision frameworks that explicitly aim to combine molecular, imaging, and clinical data for patient stratification and decision support, rather than encompassing all forms of single-modality or workflow-level AI applications in healthcare. The complete search strings for both databases are provided in Supplementary Table S1. Only articles and reviews published in English and indexed in SCI-E and SSCI were included. After applying unified inclusion/exclusion criteria and standardized data-cleaning procedures, 810 records from WoSCC (363 Articles and 447 Reviews) and 999 records from Scopus (420 Articles and 579 Reviews) were retained for subsequent bibliometric analyses and cross-database consistency assessment. Cross-database validation was conducted at the level of structural and thematic concordance rather than record-level equivalence, thereby preserving database independence while enabling comparative robustness assessment.
Data cleaning, screening, and preprocessing
The data collection and screening workflow is illustrated in Figure 1. Literature retrieval, screening, and preprocessing were conducted independently and in parallel for WoSCC and Scopus to preserve database-specific structures prior to cross-database comparison. In WoSCC, the initial search yielded 850 records. After restricting the publication period to 2015–2025, 842 records remained. Language filtering (English only) further reduced the dataset to 838 records. During document type screening, Early Access records, Book Chapters, Proceeding Papers, and other publications deemed irrelevant to the study topic or duplicate were excluded, resulting in 810 eligible publications, including 363 Articles and 447 Reviews. In Scopus, the initial search identified 1285 records. After applying the same temporal restriction, 1250 records remained, and language filtering yielded 1218 records. Subsequently, Early Access records, Book Chapters, Proceeding Papers, and other non-eligible document types were excluded, leading to a final dataset of 999 publications, comprising 420 Articles and 579 Reviews. All publications ultimately included in the analysis were independently screened by two authors at the title and abstract level to confirm relevance to AI-enabled precision medicine. In cases of disagreement, a third author adjudicated and consensus was reached through discussion. Metadata preprocessing was performed to ensure analytical consistency. Author names and institutional affiliations were standardized to resolve spelling variants, abbreviations, and affiliation fragmentation. Keywords were normalized through case unification, synonym merging, and plural-singular harmonization to reduce semantic redundancy. Detailed keyword normalization rules and synonym mapping procedures are provided in Supplementary Methods. Deduplication was applied solely for metadata standardization, not to replace database-stratified analyses. Duplicate records were identified primarily using digital object identifiers (DOIs), with title + publication year + first author used as supplementary criteria when DOIs were unavailable. Potential duplicate records were further manually inspected by two authors, and discrepancies were resolved by a third author through discussion. Importantly, deduplication did not substitute the parallel analytical framework; WoSCC and Scopus datasets were retained separately for subsequent bibliometric analyses and cross-database consistency assessment. All screening and preprocessing steps were completed independently within each database, ensuring transparency, reproducibility, and robustness against database-specific indexing bias (Figure 1).

Parallel dual-database bibliometric framework with cross-validation for AI-driven precision medicine research.
Bibliometric and network visualization analyses
This study employed a comprehensive bibliometric and science-mapping framework to characterize the structural features and evolutionary dynamics of AI-driven precision medicine. Indicators included annual publication output, cumulative citation trends, and measures of productivity and academic influence across countries/regions, authors, institutions, and source journals. Collaboration patterns were examined through co-authorship analyses at the country, institutional, and author levels. Journal and reference co-citation analyses were performed to delineate the intellectual base of the field, while keyword co-occurrence networks, thematic clustering, and temporal evolution analyses were used to identify major research hotspots and their developmental trajectories. Analyses were primarily conducted in R (version 4.4.2) using bibliometrix (version 4.3.2) to extract publication and citation trends, quantify productivity and impact, identify core journals based on Bradford's law, and assess thematic evolution. VOSviewer (version 1.6.20) was used to construct co-authorship, co-citation, and keyword co-occurrence networks with association-strength normalization and minimum frequency thresholds (e.g., keyword occurrence ≥5). The minimum keyword occurrence threshold was selected as a commonly adopted empirical setting in exploratory bibliometric analyses to balance information retention with noise reduction and to enhance the interpretability and stability of network visualizations. CiteSpace (version 6.3.R5) supported knowledge-graph construction, burst detection, and 1-year time-sliced visualizations, with g-index (
Statistical analysis
All analyses were conducted within a descriptive bibliometric framework to map structural configurations and temporal patterns; no regression modeling, trend forecasting, or hypothesis-driven statistical analyses were performed to avoid model-induced extrapolation bias. To make cross-database validation explicit and measurable, we further quantified agreement between WoSCC and Scopus using rank-based correlations and set-overlap metrics. Specifically, Spearman's rank correlation coefficients (
Results
Temporal dynamics and disciplinary landscape of the literature
As shown in Figure 2A, the literature exhibited a sustained increase in both publication output and citation impact from 2015 to 2025, with a clear stage-wise evolution. During the early period (2015–2018), publication activity remained relatively modest, corresponding to an initial exploratory phase. From 2019 onward, annual outputs increased markedly and entered an accelerated growth phase after 2021, maintaining high levels through 2024–2025. In parallel, cumulative citation trajectories rose steadily over time, indicating the continuous consolidation of academic influence and sustained scholarly attention. The total volume of retrieved records differed between databases, with 810 publications indexed in the WoSCC and 999 in Scopus. This difference primarily reflects Scopus's broader journal coverage, faster indexing updates, and greater inclusion of interdisciplinary sources, whereas WoS places stronger emphasis on a more selective core journal framework. In terms of disciplinary composition (Figure 2B), publications were distributed across medicine, computer science, bioinformatics, and engineering, underscoring the inherently interdisciplinary nature of the field. Over time, the disciplinary focus gradually shifted from predominantly algorithmic or technology-oriented studies toward more integrative research frameworks emphasizing multi-source data integration and clinical relevance.

Annual scientific production and disciplinary distribution of the literature across dual databases. Blue represents the Web of Science Core Collection (WoSCC), and green represents Scopus. (A) Annual publication output and cumulative citation trends from 2015 to 2025 based on records retrieved from WoSCC and Scopus. Bar charts indicate the number of publications per year, while line plots depict cumulative citation counts. (B) Distribution of publications across major subject categories indexed in WoSCC and Scopus. Horizontal bar charts display the publication volume within each disciplinary category.
Global collaboration networks and national research contributions
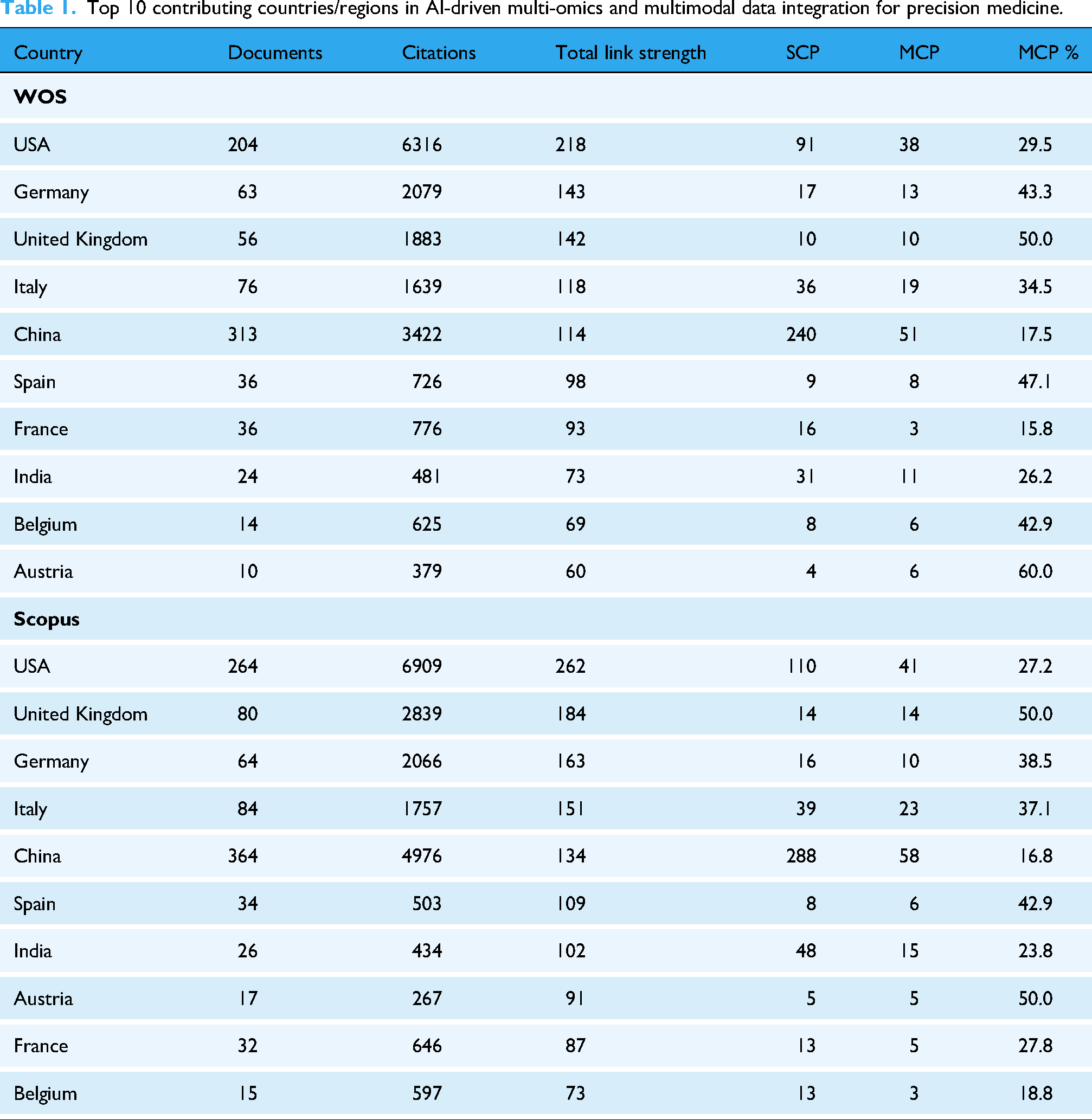
Within the collaboration networks (Figure 3A–C), the United States and China occupy clear hub positions, maintaining dense and stable collaborative ties across regions. This centrality is supported by their substantial research capacity in AI infrastructure, access to large-scale health and biomedical data resources, and well-established translational platforms linking methodological innovation with clinical implementation. Across WoSCC and Scopus, a highly overlapping set of leading countries is identified, indicating strong structural concordance in the global research landscape. Consistent with this structural prominence, Table 1 shows that in the WoS dataset the United States contributed 204 publications with 6316 citations, whereas China produced a higher volume of 313 publications but accumulated 3422 citations. A similar pattern is observed in Scopus, where the United States recorded 264 publications and 6909 citations, compared with China's 364 publications and 4976 citations. These figures indicate that publication output and citation impact are not strictly proportional, reflecting differences in citation structures, research maturity, and international visibility. European countries, particularly the United Kingdom, Germany, and Italy, form a tightly connected collaboration cluster, bridging research activities between Europe, North America, and Asia. Countries such as Canada, Australia, Spain, and France further contribute to network connectivity. Temporally (Figure 3D), leading countries exhibit steadily rising publication trajectories, with accelerated growth aligning with the expanding clinical integration of data-driven methodologies and the strategic prioritization of digital health research globally. Rank-based cross-database comparison demonstrated high concordance for leading countries between WoSCC and Scopus (Spearman's

Global collaboration patterns and national research productivity. (A) Circular visualization illustrating international collaboration relationships among countries and regions, with ribbons representing collaborative linkages between countries. (B) Country-level collaboration network based on co-authorship analysis, highlighting the structural relationships among major contributing countries. (C) World map showing the global distribution of research output and international collaboration patterns across countries. (D) Temporal trends in national research productivity from 2015 to 2025 for major contributing countries.
Top 10 contributing countries/regions in AI-driven multi-omics and multimodal data integration for precision medicine.
Author- and institution-level collaboration architecture and knowledge hubs
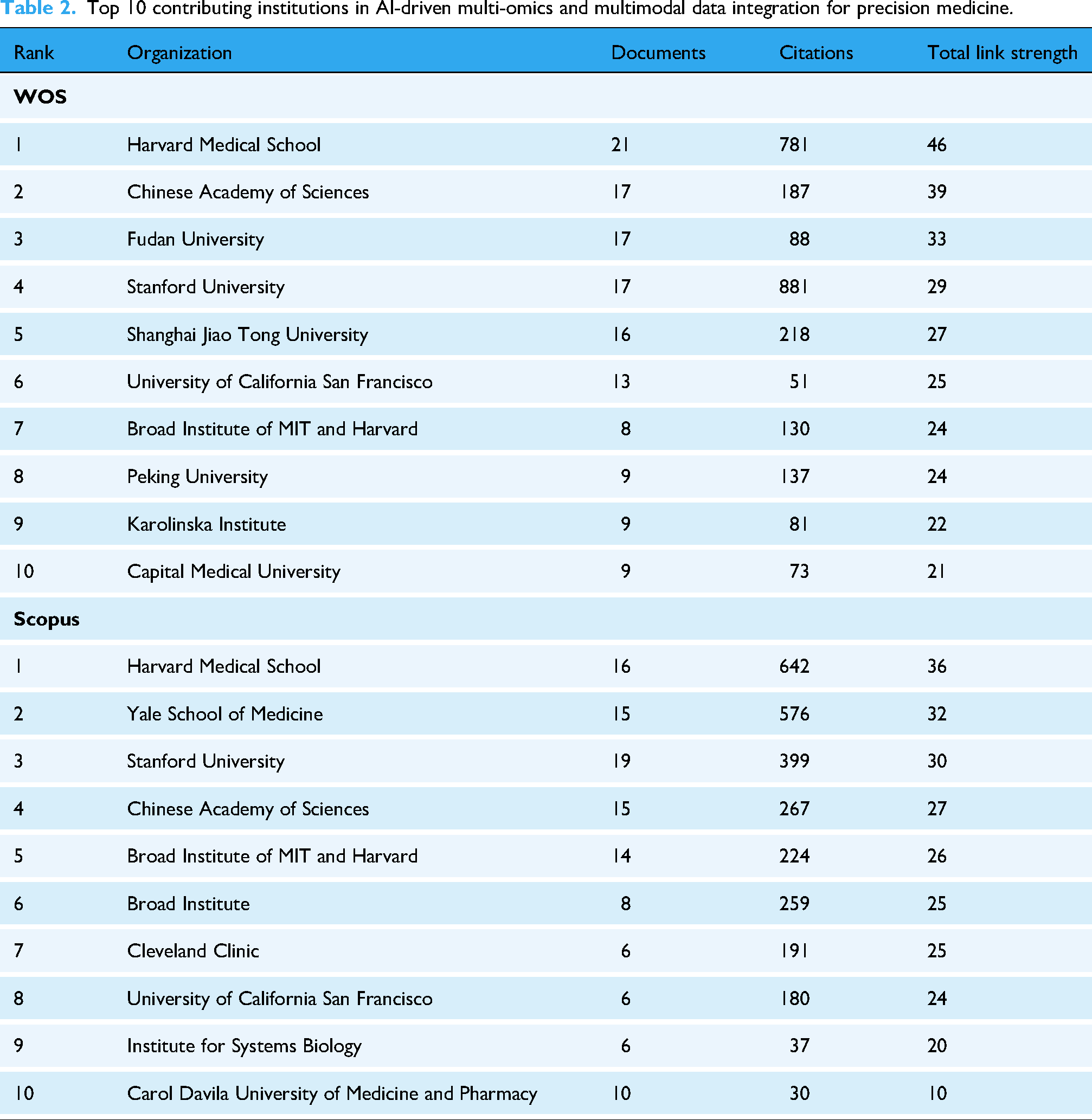
At the author level, the co-authorship networks derived from WoSCC and Scopus (Figure 4A and B) display similarly fragmented and multi-clustered structures rather than a single, unified core, indicating consistent collaboration patterns across databases. Several highly connected authors, including Mersha Tesfaye B., Lin Eugene, Proper Steven P., Rothenberg Marc E., and Hershey Gurjit K., appear as local hubs within specific collaborative clusters. However, these clusters remain partially separated and reflect thematic intersections between clinical research domains and data-driven methodologies, rather than a stable, field-wide methodological leadership. The temporal distribution of author productivity (Figure 4C) further indicates that most contributors exhibit episodic or project-driven publication patterns, with intensified activity in recent years, consistent with the rapid expansion and interdisciplinary adoption of AI-enabled multi-omics research. In contrast, institution-level analyses reveal a markedly more coherent and stable collaborative architecture. The institutional collaboration network and chord diagram (Figure 4D and E) identify Harvard Medical School, Stanford University, the Chinese Academy of Sciences, Fudan University, Shanghai Jiao Tong University, and Zhejiang University as central hubs linking geographically and disciplinarily diverse research teams. These institutions consistently rank among the top contributors in both databases (Table 2), underscoring their pivotal role in providing large-scale data resources, computational infrastructure, and translational platforms that support AI-driven precision medicine. Minor discrepancies in institutional ranking positions between databases are likely attributable to differences in affiliation normalization and indexing coverage rather than substantive changes in research leadership, further supporting strong cross-database concordance at the structural level. Rankings of leading institutions showed moderate concordance between WoSCC and Scopus (Spearman's
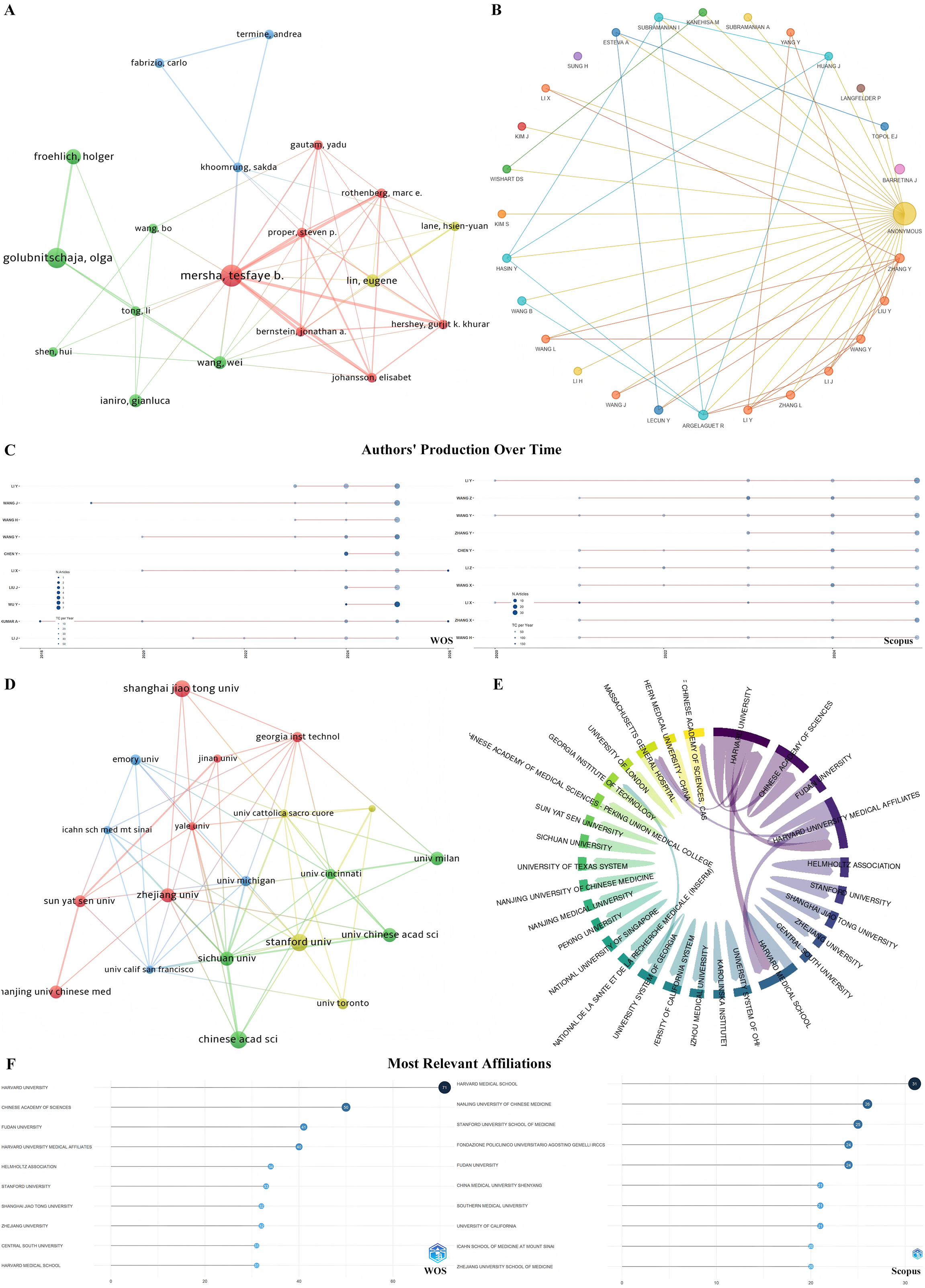
Author- and institution-level collaboration patterns and productivity profiles. (A-B) Author collaboration networks constructed from co-authorship relationships, illustrating major author clusters and collaborative structures within the field. (C) Temporal distribution of publication output for representative authors over time. (D) Institutional collaboration network depicting co-affiliation relationships among leading research organizations. (E) Chord diagram visualizing collaborative linkages among core institutions. (F) Ranking of the most relevant affiliations based on publication output and citation performance.
Top 10 contributing institutions in AI-driven multi-omics and multimodal data integration for precision medicine.
Intellectual foundations and source journal structure of the field
In the journal co-citation network (Figure 5A), multidisciplinary flagship journals—
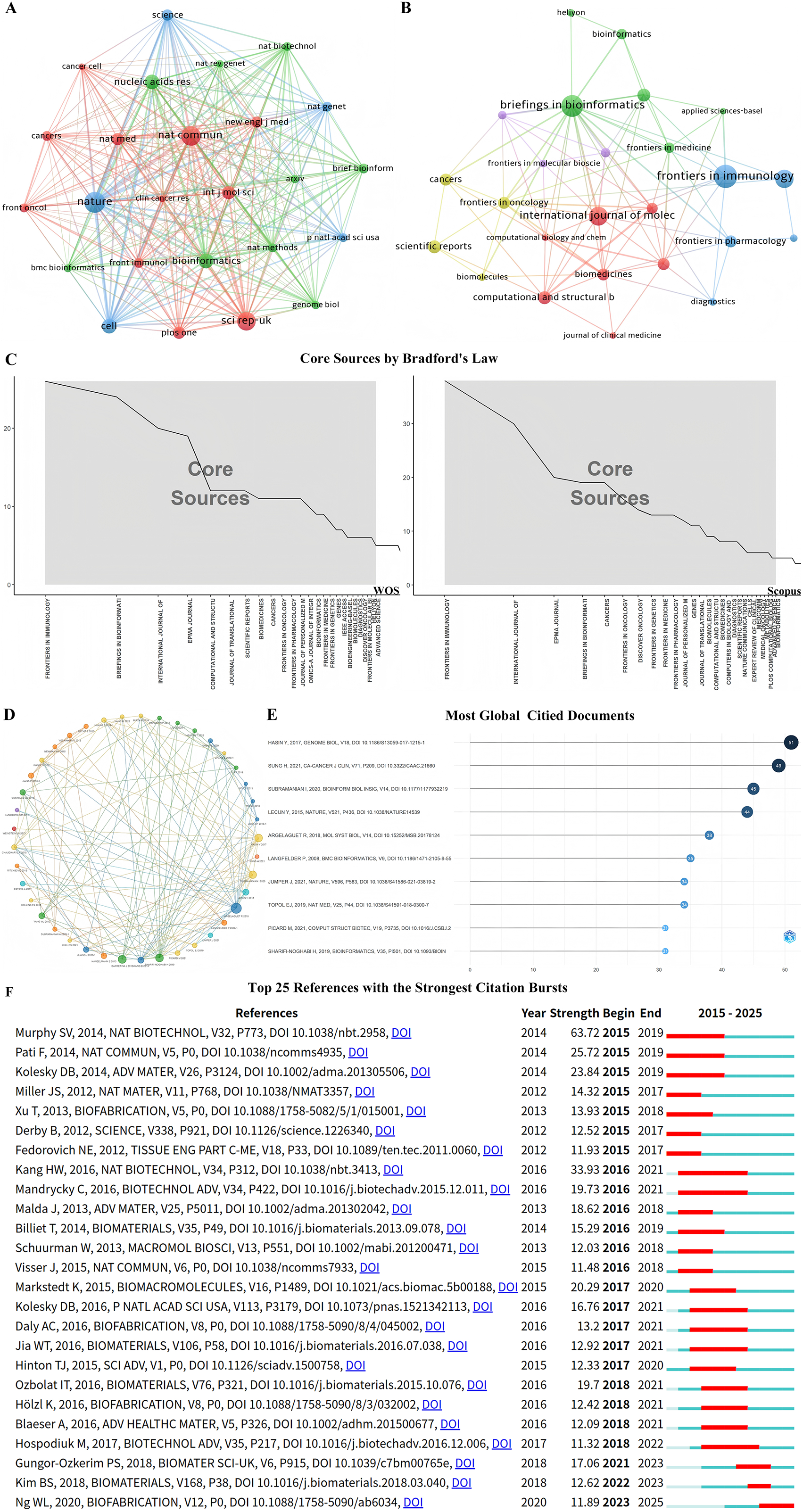
Source journals, co-citation patterns, and knowledge structure of the literature. (A-B) Journal co-citation networks constructed from the analyzed dataset, illustrating the intellectual connectivity and knowledge relationships among major source journals. (C) Core source journals were identified based on Bradford's law, with journals ranked by publication output and the cumulative distribution curve distinguishing core and peripheral sources. (D) Reference co-citation network of the included publications, highlighting major co-citation clusters that collectively define the knowledge base of the field. (E) Most globally cited publications within the dataset, with bars representing total global citation counts. (F) Top 25 references with the strongest citation bursts from 2015 to 2025, where red segments indicate periods of intensified scholarly attention.
Top 10 contributing source journals in AI-driven multi-omics and multimodal data integration for precision medicine.

Top 10 most influential documents in AI-driven multi-omics and multimodal data integration for precision medicine.

Emerging research frontiers and evolution of keyword hotspots
As shown in Figure 6A, keyword co-occurrence networks derived from the two datasets display comparable core structures. Central hubs consistently include AI, machine learning, precision medicine, personalized medicine, and multi-omics, which are densely connected with data- and methodology-related terms such as genomics, proteomics, radiomics, biomarkers, and data integration. This configuration indicates that the field has progressed beyond isolated algorithmic applications toward a data-fusion-oriented research paradigm. Thematic clustering analysis in Figure 6B reveals functional stratification across research topics. In addition to dominant clusters centered on AI and precision or personalized medicine, clinically oriented themes, including liquid biopsy, patient stratification, and tumor microenvironment, are tightly linked to multi-omics- and genomics-driven clusters. These associations underscore a growing emphasis on translating integrated molecular and imaging data into clinically meaningful contexts, consistent with digital health applications involving real-world data environments and decision-support frameworks. In addition, explicit quantification of representative digital-health- and clinical-implementation-related keywords demonstrates stable and substantial representation across both databases (Supplementary Table S2). For example, AI appears 219 times in WoS and 233 times in Scopus, machine learning shows 230 occurrences in WoS and 232 in Scopus, and precision medicine exhibits similarly high visibility (WoS: 232; Scopus: 261). Collectively, these quantitative patterns support the sustained embedding of AI-driven precision medicine within digital-health infrastructures and clinically oriented research contexts. From a temporal perspective, the keyword evolution arc in Figure 6C demonstrates a clear transition between 2020 and 2025. Early emphasis on generic algorithmic concepts, including machine learning and deep learning, progressively converges toward themes emphasizing data integration, multi-omics integration, and interpretable biomarkers. Exploratory temporal trend visualization based on keyword frequencies in Figure 6D indicates sustained prominence of these topics in recent years. Consistently, the hotspot evolution heatmap in Figure 6E confirms a shift from algorithm-dominated phases to an integrated landscape centered on biologically interpretable and clinically implementable precision medicine frameworks. Top-ranked author keywords showed high cross-database overlap between WoSCC and Scopus (Jaccard similarity ≈ 0.76), indicating stable core thematic vocabularies across databases (Supplementary Table S3).

Keyword co-occurrence, thematic clustering, and temporal evolution of research hotspots. (A) Keyword co-occurrence networks constructed from the study datasets, illustrating the structural relationships among high-frequency keywords. (B) Keyword clustering landscape revealing major thematic communities derived from the keyword network. (C) Keyword evolution arc diagram (2020–2025) depicting temporal transitions and continuity among representative keywords across consecutive years. (D) Exploratory temporal trend visualization based on keyword frequencies, with confidence intervals for selected high-frequency keywords, showing observed patterns and smoothed trajectories. (E) Heatmap of research hotspot evolution across three time periods, with color intensity representing relative keyword density.
Discussion
Evolutionary milestones of AI-driven precision medicine
Integrating the findings derived from the dual-database cross-validated bibliometric analyses, this digital-health-oriented synthesis reveals that the evolution of AI-driven precision medicine over the past decade follows a coherent, stage-wise trajectory that is concisely crystallized by the milestone publications summarized in Figure 7. Rather than representing isolated breakthroughs, these landmark studies collectively trace the shifting intellectual and translational priorities of the field, which are also evident in the temporal patterns of keywords, institutional leadership, and national contributions identified in our results. During the early phase (2015–2017), research activity was primarily oriented toward methodological feasibility and data source expansion, as exemplified by Andreu-Perez et al. (2015,

Temporal trajectory of milestone publications shaping the conceptual and methodological evolution of the field.
A three-axis conceptual framework for mechanism-informed and clinically actionable precision medicine
Building on the hotspot evolution and knowledge-base structures identified in our results, the current thematic frontier is no longer defined by isolated algorithmic performance, but by how multimodal evidence can be organized into mechanistically coherent and clinically actionable frameworks. A first mechanistic axis centers on multimodal aggregation toward biologically grounded inference: systems-level integration is increasingly framed as a means to recover disease programs (modules, pathways, and context-specific regulatory circuits) rather than merely improve prediction. Network-medicine syntheses and integrative analytics 33 provide the conceptual scaffolding for mapping heterogeneous signals onto shared biological mechanisms, while patient-derived experimental systems such as hiPSC-based multi-omics 34 enable mechanistic interrogation in genetically defined backgrounds. This trajectory is reinforced by the continued prominence of metabolism-linked omics as an interpretable layer connecting phenotype to intervention, and by precision-medicine frameworks that explicitly position biomarkers and systems biology as pillars for translational stratification.35,36 A second axis highlights explainable AI as the bridge from prediction to causal interpretation. Recent work emphasizes that “explanation” must be constrained by biological priors and validated against mechanistic knowledge, 37 motivating architectures that embed pathway structure or biologically visible representations to improve interpretability and hypothesis generation. 38 Explainability is increasingly evaluated by its ability to clarify clinically relevant mechanisms rather than provide post hoc narratives.39,40 A third axis emphasizes research attention to decision-support-oriented frameworks, consistent with our observed shift toward clinically operational themes (e.g., patient stratification, decision support, and real-world integration). Imaging-driven clinical AI has evolved into a prominent research substrate for multimodal precision workflows, 41 with contemporary studies directly linking interpretable model outputs to therapy response assessment. 42 In parallel, hybrid clinical decision support systems that combine rule-based logic with AI components 43 address auditability and deployment constraints, while emerging LLM-agent paradigms extend precision care into longitudinal management settings, emphasizing scalable, patient-facing decision support. 44 Together, these axes illustrate a field-wide transition toward mechanism-informed integration and increasing translational orientation within digital health and clinical implementation contexts—precisely the pattern reflected across our co-citation structures, keyword clusters, and temporal hotspot shifts. It should be noted that the following forward-looking perspectives (e.g., organoid-based modeling, circRNA vaccines, and material-enabled delivery strategies) are extrapolated from convergent bibliometric trends rather than directly derived from quantitative analyses, and are intended to highlight plausible future directions rather than validated conclusions. In addition, a brief sensitivity consideration regarding construct boundaries is warranted. If alternative query strategies were expanded to place greater weight on imaging-only, EHR-driven, or workflow-centered precision AI applications, the relative prominence of implementation-oriented clusters would likely increase. However, we anticipate that the overarching structural backbone—characterized by multimodal integration, mechanism-informed modeling, and clinically executable trajectories—would remain qualitatively preserved, as these elements constitute the common organizing logic across heterogeneous precision-AI paradigms.
Implications for digital health practice and policy
Beyond conceptual synthesis, these convergent bibliometric patterns also carry direct implications for digital health practice and policy. The convergent trends identified in this bibliometric synthesis have direct implications for how AI-enabled precision medicine is operationalized within digital health infrastructures and clinical care pathways. First, the growing prominence of multimodal integration and explainable AI highlights the central role of hospital-based clinical decision support systems (CDSS) as a primary implementation environment. 45 Rather than functioning as standalone prediction engines, future CDSS should be designed as integrative platforms capable of aggregating multi-omics, imaging, and electronic health record data while providing mechanistically interpretable outputs that clinicians can interrogate. 46 Actionable priorities include embedding pathway-level explanations, uncertainty estimates, and traceable data provenance into CDSS interfaces to support auditability, regulatory compliance, and clinician trust. Second, the concentration of research hotspots around oncology, immunotherapy, and patient stratification underscores the importance of multidisciplinary tumor boards as a key site of digital-health translation.47,48 AI systems that integrate genomic, radiomic, and pathological features can serve as decision-support companions during case discussions, enabling structured comparison of therapeutic options and risk profiles. 49 From a policy perspective, standardizing data-sharing protocols, model-reporting formats, and validation benchmarks across institutions will be essential to ensure equitable deployment and cross-center interoperability.50,51 Practically, health systems should prioritize co-design of AI tools with oncology clinicians to align algorithmic outputs with real-world decision workflows. Third, the increasing linkage between AI-driven precision medicine and longitudinal care models points to remote monitoring and mHealth ecosystems as critical components of learning health systems.52,53 Wearable sensors, patient-reported outcomes, and home-based diagnostics can continuously feed multimodal data streams into adaptive models that update risk stratification and treatment recommendations over time.54–56 Actionable steps include integrating remote data pipelines with hospital information systems, establishing governance frameworks for continuous model updating, and incorporating feedback loops that link patient outcomes to model refinement. Collectively, these measures position digital health not merely as a delivery channel, but as an enabling infrastructure for continuously learning, precision-oriented care. Beyond technical and infrastructural considerations, effective clinical execution of AI-enabled precision medicine also depends critically on human factors and governance design. 57 Emerging scholarship argues that AI can function as a “moral technology” only when governance safeguards ensure that efficiency gains are reinvested into patient-clinician interaction and relational care, 58 rather than being captured solely as increased throughput or productivity. From this perspective, implementation strategies should prioritize workflow integration that augments clinical reasoning, preserves professional judgment, and protects time for communication and shared decision-making. 59 Embedding such human-centered and governance-aware principles alongside technical infrastructures is essential for translating AI-enabled precision medicine into clinically meaningful and ethically sustainable practice.
Future directions for digital-health-enabled precision medicine
Future research should advance from algorithm-centric integration toward biologically grounded and clinically oriented pathways, with digital health ecosystems and learning health systems serving as central implementation environments. Microenvironment-aware modeling, tissue-level scaling of multi-omics signals, and embedding multimodal models into clinical pathways through prospective validation and real-world integration will be essential. Standardized benchmarks, cross-cohort generalization testing, uncertainty-aware reporting, and privacy-preserving analytics further support robust and scalable deployment. Collectively, these routes translate the field's convergent trends into actionable precision medicine, aligning microenvironmental insight, tissue-scale interpretation, and clinically oriented translation within learning health system loops.
Limitation
Despite the methodological rigor and comprehensive analytical framework adopted in this study, several inherent limitations of bibliometric research warrant consideration. First, the analysis was restricted to English-language publications indexed in the WoSCC and Scopus, which may underrepresent relevant studies published in other languages or regional journals, reflecting a common language and database coverage bias. Second, differences in journal inclusion policies, subject classification schemes, and metadata standards across databases may introduce residual heterogeneity that cannot be fully eliminated, even with conceptually aligned search strategies and cross-database consistency checks. Third, although multi-omics and multimodal integration emerged as prominent and rapidly expanding themes in our bibliometric analyses, precision medicine also encompasses imaging-based, clinical-variable-driven, electronic health record-based, and other omics-free approaches that may be comparatively under-represented due to terminology heterogeneity, indexing practices, or clustering thresholds. Accordingly, the thematic prominence of multi-omics in this study reflects patterns within the indexed literature rather than a normative prioritization of specific methodological pathways. Fourth, bibliometric approaches are designed to map research structures, knowledge linkages, and temporal trends at a macro level, rather than to evaluate algorithmic performance, biological mechanisms, or clinical efficacy. In addition, raw citation counts may be influenced by self-citation practices and recency effects, potentially favoring earlier publications or highly networked research groups. To mitigate overinterpretation, our analyses emphasize structural concordance, co-citation networks, and thematic evolution rather than absolute citation-based ranking. Fifth, citation-based indicators inherently lag behind real-world technological deployment. In particular, the rapid clinical uptake of large language models and agent-based systems in decision support, clinical documentation, and patient interaction may not yet be fully reflected in publication and citation patterns, leading to potential underestimation of their current practical impact. Accordingly, the findings should be interpreted as a structural overview of the field's evolution, not as direct validation of specific methods or therapeutic strategies. Boundaries underscore bibliometrics’ complementary role clearly.
Conclusion
This dual-database cross-validated bibliometric analysis systematically delineates the stage-wise evolution of AI-driven precision medicine from 2015 to 2025. Our findings demonstrate a clear paradigm shift from early algorithm-centric exploration toward a framework oriented around multimodal and multi-omics integration, biological interpretability, and clinical executability, with institution-level collaboration networks serving as key drivers of high-impact research. Methodologically, maintaining database independence and explicitly assessing cross-database concordance reduced indexing bias and strengthened the robustness of the inferred trends. Based on these results, future efforts should prioritize mechanism-constrained multimodal integration, advance interpretable models that align with biological and clinical logic, and accelerate the deployment of AI systems within real-world clinical pathways. Emphasis on tissue-scale and microenvironment-aware applications will be essential to translating precision medicine from computational promise into clinically actionable practice.
Supplemental Material
sj-pdf-1-dhj-10.1177_20552076261436251 - Supplemental material for From algorithms to clinical execution: A cross-validated knowledge atlas of AI-enabled precision care (2015–2025)
Supplemental material, sj-pdf-1-dhj-10.1177_20552076261436251 for From algorithms to clinical execution: A cross-validated knowledge atlas of AI-enabled precision care (2015–2025) by Boxiang Zhang, Lucy Yue Lau, Kuan Wang, Kejin Li, Boyang Li, Zehao Hong, Richard J. Aldridge, Elizabeth A. Whitmore, Zeliang Zhao and Yi Chen in DIGITAL HEALTH
Supplemental Material
sj-docx-2-dhj-10.1177_20552076261436251 - Supplemental material for From algorithms to clinical execution: A cross-validated knowledge atlas of AI-enabled precision care (2015–2025)
Supplemental material, sj-docx-2-dhj-10.1177_20552076261436251 for From algorithms to clinical execution: A cross-validated knowledge atlas of AI-enabled precision care (2015–2025) by Boxiang Zhang, Lucy Yue Lau, Kuan Wang, Kejin Li, Boyang Li, Zehao Hong, Richard J. Aldridge, Elizabeth A. Whitmore, Zeliang Zhao and Yi Chen in DIGITAL HEALTH
Supplemental Material
sj-docx-3-dhj-10.1177_20552076261436251 - Supplemental material for From algorithms to clinical execution: A cross-validated knowledge atlas of AI-enabled precision care (2015–2025)
Supplemental material, sj-docx-3-dhj-10.1177_20552076261436251 for From algorithms to clinical execution: A cross-validated knowledge atlas of AI-enabled precision care (2015–2025) by Boxiang Zhang, Lucy Yue Lau, Kuan Wang, Kejin Li, Boyang Li, Zehao Hong, Richard J. Aldridge, Elizabeth A. Whitmore, Zeliang Zhao and Yi Chen in DIGITAL HEALTH
Supplemental Material
sj-docx-4-dhj-10.1177_20552076261436251 - Supplemental material for From algorithms to clinical execution: A cross-validated knowledge atlas of AI-enabled precision care (2015–2025)
Supplemental material, sj-docx-4-dhj-10.1177_20552076261436251 for From algorithms to clinical execution: A cross-validated knowledge atlas of AI-enabled precision care (2015–2025) by Boxiang Zhang, Lucy Yue Lau, Kuan Wang, Kejin Li, Boyang Li, Zehao Hong, Richard J. Aldridge, Elizabeth A. Whitmore, Zeliang Zhao and Yi Chen in DIGITAL HEALTH
Supplemental Material
sj-docx-5-dhj-10.1177_20552076261436251 - Supplemental material for From algorithms to clinical execution: A cross-validated knowledge atlas of AI-enabled precision care (2015–2025)
Supplemental material, sj-docx-5-dhj-10.1177_20552076261436251 for From algorithms to clinical execution: A cross-validated knowledge atlas of AI-enabled precision care (2015–2025) by Boxiang Zhang, Lucy Yue Lau, Kuan Wang, Kejin Li, Boyang Li, Zehao Hong, Richard J. Aldridge, Elizabeth A. Whitmore, Zeliang Zhao and Yi Chen in DIGITAL HEALTH
Footnotes
Acknowledgements
The authors thank VOS viewer, CiteSpace, Scimago Graphica, BiblioVista and R-bibliometrix techniques for their support.
Ethical approval
As this study is a bibliometric analysis based on previously published data, no human participants, animals, or clinical samples were directly involved. Therefore, ethical approval was not required.
Consent to participate
Not applicable. This study did not involve human participants or patient data.
Consent to publish
All authors have read and approved the final version of the manuscript and consent to its publication in this journal.
Author contributions
CRediT: Boxiang Zhang: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Software, Writing - original draft; Lucy Yue Lau: Conceptualization, Data curation, Methodology, Software; Kuan Wang: Formal analysis, Investigation, Software; Kejin Li: Investigation, Software, Methodology; Boyang Li: Formal analysis, Investigation, Methodology; Zehao Hong: Formal analysis, Investigation, Methodology; Richard J. Aldridge: Data curation, Formal analysis, Investigation; Elizabeth A. Whitmore: Investigation, Methodology, Software; Zeliang Zhao and Yi Chen: Conceptualization, Data curation, Formal analysis, Methodology, Software, Supervision, Visualization, Writing - review & editing.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Xinjiang Uygur Autonomous Region, (grant number 2022D01C297).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and materials
All data generated or analyzed during this study are included in this published article and its supplementary information files. The raw dataset used for bibliometric analysis (Bibliometrix-Export-File.xlsx), which contains the list of included articles and relevant metadata, is provided as a supplementary file to enhance transparency and replicability.
Supplemental material
Supplemental material for this article is available online.
