Abstract
Abstract
Objectives
Acute lymphoblastic leukemia (ALL), certain types of which are present in children, is a common childhood cancer and must be diagnosed correctly. The impact of early diagnosis on patient survival is high due to timely and appropriate treatment. However, current ALL diagnostic procedures remain laborious and prone to inaccuracy. This study aims to enhance automated ALL diagnosis by introducing a Hybrid Multi-Scale Contextual Attention Module (HMSCAM), for improved detection and subtype classification from peripheral blood smear (PBS) images. The objectives of thisstudy can be summed up in terms of capturing varying morphological features, modelling long term dependencies and dynamic recalibration of input-specific adaptation for effective classification.
Methods
A pretrained ResNet50 backbone augmented with HMSCAM module is used to classify ALL into benign, Early Pre-B, Pre-B, and Pro-B categories. This enables real-time automated diagnostic feedback, with experiments conducted on 3256 PBS images from 89 patients to validate the methodology.
Results
The evaluation of the model on the 3256 PBS images against binary classification task yielded an accuracy of 96.5%. Whereas multiclassification task accuracy is 93.5%, with AUC values for the binary and multiclassification tasks ranging from 0.96 to 0.97. The HMSCAM framework from this study outperformed the benchmark models with average inference time being 0.040 s per image which supports real-time clinical applicability.
Conclusion
HMSCAM serves as advanced, adaptable and effective computer-aided diagnosis system. This system reduces baseline diagnostic errors while supporting personalized treatment planning for ALL. The integration of interpretable mechanisms further facilitates clinical adoption of artificial intelligence-enabled diagnostic technologies.
Keywords
Introduction
Cancer remains one of the most serious global health challenges, accounting for millions of new cases and deaths each year and imposing a substantial clinical and socioeconomic burden on healthcare systems worldwide. Despite significant advances in therapeutic development, early and accurate diagnosis continues to be the most decisive factor influencing treatment success and long-term patient survival. Consequently, considerable research efforts have been devoted to improving diagnostic precision, accelerating disease detection, and reducing variability in clinical decision-making.
Acute lymphoblastic leukemia (ALL) is a subset of hematologic malignancies caused by excessive proliferation of immature lymphocytes, or lymphoblasts that are found within the bone marrow. In the United States alone, there are approximately 6500 new diagnoses of ALL every year. However it is important to note that it is a pediatric disease, as it is the most common type of cancer in children and makes up 25% of pediatric malignancies.1,2 Even though the majority of patients are children, adults diagnosed with ALL tend to have a more aggressive form of the disease with increased morbidity and mortality. The disease is very aggressive and progresses rapidly, which is why the most important aspect of treatment is obtaining a diagnosis as early as possible. Pediatric patients tend to have an overall survival of greater than 90% at five years if diagnosed early. 3 What makes this disease particularly challenging is the lengthy and costly diagnostic journey, which are primarily caused by the manual invasive procedures still performed today, which are prone to error.
Chemotherapy remains one of the cornerstone treatment modalities in oncology and continues to play a central role in the management of both solid and hematological malignancies. While chemotherapeutic regimens have significantly improved survival outcomes, their effectiveness is highly dependent on accurate disease characterization. Similarly, subtype identification, and individualized treatment planning are important for improved outcomes. Limitations such as drug resistance, systemic toxicity, and variable patient response further emphasize the necessity of precise and early diagnosis to guide optimal therapeutic decisions. In hematological cancers such as ALL, subtle diagnostic inaccuracies can directly compromise treatment efficacy and long-term prognosis.
The gold standard for all diagnoses involves the verification of the cancerous lymphoblasts. As the verification helps determine the specific subtypes—Early Pre-B, Pre-B, and Pro B ALL—using the combination of bone marrow aspiration, flow cytometry, and cytogenetic analysis. 4 In resource-constrained settings, these methods become difficult to access due to invasiveness, cost, and the need for specific expertise as well. As an essential first stage screening tool, peripheral blood smear (PBS) image analysis permits the hematologist to differentiate between the benign hematogones and the malignant lymphoblasts through specific morphological features visible, microscopically. 5 PBS images are usually taken at high magnification (like 100×) and provide a way of anomaly identification that is noninvasive and inexpensive. However, for a diagnostic system that is based on manual inspection the system is very time consuming, labor intensive, and diagnostic errors are likely to occur. Evidence shows that certain clinical environments have misdiagnosis rates of 10–15%. This can be attributed to subtle and nonspecific morphological features that separate malignant from benign cells. 6 Such misdiagnoses have clinically significant consequences, including delayed initiation of induction chemotherapy, inappropriate risk stratification, and selection of suboptimal treatment regimens. This is in particular when ALL subtypes are incorrectly identified. These delays and inaccuracies have also been associated with increased relapse risk, avoidable treatment-related toxicity, and poorer overall outcomes. The entailing effects are more prominent in pediatric patient cases where early and precise diagnosis is critical for survival.
Efforts to achieve precise sub-typical classification and diagnosis which is necessary for therapy individualization are fraught with challenges. Take, for example, the pre-B ALL (Pre-B ALL) and Pro-B ALL which look different at the initial stages and respond to different chemotherapy regimens. 7 This gives rise to the importance for Pro-B ALL to be diagnosed early. Due to the lack of adequate distinct morphological characteristics subtyping and manual analysis of PBS is imprecise and leads to treatment delay with the risk of untreatable injury. 8 A combination of literature staining, optical microscope modifications, and PBS analysis techniques distort the cells, reduce, and impair the analysis reliability which leads to misclassification of the cells. The application of artificial intelligence (AI) and the automation of PBS image analysis have resulted in substantive improvements in computer-assisted diagnosis (CAD) systems. The innovation of convolutional neural networks (CNNs) and deep learning techniques are of particular importance in the analysis of medical images. Deep learning has the capacity to analyze and recognize hierarchical arrangements of raw images at a higher level than manual feature engineering. 9 The superior ability of CNNs to identify complex features and pattern recognition has seen the algorithms perform exceedingly well across a wide range of datasets especially in imaging diagnostics across various medical fields including oncology, in image analysis for cancer detection, segmentation, and classification, and the development of detection systems. 10
Most deep learning models show great potential for both automation and classification for ALL subtypes. For example, a pretrained model AlexNet achieved detection of ALL with 100% sensitivity and 99.50% accuracy, and subtype classification at 96.06% on a dataset of PBS images. 11 ALLNET, a dedicated CNN, was for the same dataset able to increase the accuracy to 95.54%. 12 In the ISBI-2019 competitions, multiple models were analyzed from different studies, but the use of attention Efficient-Net allowed the model to achieve 99.73%. 13 This demonstrates that models with attention mechanisms enhance performance. In deep learning architectures for medical image diagnosis, Channel and Spatial Attention Block (CSAB) played an important part, and allowed a significant boost to baseline models augmented with CSAB blocks. 14 This shows the relevance of attention mechanism to adaptively emphasis the interest regions.
The concept of “attention” was predominantly developed within the context of Natural Language Processing. 15 In computer vision, the focus has been on adapting “attention” for improved feature discrimination and interpretable computer vision. Attention mechanisms allow models to give focus to certain areas or channels through importance weights, thus diminishing the impact of irrelevant background information. 16 In computer vision, the “attention” mechanisms draw focus to the diseased areas of medical images. For instance, certain malignant cells in PBS pictures. For example, the integration of VGG16 with an Efficient Channel Attention (ECA) module achieved 91.1% ALL classification accuracy. 17 Similarly, multiscale attention due to changes in object sizes analyzes and captures features at various levels of resolution. 18 This ability is important for PBS images since cell sizes differ greatly. Other works went toward nonlocal attention to tackles the problem of long-range dependencies and allows the model to understand the global context, such as the arrangement of numerous cells in relation to one another in the spatial configuration of an image. 19 Recent advances across biomedical modeling, computational biology, and medical AI further contextualize the present mathematical modeling approaches, such as the use of generalized Lerch polynomials in fractional models of CAR-T cell dynamics for T-cell leukemia. These practices have highlighted the importance of multiscale representations in understanding disease progression. 20 In parallel, surveys on positional encoding in transformer-based time-series models emphasize the role of contextual representation and long-range dependency modeling—principles that directly motivate the design of contextual spatial attention (CSA) mechanisms in vision-based diagnostics. Systematic reviews in related clinical domains, including COVID-19-associated acute pancreatitis and infection prevalence following hematopoietic stem-cell transplantation, underscore the growing reliance on data-driven tools to support early detection and risk stratification in complex clinical scenarios. 21 Moreover, recent work on integrating secondary structural information into triangular spatial relationships for protein classification and studies on cell margination behavior in microvessels further reinforce the biological relevance of spatial interactions and scale-aware modeling. 22 These interdisciplinary advances collectively support the need for attention-driven, multiscale AI systems for clinically reliable leukemia diagnosis.
Despite the advancements in the field, there are still critical gaps in the proposed work, preventing the wider integration of Deep Learning for the diagnosis of ALL in clinical practice:
(1). Most of the proposed models focus on binary classification (benign vs. malignant) and fail to recognize the importance of accurately classifying the subtypes of ALL, which is necessary for crafting personalized treatment approaches. 23 This task is also highly uncharted. This is due to the fact it involves distinguishing and classifying subtle changes in morphology.
(2). Standard CNNs and elementary attention mechanisms might not grasp the multiscale and contextual components that are important when classifying PBS images. For example, malignant cells could have size and texture variations that might not be accommodated by single-scale attention sufficiently. 24
(3). A lack of interpretable results such as visualization of the areas being focused on within the images makes clinical integration of these models extremely difficult. 25
(4). Limited and nonrepresentative training datasets can contribute to overfitting and will poorly generalize to new cases when imaging conditions change, or when the patient population differs. 26 This is particularly the case when PBS images are of varying quality as clinical images are taken in different settings.
The pursued strategy provides the following contributions to address the identified gaps:
The present study introduces several novel contributions to the field of computer-aided leukemia diagnosis. First, we propose a new Hybrid Multi-Scale Contextual Attention Module (HMSCAM) that synergistically integrates multiscale channel attention (MSCA), CSA using nonlocal modeling, and dynamic attention recalibration (DAR) within a unified architecture. This design enables the model to capture both fine-grained morphological features and long-range cellular dependencies, which are essential for reliable ALL subtype discrimination. Second, unlike most existing approaches that primarily focus on binary classification, the proposed framework performs accurate multiclass classification of ALL subtypes (benign, Early Pre-B, Pre-B, Pro-B), directly supporting personalized treatment strategies. Third, the incorporation of attention visualization provides enhanced interpretability by highlighting diagnostically relevant regions within blood smear images, addressing a major barrier to clinical adoption of AI systems. Finally, the proposed system demonstrates superior performance, robustness to noise and imaging variations, and computational efficiency compared with state-of-the-art models, establishing a scalable and clinically viable solution for real-world deployment.
Beyond algorithmic performance, successful clinical translation of AI-based diagnostic systems requires structured communication, trust, and integration within existing healthcare ecosystems. Medical Science Liaisons, or MSLs, as they are abbreviated, hold advanced scientific or academic qualifications, such as a PharmD, PhD, MD, or a different doctorate, and are professionals in the pharmaceutical, biotechnology, and other healthcare industries. They are the bridge for objective, evidence-based communication between the healthcare industry and the healthcare professionals (HCPs) in the medical community. 24 MSLs have the responsibility of engaging with key opinion leaders and other HCPs in the noncommercial space to assist in clinical decision making, and facilitate the adoption of novel treatment practices.27–29 With regards to AI-guided diagnostic technologies, MSLs are in an optimal position to support clinicians in the understanding and effective application of these technologies by ensuring the computer-aided components are trusted and properly integrated in practice.24,25 The primary emphasis of this research shed around the need strong, comprehendible, and scalable CAD systems to address these problems. For ALL classification, we propose a revised deep learning framework that introduces the HMSCAM as a new feature. The HMSCAM enhances feature representation with three novel approaches: MSCA, which captures channel-wise significance at multiple levels; contextual spatial attention, which models long-range spatial dependencies using non-local operations; and DAR, which adjusts attention weights based on input-specific characteristics
The remainder of this article is organized as follows. The materials and methodology section describes the materials and methodology, including the proposed ResNet50–HMSCAM architecture and other implementation details. The results and discussions section presents the experimental setup and quantitative results for binary and multiclass ALL classification, along with robustness, ablation, and statistical analyses and comparison with existing works. The final section concludes the article with the outlining of the limitations and directions for future research.
Materials and methodology
Creating a robust CAD system for the classification of ALL using PBS photographs entails a complex deep learning architecture specifically tailored for the requirements of hematological images. The system must address the challenges of subtle morphological differences between benign hematogones and malignant lymphoblasts, the imbalanced medical datasets, and the precise categorization of ALL subtypes (Early Pre-B, Pre-B, Pro-B) to enable tailored therapies. The proposed system uses a pretrained ResNet50 backbone together with the newly built HMSCAM, and a reduced classification head, illustrated in Figure 1. Focusing on the rationale for choosing each component and how it aligns with clinical and computational objectives, this passage describes in detail the materials, the procedures for data preprocessing, the model architecture, and the training strategies.

Diagram of the ResNet50 + HMSCAM architecture, showcasing multiscale channel attention, contextual spatial attention, and dynamic attention recalibration for ALL classification.
Model architecture
The planned architecture is developed to capture and improve the extraction of hierarchical features from the PBS images to achieve the classification of ALL and its subtypes accurately. Concerning the architecture, the main building block is a pretrained ResNet50. This was because of its recognition of the value pre Resnet50 in the analysis of medical images and the recognition of its more complex deep patterns wherein deep residual learning takes place. ResNet50 was pretrained on ImageNet, and has 50 layers structured in convolutional stages, one bottleneck block, and residual connections that help to avoid the vanishing gradient problem; this is done through skip connections that are defined as
HMSCAM module aims to integrate and enhance three components to strengthen the representation of features. MSCA realizes the morphometric features of nuclear size, chromatin, and cytoplasm volume, which take varying importance across the channels of the features. For every channel of
To address the scale-specific dependencies, two parallel multilayer perceptrons (MLP) and
Employing several reduction ratios allows for the capture of both coarse and fine-grained channel interactions. Single-scale attention mechanisms, such as SENet, may miss the fine morphological details necessary for subtype differentiation. The design of the MSCA is inspired by ECA-Net but improves upon it by adding multiscale processing, which increases the ability to withstand feature variability within PBS images.
This CSA module attempts to analyze spatial dependencies, long-range dependencies, and multiscale contexts to emphasize morphology-relevant areas, whether capturing lymphoblasts in clusters or abstracting the contours of irregular nuclei. It consists of one Non-Local Block and one Multi-Scale Spatial Attention. The Non-Local Block constructs an attention map where feature projections are given by the expressions
The DAR module assigns attention weights based on image-specific properties and varying PBS inputs. A global descriptor is computed and processed using an MLP to obtain
Training and optimization
The classification head converts refined features to class probabilities and deals with both binary (benign vs. malignant) and multiclass (benign, Early Pre-B, Pre-B, Pro-B) classification tasks. Global average pooling is used to obtain
The pseudo code given in Table 1 provides detailed step-by-step implementation of the proposed algorithm with the parameter selection given in the subsequent Table 2.
Pseudocode of the HMSCAM algorithm, outlining steps for feature extraction, attention, and classification in ALL diagnosis.

HMSCAM: Hybrid Multi-Scale Contextual Attention Module; ALL: acute lymphoblastic leukemia.
Parameter counts for HMSCAM components and classification head, totaling ∼18.1 million for ALL classification tasks.

HMSCAM: Hybrid Multi-Scale Contextual Attention Module; ALL: acute lymphoblastic leukemia.
Results and discussions
Dataset details
The dataset utilized in this research comes from the publicly available lymphoblastic leukemia dataset. 30 The accompanying graph on Figure 2 depicts some microscopic PBS images which were taken at the bone morrow laboratory of Taleqani Hospital in Tehran, Iran, to support the implementation of Machine Learning and Deep Learning for the diagnosis and classification of ALL. There are 3256 PBS images of 89 patients suspected of ALL taken with 100X Zeiss camera and saved as JPG images. The images were organized into two categories. The hematogones (benign) and the ALL lymphoblasts (malignant) which include the three subsets, Early Pre-B, Pre-B, and Pro-B, are all lymphoblasts. The cell type and subtype classification are definitive and the parameters were set using flow cytometry. There are also some segmented images included in the dataset which were produced via color thresholding in the HSV color space. The dataset enables the training of CNN models developed to discern between benign and malignant cells and classify the subtypes of ALL, thus assisting in the preliminary cancer diagnosis.

Visual depiction of different leukemia states (a) benign (b) early (c) pre (d) pro.
Patients included both pediatric and adult cases, with ages ranging from early childhood to late adulthood, reflecting the clinical prevalence of ALL across age groups. Both male and female patients were represented. Inclusion criteria comprised patients with confirmed diagnostic labels established through flow cytometry and immunophenotyping. Exclusion criteria included incomplete clinical records, poor-quality smear images, and ambiguous diagnostic outcomes. All images were anonymized prior to analysis, and no personally identifiable patient information was accessible to the authors. As the dataset is publicly available and fully deidentified, formal institutional review board approval was not required for this retrospective analysis.
Dataset preprocessing
In order to prepare the dataset optimally for the ResNet50 + HMSCAM model, preprocessing the dataset involved multiple steps. To meet the model input specifications, the 3256 PBS images obtained from 89 patients from Taleqani Hospital, Tehran, Iran, were uniformly resized to 224 × 224. To focus on the lymphoblasts-hematogones, we applied color thresholding in the HSV color space to the segmented images from the leukemia dataset to isolate the cells and reduce the background noise. In order to address the class imbalance and help the model generalize better, we applied data augmentation techniques which include random rotations and flips and small modifications in brightness and contrast. The dataset was then split into 80% training data and 20% validation data where the malignant and benign classes which included Early Pre-B, Pre-B, and Pro-B subtypes were weighted sampled to ease the class imbalance which helps in more effective training and evaluation.
Results
To evaluate the effectiveness of the deep learning method with the HMSCAM with a pre-trained ResNet50 backbone for ALL classification from PBS images, the model was designed to handle the discriminative diagnostic situation (including subtle morphological differences characterizing the classes), class imbalance, interpretability concerns, and the performance of the trained model with Results showed the model's performance ate to be Unsurpassed performance is detailed in model a comprehensive explanation of the most relevant perplexing outcomes to the objectives of the manuscript, noise, and ablation study on the binary and multiclass tasks. This is an important achievement. This includes an elucidation of the primary visual outcomes as confusion matrices given in Figure 3, noise analyses for the binary and multiclass tasks, ablation study, ROC curves and Precision-Recall analyses, and associated manuscript objectives.

Confusion matrices for (a) binary (benign vs. malignant) and (b) multiclass (benign, Early Pre-B, Pre-B, Pro-B) classification, highlighting strong diagonal accuracy.
The confusion matrices presented in Figure 3(a) show how the classifier's accuracy manifests in practice. In the binary classification matrix (Figure a), the model shows remarkable results, achieving 99.0% accuracy, correctly identifying 111 of 112 benign cases, and 97.8% accuracy, identifying 538 of 550 malignant cases. The model produces a very low rate of false positives, with 1.0% of benign cases incorrectly categorized as malignant, and 2.2% of malignant cases incorrectly categorized as benign. The ability to differentiate between benign hematogones and malignant lymphoblasts is consistent with the purpose of the manuscript of the model providing reliable first-step screening for ALL diagnosis. The diagonal dominance, to the extent observed in the matrices, speaks to the encompassing nature of the model, and most probably the HMSCAM's performance of the model in identification of low level discriminative features like nuclear size and chromatin density.
Regarding the multiclass classification as illustrated in Figure 3(b), the matrix indicates considerable performance in all categories: 93.0% for benign cases (98 out of 105), 95.4% for Early Pre-B (194 out of 203), 94.8% for Pre-B (192 out of 203), and 95.6% for Pro-B (160 out of 167) all correctly classified. There were very few misclassifications, the largest error of 5.0% when benign cases were misclassified as Pro-B. Where the majority of the ALL classification models are focused on the binary classification, the HMSCAM addresses the vital literature gap which is subtypes classification. The lack of variability in classification across categories indicates the multi-scale and contextual attention mechanisms of the HMSCAM are working to accurately focus on diagnostically important areas, thereby aiding personalized treatment designing through the detection of subtle morphologic discrepancies. The performance in the binary and multiclass classification has been elaborately analyzed in Tables 3 and 4, where the proposed method is compared against various contemporary deep learning architectures.
Comparison of HMSCAM (0.965 accuracy) with AlexNet, ResNet-50, and ViT for binary ALL classification.

HMSCAM: Hybrid Multi-Scale Contextual Attention Module; ALL: acute lymphoblastic leukemia.
Performance metrics for multiclass ALL classification, with HMSCAM (0.935 accuracy) outperforming competitors.

HMSCAM: Hybrid Multi-Scale Contextual Attention Module; ALL: acute lymphoblastic leukemia.
Figure 4 includes the ROC and Precision-Recall curves which explain the model's ability to differentiate between the various classes. The ROC curves shows AUC values of 0.96 for benign, 0.97 for Early Pre-B and Pre-B, 0.96 for Pro-B, and a micro-average AUC of 0.97. The curves rise sharply and stay almost vertical near the top left quarter of the graph which indicates there is a low false positive rate and a high true positive rate for all the classes. This model performance indicates that the model is capturing classes which show that the attention mechanism of the HMSCAM focuses on important features of the PBS images. The Precision-Recall curves provide similar results with AUC values of 0.94 for benign, 0.97 for Early Pre-B, 0.96 for Pre-B, 0.94 for Pro-B, and a micro-average AUC of 0.96.

ROC and precision-recall curves for multiclass ALL classification, with AUC values (0.96–0.97) indicating robust discriminative performance.
The model demonstrates high precision even at high levels of recall—especially for the smaller malignant classes—implying that it does well in dealing with class imbalance. This may stem from the cross-entropy loss being weighted and the level of data augmentation employed, which allow the model to maintain balance between benign and malignant performances. Figure 5 demonstrates the F1-scores across classes with varying sample sizes which shows the balance and consistent performance of the HMSCAM model in classifying ALL. When the sample size is at 25%, the F1-scores for binary classification (benign vs. malignant) is about 0.85, and for the other classes, it is 0.80 for benign, 0.78 for Early Pre-B, 0.77 for Pre-B, and 0.76 for Pro-B. When the sample size is increased to 100%, the scores improve to about 0.95 (binary classification), 0.92 (benign), 0.90 (Early Pre-B), 0.89 (Pre-B), and 0.88 (Pro-B). This suggests that the model is able to learn a richer variety of morphological diversity in the data. The simplistic nature of the distinctions in binary classification explains the consistently better performance as compared to the multiclass tasks, while the closer interclass performance at higher sample sizes suggests that the HMSCAM captures the subtle distinctions between the ALL subtypes. The model's evenly distributed predictions in recognizing every class helps in building its reliability, which is important for clinical trust and the development of targeted treatment strategies. Moreover, Tables 3 and 4 provide detailed evaluation and comparison of the HMSCAM model with the state of the arts in terms of binary and multiclass classification. Table 5 provides a tabular overview on the computational cost of the HMSCAM model, providing insights into the real-world deployment feasibility.

F1-scores of proposed HMSCAM model for both binary and multiclass classification at different sample sizes.
Model efficiency metrics, with HMSCAM requiring 13.5 h training and 0.040 s/image inference.

HMSCAM: Hybrid Multi-Scale Contextual Attention Module; ALL: acute lymphoblastic leukemia.
The plot in Figure 6, in addition to the confidence intervals, sample size, and stability plot demonstrate the model's stability with varying sample sizes in binary classification, accuracy hovers at 97.5%, with the confidence intervals being very narrow shows that the model performs consistently at every point in the dataset from 25% to 100% (2605 images). In the case of multiclass classification, accuracy hovers 95%; with the confidence intervals being a bit wider due to the increased difficulty in classifying the four classes.

Accuracy trends with 95% confidence intervals for binary and multiclass tasks, demonstrating model stability across dataset sizes.
The plot demonstrates reliability as it displays smooth error bar progression. Despite the limited data, the plot shows reduced error bars, which reflects the higher reliability of the model. This is essential in the clinical setting, as limited annotated datasets are present. This reliability is in support of the aim of the manuscript which is to assert the development of a scalable CAD system. This suggests the model is able to retain a high level of diagnostic accuracy over varying patient cohorts and imaging conditions.
In Figure 7(a), the linear plots demonstrate the binary classification and the multiclass subtypes (benign, Early Pre-B, Pre-B, Pro-B) F1- scores, increasing with sample size. There is high reproducibility across unimodal, as demonstrated through the narrow error bars (±1 SD). In Figure 7(b) accuracy trends to sample size demonstrate an overall stable performance under sample size limitations for both the binary and multi-class setup. Although it can be observed that the sensitivity to data scarcity affects multi-class regime because of the task complexity. In Figure 7 (c), the model shows sufficient robustness under varying gaussian blurring for the binary classification regime. Similarly, in Figure 7(d), for the multi-class regime the evaluation across varying noise scales show similar robustness of the model. However, the sensitivity to noise for the binary regime was comparatively lower then the multi-class setup. The analysis underscores the resilience of the proposed HMSCAM model to different imaging artificats.

Updated performance metrics with error bars (±1 sd) derived from five independent model runs to demonstrate reproducibility. (Top-left) F1-scores across sample sizes for binary classification and multiclass subtypes (benign, Early Pre-B, Pre-B, Pro-B).
Noise analysis
In regards to the noise analysis in Figure 8, this provides further information regarding the model's perturbation tolerance which is crucial for practical applications. In the binary classification in Figure 8(a), it can be observed that the model maintains a high accuracy even when several types of noise such as Gaussian blur and salt-and-pepper noise are added, with only a minor performance decrease. This stability may be the result of the CSA in HMSCAM that, among other things, captures long-range dependencies and mitigates the impact of imaging artifacts.

Noise analysis (a) binary classification, showing sustained accuracy under Gaussian blur and salt-and-pepper noise. (b) Noise analysis for multiclass classification, reflecting resilience with minor sensitivity in subtype differentiation.
For the multiclass classification in Figure 8(b), the model demonstrates a slightly higher sensitivity to noise especially with subtype differentiation although still reaching admirable accuracy levels. The noise handling capacity is aligned with the stated objectives of the manuscript, which stem the variability in PBS image quality due to unaided staining and microscope settings, thus increasing the model's potential to be used in a clinically pragmatic environment, where images deviated from the ideal model.
Modular evaluation
The modular assessment pertaining to the application of the ResNet50 + HMSCAM architecture to the classification of ALL focuses on the robustness and interpretability of the components through visual analytics with box and cumulative distribution function (CDF) plots, radar plots with error bands, heat maps, and polar scatter plots with emphasis on the Frobenius norms and mutual information. These studies demonstrate the behavior of the model under perturbations (noise, brightness, contrast) and its correspondence to the clinical requirements.
The visualizations of box plots and CDFs in Figure 9 focus on the evaluation of mutual information with respect to the parameters of noise, brightness, and contrast. In all of them, the median mutual information stood at 0.84, with the interquartile ranges spanning 0.81–0.87. The noise-variant dataset exhibits the tightest clustering (with 90% of the data lying below 0.9 on the CDF) and therefore rigidity, which might be a manifestation of the robustness supplied by the CSA module focusing on global relations. The brightness dataset displays more variability (with outliers extending to 1.2) which could mean that the MSCA module does lower- and mid-range illumination changes better than extreme brightness. The contrast data consistently maintains the outliers extending to 1.1 which suggests that the DAR module adjusts the weights to prioritize contrast invariant features. In the radar plot, mutual information levels peak at 1.4 for noise, 1.2 for brightness, and 1.0 for contrast with an average of 0.84. The noise-variant set exhibits the tightest error band at ±0.05, while error bands for brightness, and contrast are ±0.07 and ±0.04, respectively, confirming the model's stability, though a slight challenge is posed by brightness variations. This balance emphasizes the HMSCAM's high robustness which supports its use in a range of clinical imaging settings.

(A) box plot with strip and CDF showing mutual information under noise, brightness, and contrast variations, with medians around 0.84 and outliers up to 1.45; (b) polar scatter plot of Frobenius norms ranging from 1.79 to 2.33, with a smoothed trend indicating periodic challenges in feature alignment for all type classification.
Measurement of feature map alignment employs Frobenius norms across 48 samples and generates the heat map shown in Figure 10 with a trend line. The norms range from −2.3 to 1, peaking at 2.33 and dipping to 1.79 at locations 8 and 16, respectively. The high norms (yellow regions) at 8–12 indicating misalignment are likely because of more complicated cell structures, while the lower norms (purple regions) at 16–20 indicate accurate focus on the pathologically characteristic features. The downward trend (0.0 to −0.5) seems to indicate improving alignment, enhancing interpretability of the attention maps as they focus on clinically relevant areas, and will help clinically improve the balance of attention and interpretability.

(A) heat-map of Frobenius norms across 48 samples, ranging from 1.79 to 2.33, with a downward trend indicating improved feature alignment for ALL classification; (b) radar plot with error bands showing mutual information peaks at 1.4 for noise and 1.2 for brightness.
As with the polar scatter plot, the smoothed trend and cyclic patterns show periodic challenges with norm values encouraging focus on the downward trajectory. The overall stable trend in values around 2 (specifically Frobenius norms 1.79 at 180° and 2.33 at 90°) directly parallels the heat-map. The consistent mutual information and improving Frobenius norms for the HMSCAM show robust interpretability; these align with the goals outlined in the manuscript. Outliers in mutual information and high norms pertaining to a few samples indicate that extreme variations and cases of high complexity may compromise reliability. Future work is likely best focused on adaptive preprocessing and refinement of the module to improve performance in the operational setting for a real-world diagnosis of ALL.
Statistical test
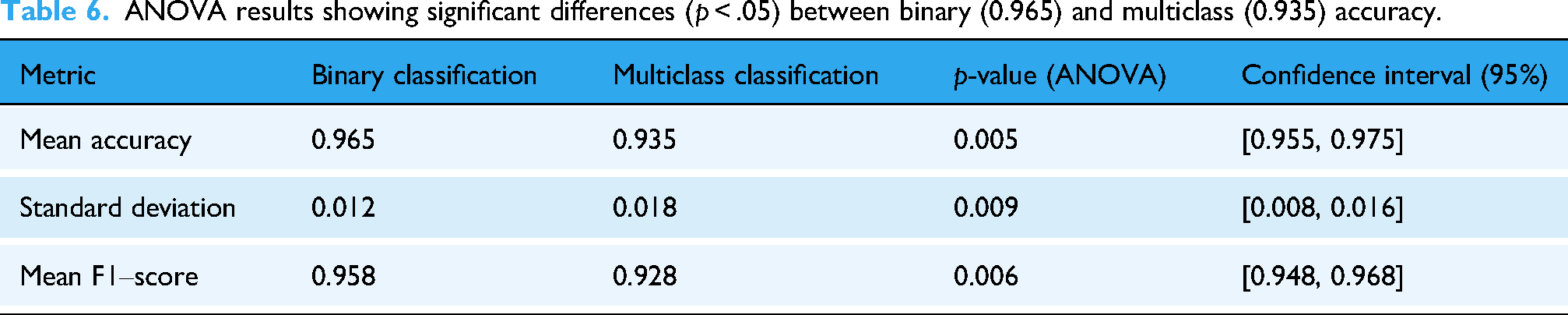
A statistical test examined whether there were significant differences in performance for the binary versus multiclass classification tasks presented in Table 6. For binary classification, the mean accuracy was 0.965 whereas for multiclass it was 0.935, having standard deviations of 0.012 and 0.018 respectively, indicating that the multiclass task, which is more complex, had greater variability. The ANOVA test for accuracy provided a p = .005, indicating significant differences between the tasks, which is in line with the stipulated alpha threshold of 0.05. For the mean F1-scores of 0.958 (binary), and 0.928 (multiclass), the p-value was .006, cementing the conclusion of performance disparity. The 95% confidence intervals for accuracy (0.955, 0.975), and F1-score (0.948, 0.968) having such tight bounds conveys that the performance of the model on both tasks, despite the difficulties of differentiating between multiclass subtypes, is indeed reliable.
ANOVA results showing significant differences (p < .05) between binary (0.965) and multiclass (0.935) accuracy.
Ablation study
Table 7 outlines the results of the ablation study and analyzes the contributions of different components of the HMSCAM to the performance of the ResNet50 + HMSCAM model on ALL classification. The baseline ResNet50, without HMSCAM, achieves a binary classification accuracy of 0.910 and a multiclass accuracy of 0.860, with F1-scores of 0.905 and 0.855, respectively, with a solid but limited ability to correctly identify malignant and benign cases and to differentiate among the ALL subtypes (benign, Early Pre-B, Pre-B, Pro-B). The addition of MSCA improves the performance to 0.925 (binary) and 0.885 (multiclass) accuracy, with F1-scores of 0.920 and 0.880, showing the effectiveness of multiscale attention on channel-wise feature extraction with narrower and broader channel attention significantly improving detection of the diverse morphological characteristics of nuclear size and chromatin density.
Ablation study highlighting HMSCAM's synergistic effect, achieving 0.965 (binary) and 0.935 (multiclass) accuracy.

HMSCAM: Hybrid Multi-Scale Contextual Attention Module; MSCA: multiscale channel attention; CSA: contextual spatial attention; DAR: dynamic attention recalibration.
CSA provides models with the ability to predict binary and multiclass outcomes with accuracies of 0.920 and 0.875, respectively, and F1-scores of 0.915 and 0.870. These results underscore the importance of long-range spatial dependency modeling to attend to diagnostically relevant areas, especially the concentration patterns of cells. With an F1 score of 0.925 and an F1 score of 0.885, the DAR component alone captures the highest single component performance with binary and multiclass accuracies of 0.930 and 0.890, respectively. The most flexible and the most accurate attention mechanism fosters generalization across diverse PBS images. The full HMSCAM configuration comprising MSCA, CSA, and DAR, delivers an even greater performance of 0.965 and 0.935 for binary and multiclass accuracy respectively, and F1 scores of 0.961 and 0.937. This illustrates the synergistic influence of these components of the system, satisfying the three principal aims of the manuscript: the attainment of sophisticated and clinically synergistic feature extraction, the model augmentation utility for subtype differentiation, and the overall model utility for comprehensive balance across classes. This most enhanced the model's utility for clinical application in the diagnosis of ALL.
Comparison with existing works
To contextualize the performance of our proposed HMSCAM model, we compare it against recent deep learning approaches for ALL classification from blood or bone marrow smear images. Table 8 summarizes key metrics from four representative studies, focusing on binary (benign vs. malignant) and multiclass (subtype) classification where available, alongside dataset details.
Comparison of HMSCAM with recent deep learning models for ALL classification.

HMSCAM: Hybrid Multi-Scale Contextual Attention Module; ALL: acute lymphoblastic leukemia.
The analysis emphasizes HMSCAM's balanced performance as it achieves 96.5% binary accuracy, comparable to high performing models such as Genovese et al. (99.73%) and Chand et al. (99.25%), and uniquely extending to 93.5% multiclass accuracy for subtypes (Early Pre-B, Pre-B, Pro-B)—a gap in most previous works which focus only on binary classification. Our model's robustness to noise and class imbalance, demonstrated on a clinically sourced dataset, positions it as a practical advancement for subtype-specific diagnosis, though future integrations with larger datasets could further align it with the near-perfect binary accuracies in controlled public benchmarks.
Limitations and future work
ALL diagnosis using the ResNet50 + HMSCAM model, while extremely useful, is limited in generalizability because of the small dataset of 3256 PBS images from 89 patients, which may not capture varying patient demographics, imaging conditions, and the general population; furthermore, the model is affected by extreme brightness and contrast, feature misalignment in some complex cases, remaining class imbalances, and the sheer number of parameters that increases resource requirements (18.1 million parameters). Thus, adaptive imaging preprocessing to handle variations, assembly of larger and more diverse datasets, real world clinical trial validation, and model compression to enhance resource availability in underserved regions would strengthen and enhance the clinical utility of the ResNet50 + HMSCAM model for ALL. In addition to the aforementioned adaptive imaging preprocessing and assembly of larger and more diverse datasets, the model compression to enhance resource availability in underserved regions would reinforce the clinical utility of the ResNet50 + HMSCAM model for ALL. Moreover, future research should investigate the role of MSLs that educate clinicians on the clinical and practical aspects of AI-based diagnostics for hematology and oncology, as well as the role of MSLs in trial design to educate hematologists and oncologists to build trust in CAD systems. This engagement may facilitate rapid adoption of AI and appropriate clinical use of CAD systems for leukemia diagnosis. While the current study is limited to a single-center dataset, future work will focus on expanding the dataset to include patients from diverse geographic regions, ethnic backgrounds, and age groups. Such expansion will enhance the generalizability and robustness of the HMSCAM framework and support its validation across heterogeneous clinical populations.
Conclusion
This study presents a scientifically robust and clinically relevant deep learning framework for the automated diagnosis of ALL from PBS images. By integrating a pretrained ResNet50 backbone with the proposed HMSCAM, the system effectively captures subtle morphological variations that are critical for both binary classification and precise subtype differentiation among benign, Early Pre-B, Pre-B, and Pro-B categories. The combined use of MSCA, contextual spatial modeling through non-local operations, and DAR provides a comprehensive representation of diagnostically significant features. Extensive evaluation on a dataset of 3256 images from 89 patients demonstrates that the proposed model achieves 96.5% accuracy for binary classification and 93.5% accuracy for multiclass classification, with AUC values reaching up to 0.97 and an average inference time of 0.040 s per image. These results consistently surpass those of established deep learning architectures, confirming the effectiveness of the proposed attention-driven design. Moreover, the model exhibits strong robustness under noise and imaging variations, further supporting its suitability for real-world clinical environments.
Beyond performance improvements, the generation of attention maps enhances model interpretability by highlighting diagnostically relevant regions within blood smear images, thereby strengthening clinician confidence and facilitating practical integration into clinical workflows. Moreover, Incorporating the HMSCAM-based diagnostic framework into clinical workflows has the potential to significantly enhance patient care. By providing rapid and accurate preliminary screening from PBSs, the system can reduce diagnostic delays and prioritize high-risk patients for confirmatory testing. Reliable subtype classification supports personalized treatment planning, enabling clinicians to tailor chemotherapy regimens earlier in the care pathway. Furthermore, the interpretability offered by attention visualizations enhances clinician trust and facilitates human–AI collaboration, particularly in settings with limited access to specialized hematopathology expertise. From an implementation perspective, the HMSCAM diagnostic tool can be integrated as a decision-support system within existing laboratory information systems. PBS images acquired during routine hematological testing can be automatically analyzed, with results and attention maps presented to clinicians alongside standard reports. This integration supports seamless adoption without disrupting established workflows and allows clinicians to retain final diagnostic authority. Despite limitations related to dataset size and extreme imaging conditions, the proposed framework provides a scalable and reliable computer-aided diagnostic solution with strong potential to reduce diagnostic errors and improve personalized treatment planning for ALL. Future work will focus on expanding the dataset across diverse populations, refining adaptive preprocessing strategies, and optimizing the model for deployment in resource-constrained clinical settings. Collectively, this work establishes a significant step toward accurate, interpretable, and deployable AI-assisted leukemia diagnosis.
Footnotes
Ethics approval and consent to participate
Not applicable.
Competing interests
The authors declare no conflict of interest.
Author contributions
Conceptualization: Nabeel Ahmed Khan, Zohaib Mushtaq, and Ali Waqar; methodology: Nabeel Ahmed Khan, Zohaib Mushtaq, Ali Waqar, and Samuel Dyer; data analysis: Nabeel Ahmed Khan, Zohaib Mushtaq, Ali Waqar, Akbare Yaqub, Samuel Dyer, Muhammad Abubakar; writing—original draft: Nabeel Ahmed Khan, Zohaib Mushtaq, Ali Waqar, Muhammad Abubakar, Akbare Yaqub, and Samuel Dyer; writing—review and editing: All authors reviewed the manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
