Abstract
Objective
The objective of this retrospective study is to compare three learning approaches for blood transfusion (BT) prediction after intensive care unit (ICU) admission: local learning (LL), federated learning (FL), and centralized learning (CL) across five ICU databases (eICU Collaborative Research Database, Medical Information Mart for Intensive Care IV, High-Resolution Intensive Care Unit Dataset, Amsterdam University Medical Center Database, University Hospital of Augsburg).
Methods
As machine learning model we used XGBoost and included 15 clinical variables. The prediction task consists of a 3-h observation window, followed by a 2-h prediction window. We evaluated the models using internal and external validation with area under the receiver-operator curve, area under the precision-recall curve, PPV, Brier score and F1 score.
Results
CL consistently outperformed FL and LL in both internal validation (AUPRC range: 0.73–0.95 (CL) vs 0.63–0.96 (FL) and 0.69–0.96 (LL)) and external validation (AUPRC range: 0.61–0.89 (CL) vs 0.45–0.91 (FL) and 0.37–0.90 (LL)). FL showed variable performance across datasets.
Conclusions
The complexity of the multivariable clinical prediction of BTs may create substantial challenges for FL effectiveness, particularly under high data heterogeneity conditions that are common in healthcare.
Keywords
Introduction
Blood transfusions (BTs) represent critical interventions in intensive care units (ICUs), with accurate prediction of transfusion needs essential for optimal patient blood management and resource allocation. Recent advances in machine learning (ML) for BT prediction have demonstrated remarkable progress, with sophisticated models achieving area under the receiver-operator curve (AUROC) values exceeding 0.97 and F1 scores of 0.89 in large-scale studies. 1 However, current ML approaches face a fundamental challenge: models developed and validated within single institutions demonstrate limited generalizability when applied to external healthcare settings.
This limitation comes from institutional differences in patient populations, clinical practices, data collection procedures, and documentation standards that create distributional heterogeneity across healthcare systems. The federated learning (FL) framework offers a potential solution to this challenge in collaborative ML in healthcare. Recent studies have demonstrated that FL can achieve performance comparable to centralized approaches with AUROC values consistently exceeding 0.90 across diverse healthcare applications.2,3 The theoretical appeal of FL lies in its ability to leverage diverse datasets from multiple institutions without requiring data sharing, potentially addressing both privacy concerns and generalizability challenges simultaneously.
Despite this promise, recent research reveals significant challenges in applying FL to healthcare data. Studies have shown that FL performance can degrade by 10–55% under highly skewed conditions typical of medical data heterogeneity. 2 The non-independent and identically distributed (non-IID) characteristics of healthcare data can fundamentally challenge FL performance, particularly for complex clinical prediction tasks involving multiple interacting variables.
To date, BT prediction in ICUs and FL represent two important but largely separate research areas, with no studies investigating how FL can be applied to BT prediction in multiinstitutional settings. Moreover, existing research in this domain is limited to single institutions. Sheikhalishahi et al. 4 confined their analysis to Amsterdam University Medical Center Database (UMCdb), preventing generalizability assessment across healthcare institutions. Similarly, Mitterecker et al. demonstrated performance only on single-institution data from the Western Australian PBM system. 5
Studies that attempted external validation reveal additional limitations. Kang et al. 6 observed performance degradation across Medical Information Mart for Intensive Care IV (MIMIC-IV) and eICU Collaborative Research Database (eICU-CRD) datasets but did not explore collaborative learning approaches. Levi et al. 7 noted substantial class distribution differences between datasets (48% vs 26%) but did not investigate federated approaches. Rockenschaub et al. 8 identified that only 23.9% of ICU scoring systems used external validation, with 83.3% relying on MIMIC-IV and eICU-CRD, but did not examine FL solutions. Moor et al. 9 demonstrated substantial performance drops (0.846 to 0.761 AUROC) across international sites but did not investigate FL.
In terms of FL literature, existing studies suffer from methodological limitations. Teo et al. 10 revealed that only 5.2% of FL studies in healthcare represented real-world applications, the majority used artificially split datasets that fail to capture authentic clinical practice differences and thereby overestimate model generalizability. Schwinn et al. 11 applied FL across five genuine ICU databases but focused on a simple SpO2 prediction involving a single variable, leaving unexplored how prediction complexity affects FL performance.
In summary, current research has developed effective single-institution transfusion prediction models and documented the challenges of cross-institutional validation. Although FL applications in healthcare have demonstrated feasibility, they have largely been limited to simple prediction tasks. A critical gap remains: addressing multiinstitutional data heterogeneity through collaborative learning approaches for complex clinical predictions.
Objective
To address this gap, this study applies FL to a complex clinical prediction task using data from multiple international databases rather than artificial splits of a single dataset, thereby investigating FL under conditions of data heterogeneity that closely mirror real-world deployment scenarios. Specifically, the aim of this retrospective study is to predict the need for BTs among a general ICU patient cohort after admission to the ICU using data from five distinct hospital databases. We hypothesize that FL will outperform local learning (LL) in the external validation setting.
Methods
We compare three different approaches for predicting BTs in ICU patients: training models locally at each hospital (LL), using FL to train across sites without sharing data, and combining all data for centralized training (centralized learning (CL)). To evaluate these approaches, we used five ICU databases from different countries and conducted both same-site and cross-site validation to understand how well each approach generalizes to new data. The following subsections describe the data sources, preprocessing steps, model development, and evaluation strategy.
Data
In this retrospective study, we used four publicly available and deidentified databases and one ICU dataset from the University Hospital of Augsburg (UKA). The four public ICU databases are: (1) the eICU Collaborative Research Databse (eICU-CRD) 12 from the United States with a data collection period from 2014 to 2015; (2) the Medical Information Mart for Intensive Care (MIMIC-IV) 13 , which originates from the Beth Israel Deaconess Hospital in Boston, Massachusetts (USA), with data collected between 2008 and 2019. (3) the High-Resolution Intensive Care Unit Dataset (HiRID) 14 from Switzerland, where data collection was carried out between 2008 and 2016; and (4) the Amsterdam UMCdb 15 from The Netherlands with data collected from 2003 to 2016. For the UKA, the data was collected between 2010 and 2023.
Data preprocessing
We constructed a general BT cohort by limiting the population to adult patients (age >18 years, height <220 cm and weight <150 kg). Statistical outliers were removed by replacing values below the 2nd percentile and above the 98th percentile with NaN for each feature.
Starting from the raw clinical records, patient data was loaded as time-stamped clinical measurements. For each patient, we selected demographic information (age, gender, height, weight) and 15 key physiological measurements, including hemoglobin levels, blood pressure, heart rate, and renal function markers. For a complete list of the predictors included, see Table 1 and the Supplemental Materials S1, S2, and S3. The feature selection is based on a previous study on the prediction of BTs in the ICU. 7 This initial feature set was further reduced by excluding highly correlated variables to reduce redundancy. To take varying measurement frequencies in the different databases into consideration, we included only one measurement per hour per patient. Data were then transformed from a long format to a wide format to create uniform feature representations across patients with variable numbers of measurements. This transformation involved pivoting time series variables to create columns representing sequential measurements while incorporating static demographic patient characteristics, such as age and gender, among other features. The resulting matrix contained one row per patient, with columns for each type of measurement and time point. Since XGBoost handles missing values internally, the model does not require imputation of missing values.
Demographic characteristics of all database cohorts—All patients.
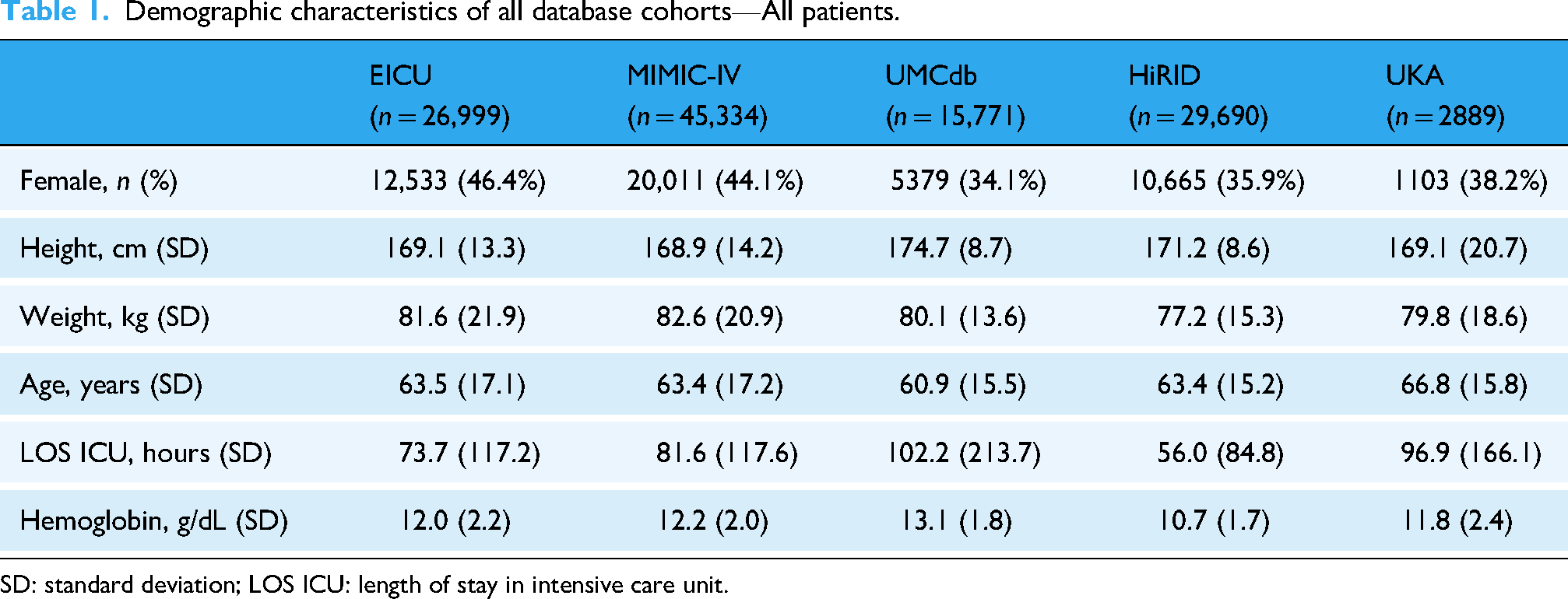
SD: standard deviation; LOS ICU: length of stay in intensive care unit.
Observation and prediction window: We considered a 3-h observation window based on the literature and clinical input from clinicians. In other words, the model uses clinical data from this 3-h window to predict whether a BT will be required within the subsequent 2 h (the prediction window). The observation window spanned 5 to 2 h before the transfusion event. For nontransfused patients, we probabilistically sampled observation window lengths to match the temporal distribution patterns of transfused patients, thereby minimizing potential biases from group-specific observation lengths. See Figure 1 on the right for a summary.

Overview of the study design. Left panel: the three learning paradigms compared—local learning (LL), federated learning (FL), and centralized learning (CL). Right panel: the observation window (5 to 2 h before transfusion) and prediction window (2 h) used for model training and evaluation.
Target variable: The target variable BT was derived from the “Packed blood cell transfusion” indicator in the source data. The target variable was labeled as 0 “no BT” with “Packed red blood cell transfusion” not listed as a procedure or 1 “BT” with “Packed red blood cell transfusion” as a procedure with a value >0. For the sake of performance metrics being comparable across datasets, we harmonized the prevalence of BT cases to 20%. This aligns with the average prevalence of 24% reported by Raasveld et al. 16 and Vincent et al. 17 for BTs in ICUs. Prevalence harmonization was implemented by randomly subsampling controls to increase or decrease prevalence in each dataset as needed to match the set harmonized prevalence of 20%. Additionally, we applied a minimum data completeness threshold of 20% to exclude patients with insufficient measurements, ensuring adequate data quality for model development.
Learning paradigms
We compared three learning paradigms to investigate how training strategy affects model generalizability (Figure 1, on the left side). In LL, each institution trains a model exclusively on its own data, reflecting the current standard in clinical ML development. While LL preserves data privacy, it limits training to local patient populations, potentially restricting external generalizability. FL enables collaborative model training without requiring raw data exchange. Following the Federated Averaging algorithm, 18 participating sites train models locally and share only model parameters with a central server, which aggregates updates to produce a global model. This iterative process allows learning from diverse populations while data remains at source institutions. CL pools all institutional datasets at a single location for training. While CL maximizes data volume and diversity, it requires data transfer across institutional boundaries, facing legal and ethical hurdles due to privacy regulations. CL serves as a theoretical benchmark representing maximum data availability.
Machine learning
As an ML model XGBoost 19 was used with a 70/20/10 train-test-validation split. Following standard ML practices, the training set was used for model fitting, the validation set for hyperparameter optimization, and the test set was only used for final performance evaluation. Hyperparameter tuning was conducted empirically. For the data integration process we used our custom Python-based tool set. This setup is currently in preparation for publication. The Observational Medical Outcomes Partnership Common Data Model was used to standardize the data from the different databases.
For the FL setting specifically, model training was configured with 100 global rounds and two local epochs. While Mehrjou et al. 20 found in a study based on a single database that a large number of local epochs improves model performance, it has also been shown that, due to non-IID data, many local updates may lead to degraded performance. 21 Therefore, a configuration with few local epochs and a high number of global rounds is preferable in a setting with multiple heterogeneous databases as it helps the FL models converge efficiently without excessive divergence from the joint global optimum. The Flower framework 22 was implemented in a dockerized environment to simulate a federated scenario.
Statistical analysis
For demographic comparisons, continuous variables are reported as mean ± standard deviation (SD). Statistical significance between the BT and nontransfusion (non-BT) groups within each database was assessed using the Mann–Whitney U test for continuous variables and the χ2-test for categorical variables. To quantify differences between the databases, the Kruskal–Wallis test was used as the nonparametric alternative to the ANOVA. For the transfusion timing, the Kruskal–Wallis test was followed by Dunn's test with Bonferroni correction for multiple testing. All statistical tests were two-tailed. Results with p < .001 were considered to be significant.
Evaluation
The model performance was evaluated across three training paradigms: LL, FL, and CL. Our analysis relied on complementary evaluation metrics to capture different aspects of predictive performance. Precision reflects the proportion of patients predicted to require a BT who actually required one. This is also known as the positive predictive value. The F1 score, defined as the harmonic mean of precision and recall, provides a balanced measure of the model's ability to correctly identify patients requiring transfusion while minimizing missed cases and false alarms. The AUROC and the area under the precision-recall curve (AUPRC) both evaluate discriminative ability, that is, the model's capability to separate patients requiring BTs from the patients who do not, but the metrics differ in their response to class imbalance. The AUROC treats both classes equally and may produce optimistic estimates when the majority of patients do not require transfusions, as it is influenced by the large number of correctly identified negative cases. AUPRC addresses this concern by focusing on the precision-recall tradeoff. By emphasizing performance of the positive class, AUPRC provides a more clinically meaningful evaluation when the primary goal is to identify patients requiring intervention, i.e. the positive minority class. Finally, we report the Brier score to evaluate probabilistic calibration, which indicates how closely predicted probabilities align with the observed outcome frequencies. Values closer to 0 indicate better calibration, while higher values indicate poorer calibration. All of the above evaluation metrics range between 0 and 1. FL results are aggregated using macro averages, where appropriate.
Two evaluation schemes were implemented: internal and external evaluation. For internal evaluation, the LL models were trained and tested on the same database, the FL model trained on all five databases was tested on each database separately, and the CL model trained on joint data of all five databases was tested on each database's test set individually. For external validation, LL models trained on individual databases were evaluated on the remaining four databases, while FL and CL models were trained on four databases and evaluated on the held-out fifth database, rotating through all databases as test sets in a leave-one-site-out cross-validation approach.
Results
This section presents the performance evaluation of BT prediction models across five ICU databases, comparing LL, FL, and CL. First, we describe the characteristics of the cohort and transfusion patterns across the databases. Next, we report the internal validation results. Finally, we evaluate the external validation performance to assess cross-site generalizability.
Cohort characteristics
Table 1 describes the general patient characteristics of the five ICU databases. All variables shown in Table 1 differed significantly between databases (p < .001). For a more detailed comparison of the cohort characteristics of transfused and nontransfused patients separately, see Supplemental Table S1. In total, the databases include 120,683 patients. Patients who received BTs exhibited significantly lower baseline hemoglobin levels (9.0–10.3 g/dL) than nontransfused patients (11.1–13.4 g/dL), consistent with the clinical indication for transfusion therapy (all p < .001). Additionally, BT patients had longer ICU stay in all databases. The most pronounced difference was observed in UMCdb, with 265.9 h versus 60.4 h (p < .001).
Beyond these within-database patterns, substantial between-database heterogeneity was observed in transfusion timing. The timing of transfusions with respect to ICU admission varied substantially between databases (see Table 2), with median transfusion times ranging from 9.3 h in HiRID to 35.5 h in UKA. All pairwise comparisons except for the MIMIC-IV/UMCdb pairing were significant (p < .001). This heterogeneity in transfusion timing creates significant variation in the pretransfusion observation window available for predictive modeling. Some databases provide limited early ICU data (e.g. HiRID: IQR 5.2–25.4 h), while others offer extended pretransfusion monitoring periods (e.g. UKA: IQR 11.2–124.6 h). This temporal variation may impact the performance of transfusion prediction models across different healthcare systems. For a more detailed overview of the clinical features, see Supplemental Tables S2 and S3.
Timing of blood transfusion events relative to ICU admission across databases.
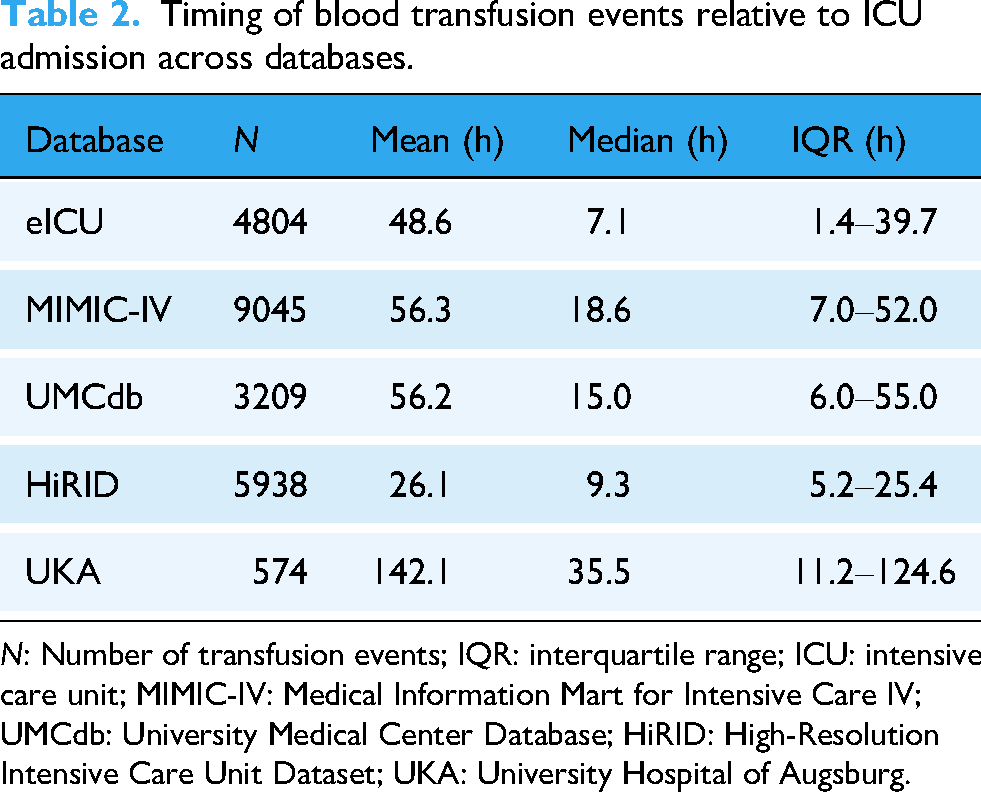
N: Number of transfusion events; IQR: interquartile range; ICU: intensive care unit; MIMIC-IV: Medical Information Mart for Intensive Care IV; UMCdb: University Medical Center Database; HiRID: High-Resolution Intensive Care Unit Dataset; UKA: University Hospital of Augsburg.
Internal validation
We first evaluated internal model performance where training and testing occurred on the same site. As seen in Figure 2, CL achieved higher performance scores than both FL and LL on all datasets when the models were trained and tested on the same site. See also Supplemental Table S4 for detailed results. CL consistently obtained the highest AUPRC scores across all datasets. FL demonstrated variable performance depending on the dataset, with particularly low performance on HiRID. LL showed more stable performance than FL but consistently achieved lower scores than CL across all sites.
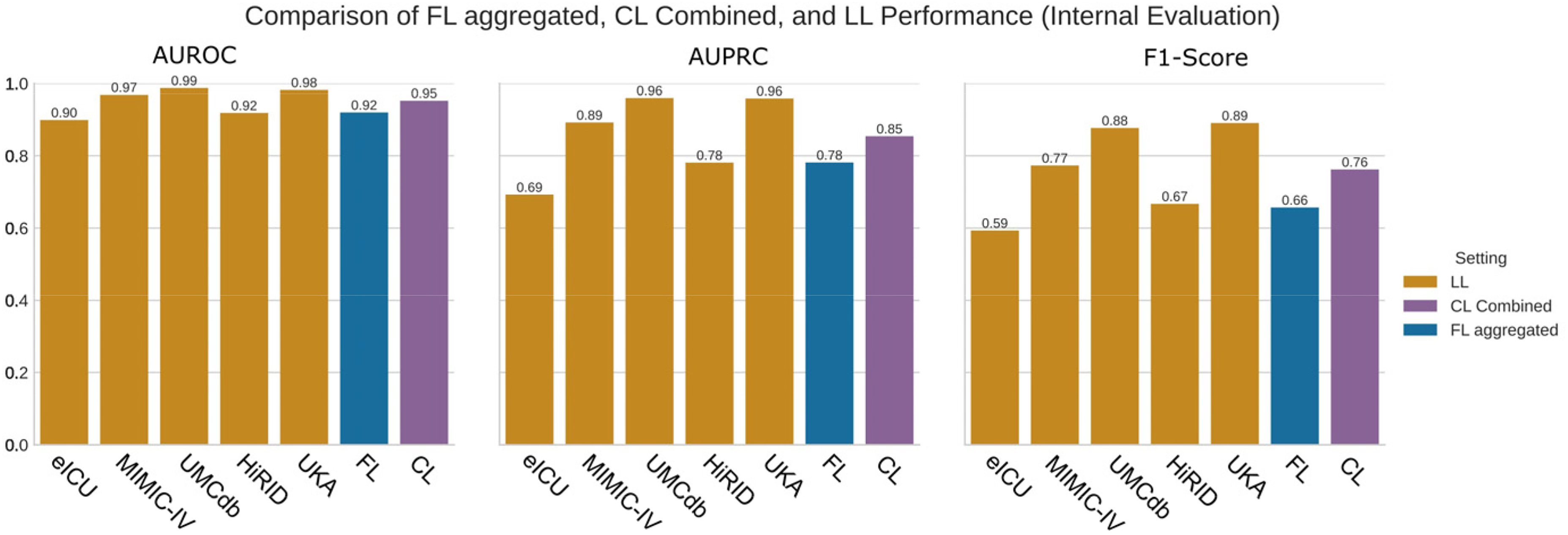
Comparison of the area under the receiver operator curve (AUROC), area under the precision-recall curve (AUPRC) and F1 score for local learning (LL), federated learning (FL), and centralized learning (CL). LL is trained and evaluated on the same database. CL is trained on a pooled, combined dataset and evaluated on a combined test set. FL is trained federated on all databases and evaluated on each database separately. FL is shown in an aggregated form, i.e. as average of the individual models.
F1 score comparisons followed similar patterns, with CL maintaining superior precision-recall balance across all datasets. The performance differences between methods were most pronounced on HiRID, where FL showed substantially lower F1 scores, around 0.41. On other datasets, the differences between methods were less dramatic, although CL consistently maintained the highest scores. In summary, internal validation F1 scores ranged from 0.41 to 0.89 across all combinations of dataset and learning approach, with CL consistently achieving the highest performance, followed by LL, and FL showing the most variable results. For a more detailed graphical comparison between FL, LL, and CL, see Supplemental Figures S1 and S2. To improve model interpretability, we computed SHAP values for the CL model. The resulting feature importance summary is provided in Supplemental Figure S3.
External validation
Having established baseline performance through internal validation, we next evaluated how well models generalized to unseen data. Figure 3 shows the AUPRC results of the external validation. For more detailed results, see Supplemental Tables S5 and S6.
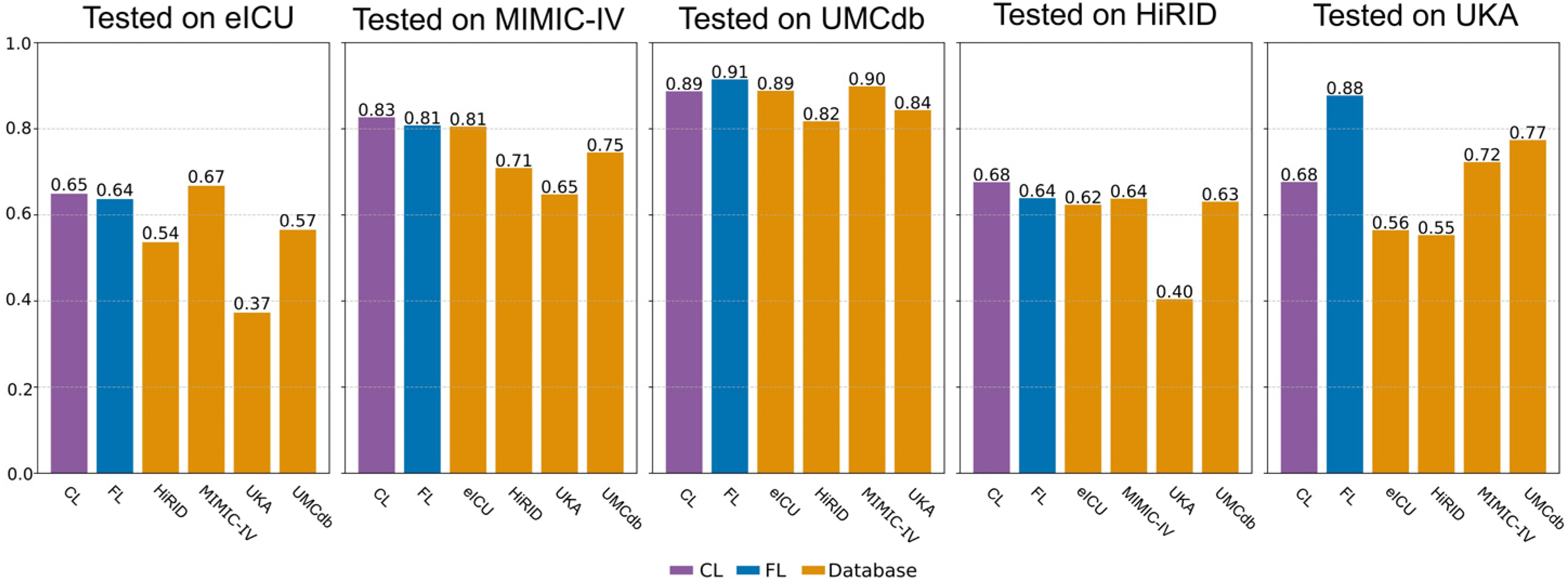
Area under the precision-recall curve (AUPRC) for the external validation setting. On the y-axis the database or method on which the model has been trained is shown.
As shown in Figure 3, CL generalized most effectively to unseen datasets, while FL exhibited inconsistent performance and LL failed to adapt across sites. Models tested on UMCdb achieved consistently high scores across most training datasets, with values frequently exceeding 0.80. In contrast, external validation performance on HiRID showed poor performance regardless of training approach, while UKA showed more variation depending on the training dataset. Specifically, F1 scores on HiRID ranged from 0.38 to 0.43 (see Supplemental Table S5), whereas testing on UKA yielded more variable results, with some training approach and training dataset configurations achieving F1 scores above 0.80. Overall, external validation performance demonstrated substantial variation depending on the specific dataset pairing, with certain combinations consistently outperforming others regardless of the learning approach. External validation AUPRC values ranged from 0.37 to 0.91 across all combinations of training and testing data and learning approaches. Across most training-testing combinations, CL achieved the highest performance, followed by FL, with LL showing the lowest performance in most scenarios. In summary, LL models demonstrated substantial limitations in cross-site prediction, CL consistently delivered robust performance across datasets, and FL showed inconsistent results across sites and metrics. Notably, FL did not consistently outperform LL in the external validation setting, contrary to our hypothesis.
Comparison of internal and external validation
To directly compare generalization performance across learning approaches, we examined the gap between internal and external validation results. Internal validation performance showed CL consistently achieving the highest AUPRC and F1 scores (see Figure 3 or Supplemental Tables S5 and S6) across most datasets, with FL demonstrating intermediate performance and LL showing the most variable results. A notable exception was the UKA dataset where FL substantially outperforms both CL and LL, achieving the highest internal validation scores across all datasets and learning approaches. In contrast, external validation revealed that CL and FL demonstrated comparable performance, both substantially outperforming LL across most training-testing combinations. LL consistently showed the poorest external validation performance, indicating limitations in cross-site generalization. Dataset-specific patterns emerged: UMCdb consistently served as the best-performing test dataset across most training scenarios, while HiRID as a test dataset demonstrated the poorest external validation performance regardless of the learning approach employed. The choice of training dataset also impacted external validation performance, with some datasets enabling robust generalization across multiple test sites while others showed poor cross-site transferability. These performance differences between learning approaches were more pronounced in external validation than in internal validation, highlighting the varying generalization capabilities of each approach. This pattern is summarized in Figure 4. The figure demonstrates that while all three learning approaches achieved similar internal validation performance (AUPRC: 0.81–0.85), external validation revealed substantial performance degradation across all methods. Nevertheless, CL and FL outperformed LL on average in cross-site generalization (AUPRC: 0.74 vs 0.66). To provide a more complete picture of model performance, we also assessed calibration via the Brier score and precision (Supplemental Figures S4 and S5). The Brier score analysis revealed that FL exhibited consistently poor calibration across both validation settings (0.28–0.29), whereas LL and CL achieved substantially better calibration, particularly in internal validation. Interestingly, despite this poor absolute calibration, FL showed the lowest standard deviation and the smallest performance drop between internal and external validation, suggesting more stable predictions across sites. Precision followed similar patterns to the metrics reported above, with CL achieving the highest values in both internal (0.73 ± 0.11) and external (0.65 ± 0.24) validation. This stability pattern was again evident for FL, it's precision remained relatively constant between validation settings (0.59 to 0.56), indicating that while its overall precision was lower, it generalized more consistently than LL, which dropped substantially from 0.68 to 0.50.

Comparison of internal and external validation performance across learning approaches. External validation (circles, solid lines, left) represents cross-site generalization performance with each dot showing AUPRC when a model trained on one dataset is tested on a different dataset. Internal validation (squares, dashed lines, right) shows performance when models are trained and tested on the same dataset. Range lines indicate minimum and maximum performance, while diamonds mark mean performance.
Discussion
We studied a general ICU cohort, rather than specific patient subgroups, based on clinical decision support considerations: algorithms developed for narrow populations face integration barriers in real-world ICU settings, where patient heterogeneity is the norm. A model applicable to diverse patients minimizes the friction clinicians’ face in deciding whether AI-assisted decision support applies to a given case. Additionally, we investigated whether increased model complexity when predicting BT requirements, involving multiple interacting clinical variables, impacts FL performance in external validation compared to models trained locally or centrally.
Our results reveal a nuanced picture. This study demonstrates that in the internal validation setting, CL consistently outperforms both FL and LL approaches for BT prediction across five ICU databases, while LL frequently outperforms FL, challenging the assumption that FL improves upon local models for complex clinical prediction tasks. Our findings contribute to the rapidly expanding field of ML in transfusion medicine. A recent review 23 identified 93 articles (56% published within three years), with 58% focusing on transfusion prediction. The review highlighted limited generalizability as a persistent challenge, which our multiinstitutional FL with external validation addresses. Consistent with existing LL studies, our internal validation results support the use of ML for BT prediction. Sheikhalishahi et al. achieved robust performance using XGBoost on UMCdb data (AUROC up to 0.85), while Rafiei et al. reported exceptional single-institution results with AUROC of 0.97, accuracy of 0.93, and F1 score of 0.89.1,4 However, these single-center approaches did not address challenges that arise in multiinstitutional settings, e.g. generalizability and statistical data heterogeneity. This present study contributes to improve the understanding of the role of FL in healthcare. A systematic review 10 of 612 FL healthcare studies found only 5.2% involved real-life applications, with most remaining proof-of-concept studies. The review identified considerable barriers to clinical translation, consistent with our findings that FL faces substantial challenges in complex clinical prediction tasks like BT prediction.
Comparative studies in healthcare FL provide perspective on our findings. Vaid et al. 24 showed federated models achieved comparable performance to centralized models for COVID-19 mortality prediction, but their data originated from the same health system, limiting heterogeneity compared to our international datasets. Levi et al. 7 described cross-database validation challenges for bleeding prediction, noting substantial class imbalance differences between datasets. Our external validation confirms these challenges: while CL maintains robust performance across sites, FL exhibits larger variability, particularly when testing on datasets with substantial distributional differences.
Statistical data heterogeneity and performance impact
The observed performance patterns align with measured data heterogeneity, supporting theoretical predictions about FL performance under non-IID conditions. Our KullbackLeibler Divergence analysis revealed strongest distributional differences between HiRID and other datasets, 25 which is consistent with FL's poor external validation performance on HiRID (F1 scores of 0.38–0.43 across all settings). This aligns with established theory that FL benefits from small, local datasets with IID data, while pathologically non-IID distributions favor local models. 26 The superior internal validation performance of LL compared to FL on individual datasets (particularly evident on HiRID, where LL achieved an AUPRC of 0.78 vs FL's 0.63) supports the theoretical expectation that local models perform better when evaluated on the same data they were trained on, especially under non-IID conditions. 26 Theoretical insights indicate that in non-IID scenarios, global model convergence relies predominantly on IID clients, while non-IID clients diminish both model accuracy and convergence speed. 27 However, existing optimization algorithms assume IID data, making them less suitable for FL's heterogeneity. Healthcare-specific challenges arise from differences in patient populations, clinical practices, and data collection procedures across institutions, 28 while mechanistically, FL's performance degradation stems from weight divergence that increases with greater distributional heterogeneity. 29 FL performance can degrade by 1055% under highly skewed medical data conditions, 2 suggesting that the heterogeneity inherent in multiinstitutional ICU data creates considerable non-IID conditions that challenge the effectiveness of FL in complex clinical prediction tasks such as BT prediction.
Complexity effects on FL performance
The contrast between our findings and simpler FL applications highlights how prediction complexity impacts FL effectiveness. While previous work on single-variable SpO2 prediction achieved FL performance matching CL, 11 our BT prediction task involving 15 clinical variables with complex interactions introduces substantially greater heterogeneity across institutions. This increased task complexity poses fundamental challenges for FL in diverse healthcare settings, explaining the suboptimal performance compared to centralized approaches. Sadilek et al. support this complexity-performance relationship, demonstrating that FL maintains equivalent performance to centralized models for simple tasks, but performance equivalence becomes harder to maintain as task complexity and data heterogeneity increase. 30
External validation context
External validation of clinical AI models typically relies on limited datasets, with most studies using only MIMIC-IV and eICU-CRD. 8 Our study addresses this limitation by incorporating five diverse international ICU datasets, enabling systematic analysis of cross-institutional performance patterns and statistical heterogeneity challenges. The performance degradation observed across all learning paradigms on external test sets, particularly HiRID, demonstrates the complexity of developing universally applicable clinical prediction models in heterogeneous healthcare environments. Importantly, CL maintained more robust performance across institutional boundaries compared to FL approaches, highlighting how data heterogeneity poses greater challenges for FL in complex clinical prediction tasks. These findings support recent calls for more nuanced approaches to clinical validation, where external validation datasets should be carefully chosen to reflect anticipated use cases rather than assuming universal generalizability. 31
Clinical decision support integration and future directions
Recent advances in clinical decision support systems have demonstrated substantial clinical impact with a 26% reduction in blood product costs through real-time EHR-integrated alerts, 32 underlining the importance of developing BT prediction models. However, our results suggest that FL's privacy-preserving advantages in healthcare are challenged by statistical data heterogeneity in complex clinical prediction tasks. Current FL algorithms like FedAvg use size-based weighted averaging that fails to account for data quality variations across institutions. 27 The consistently superior CL performance in our validation demonstrates that centralized approaches with pooled data remain more viable under present conditions. Future healthcare FL implementations require sophisticated client contribution assessment methods that consider not only data volume but also data quality as data completeness and accuracy impact classification model performance. 33
Conclusion
CL outperformed FL and LL consistently in predicting BTs across five ICU databases. For external validation, FL was slightly better than LL. The performance of the external validation corresponded inversely to the statistical data heterogeneity between training and test datasets. The more pronounced the distributional differences, the worse the external validation performance. Unlike simpler, single-variable applications, which have demonstrated clearer FL success, the complexity of this multivariable clinical prediction task poses substantial challenges to the effectiveness and generalizability of FL.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076261428383 - Supplemental material for Predicting blood transfusion after ICU admission in five databases: A comparison of three machine learning paradigms
Supplemental material, sj-docx-1-dhj-10.1177_20552076261428383 for Predicting blood transfusion after ICU admission in five databases: A comparison of three machine learning paradigms by Johanna Schwinn, Seyedmostafa Sheikhalishahi, Matthaeus Morhart, Iñaki Soto Rey, Philipp Simon, Mathias Kaspar and Ludwig Christian Hinske in DIGITAL HEALTH
Footnotes
Acknowledgments
We would like to express our sincere gratitude to the institutions that kindly provided database access: Beth Israel Deaconess Medical Center (MIMIC-IV), Universitätsklinikum Augsburg and Bern University Hospital (HiRID), Amsterdam University Medical Center, and the critical care units contributing to the eICU Collaborative Research Database.
Ethical considerations
This study was conducted according to the institutional ethical guidelines. Ethical approval for the UKA data was obtained from the responsible Ethics Committee at the Ludwig-Maximilians-Universitat München (Reference: 23-0969, Date: 05 January 2024). Thë eICU-CRD and MIMIC-IV databases are publicly available, deidentified datasets that can be accessed through PhysioNet after completing the required training courses. The HiRID and UMCdb datasets were accessed under preexisting institutional agreements for secondary data analysis. All datasets contain deidentified patient data, complying with applicable privacy regulations. No further patient consent was mandatory, as all data were anonymized before analysis.
Use of AI
The AI tool claude.ai was used to assist with language editing, proof-reading, section feedback and to improve the natural flow of the text. Additionally, claude.ai was used to format LaTeX tables. DeepL supported grammatical refinement and word selection.
Author contributions
Johanna Schwinn: conceptualization, data curation, formal analysis, investigation, methodology, validation, writing—original draft, and writing—review and editing; Seyedmostafa Sheikhalishahi: investigation, writing—review and editing, and supervision; Matthaeus Morhart: data curation, and writing—review and editing; Iñaki Soto Rey: funding acquisition, and writing—review and editing; Philipp Simon: writing—review and editing; Mathias Kaspar: investigation, methodology, project administration, supervision, writing—review and editing; Mathias Kaspar and Ludwig Christian Hinske contributed equally to this work; Ludwig Christian Hinske: conceptualization, funding acquisition, methodology, resources, project administration, writing—review and editing; all authors have read and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the German Ministry of Education and Research, BMBF (Grant No. 01ZZ2005).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
