Abstract
Objective
Ulcerative colitis (UC) endoscopic image classification presents challenges owing to imbalanced medical imaging data, particularly when the clinical importance of accurate positive predictions increases with disease severity. This study proposes a multi-task learning (MTL) framework inspired by the coarse-to-fine processing mechanism of the human brain to address these challenges.
Methods
The proposed MTL framework was evaluated using endoscopic images of UC, focusing on its ability to classify disease stages with an emphasis on the accurate detection of advanced cases.
Results
Our findings demonstrate that the proposed framework effectively mitigates the limitations posed by imbalanced datasets, particularly by enhancing classification performance in severe disease stages. Notably, DenseNet121 exhibited significantly superior performance compared to other backbones and achieved an additional performance gain in identifying Mayo endoscopic scores of 2 and 3 following joint-loss optimization. Additionally, MobileNet-v3-large, despite being a lightweight model, demonstrated notable gains under the proposed optimization scheme, highlighting the versatility of the framework across architectures with different computational complexities.
Conclusion
MTL-based computer-aided diagnosis enables conservative and accurate identification of patients in critical stages, supporting timely and appropriate treatment decisions while reducing the risk of underdiagnosis and delayed care. Furthermore, our results highlight the potential of MTL in overcoming data imbalance issues. Future studies should explore integrating multiple convolutional neural network-based models to further boost classification accuracy.
Keywords
Introduction
Ulcerative colitis (UC) is a chronic inflammatory bowel disease characterized by periods of remission and exacerbation throughout a patient's lifetime. 1 Its clinical manifestations include rectal bleeding, diarrhea, and abdominal pain. The disease typically involves the rectal mucosa and is characterized by a continuous pattern of lesions. Traditionally, the therapeutic goal for patients with UC has been defined as clinical remission, which focuses on symptom alleviation. Recent studies have highlighted the importance of mucosal healing as a critical therapeutic objective.2–4 Mucosal healing has been shown to influence the selection of therapeutic agents, reduce hospitalization rates, decrease the frequency of surgical interventions, and improve overall patient prognosis. 5
Mucosal healing is defined as the complete resolution of visible inflammatory findings on endoscopic examination. Among the various endoscopic scoring systems, the Mayo endoscopic score (MES) is one of the most widely used tools owing to its simplicity and extensive validation in a previous study. 6 It offers a kappa coefficient of 0.5, indicating relatively fair inter-observer agreement, and evaluates features such as vascular pattern, ulceration, and bleeding to assign a score ranging from 0 to 3. 7 Although MES is a clinically practical and relatively efficient quantitative metric, it remains subject to subjective interpretation and the expertise of the endoscopist, resulting in variability based on operator proficiency. 8 To address this issue, researchers and practitioners have developed computer-aided diagnosis (CAD) systems that aim to reduce subjectivity and improve assessment consistency.9–11
Recent studies have explored the application of artificial intelligence (AI) models trained on large-scale endoscopic datasets to CAD systems designed for both endoscopic12–14 and surgical procedures.15–17 Although some studies have used proprietary datasets to train AI models, the difficulty in acquiring large-scale datasets has led many researchers to utilize publicly available labeled images for ulcerative colitis (LIMUC) datasets for training and evaluating AI models. A recent study introduced a learning framework for classification models that goes beyond merely comparing model performance. Regression-based deep learning approaches have demonstrated enhanced performance, offering a promising solution for overcoming the inherent variability associated with human interpretation. 18 The architecture of the learning model and the framework for the learning method can contribute to superior performance, even when using the same model. Nevertheless, the proposed learning framework, which requires performance improvements and justification for its explainability, raises theoretical concerns regarding the consistency of the model structure, particularly in regression-based models that classify outcomes by applying a specific threshold to the final continuous output value.
First, the increased likelihood of misclassifying a class arises from minor errors near the threshold, which could exacerbate the problem, particularly when the data are densely distributed. Second, if continuous scores are concentrated within certain ranges of the dataset, certain classes—particularly those near the threshold—may become underrepresented or overrepresented, potentially resulting in imbalances in model performance. Third, class boundaries may not always be clearly defined in clinical data. For example, if the boundary between MES 1 and MES 2 is ambiguous, a regression-based approach might struggle to make clinically meaningful decisions. To address these issues, an alternative framework based on the multi-task learning (MTL) approach is validated, offering improved explainability and superior performance. The remission state is first classified in this framework, followed by a detailed subclassification of both the remission and active states. This approach addresses the challenges observed in previous studies and provides a hierarchical and clinically meaningful classification process.
MTL typically employs a shared network architecture with separate branches for each task, enabling the model to generalize more effectively by leveraging the complementary nature of the tasks. It facilitates the learning of shared representations across tasks while simultaneously capturing task-specific features. The key idea is to leverage the shared information between tasks to improve the overall performance, particularly when individual tasks do not have sufficient data for effective learning. By leveraging the shared information between different MES classes, MTL can improve the accuracy and robustness of the model. Therefore, MTL can be particularly suitable for clinical settings where large-scale learning datasets are not readily available or where there is an imbalance in the dataset between classes. In medical image analysis, MTL has been successfully applied in various domains. 19 Prominent examples include training the model to perform both segmentation and classification of tumors 20 or organs. 21 These examples highlight the potential of MTL in enhancing the performance of deep learning models in medical image analysis, making it particularly relevant for complex tasks, such as the estimation of MES in UC. Considering the similarity between the MTL approach and human object perception and classification processes, such as when an operator first examines an endoscopic image, determines whether the severity of colitis is significant, and then further assesses the degree of severity, evaluating MES classification performance using an MTL learning framework is justified.
Moreover, to enhance the performance of the model during training and in alignment with the second objective of this study, joint-loss function optimization was explored by fine-tuning the model with a pre-trained network on the same task using the F1 score. This approach enables the model to use previously learned features from a broader dataset, thereby enhancing its ability to generalize and detect subtle patterns. This technique can potentially enhance the precision and recall of the model, particularly for underrepresented classes. Furthermore, this may enable the model to focus on subtle features that are critical for classification. Such a strategy not only improves the overall performance of the model but also helps maintain the robustness of the framework across diverse class distributions, which remains a persistent challenge and is directly addressed in this study. In addition, most previous medical image classification studies have employed ordinal regression approaches to address the multiclass nature of the task, leveraging the inherent order among class labels, particularly in relation to disease severity.22–24 Although these methods reflect the ordinal structure, they often rely on rigid label ordering and threshold-based outputs, which may introduce challenges in interpretability and lead to performance instability, particularly in the presence of class imbalance.25,26
Therefore, this study aimed to identify key features predictive of disease outcomes by proposing an efficient deep learning framework based on the MTL approach, which closely mimics the human coarse-to-fine reasoning process. Specifically, the proposed framework first performs a global assessment of endoscopic images to distinguish broad categories, such as the presence or absence of inflammation (coarse level). Subsequently, it conducts a more detailed analysis to quantify the severity and pattern of inflammation (fine level). This hierarchical approach reflects how clinicians intuitively process medical images, starting with general impressions and refining their evaluations through focused observations. Using this method, this study aims to contribute to the development of personalized and timely treatment strategies.
Methods
Proposed framework
The systematic framework for classifying MES from endoscopic images of patients with UC is illustrated in Figure 1. It begins by exploring the datasets used and the preprocessing techniques applied to the images, including noise reduction and normalization, to enhance image quality. Following preprocessing, the images are labeled for MTL learning and split into training (85%) and testing (15%) datasets. The training dataset is subjected to 10-fold cross-validation, and class-wise F1 scores from each fold are used to fine-tune the model through joint-loss optimization. After training, the fine-tuned model is evaluated using the held-out test dataset to assess its final performance.

Schematic of MTL framework for UC MES classification. UC: ulcerative colitis; MTL: multi-task learning; MES: Mayo endoscopic score.
Dataset
The LIMUC dataset is a comprehensive collection of 11,276 endoscopic images from 564 patients who underwent 1043 colonoscopy procedures. The images were obtained using a Pentax EPK-i video processor and a Pentax EC-380LKp video colonoscope with a resolution of 352 × 288 pixels.
The images in this dataset were selected at different time points during colonoscopy, and there was no spatial relationship between the images from the same patient, resulting in a high level of heterogeneity. In addition, images deemed unsuitable for evaluation owing to the presence of intestinal contents, artifacts, or other factors were excluded, thereby ensuring the overall quality of the data. Furthermore, to enhance the reliability of the ground truth and minimize inter-observer variability, each image was independently assessed by two experienced gastroenterologists using the MES system. In cases where the two reviewers disagreed, a third independent expert, blinded to the previous evaluations, provided an additional assessment, and the final MES label was determined by majority voting. Although the public dataset did not include quantitative measures of inter-observer agreement, this consensus-based labeling process was adopted in prior studies to reduce subjectivity and improve the consistency and reliability of the annotations used for model development.
The final dataset consisted of images labeled as MES 0 (no inflammation), MES 1, MES 2, and MES 3, representing increasing severity of inflammation. The dataset distributions were as follows: MES 0–6105 images (54.14%), MES 1–3052 images (27.07%), MES 2–1254 images (11.12%), and MES 3–865 images (7.67%).
The LIMUC dataset is publicly available, making it the largest publicly labeled dataset for UC endoscopic images. This dataset serves as a valuable resource for developing and evaluating algorithms aimed at improving UC diagnosis and treatment. Given its comprehensive labeling process and diverse image collection, LIMUC offers an ideal platform for training machine learning models in the medical image analysis domain.
Image preprocessing
Image normalization adjusts pixel intensity values to standardize their distribution across channels, thereby enhancing data uniformity for model training. During preprocessing, endoscopic image datasets collected under varying conditions, such as differences in illumination, reflections from internal fluids, and variations in camera distance or angle, often exhibit inconsistencies in intensity values and object scales. To address these issues, the mean and standard deviation of each channel in the training dataset were calculated and the images were normalized accordingly. Additionally, data augmentation techniques, including random horizontal flipping and random rotation within −180° to 180°, were employed to simulate diverse imaging conditions and improve model robustness. Figure 1 shows the normalized endoscopic images obtained after preprocessing.
Hierarchical labeling for image classification
This study used a hierarchical labeling framework to classify the endoscopic images of UC to improve the precision of disease severity and remission status prediction. Our methodology involved a three-level classification system: (1) binary classification differentiating between remission and active disease, (2) finer classification within the remission group (MES 0 vs. MES 1), and (3) detailed categorization within the active disease group (MES 2 vs. MES 3). This hierarchical approach enables the model to learn specific characteristics at each level, leading to more accurate and granular predictions. This structured classification scheme offers significant potential for enhancing clinical decision making, providing a more detailed understanding of disease progression, and supporting timely and effective treatment strategies.
Image classification model
The MTL framework was used to classify colonoscopy images hierarchically to identify the remission and active states of UC. The proposed model performs binary classification at the primary task level and classifies remission and active states into subcategories. To validate its effectiveness, ResNet18 was implemented as the baseline feature extractor, and its performance was compared with that of state-of-the-art architectures, including ResNet50, DenseNet121, Inception-v3, and MobileNet-v3 Large, which have been employed in similar studies on UC endoscopic image classification. The MTL architecture employed in this study integrates a shared feature extractor and task-specific classification head. The shared feature extractor leverages convolutional layers from various backbone networks—ResNet18, ResNet50, and DenseNet121—pre-trained on the ImageNet dataset to learn common representations from the input images. For each backbone model, the final fully connected layer was removed and replaced with an identity layer, enabling the extraction of task-agnostic feature vectors with varying dimensions (512 for ResNet18 and 2048 for ResNet50). These feature vectors were subsequently used as inputs for the three task-specific classification heads.
The first task, primary classification, involved a binary classification problem to distinguish remission from active states. The second task, remission subclassification, was an ordinal classification task designed to further categorize the remission states into finer subcategories. Finally, the third task, active subclassification, employed ordinal classification to differentiate and categorize active states into specific subcategories. Together, these tasks formed a comprehensive approach to address the complexities of multi-label classification in the dataset.
Fine-tuning using joint-loss optimization
To optimize the multi-task model, the joint-loss function, which is defined as the sum of the task-specific losses, was employed:
The training and testing datasets were split at an 85:15 ratio, and the training dataset was further evaluated using 10-fold cross-validation to ensure robust model validation. Stratified sampling was applied during the dataset split to preserve the class distribution across all tasks.
For remission and active subclassification, masking techniques were applied during the loss calculation to exclude invalid labels (−1) from the computation. This approach ensures that the loss function accurately captures the hierarchical and ordered nature of these classification tasks, where −1 indicates that a sample is irrelevant to a specific task. For example, patients not exhibiting active disease were assigned −1 for the active subclassification task.
After handling task irrelevance through masking, we proceeded to model training. Following data augmentation to improve the robustness, the models were trained using the Adam optimizer with a learning rate of 0.0002. Early stopping was applied to prevent overfitting, and task-specific metrics, such as accuracy, were computed solely for valid data points.
Evaluation metrics
The performance of the model was assessed using several evaluation metrics, including accuracy, precision, recall, F1-score, and specificity, derived from the true-positive, false-positive, true-negative, and false-negative values.
For the primary classification task, which involved distinguishing remission states from active states, precision, recall, F1-score, and support were calculated to evaluate the performance of the model across the two classes. Additionally, confusion matrices were generated for each classification task to provide detailed insights into the classification behavior of the model. For the ordinal classification tasks, namely, remission and active-state subclassifications, precision, recall, F1-score, and support were computed similarly, with individual metrics evaluated for each subcategory. Additionally, the macro-averaged accuracy, precision, recall, and F1-score were calculated to summarize the overall performance across all subcategories of each task. These metrics provide a comprehensive measure of the ability of the model to accurately classify primary and subcategory labels. Furthermore, the specificity was computed to evaluate the ability of the model to identify true positives and negatives across all classification tasks.
Results
This section presents the most suitable model for UC MES MTL classification, focusing on the evaluation metrics for each deep learning model and performance changes following loss function optimization based on initial learning and the F1 score. Additionally, it provides a comparative analysis between DenseNet121, which was the top-performing model in previous studies, and the model trained using the proposed framework on the same dataset. All the models utilized the Adam optimizer with a learning rate of 0.0002 and no weight decay (0.0). Early stopping was set to a threshold of 25 epochs to prevent overfitting. The training was conducted over 200 epochs, with learning rate scheduling applied; if there was no improvement in validation loss for 15 epochs, the learning rate was reduced by a factor of 0.2.
Python (3.10.12) was employed for training and evaluating the system on a machine equipped with an Intel Xeon CPU @ 2.20 GHz, running 64-bit Linux. The machine featured 83.48 GB RAM and was enhanced with an NVIDIA A100-SXM4-40GB GPU with CUDA version 12.2 for efficient computation. For dataset preprocessing and image construction, we used NumPy (2.0.2) for numerical operations, Pandas (2.2.2) for data management, and OpenCV (4.11.0) and PIL (Pillow 11.2.1) for image loading and transformation. Custom dataset handling was implemented via PyTorch's torch.utils.data.Dataset class. We also utilized Matplotlib (3.10.0) for plotting and visualizations, and SciPy (1.15.3) for statistical analysis (i.e. Wilcoxon rank-sum test). Model training and evaluation were conducted with PyTorch (2.6.0 + cu124).
Class-wise evaluation metrics
This section presents the precision, recall, and F1 score for MES classes derived from the UC datasets using the MTL framework before joint-loss optimization (α = β = γ = 1/3 in Equation (1)). Additionally, the convolutional neural network (CNN)-based model for the MTL framework was compared across various performance metrics, as presented in Table 1. The accuracy of each classifier in the MTL framework was greater than 0.9, except for Inception-v3.
10-fold CV accuracy results of MTL classification models for UC scores.

Best value.
UC; ulcerative colitis; MTL: multi-task learning.
Furthermore, as shown in Figure 2, DenseNet121 consistently outperformed other models across all MES classes in terms of precision, recall, F1-score, and specificity. Statistically significant differences were observed among most model comparisons, as indicated by the asterisks, suggesting the robustness of DenseNet121 within the MTL setting.
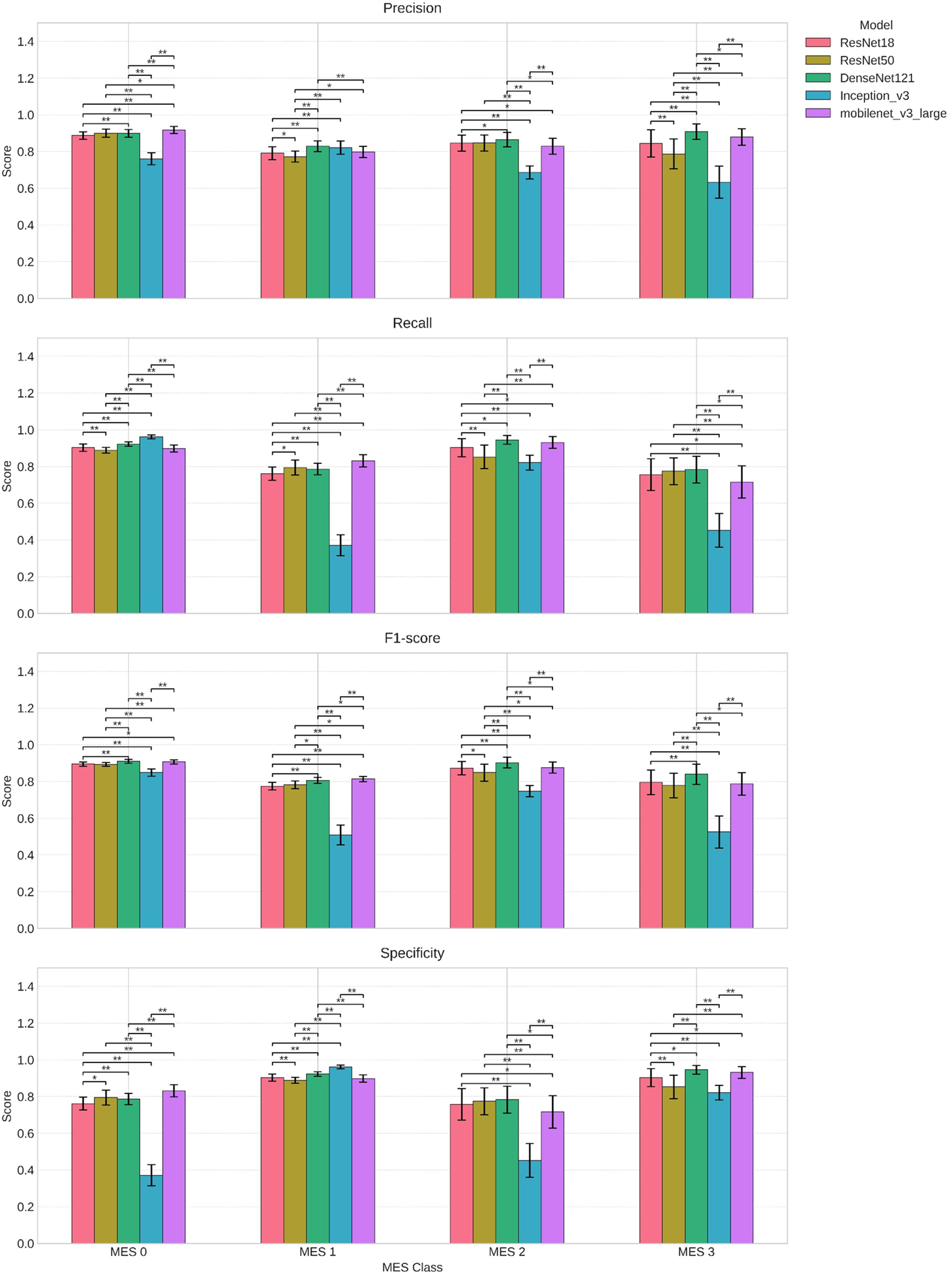
Class-wise performance across evaluation metrics during 10-fold cross-validation for models.
F1-based loss function optimization
This section proposes increasing the weight of the loss function for the class with a lower F1 score (based on the inverse of the F1 score) to enhance class-specific learning performance through fine-tuning. Specifically, the loss function is adjusted by applying a weight factor proportional to the inverse of the F1 score for each class. This adjustment prioritizes lower performance classes, directing the model's attention toward improving the sensitivity of the underperforming categories. As shown in Figure 3, although performance changes varied across the models, an overall shift in the performance of each model with respect to the imbalanced classes was observed. By emphasizing these classes during fine-tuning, the performance disparity across different classes was partially alleviated, resulting in a modestly balanced classification system.

Performance change after F1-based loss function optimization.
Comparison of macro-performance with previous results based on Densenet121
This section compares the proposed framework with the findings of a recent study that used the same LIMUC dataset. Polat et al. applied a regression-based approach using five different CNN architectures, with DenseNet121 yielding the best performance. 18 As presented in Table 2, the same model was used, and the proposed learning framework, along with F1-based joint-loss optimization, was applied. The performance of each class was then compared. Consequently, the framework proposed in this study demonstrated a balanced improvement in the classification performance of MES 2 and MES 3, which were characterized by relatively small sample sizes.
F1-score differences across MES classes using the same backbone (DenseNet121).
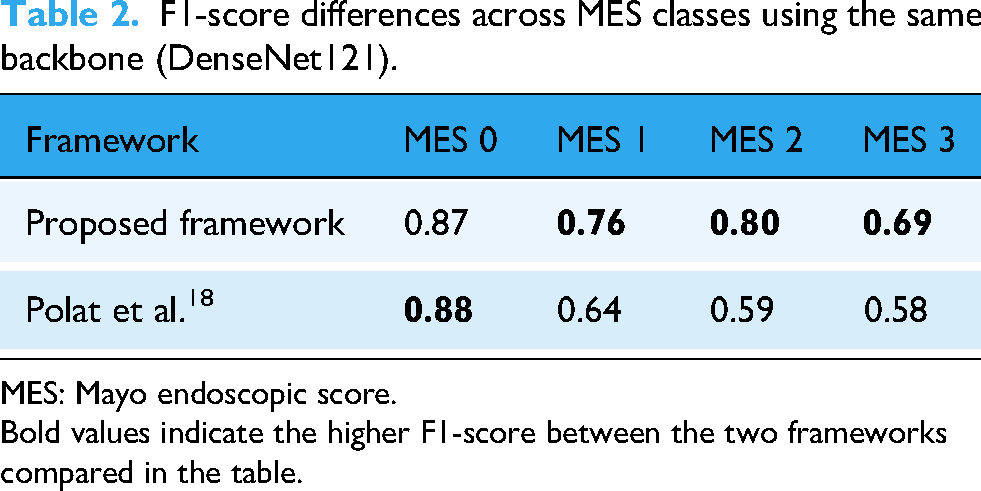
MES: Mayo endoscopic score. Bold values indicate the higher F1-score between the two frameworks compared in the table.
Discussion
Endoscopy is the preferred modality for UC assessment because it provides real-time visualization of the bowel wall characteristics and can effectively diagnose various conditions. Studies focusing on UC prognosis have highlighted its ability to capture key features, such as bowel wall thickness, vascularization, and mucus, which indicate disease severity. Researchers have reported encouraging results regarding predictive performance and clinical applicability by integrating these imaging features into machine learning models. Various methods for diagnosing UC and predicting its prognosis have been extensively studied, focusing on combining medical imaging and deep learning techniques to develop automated prediction systems.27–30 Deep learning architectures based on CNNs have demonstrated remarkable efficacy in classifying imaging data by detecting subtle patterns in imaging features. 31 Additionally, various methodologies have been proposed to improve the accuracy of MES prediction using UC images.
To address the challenges posed by imbalanced medical imaging data, we developed an MTL framework and evaluated its performance across various backbone networks. Our results showed that DenseNet121 delivered strong performance across most metrics and classes. In addition, it exhibited the most substantial gains after the application of joint optimization, particularly in the F1-scores for MES 2 and MES 3. Furthermore, our study demonstrated that the proposed framework, when using the same backbone (i.e. DenseNet121), significantly improved F1-scores for the active-state classes, which are clinically important yet severely underrepresented owing to class imbalance, as compared to a recent study. 18 These findings indicate that our optimization strategy is particularly effective in enhancing the detection performance for low-prevalence but clinically significant severe UC states by improving both the precision and recall.
Regarding MES 2 and MES 3, which represent the active stages of UC, ensuring the accurate measurement of disease severity is essential for timely intervention. MES 3 is characterized by extensive inflammation and substantial tissue damage, underscoring the critical need for accurate identification for prompt and appropriate treatment. In this context, a positive classification (indicating the presence of severe disease) is a safer and more reliable approach, ensuring that patients at this stage are not overlooked and receive the necessary therapeutic intervention without delay. 32 Conversely, a negative classification for MES 3 could lead to an underestimation of the disease severity, potentially resulting in treatment delays and deterioration of the patient's condition. Therefore, for MES 3, positive classification is crucial to ensure that patients are diagnosed correctly and promptly, ultimately enhancing clinical outcomes and patient management.
The interpretability of our model was supported by Grad-CAM visualizations, which revealed that the proposed framework consistently attended to diagnostically meaningful features. As shown in Figure 4, the model primarily focused on the inflamed peripheral mucosal regions rather than the central lumen when predicting MES 3 cases. This alignment with clinical observations 33 suggests that our framework not only performs well but also provides insights that are explainable and potentially clinically actionable.

Grad-CAM visualizations illustrating model attention during MES 3 prediction. MES: Mayo endoscopic score.
In our study, the effects of the F1-based joint-loss optimization varied across models. Model-wise, ResNet18, DenseNet121, and Inception-v3 exhibited notable improvements in both the F1 score and recall for MES 3, suggesting that these architectures responded well to the re-weighted optimization strategy. ResNet50 and MobileNet-v3-large exhibited modest changes across the performance metrics. This finding suggests that, for certain architectures, the hierarchical MTL framework alone may offer sufficient robustness to distinguish UC states under class imbalance, indicating that additional optimization strategies may not always be necessary.
MobileNet-v3-large, despite being a lightweight model, exhibited relatively strong baseline performance even without joint-loss optimization and showed notable improvements following joint optimization, particularly in differentiating the remission-active boundary between MES 1 and MES 2, a task known to be challenging even for expert endoscopists. This suggests that robust performance can be achieved not only through complex, heavy architectures but also through compact models when guided by a carefully designed learning framework. Given its computational efficiency, MobileNet-v3-large demonstrates particular promise for deployment in resource-constrained settings, highlighting both the versatility and practical clinical utility of our proposed strategy.
Recent studies have proposed diverse strategies to improve UC severity prediction. A ResNet50-based system 29 was developed to improve spatial coverage by scoring inflammation across segmented regions of the colon in full-length endoscopic videos. Turan and Durmus 34 introduced UC-NfNet with synthetic data augmentation to address inter- and intra-observer variabilities in MES grading. More recently, Qi et al. 35 developed a transformer-based model (UC-former) that achieved near-perfect classification accuracy across all MES categories, outperforming human endoscopists.
Although these approaches offer important advances in model architecture, data augmentation, and spatial coverage, they generally treat MES prediction as a single-task problem and do not directly address class imbalance at the training level. In contrast, our study introduces a novel hierarchical label design and MTL framework that explicitly incorporates the ordinal nature of MES and applies F1-based joint-loss optimization to improve sensitivity for underrepresented, yet clinically critical, active-state classes (MES 2 and MES 3). This approach provides a flexible and interpretable training structure that complements existing architectural advances, while offering a new direction in learning formulations for imbalanced clinical data. Collectively, these findings establish a foundation for more equitable and clinically meaningful AI-based UC assessment strategies.
While our model demonstrates strong classification performance, its clinical utility extends beyond metrics. Although AI is gaining attention in medical education, a recent study indicated that healthcare professionals and trainees often lack sufficient knowledge and confidence to effectively engage with AI systems. 36 This gap poses a barrier to clinical adoption and highlights the need for interpretable and user-aligned tools.
Recent studies have shown that AI models can improve diagnostic consistency and workflow scalability when seamlessly integrated with clinical infrastructures.37–39 In practice, such a system can be embedded into a picture archiving and communication system (PACS) for real-time MES scoring during colonoscopy procedures, providing endoscopists with immediate and objective feedback. Furthermore, integration with electronic health record (EHR) systems would enable the automatic documentation of disease activity, thereby supporting longitudinal tracking and personalized treatment adjustments. For instance, accurate MES classification can inform therapeutic decisions, such as escalation to biologics for MES 2–3 or de-escalation in MES 0–1.
From a regulatory perspective, considerations, such as explainability and reliability, remain critical for deployment in clinical environments. Our model addresses this challenge by providing intuitive visual feedback through Grad-CAM, making it more accessible to clinicians with limited AI expertise, and thus more likely to be trusted and adopted in practice. Previous studies have emphasized that explainable AI approaches are essential for securing clinician trust and ensuring patient safety in clinical settings. 40 Our use of Grad-CAM visualizations, which highlight class-discriminative regions in endoscopic images, allows clinicians to qualitatively assess the focus of the model and enhance its transparency. Furthermore, the use of a lightweight architecture, such as MobileNet-v3, validated in our study to deliver strong performance, directly addresses concerns related to reliability, scalability, and real-time deployability. Its computational efficiency and operational stability make it particularly suitable for integration into clinical infrastructures, such as PACS and EHR systems, while also supporting compliance with regulatory expectations for consistent behavior and model transparency.
Limitations and prospects
Despite the significance of the findings, the following limitations suggest directions for future research. First, the dataset exhibited a significant class imbalance, with an underrepresentation of classes such as MES 3, which complicates the achievement of consistent performance across all categories. Second, despite the involvement of multiple reviewers, image classification remained partially subjective, potentially introducing bias. Third, this study relied on 2D endoscopic images, which may not fully capture the complexity of 3D anatomical structures, thereby limiting its clinical applicability. Fourth, while the study compared its framework with prior works, it did not incorporate recent state-of-the-art models, such as transformer-based architectures or hybrid approaches. Fifth, the framework exhibited reduced specificity for critical classes, which could affect its reliability in distinguishing between severe cases. Sixth, the study did not thoroughly explore the explainability of the model outputs, which is a crucial factor in clinical settings. Furthermore, it did not assess the performance of the model in real-time scenarios, which is essential for its clinical adoption. Seventh, the performance heavily depended on preprocessing techniques; however, no analysis has been provided on how changes in preprocessing may affect the results. Eighth, although the model demonstrated promising performance on the single-center dataset, its generalizability to external settings has not yet been assessed.
Future studies can be categorized into four key areas. First, data and model enhancement should address class imbalance through advanced data augmentation or synthetic data generation, integrating 3D endoscopic or multi-modal data for better anatomical representation, and incorporating state-of-the-art models, such as transformers. Although our current study focused on conventional CNN-based backbones to enable a fair comparison with prior UC studies, future validation of transformer-based or hybrid architectures is essential to confirm the generalizability of our framework. Additionally, improving the specificity of critical classes through strategies such as focal loss remains a key direction. Second, the subjectivity of image classification can be reduced by using objective evaluation methods involving a diverse set of reviewers. Third, model generalizability and clinical applicability should be addressed by validating the framework with multi-center or multi-device datasets and evaluating its robustness on full-length endoscopy videos. Incorporating domain adaptation techniques and calibration strategies may improve its reliability in diverse clinical settings. Fourth, analyzing the impact of preprocessing techniques on model outcomes is crucial for optimizing and ensuring clinical adoption.
Conclusion
In conclusion, this study suggests that MTL can serve as an effective learning framework for classifying UC endoscopic images, particularly when combined with F1-based joint-loss optimization. This approach appears to be beneficial in addressing the challenges associated with imbalanced data, although further validation is warranted. The results also indicate that different CNN-based models exhibit varying strengths across distinct MES classes. Based on these findings, future studies could explore the potential of integrating multiple models within an MTL framework to leverage their complementary capabilities. Such an approach could contribute to improving classification performance and offer a more comprehensive strategy for supporting UC diagnosis.
Footnotes
Ethical considerations
This study used publicly available datasets that are fully anonymized and do not contain any personally identifiable information. Therefore, ethical approval was not required.
Author contributions
Conceptualization was done by JL and EK; methodology done by EK; software done by JL; validation done by EK; formal analysis done by JL; investigation done by JL and EK; resources done by EK; data curation done by EK; writing—original draft preparation done by JL; writing—review and editing done by EK; visualization done by JL; supervision done by EK; project administration done by JL and EK; funding acquisition done by JL and EK.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported in part by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (RS-2023-00275579), and in part by the research fund of Hanyang University (HY-202400000003726).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.

