Abstract
Objective
The study aims to present an active learning approach that automatically extracts clinical concepts from unstructured data and classifies them into explicit categories such as Problem, Treatment, and Test while preserving high precision and recall and demonstrating the approach through experiments using i2b2 public datasets.
Methods
Initially labeled data are acquired from a lexical-based approach in sufficient amounts to perform an active learning process. A contextual word embedding similarity approach is adopted using BERT base variant models such as ClinicalBERT, DistilBERT, and SCIBERT to automatically classify the unlabeled clinical concept into explicit categories. Additionally, deep learning and large language model (LLM) are trained on acquiring label data through active learning.
Results
Using i2b2 datasets (426 clinical notes), the lexical-based method achieved precision, recall, and F1-scores of 76%, 70%, and 73%. SCIBERT excelled in active transfer learning, yielding precision of 70.84%, recall of 77.40%, F1-score of 73.97%, and accuracy of 69.30%, surpassing counterpart models. Among deep learning models, convolutional neural networks (CNNs) trained with embeddings (BERTBase, DistilBERT, SCIBERT, ClinicalBERT) achieved training accuracies of 92–95% and testing accuracies of 89–93%. These results were higher compared to other deep learning models. Additionally, we individually evaluated these LLMs; among them, ClinicalBERT achieved the highest performance, with a training accuracy of 98.4% and a testing accuracy of 96%, outperforming the others.
Conclusions
The proposed methodology enhances clinical concept extraction by integrating active learning and models like SCIBERT and CNN. It improves annotation efficiency while maintaining high accuracy, showcasing potential for clinical applications.
Keywords
Introduction
Daily, a huge amount of data is generated in the clinical domain from clinical reports, scientific research, and clinical databases such as electronic health record (EHR). 1 These data, mostly unstructured, contain hidden information and knowledge that might help solve clinical questions about patient health conditions, clinical reasoning, and inferencing. However, extracting information from unstructured data is a challenging task due to data heterogeneity, variability, and ambiguity. To achieve the goal of meaningful use, transforming routinely generated clinical reports, scientific research, and EHR data into actionable knowledge requires systematic approaches. 2
Researchers have proposed and utilized different techniques and methodologies to extract hidden information and convert it into actionable knowledge by performing autonomous computational extraction. 3 In the clinical domain, Information Extraction (IE), a sub-field of natural language processing (NLP), pertains to the automated extraction of predefined clinical concepts from unstructured clinical text. This process, known as clinical concept extraction, encompasses both concept mention detection and concept encoding. The named entity recognition (NER) techniques as referenced in Navarro et al. 4 are usually employed for detecting concept mentions within the broader domain. The NER approach effectively identifies concept mentions in clinical textual data, encompassing categories such as “problem” (signs or symptoms, findings, disease or syndrome, etc.), “treatment” (organic chemicals, diagnostic procedures, and/or pharmacological substances), and “test” (laboratory procedures and clinical attributes). 5 Concept encoding aims to map the mentions to concepts in standard terminologies or those defined by downstream applications.6,7 Concept extraction has been adopted to extract clinical information from text for a wide range of applications, ranging from supporting clinical decision-making to improving the quality of care, achieving better clinical outcomes, and providing time and budget-constrained services to the community. 8
Methods for developing clinical concept extraction and classification applications have been largely adopted from the general NLP domain 9 and can typically be distinguished into rule-based approaches and statistical approaches with four categories: rule-based, traditional machine learning (ML) (non-deep-learning variants), deep learning (DL), or hybrid approaches. The main ingredient of a rule-based system is knowledge-based, relying on rules created by domain experts, and is considered highly efficient in exploiting language-related knowledge characteristics. 10 Similarly, rule-based approaches play an important role in preparing the initial level of annotated data for data-driven approaches. For this purpose, a lexical-based approach is extensively used to identify relevant semantic information for explicit terminology such as “cancer,” whose semantic type is “Neoplastic Process” in Unified Medical Language System (UMLS). Furthermore, rules can be applied to explicitly categorize the biomedical term “cancer” as a clinical concept under “Problem” based on semantic types. Likewise, rule-based methods are effective in clinical settings due to their specialized language properties. However, it can be laborious developing a system that requires both technical NLP experts and clinical specialists to work together. Moreover, the final applications may have limitations in terms of portability and generalization beyond the scenario for which they were intended. 11
To overcome the rule-based clinical information extraction system, ML and DL have been proven to be efficient in the clinical practice setting for clinical information extraction and classification. However, an effective supervised ML or DL model needs human involvement to annotate a huge set of training data. Furthermore, annotating data manually needs a domain expert that requires significant time to do so, which is tedious and expensive. The annotation problem is the primary focus in the medical domain, and expert knowledge is needed for accurate annotation. The other popular methods, such as crowd sourcing, are not suitable for creating labeled clinical training data because of the sensitive nature of the domain and expert requirements. Also, findings of a systematic review 12 show that most datasets used in training ML models for text classification consist of mere hundreds or thousands of records because of annotation blockades.
The manual annotation process issues have been resolved by modern orthogonal approaches such as active learning (AL) and transfer learning (TL), which are utilized as machine-assisted pre-annotations. 13 AL provides a subset of high-value training samples by reducing the huge amount of data required for labor-intensive data annotation without losing quality. 14 The selection of samples is iterative, starting with a high-quality manually annotated subset of samples to automatically generate another subset of annotations, thus increasing the subset to annotated text to use in the subsequent iterations of the process. 13
A hybrid learning (unsupervised and supervised) approach is followed to perform an active learning process. A hybrid learning approach potentially offers the advantages of both supervised and unsupervised learning while minimizing their respective weaknesses. An unsupervised approach comprises a rule-based system with a domain-specific lexicon or knowledge base such as UMLS. 15 Such a system aids in the preparation of clinical concept extraction and annotation with a certain level of accuracy at the initial stage. 16 In other words, rule-based systems are used for feature extraction, where the outputs become features used as input for the machine learning system. While developing machine learning methods, required label data that can be generally acquired from a rule-based approach. The applications of hybrid systems include automatic de-identification of psychiatric notes 17 and detection of clinical note sections. 18
In addition, employing a hybrid learning approach for automatic data annotation can be performed on unannotated or unstructured data. Considering this, one of the popular approaches for active learning is embedding similarity in a high-dimensional space, while keeping a domain expert in loop. A word or sentence embedding similarity can be measured within some threshold value among annotated data and unannotated data. A DL and transformer-based architecture models such as ELMO 19 and BERT 20 can be leverage to generate contextual word embedding for annotated and unannotated data in the same domain. Subsequently, similarity between these embeddings is then measured, and a domain expert is involved to validate the classification of information based on the embedding similarity. Notably, identifying prominent embedding similarity threshold value is main ingredient that leads to proper data classification and annotation. Though, various similarity indexes can be used, such as cosine similarity, Euclidean Distance, etc. This approach facilitates the automatic classification of similar concept into explicit categories such as Problem, Treatment, and Test. The similarity process and indexes used are discussed in detail in the section “Proposed methodology”. We believe that implementing this approach in a clinical practice setting can enable domain experts and machine learning specialists to automatically generate annotated data without requiring human intervention, thereby saving time and effort. Also, it allows clinical domain and machine learning experts to perform NLP operations such as finding similarity between two concepts that indicate the same disease from two different clinical results documents, searching similar concepts in a clinical document, recommending similar treatments for similar diseases, etc.
Our research involves analyzing previous approaches and methodologies for automating information extraction and classification across diverse domains. We introduced a new method tailored for clinical practice settings. This method automatically extracts, classifies, and annotates clinical concepts from unstructured clinical documents. Similarly, we eliminate the need for domain expertise and reduce the time required, thus accelerating the training of downstream AI-based models.
The proposed methodology is composed of three modules: (a) label data preparation module (M1); (b) active transfer learning (ATL) module (M2); and (c) DL and LLM models for automatic concept classification and annotations (M3). Similarly, an I2b2 2010 challenge dataset is used to evaluate the proposed methodology, consisting of discharge summaries from Beth Israel Deaconess Medical Center, i2b2 Test data, and Partners Healthcare (which consists of discharge summaries).
16
Label data preparation module (M-1): The module (M1) served for preparing label data required in the active learning process at the initial stage. Initially, we applied preprocessing to clean the data. The syntax and morphological-based approach are introduced to identify the biomedical term boundaries. Further, word-to-lexicon matching is applied to annotate the clinical terminologies with semantic information leveraging UMLS Metathesaurus. Finally, handcrafted rules are applied to classify the semantic annotated clinical terms into Problem, Treatment, and Test categories with an acceptable accuracy as shown in Figure 1 (module 1). ATL module (M-2): In this module (M-2), we acquired only true label instances or clinical concepts from (M-1) of explicit concept categories. Leveraging variant BERT base models like BERTBase, DistilledBERT, SCIBERT, and ClinicalBERT, we construct word embedding models for known or reference concept categories such as Problem Model, Treatment Model, and Test Model, as illustrated in Figure 1 (M-2). Furthermore, BERT base variant models are utilized to generate candidate embeddings for tests or unlabeled concepts. A cosine similarity index is employed to assess embedding similarities between candidate and reference concepts. Subsequently, a candidate concept is assigned to a specific concept category if it exhibits a high similarity score with the known concept category models. This categorization process iterates until all candidate concepts have been categorized. Thereafter, we utilized a domain expert to manually assess the predicted concepts based on embedding similarity. Ultimately, we acquired sufficient labeled concepts through the ATL process (M-2). DL and LLM models (M-3): Concurrently, we trained DL models over contextual word embeddings generated by various BERT-based variants LLMs. The aim of incorporating DL models is to streamline and enhance the clinical concept annotation process, as depicted in Figure 1 (M-3). The DL models utilized in our study include recurrent neural networks (RNNs), long short-term memory (LSTM) networks, bidirectional LSTM (BiLSTM) networks, gated recurrent units (GRUs), and convolutional neural networks (CNNs). Additionally, we compared the performance of these DL models with that of large language model (LLM) in clinical concept classification. Moreover, we conducted parameter tuning for both DL and LLM models during training to improve accuracy and minimize training loss.

The proposed system workflow towards clinical concept classification consists of three modules: (a) label data preparation; (b) active transfer learning; (c) deep learning and large language model (LLM).
The rest of the article is structured as follows: In the second section, related work is presented related to clinical concept extraction and classification and various approaches are discussed in this section. Thereafter, we discussed the proposed methodology in the third section, depicting scenarios and workflows towards clinical concept extraction and classification. Afterward, evaluation matrices and datasets are illustrated in the fourth section. In the fifth section, experimental setup design, results and analysis of proposed methodology for clinical concept extraction and classification are illustrated. Furthermore, in the sixth section, we provide a detailed discussion that offers a complete overview of the proposed work and its impact on the clinical domain. Finally, in the seventh section, we conclude the study by addressing a few limitations and suggesting future work to guide readers and researchers in this field.
Literature review
In the previous section, we discussed four types of approaches that are utilized for clinical concept extraction. In this section, we will further discuss the effort that is made toward the clinical concept extraction while employing the following approaches: such as Rule Based, ML, DL, LLM, TL, and hybrid approaches.
Rule-based approaches
Rule-based concept extraction approaches use an extensive collection of rules and keyword-based characteristics to detect predetermined patterns in text. 21 The rule-based method has been widely used in many clinical applications because of its simplicity and tractability, which refers to its efficacy in incorporating domain-specific information. An early endeavor in clinical concept extraction, known as the Medical Language Processing Project, was derived from the Linguistic String Project. Its objective was to extract symptoms, medications, and potential side effects from medical records. This was accomplished by utilizing a semantic lexicon and a vast set of rules. 22 An inherent advantage of using rule-based methodologies is the ability to obtain dependable outcomes promptly and at a minimal expense, since it obviates the need for laboriously annotating a substantial number of training instances. 9 The effective use of rules and well-curated dictionaries might lead to a very favorable performance, depending on the individual tasks at hand. Several tasks have used rule-based matching techniques with various degrees of effectiveness.23,24 For instance, in the 2014 i2b2/UTHealth de-identification challenge, the top four teams, including the winning team, used rule-based techniques. 25 Similarly, the 2009 i2b2 medicine challenge identified 10 rule-based systems among the top 20 systems. 26 In the i2b2/UTHealth Cardiac risk factors challenge, Cormack et al. 27 showed that a system based on pattern matching may reach competitive performance by using various lexical resources. Furthermore, using pre-existing resources such as clinical criteria, guidelines, and clinical corpora may significantly decrease development efforts. A widely used approach is to use well-curated clinical dictionaries and knowledge base. The dictionary functions as a specialized knowledge repository for a specific field or task, allowing for easy modification, updating, and aggregation. 28 Established medical terminologies and ontologies, such as UMLS Metathesaurus, 15 Medical Subject Headings (MeSH), and MEDLINE, 5 have been used in clinical information extraction activities due to their comprehensive collection of well-defined concepts associated with numerous words. Although there are advantages, the lexicon or dictionary-based approach also has limitations. These include the difficulty of creating general rules that apply to the entire problem or system, the inability to capture complex semantic relationships between words, and the challenge of handling name entities, particularly in dynamic specialized domains.
ML-based approaches
To overcome the challenges with rule-based approaches, a cutting-edge machine learning method is used for the purpose of clinical concept categorization. ML is able to acquire patterns without the need for explicit programming by learning the correlation between input data and labeled outputs. 29 Commonly used conventional machine learning methods for clinical concept extraction include conditional random fields (CRFs), support vector machines (SVMs), structural support vector machines (SSVMs), logistic regression (LR), Bayesian model, and random forests. Whereas CRFs and SVMs are the predominant models for clinical concept extraction. 30 Because they offer a combination of sequential modeling capabilities, robustness, interpretability, and availability that make them well-suited for clinical concept extraction tasks. Similarly, CRFs may be seen as an extension of LR specifically designed for analyzing sequential data. On the other hand, SVMs use diverse kernels to convert data into a hyperspace that is more readily distinguishable. Further, SSVMs are a technique that combines the benefits of both CRFs and SVMs. 30 In their study, Tang et al. conducted a comparison between SSVMs and CRF utilizing the datasets from the 2010 i2b2 NLP challenge. They found that the SSVMs outperformed the CRFs when employing the same features, as seen by their improved performance. In addition, Wang and Akella 31 used NLP features, including semantic, syntactic, and sequential aspects, as input for a supervised classical machine learning model in order to extract mentions of disorders from clinical notes. Nevertheless, ML models need labeled data. Manually preparing annotated data requires domain expert and is a laborious operation that consumes a significant amount of time and resources. In the same way, ML models may not always achieve optimal performance on labeled data, since they might suffer from overfitting or underfitting. This often necessitates the use of complicated feature engineering techniques.
DL-based approaches
Similarly, DL is efficiently used in the clinical field for a range of activities. In fact, it is a specific area within ML that emphasizes the automated acquisition of features across many layers of abstract representations. 32 The algorithms mostly revolve on neural networks, including RNNs, CNNs, and transformers. Notably, DL differs from classic machine learning approaches by reducing the need for manually designing explicit data representations like bag-of-words or n-grams. A significant number of deep learning applications in concept extraction have used either modified versions of RNNs or CNNs. CNNs use convolutional filters to capture spatial correlations in the input data and pooling layers to reduce computational complexity. As a result, CNNs have shown to be very effective for computer vision tasks, they may encounter challenges when it comes to identifying long-range dependencies that are often present in text. 33 In contrast, RNNs are a kind of neural networks that specifically represent connections in a sequence. This makes RNNs particularly well-suited for tasks that involve capturing long-term dependencies.34,35 Since, conventional RNNs are constrained in their ability to represent text owing to the vanishing gradients issue, which limits their capacity to capture long-range dependencies between words. Because of that, models such as LSTM and GRU have been devised to tackle this problem by segregating the gradient propagation and controlling it via “gates.” Although shown to be efficacious, these methods simply mitigate the problem rather than fully resolve it, since they are still constrained to sequence lengths ranging from tens to hundreds of words. 34
LLM-based approaches
Moreover, the process of training these models requires significant computational resources and is challenging to parallelize because of the sequential nature of weight training. Recently, the transformer architecture has been offered as a solution for several of these issues. The transformer design eliminates the need of processing text sequentially by simultaneously processing the full sequence using matrix multiplications. This enables the network to remember the significant elements in the sequence. 36 Actually, long sequences need significant memory resources for training. To accommodate extended sequences of text without overwhelming memory limits, it is necessary to break up the sequence into smaller chunks and add additional layers to the model. In that case, Transformers are capable of accurately representing associations between words that are far apart, and they are much more efficient in terms of processing resources when compared to variations of RNNs. Architectures such as ULMFit, ELMO, BERT, and GPT have shown substantial improvements in the performance of state-of-the-art natural language processing workloads. As a result, researchers have constructed huge models based on Transformers, such as ClinicalBERT, SCIBERT, and BIO-BERT. These models have shown promising performance in the medical and clinical field. Similarly, these models have been employed for various tasks in the clinical domain, including the de-identification of personal health information, 7 the identification and de-identification of medical risk factors associated with coronary artery disease from diabetic patient records, and the extraction of medical problems.7,37
TL-based approaches
TL aims to transfer the existing knowledge from a large, well-trained model and apply it to a new model at its beginning stage. Subsequently, the novel prototype gradually acclimates to the given job. Similarly, TL offers optimized initialization that enhances performance in downstream tasks, particularly when the dataset for the downstream job is limited in size. Moreover, the researcher has come up with a method called ATL to enhance the effectiveness of TL. This method involves a domain expert who verifies the results acquired by a LLM. Such ATL techniques have been used in a clinical setting to reduce the laborious task of data annotation and improve the effectiveness of model classification with a small number of labeled sample sets.14,38 For instance, Li et al. 39 adopted ATL to decrease the need for annotations in the de-identification procedure. They achieved this by integrating actual clinical trials with i2b2 datasets, demonstrating that trained models performed better than the typical passive learning framework. Similarly, Tomanek and Hahn 40 also investigated the effect of ATL on reducing the time needed for annotating data for the extraction of entities such as person, organization, and place. It was shown that the ATL method reduces data annotation time and cost by up to 33% compared to the baseline. Further, Chen 41 performed a simulation study to reannotate a portion of the i2b2/VA 2010 dataset from the concept extraction challenge. Their findings demonstrated that the query technique based on ATL significantly decreased the amount of data required for human annotation in comparison to the baseline.
Likewise, the word embedding similarity approach in the ATL environment is proposed by the author for causality mining in the clinical text. 42 In this study, the author applied a BERT model to generate embedding vectors utilizing training data obtained from SemEval Task 8. 43 Then, a word embedding similarity operation was conducted using a similarity threshold value to compare the training and test data embedding vectors, leading to automated classification. Though TL enables leveraging knowledge from one domain to another, applying it in healthcare may result in suboptimal performance without careful adaptation due to domain specificity.
Hybrid approaches
Conversely, hybrid approaches provide exceptional support towards solving complex tasks in the clinical domain. By integrating both rule-based and machine learning methodologies into a single system, possibly providing the benefits of each while reducing their individual limitations. There are two primary hybrid techniques, referred to as terminal hybrid approaches and supplementary hybrid approaches. 5 In a terminal hybrid technique, rule-based systems are used to extract features, which are then utilized as input for the machine learning system. The machine learning system then serves as the final stage in order to choose the most optimum features. For example, Wang and Akella 44 used NLP attributes, including semantic, syntactic, and sequential aspects, as input to a supervised classical machine learning model. Their objective was to extract mentions of disorders from clinical notes. Moreover, hybrid systems were applied for automatically removing personal information from psychiatric notes 17 and identification of sections in clinical notes. 18 In contrast, supplemental hybrid approaches use ML techniques to address shortcomings in the extraction of entities that exhibit subpar performance when extracted only using rule-based methods. In this study, 45 research infused a supplemental hybrid system with a user interface to facilitate interactive concept extraction. Owing to that, Meystre et al. 46 utilized a conventional machine learning classifier to extract medications for congestive heart failure as an additional component to the rule-based system. This system extracted references and values of left ventricular ejection fraction, along with other concepts, to evaluate treatment performance measures. However, the coordination between rule-based and ML/DL models remains unsophisticated, resulting in a lack of smooth integration or performance boost.
Through the adoption of various methodologies, we have formulated a pipeline for the automated classification of clinical concepts. Our approach builds on the limitations of existing models by addressing key challenges identified in prior work. We mitigate the rigidity of rule-based approaches by incorporating the UMLS Metathesaurus for more structured and semantically rich data preparation. The reliance on large annotated datasets for DL models is reduced through ATL, which leverages state-of-the-art LLMs while keeping domain experts involved in the loop, ensuring clinical accuracy and reducing error propagation. This approach overcomes the challenges of domain generalization seen in traditional ML and TL models. Additionally, we enhance the integration of rule-based and ML techniques through a novel hybrid mechanism that uses rule-based preprocessing and DL/LLM-based classification, improving the overall adaptability and precision of classifying clinical concepts into Problem, Treatment, and Test categories. This amalgamation enhances the adaptability and effectiveness of the classification process, showcasing the coordination between rule-based and ML techniques in the realm of clinical concept classification.
Proposed methodology
The proposed methodology comprises of three main modules, such as initial level label data preparation module (M1), ATL module (M2), and DL and LLM (M3). Figure 1 provides an overview of the clinical concept classification system, illustrating the interdependence of each module, with each successive module relying on the output of the preceding one. We have presented the role of each module within the proposed system.
Label data preparation process (M1)
Recently, emerging Medical Language Processing techniques and DL models are playing a cornerstone role in the AI-based clinical decision support system (CDSS) system that needs label data. Preparing label data requires domain experts, and it is tedious to prepare label data manually. To save time and energy, various semantic, syntactic, and lexical-based approaches are utilized to automatically prepare label data. In the proposed methodology, we utilized a semantic-based automatic labeling approach that involves data preprocessing, word boundary detection, semantic breakdown using UMLS, and rules that categorize concepts into Problems, Treatment, and Test based on UMLS semantics. Each subprocess is explained in this section below.
Data preprocessing process
In the clinical domain, often data are available in heterogeneous and unstructured formats, resulting in distortion and ambiguity. Data preprocessing is one of the preliminary steps in the development of any AI-based CDSS system. Applied preprocessing operations such as tokenization, stop word removal, lemmatization, N-gram, and part of speech (POS) tagging are introduced below.
Let Tokenization: Each document Lemmatization: Employed the Lemmatization, base form of word obtained to enhance the meaning of ambiguous words. Most of the cases lemmatization is preferred instead of steaming to obtain precise and accurate information such as lemmatizing word “caring,” it returns “care,” while applying stemming it returns “car” which is erroneous. N-gramming: N-gram of word is applied to obtain a set of co-occurring words in a sentence as shown in equation (1). Duplication: Duplicate words are removed to reduce data dimensionality and avoid ambiguity. Such as: “ Punctuation removal: Punctuation marks such as commas, periods, question marks, and exclamation points serve grammatical purposes but may not carry substantial semantic meaning for some NLP tasks. Removing them simplifies the text, making it easier to process and analyze. Further, removing punctuation ensures that similar texts are recognized as such, even if they differ in punctuation usage during text similarity approach. POS tagging: The POS tagging using NLTK NLP library was employed and then constructed a regular expression pattern to filter only meaningful information explicitly like noun, adjective, and adverb from a list of words as shown in Reg.3.1. In Reg.3.1,


Word boundary detection
The method of detecting single or multiple neighboring terms that signify a clinical term is known as word boundary detection in the NLP domain. Multiple neighboring terms describing a clinical term may be a combination of stop words, punctuations, and digits, rendering it difficult to retrieve details. Using rules and regular expressions, we establish a protocol that smoothly defines the boundary between single or multiple adjacent terms of a clinical term. Following word boundary detection, each word is then mapped to the UMLS Metathesaurus to determine whether it assimilate to a clinical concept or not. The detailed idea diagram and workflow for word boundary detection has been discussed in this study. 16
Clinical concept identification
The most common strategy is to leverage a well-curated clinical dictionary for a clinical concept extraction. The dictionary acts as a domain or task specific knowledge base. In the proposed methodology a biomedical dictionary such as UMLS has been utilized as knowledgebase for clinical concept identification and extraction with semantic information. Clinical concept extraction is a multi-step process that includes finding terms, concept identification, and semantic type extraction. Generally, various approaches like exact or approximate term matching approach to UMLS is used to identify terms. In this study, we have utilized a combined exact plus approximate term matching approach for clinical concept extraction. The steps for mapping biomedical terminologies using UMLS are outlined below:
Finding terms: Subsequently identifying word boundary, each word is mapped to the UMLS Metathesaurus. If a word matches to UMLS, it is stored in a list data structure; otherwise, the word is discarded. Concept identification: A concept in the UMLS Metathesaurus illustrates the meaning of medical terms by different names. Metathesaurus significance lies in illustrating the predefined interpretation of each name and associating all names from all source vocabularies that have identical meanings, known as synonyms. In UMLS Metathesaurus, each concept had its own permanent and special concept descriptor, which was expressed as “name,” for example, “Coronary Arteriosclerosis.” When a new concept is added to the Metathesaurus structure, it is given a specific identifier or “ui” value, such as “C0010054.” Each term in the Metathesaurus has a single or a list of concepts available. Semantic Type Extraction: Semantic type is important in concept categorization, such as Medical Problems, Medical Treatment, and Medical Test, since it gives Metathesaurus terms an interpreted and obvious sense. For instance, the general term “Trout” has the semantic type “fish” but not “animal”? since “fish” is more closely associated with the concept of “trout” than “animal.” In the Metathesaurus, every concept is assigned at least one semantic type (STY) and can have up to five semantic types.
16
Concepts with multifaceted or inherently vague meanings may encompass more than one semantic type. For example, the concept “Febrile Convulsion” is associated with both “Finding” and “Disease or Syndrome.”
47
Clinical concept classification lexicons
Clinical concept extraction and classification is significant to transform the huge amount of unstructured clinical data into a set of actionable knowledge to improve quality of care and clinical decision-making support. After identifying the concept along with semantic information in the previous section, we constructed rules to map the semantic information of clinical phrases to the semantic dictionaries as shown in Table 1. These dictionaries facilitate precise classification of clinical concepts into explicit categories. The mapping dictionaries have been enriched to include semantic types of three distinct types of categories of concept such as problem, treatment, and test. For a deeper understanding of the rules, algorithm, and implementation process, refer to our study. 16
Clinical concept semantic types dictionaries for concept classification.

Clinical concept classification example-case-study
Figure 2 depicts a detailed scenario for clinical terms mapping to UMLS, semantic information extraction from UMLS and concept classification into a specific category utilizing semantic-enriched dictionaries. In the study, three clinical terms are chosen as examples for relevant categories explicitly such as “beta blockers,” “increased heart rate” and “heart rate.” Each clinical term is mapped to the UMLS Metathesaurus.

Clinical concept extraction and classification: an example case study scenario workflow.
In response, specific clinical terms such as “Tachycardia,” “Adrenergic beta antagonists,” and “Heart rate,” identified by concept IDs like “C0039231,” “C001645,” and “C0018810,” are obtained from the UMLS. Extracting semantic information involves inputting these concept IDs into the UMLS Metathesaurus, which yields semantic type IDs such as “T033,” “T121,” and “T201,” as well as semantic types like “Finding,” “Pharmacologic Substance,” and “Clinical Attribute,” all of which are explicitly retrieved. Semantically enriched dictionaries are tailored for specific concept categories such as Problem, Treatment, and Test as outlined in Table 1. The semantic types extracted from UMLS for exclusive clinical terms are mapped to these specialized dictionaries. When there is concordance between the extracted semantic types and those in the dictionaries, the clinical terms are effectively classified into their respective categories, as illustrated in Figure 2.
ATL module (M2)
In the clinical domain, vast amounts of unstructured data are generated daily in the form of clinical reports, EHR, and EMR (Electronic Medical Record), which contain meaningful information. Recently, DL and ML techniques have been extensively used to extract this useful information and convert it into actionable knowledge. Certainly, ML and DL algorithms require huge amounts of labeled data. Another way, ATL techniques surprisingly play a vital role in many modern ML and DL problems, particularly when data labeling is complex, time-consuming, and expensive to collect. Likewise, we have introduced an ATL approach to solve and automate the data labeling process through LLMs while keeping the domain expert in the loop, as shown in Figure 1 (M2). As a result, it significantly improves the automated data preparation process, thereby facilitating the application of DL and LLM models for the clinical concept classification task. The following section illustrates individual steps and components that assess the ATL module, as depicted in Figure 1 (M2).
True instances collection
In an AL environment, the learning algorithm is given the ability to choose the subset of available examples to be labeled next from a pool of yet unlabeled instances. A set of true instances is selected while performing a semantic-based concept classification process, as discussed in the section “Label data preparation process” (M1). The core concept of this principle is that when a ML or DL algorithm is empowered to autonomously select the data it learns from, it can achieve enhanced performance while requiring fewer training labels. In this study, domain experts manually curated high-quality true instances, and the process is detailed in the Experimental setup and results and Discussion sections. The proposed methodology leverages pretrained transformer-based language models, such as BERTBase, SCIBERT, ClinicalBERT, and DistilBERT, to generate concept embeddings from a dataset of true instances. These embeddings then serve as features for downstream classification models.
Clinical concept embedding generation
Much of NLP relies on similarity in high-dimensional spaces. Typically, an NLP solution takes a text, processes it to create a large vector or array representing the text, and then performs various transformations. Similarly, we have explored several BERT-based architecture models. These models are explicitly used to generate contextual word embeddings for clinical concepts (Problem, Treatment, and Test), as shown in Figure 1 (M2). Initially, a true labeled clinical concept, previously obtained as discussed in module (M1), is fed into these BERT-based architecture models to generate embeddings. The simplest approach that we followed was to execute these BERT-based models using the sentence-transformers library by Hugging Face.
Initially, the dataset lacked balance; consequently, we curated a balanced dataset comprising a total of 6000 clinical concepts, evenly distributed among 2000 instances each of Problem, Treatment, and Test clinical concepts. Following this, we proceeded to generate word embedding vectors using BERTBase, ClinicalBERT, SCIBERT, and DistilBERT for each concept category, namely Problem Embeddings, Treatment Embeddings, and Test Embeddings. The embedding vectors for explicit categories are saved using Pickle, a Python package commonly used for serializing and deserializing Python object structures. For example, the Problem Embeddings vector contains embeddings specific to clinical concepts related to problems, while the Treatment and Test Embeddings vectors contain embeddings relevant to treatments and tests, respectively. Furthermore, generating embedding vectors for a dataset of 6000 concepts is a computationally demanding task, typically taking several hours when executed on a CPU.
Fortunately, by leveraging the efficiency of the sentence-transformer library in Python, we enabled GPU acceleration, which substantially decreased computational time. In our experiments, we efficiently processed clinical concepts, generating embedding vectors for all variants of the BERT base models in under 10 minutes each. Consequently, the overall time for producing the embedding vectors was significantly reduced to less than an hour, thanks to the utilization of an NVIDIA GeForce RTX 2060 GPU. However, as the dataset size increases, the time and computational power required for embedding vector generation also increases, especially when employing the ATL approach.
Cosine function for embedding similarity
Cosine similarity is a metric that determines how similar two vectors (words, sentences, features) are to each other. Essentially, it is the angle between two vectors. In the same way, we have leveraged the cosine similarity algorithm to find the embedding similarity between unseen or unlabeled clinical concepts (candidate concepts) and labeled clinical concepts (known concepts). Ultimately, the unseen clinical concept is categorized into an explicit concept category based on the embedding similarity matching, as shown in Figures 1 and 3.
Presented clinical concepts embedding similarity between candidate (unlabeled) concepts and known (label) concepts and active learning process for automatically boosting labeled concepts.
According to equation (1), A and B are two vectors that compute the angular distance between them. Initially, word embedding is generated for unlabeled clinical concepts. Subsequently, we computed the word embedding similarity score between the embeddings of unlabeled clinical concepts and pretrained or labeled embeddings (Problem, Treatment, and Test Embeddings). This process yielded three cosine similarity scores against the pretrained embedding models. Finally, the unlabeled concept was categorized into explicit clinical concept categories based on a similarity threshold score of 0.83. Through experimentation, we determined the optimal similarity threshold value, which is extensively discussed in the Experimental setup and results section.
Afterward, we utilized a domain expert to manually analyze the classified concepts. The domain expert manually cross-checks the newly classified concepts with the gold standard dataset. If the label assigned to the newly classified concept matches that in the gold standard document, it is added to the explicit category data list as labeled data. Otherwise, the domain expert manually categorizes the concept. This process iterates for each batch of unseen clinical concepts, with validation by domain experts. This iterative process, which we refer to as ATL, dynamically enhances the dataset, as illustrated in Figure 1 (M-2). TL involves utilizing various BERT-based models that are pretrained on generic data, with the exception of ClinicalBERT, which is specifically trained on clinical data. Then, we utilized these pretrained models for the clinical concept classification task. Similarly, we interpret the term “Active” to mean that we engage domain experts in real-time to validate the concept classifications made by the LLM model and incorporate them into the explicit concept category list.
Threshold value identification
In Module-2 (M-2), we delve into clinical concept classification using the cosine similarity approach based on a threshold value. Selecting an appropriate similarity threshold is pivotal for accurate concept classification via embedding similarity, thus requiring comprehensive experimentation to pinpoint the optimal similarity score. To achieve this goal, we utilized unlabeled or unseen concepts and conducted embedding similarity operations to derive similarity scores and identify the best threshold value. The precision–recall curve (PRC) plays a key role in this process, aiming to maximize the area under the curve. Particularly in biased datasets, where the positive class is significantly outnumbered by the negative class, the area under the PRC (AUPRC) serves as an optimal metric for threshold selection.
Our test dataset comprises a total of 1200 clinical concept samples, evenly distributed with 400 samples for each category (Problem, Treatment, and Test). The AUPRC was employed to determine optimal threshold values for various BERT base models, and further details regarding the threshold selection results are provided in the Experimental setup and results and Discussion sections.
Clinical concept embedding similarity: case study
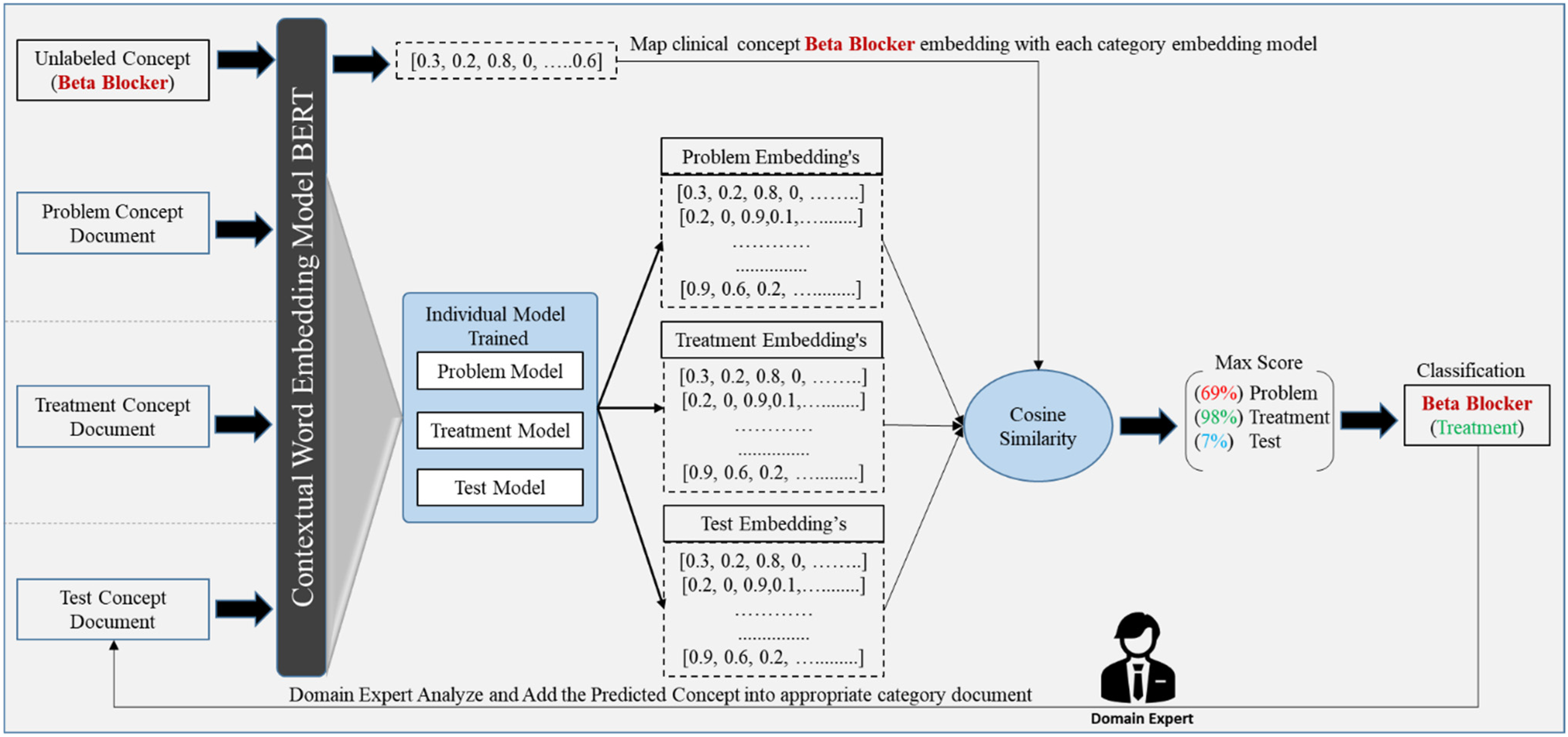
A case study has been crafted to provide a clearer and more comprehensive illustration of the clinical concept embedding similarity approach, as depicted in Figure 3. Initially, we curated a balanced dataset comprising true labeled concept documents categorized as problem, treatment, and test documents, as illustrated in Figure 3. Subsequently, each concept document underwent processing through a pretrained BERT-based models to produce contextual word embedding vectors specific to its category such as Problem, Treatment, and Test models, which encompassed explicit concept category embeddings.
Given a set of unlabeled clinical concepts like “Beta Blocker,” our objective was to categorize them into specific categories using the word embedding similarity approach. To achieve this, we utilized the BERT model to generate embeddings for the “Beta Blocker” concept. We then compared the embedding of “Beta Blocker” with the embeddings of annotated clinical concepts using cosine similarity. Consequently, the process yielded a similarity score indicating the degree of match between “Beta Blocker” and labeled concept models such as problem (69%), treatment (96%), and test (7%). By applying a
The classified concepts were further examined manually by domain experts. Following validation, these concepts were added to the explicit concept category document with labels such as “Beta Blocker” ↔ “treatment.” This process enhances the labeled data in real time. Additionally, during the embedding generation phase, we utilized various BERT-based architecture models discussed earlier. The parameters of these models were the number of transformer blocks (L): 12, hidden layer size (H): 768, and attention heads (A): 12, except for DistilledBERT, which contained the number of transformer blocks (L): 6.
DL models and LLMs (M3)
Utilizing DL and ML algorithms for clinical concept classification requires a substantial amount of labeled data. Currently, we have obtained a significant amount of labeled data through the ATL process (M2), which we have organized into explicit category documents: problem concepts, treatment concepts, and test concepts, as discussed in the section “Active Transfer Learning Module (M2)”. The size of this labeled data is explicitly stated in the section “Evaluation matrices and datasets”. These data from the three documents are then fed into various BERT-based models (ClinicalBERT, SCIBERT, and DistilBERT) to generate contextual word embeddings for the clinical concepts. These embeddings are subsequently used in downstream DL models to train them for clinical concept classification tasks.
Furthermore, there has been a rise in the utilization of deep learning classification models for text data classification problems. In our study, we employ five types of DL algorithms RNN, GRUs, LSTM, BiLSTM, and CNN for clinical concept classification, using BERT based various embeddings as input features. These models are trained to classify clinical concepts into explicit categories while maintaining high accuracy. Additionally, we leverage pretrained LLMs, such as BERTBase, DistilBERT, SCIBERT, and ClinicalBERT, to harness state-of-the-art language representations tailored to the clinical domain and assess their performance on the specific nuances of clinical text. The performance of each DL model is detailed in the section “Experimental setup and results”.
Evaluation matrices and datasets
In this section, we have discussed the empirical analysis of the proposed methodology by evaluating unstructured clinical documents provided by I2b2 National Center in 2010 NLP challenges. 48 Each process performance results are presented and discussed to highlight the need and value of the proposed methodology. Similarly, we have open-source code for clinical researchers and the health industry to run and integrate the proposed methodology into their systems or research, which is available on GitHub (https://github.com/TuriAsim/MCECS.git).
Performance measure matrices
To measure and compare system performance, generally, three indexes are used for information retrieval and extraction: precision, recall, and F1-score. Precision measures the number of valid instances in the set of all retrieved instances. Recall measures the number of valid instances in the intended class of instances. F1-score is the harmonic mean between precision and recall with β = 1 to obtain F-score. The measures can be computed through the following equation:



Datasets evaluation and preparation
We evaluated publicly available open datasets clinical datasets provided by: Partners Healthcare, and Beth Israel Deaconess Medical Center, provided by the i2b2 National Center. The clinical dataset consists of discharge summaries that have been manually annotated for three types of clinical concepts (Problem, Treatment, Test) according to the instructions granted by the i2b2/VA challenge organizers. 48 Consequently, the Beth Israel Deaconess Medical Center dataset contains 73 annotated notes, the Partners Healthcare dataset contains 97 annotated notes, and the test dataset provided by the i2b2 National Center for system evaluation contains 256 annotated notes. Overall, the i2b2 datasets comprise 426 gold standard notes, as depicted in Figure 4(A). Moreover, the datasets consist of 19,665 Problem, 14,187 Treatment, and 13,834 Test clinical concepts, cumulatively totaling 47,686 clinical concepts available in the datasets (see Figure 4(B)).

Comprehensive details of datasets and their utilization.
While evaluating the rule-based approach, we selected 20 clinical notes from each dataset. As a result, we acquired a total of 2035 clinical concepts from the rule-based approach. We then selected only true instances, totaling 500 for each concept category, to utilize them in the ATL process, as depicted in Figure 4(C). Further, while evaluating ATL approach, we acquired 1500 true instances from the rule-based approach and 3500 concepts of equal size from the gold standard data to generate training embeddings. Afterward, we chose 1200 test concepts, which were not previously seen or used, to find the threshold value and measure the LLMs’ performance towards unseen concept classification in the ATL process, as shown in Figure 4(D).
Finally, we trained the DL models and LLMs on data acquired from the rule-based approach, the active learning approach with a size of 1500, unseen data with only true instances of 885, and gold standard data with a size of 1615 (see Figure 4(E)). Overall, we utilized a dataset of 9000 clinical concepts to evaluate the DL models and LLMs in the proposed approach, with each category consisting of 3000 concepts, as shown in Figure 5(F). In the sections below, we have explicitly presented the results and performance of each approach.

Proposed approaches evaluation and experimental setup.
Experimental setup and results
The proposed methodology outlined in the section “Proposed methodology” provides a theoretical foundation for clinical concept identification and classification from unstructured clinical documents. To construct a robust implementation of this framework, it is crucial to determine the specific models and algorithms that can optimize each component individually, thereby producing high-performance intermediate results. These results can then be combined to achieve an overall optimal outcome for clinical concept classification. We conducted numerous experiments to assess the effects of a rule-based approach on initial label preparation, embedded vector generation, and similarity threshold calculation within an ATL approach. Additionally, we trained various DL models and LLMs to identify a well-balanced ecosystem that meets both our local and global optimization goals. Figure 5 depicts a detailed experimental workflow.
Initial-level labeling performance (M1)
During the label data preparation process, we conducted several experiments to assess performance based on clinical concept categorization. These experiments encompassed evaluations of individual datasets and individual concepts, as well as comparisons with alternative approaches, as elaborated below.
Individual datasets and concept-wise performance
In this section, we provide a comparative analysis of the results obtained from each dataset, examining them on both a dataset-specific and clinical concept level. We utilize the Beth, Partner, and I2b2 Test datasets for this purpose. As a result, a high precision of 83%, recall of 75%, and F1-score of 79% were measured for the Partner dataset, which was found to be better than those for the Beth and I2b2 Test datasets. The overall lowest score was calculated for the I2b2 Test dataset, with a precision of 69%, recall of 67%, and F1-score of 68%, as shown in Table 2. The arrow symbol (↑) indicates the model performance improvement in percentage on the current dataset over the previous one. For instance, the methodology applied to the Partner dataset shows an (↑8%) increase in performance compared to the Beth dataset in terms of precision, and vice versa.
Individual datasets-wise performance for clinical concepts classification.

We also computed individual concept-wise results for the proposed methodology in terms of concept classification. While performing the analysis for the individual concepts, we noticed an approximately equal and high precision of 80% for the Problem and Test categories, whereas the Treatment category gained a higher recall of 87%. The overall best performance, with an F1-score of 81%, was achieved by the Problem concept category, as shown in Table 3.
Individual concept-wise performance for concept classification.

Proposed approach vs. competitors (rule-based)
We compared our proposed approach with that of three related systems: QuickUMLS, 49 BIO-CRF, 50 and the Rules (i2b2) model. 51 The three systems were tested against the i2b2 2010 dataset for three types of concept category extraction: Problem, Treatment, and Test.
QuickUMLS employed an approximate dictionary matching approach for medical concept extraction, requiring a threshold value between 0.6 and 1.0 to select an acceptable medical concept from a collection of UMLS concepts. In our suggested methodology, we used both approximate dictionary matching and exact word matching, which resulted in 25% greater accuracy and almost 13% higher F1-score compared to QuickUMLS. However, QuickUMLS demonstrated almost 5% greater recall compared to the proposed methodology, as depicted in Table 4.
Comparative analysis among proposed approach and the competitors (QuickUMLS, BIO-CRF, Rules (i2b2)).

The Rules (i2b2) model created a simple set of rules by harvesting information from the annotated training data. This rule-based algorithm used a statistical method to categorize and extract concepts from structured and annotated data. On the other hand, our suggested rules-based methodology employed a majority vote mechanism to identify and extract concepts from unstructured clinical data. When compared to the Rules (i2b2) model, the proposed methodology yielded higher precision, recall, and F1-score, as shown in Table 4.
BIO-CRF is a medical concept extraction approach based on ML that aims to automatically identify the concept boundary and assign the concept type to them. For each medical concept, word-level and orthographic-level features were retrieved to train the BIO-CRF model. At the individual concept and dataset level, we compared the performance of the proposed approach with BIO-CRF. The proposed methodology achieved 75.76% precision and a 72.94% F1-score, which are approximately 6% and 2% higher than the BIO-CRF system, respectively. Nevertheless, BIO-CRF achieved approximately 3% higher recall than the proposed system. Overall, the proposed system performed better than the QuickUMLS, BIO-CRF, and Rules (i2b2) models, as shown in Table 4.
ATL module performance (M2)
In this section, we present the experimental results for the ATL module. The results from this module include optimal threshold value identification, individual LLMs’ performance towards concept embedding similarity, and ensemble LLMs’ performance for concept classification employing the embedding similarity approach.
Threshold value identification assessments
Our problem is a multiclass classification, so we transform the categorical class labels into a binary matrix representation as per the experiment's requirements. Ultimately, we concatenate all three concept category labels along with their similarity scores. Similarly, we also perform experiments for explicit concept models, which are presented in the article's appendix. As illustrated in Figure 6, we present the threshold scores and area under the curve (AUC) values, acquired from the evaluation of four different models: BERT, ClinicalBERT, DistilBERT, and SCIBERT, applied towards a clinical concept classification task. The threshold score indicates the decision boundary for classifying instances, while the AUC score quantifies the overall discriminative ability of an explicit model.

Threshold value identification for embedding similarity using cosine similarity approach.
Analyzing the AUC scores, we observe that all models perform reasonably well, with scores ranging from 0.73 to 0.76, as shown in Figure 6. The AUC is a crucial metric in binary classification tasks, representing the model's ability to distinguish between positive and negative instances in concept classification. In this context, an AUC score above 0.5 indicates that the models are performing better than random chance. Interestingly, ClinicalBERT, DistilBERT, and BERTBase exhibit similar AUC scores of 0.73, suggesting comparable discriminative performance on the task. In contrast, SCIBERT stands out with a slightly higher AUC score of 0.76, indicating improved discriminative ability compared to the other models.
Moving on to the threshold scores, we note that DistilBERT and ClinicalBERT have relatively higher threshold values (0.96, respectively), indicating a more conservative classification approach as shown in Figure 6. BERT and SCIBERT, on the other hand, have slightly lower threshold values (0.94 and 0.92, respectively), suggesting a relatively more lenient approach in assigning positive class labels as shown in Figure 6. The choice of threshold can be crucial in real-world applications, impacting the balance between sensitivity and specificity. A higher threshold tends to prioritize precision, reducing the likelihood of false positives but potentially increasing false negatives. Conversely, a lower threshold may result in higher recall but at the expense of precision.
We calculated the average threshold score and AUC score across the four models and found that the average threshold is approximately 0.95, while the average AUC score is approximately 0.7375. This suggests a moderate threshold level for classification and an overall moderate discriminative performance across the ensemble of models. Finally, we combined the threshold score of 0.95 and the AUC score of 0.7375 and took the average. As a result, we obtained a prominent threshold value of 0.83, consequently assisting in lower recall and precision expense towards clinical concept classification.
System performance for ATL process
Subsequently, identify the optimal threshold value which is 0.83 as shown above section. Further, all the four BERT variant-based concept embedding models trained during the AT process are evaluated individually on unseen clinical concepts. In the ATL environment, the individual LLMs exhibit varying performances towards concept classification tasks across three categories: Problem, Treatment, and Test as presented in Table 5. Each model underwent evaluation based on True Positives (TP), False Negatives (FN), False Positives (FP), True Negatives (TN), precision (P%), recall (R%), F1-score (F1%), and accuracy (A%).
Individual LLM performance towards concept classification in active transfer learning environment.

TP: true positive; FN: false negative; FP: false positive; TN: true negative; A: accuracy; P: precision; R: recall; F: F1-score.
The BERTBase model demonstrated 91 True Positives, 157 False Negatives, 74 False Positives, and 167 True Negatives, resulting in precision, recall, F1-score, and accuracy of 54.14%, 53.74%, 51.60%, and 53.33%, respectively. DistilBERT, on the other hand, exhibited 115 True Positives, 232 False Negatives, 81 False Positives, and 188 True Negatives, yielding precision, recall, F1-score, and accuracy values of 52.0%, 51.52%, 48.50%, and 49.19%, respectively. ClinicalBERT displayed 65 True Positives, 121 False Negatives, 70 False Positives, and 174 True Negatives, resulting in precision, recall, F1-score, and accuracy of 53.57%, 53.13%, 53.0%, and 55.58%, respectively. Lastly, SCIBERT recorded 143 True Positives, 260 False Negatives, 86 False Positives, and 196 True Negatives, with precision, recall, F1-score, and accuracy values of 52.72%, 52.49%, 49.20%, and 49.50%, respectively. Comparatively, ClinicalBERT demonstrated the highest precision and accuracy among the models, while BERTBase had the highest recall. The F1-scores across the models were relatively close, with ClinicalBERT achieving a slightly higher F1-score. These metrics collectively indicate nuanced differences in the LLM models’ performance, emphasizing the importance of considering various evaluation measures for a comprehensive assessment. Afterward, we incorporated a domain expert into the process to allocate the remaining concepts into proper category, ensuring a comprehensive and accurate clinical concept classification in the dynamic ATL environment.
Use the wisdom of many
Every text classification algorithm has its own strengths and weaknesses. There is no single algorithm that always works well. One way to circumvent this is by using an ensemble of multiple classifiers. The data are passed through every classifier, and the predictions generated are combined (e.g. majority voting) to arrive at a final class prediction. Owing to this, we evaluated four LLMs such as BERTBase, DistilBERT, ClinicalBERT, and SCIBERT in our study towards clinical concept classification. The UpSet analysis graph, as shown in Figure 7, provides valuable insights into the intersection and performance dynamics across different degrees for these LLMs in an ensemble model environment. Similarly, the data reveal distinct patterns and nuances, shedding light on the collaborative strengths of the models.

Upset analysis to measure the LLM performance in ensemble learning environment towards clinical concept classification.
In Figure 7, the
As we progress to the third degree, the complexity of ensemble interactions becomes evident. The intersection involving BERTBase, DistilBERT, and SCIBERT reveals a unique set of clinical concepts with a 10.6% (94) overlap. In the same way, the pairing of BERTBase, DistilBERT, and ClinicalBERT uniquely classifies concepts with a 7.8% (69) overlap. This suggests that the synergy among these three models contributes to the extraction of diverse and complex clinical information. Finally, the fourth-degree intersection, involving all four models (SCIBERT, DistilBERT, BERTBase, and ClinicalBERT), showcases a more specialized set of clinical concepts, accounting for 25.1% (222) of the total. This highlights the ensemble's ability to capture intricate medical information, leveraging the collective strengths of each model. Further exploration into additional third-degree intersections uncovers distinctive patterns, such as the 4.7% (42) and 3.2% (28) overlaps between SCIBERT, BERTBase, ClinicalBERT, and DistilBERT. This specific combination suggests a collaborative effect in extracting clinical concepts not fully captured by individual models.
To sum up, the ensemble model environment proves to be conducive to capturing a wide spectrum of clinical concepts, with varying degrees of overlap and collaboration among the four LLMs. Moreover, these findings provide nuanced insights into the intricate relationships and performance dynamics within the ensemble, offering valuable guidance for optimizing model selection in clinical concept extraction tasks. Furthermore, future research may delve deeper into fine-tuning strategies, model interpretability, and generalizability across diverse clinical datasets to further advance the effectiveness of ensemble modeling in healthcare applications.
DL and LLM models parameter tuning and performance evaluation (M3)
In our experimental environment, we leverage the Ktrain 52 framework for training LLMs, which is a streamlined interface for the TensorFlow Keras deep learning library. The Ktrain framework simplifies the processes of constructing, training, and deploying LLMs, DL, and various ML algorithms. Similarly, the TensorFlow and Keras libraries are used for training DL models for the clinical concept classification task. We utilized a final dataset comprising 9000 clinical concepts, evenly distributed with 3000 concepts in each category (Problem, Treatment, and Test), for training our classification models. The data were split with an 8:2 ratio, allocating 80% for model training and 20% for evaluation purposes. As outlined in the proposed methodology, we undertook the training and evaluation of diverse DL models, including pretrained LLMs, for the purpose of concept classification.
DL model parameter turning
Our initial approach involved applying DL models to leverage contextually generated word embeddings based on LLMs. We referred to these DL models as downstream models. Throughout the training process, which included models such as RNN, CNN, LSTM, BiLSTM, and GRU, we performed parameter tuning to identify the most optimal settings.
Table 6 provides a comprehensive overview of the tuned parameters that resulted in notable accuracy for these models. The dropout layer with a value of 2.0 is added only to the CNN model, while for the other models, we added an L1 regularization layer of 0.001 to reduce and prevent model overfitting. The purpose of adding these regularization layers to the DL models is to achieve better generalization performance on unseen data. Furthermore, in our DL model experiments, we employed the Adam optimizer, which is known for its adaptive learning rate and efficient optimization capabilities. These DL models are trained over 10 epochs, with a batch size of 32, to strike a balance between computational efficiency and generalization. Additionally, the categorical cross-entropy loss function was utilized to measure the dissimilarity between predicted and actual class distributions. This choice is particularly suitable for multiclass classification tasks, ensuring the model optimizes its parameters to minimize classification error.
Presented deep learning (DL) model and large language models (LLMs) tuning parameters utilized during training.

LLM parameter tuning
To gain a deep insight into the clinical concept classification task, we carefully choose and employed a set of hyperparameters, as shown in Table 6, for training BERT-based models, including BERTBase, ClinicalBERT, SCIBERT, and DistilBERT, using the Ktrain framework. 53 The selected hyperparameters, such as a maximum sequence length of 512, maximum features set at 10 k, embedding dimension of 768, a learning rate of 2e−5, batch size of 6, and a total of 2 training cycles, were strategically tuned to explicitly optimize the models’ performance for the clinical concept classification task. These parameters were chosen based with the aim of achieving a balance between effective learning, computational efficiency, and model generalization. Similarly, we perform ablation study to choose optimal parameter as presented in the Section “Ablation study and learning rate analysis for LLMs”. Consequently, the results of this parameter selection provide valuable insights into the fine-tuning process for BERT-based models in clinical contexts.
DL model performance
We have experimented with distinct state-of-the-art DL models (RNN, LSTM, BiLSTM, GRU, CNN) using different combinations of BERT-based word embeddings, along with their corresponding performance metrics of loss and accuracy. We found that a CNN DL model with different LLM embeddings emerged as an ideal performer in terms of achieving high training and test accuracy, as well as low training and testing loss, as shown in Table 7.
Deep learning (DL) model performance trained over large language model (LLM) contextual embeddings.

Similarly, the RNN, GRU, LSTM, and BiLSTM models demonstrated strong performance with accuracy around 87–89% for both training and testing, suggesting good learning capabilities. However, the CNN achieved the highest accuracy among BERT embeddings (BE + CNN) at 92% for training and 91% for testing, and the lowest training and testing loss of 0.22 and 0.28, respectively, emphasizing its effectiveness in capturing hierarchical features in sequential clinical data, as shown in Table 7.
Furthermore, DL models with scientific-based embeddings (SCIBERT) exhibited competitive performance, particularly with CNN (SCI_BE + CNN), which achieved remarkable accuracy of 95.3% and 92.7% on training and testing, with minimal loss of 0.14 and 0.22. Other models, like RNN, GRU, LSTM, and BiLSTM with scientific-based embeddings (SCIBERT), also performed well on both training and testing, showing improved accuracy compared to their BERT-based embedding counterparts.
Likewise, DistilBERT combined with CNN (DistilBERT + CNN) stood out with exceptional accuracy of almost 92.4% for training and 89.2% for testing, at an economical loss of 0.21 and 0.34, indicating the effectiveness of leveraging pretrained transformer-based embeddings in conjunction with convolutional layers. In contrast, LSTM, GRU, RNN, and BiLSTM models with DistilBERT embeddings also performed well, showcasing the ability of transformer-based embeddings to enhance sequential learning. The ClinicalBERT, a domain-specific contextualized embedding, contributed to improved accuracy across various models, particularly with CNN (ClinicalBERT + CNN) achieving 94.4% and 92.4% accuracy for training and testing, with a loss of 0.16 and 0.24. Despite this, the RNN, GRU, LSTM, and BiLSTM models with ClinicalBERT embeddings demonstrated notable performance, highlighting the significance of domain-specific embeddings in clinical applications. Notably, the overall CNN model with LLM embeddings stands out with the lowest training and testing loss and the highest training and testing accuracy, as shown in Table 7, highlighted in green to depict its importance.
Additionally, we calculated the time complexity for training various DL models combined with different embedding techniques, detailed analysis is provided in Table 7. To accurately assess the time complexity, we calculate the total time taken across 10 epochs. BERT embeddings (BE) combined with different models exhibit the highest time complexity, with the BiLSTM model taking the longest time at 83 seconds across 10 epochs, followed by LSTM at 72 seconds, and GRU, RNN, and CNN all within the 53 to 61 seconds’ range. The SCIBERT embeddings (SCI_BE) show a noticeable reduction in time complexity across all models compared to BERT. Specifically, SCI_BE combined with GRU achieves the fastest processing time at 32 seconds across 10 epochs, while BiLSTM, LSTM, RNN, and CNN also perform efficiently, ranging from 33 to 48 seconds over 10 epochs. In the similar way DistilBERT embeddings further reduce time complexity, with the GRU model being the most efficient at 30 seconds over 10 epochs, followed by RNN and CNN at 36 and 39 seconds, respectively. Interestingly, the combination of ClinicalBERT embeddings with GRU also reaches 30 seconds per epoch, matching the performance of DistilBERT + GRU. Across all models, GRU consistently demonstrates the lowest time complexity, regardless of the embedding type. Conversely, BiLSTM generally incurs the highest time costs, particularly when paired with BERT embeddings. This analysis reveals that while traditional BERT embeddings are powerful, they are more computationally expensive. In contrast, SCIBERT, DistilBERT, and ClinicalBERT embeddings offer significant reductions in time complexity, especially when paired with GRU and other simpler models like RNN and CNN. These findings suggest that for tasks where computational efficiency is crucial, SCIBERT or DistilBERT combined with GRU or RNN might be optimal choices (Figure 8).

Training and validation accuracy and loss curves for CNN models trained on embeddings generated from four BERT-based variants (BERTBase, DistilBERT, ClinicalBERT, and SCIBERT). The plots demonstrate the model's performance over epochs, highlighting the efforts to mitigate overfitting and ensure robust generalization. Training accuracy and loss are represented by solid lines, while validation accuracy and loss are depicted by dashed lines for each model variant.
Mitigating overfitting and ensuring robust model performance
Figure 9 illustrates the training and validation accuracy and loss curves for CNN models trained on embeddings generated from four BERT-based variants: BERTBase, DistilBERT, ClinicalBERT, and SCIBERT, over 10 epochs. These curves offer a comprehensive visual representation of the models’ performance and their ability to generalize.
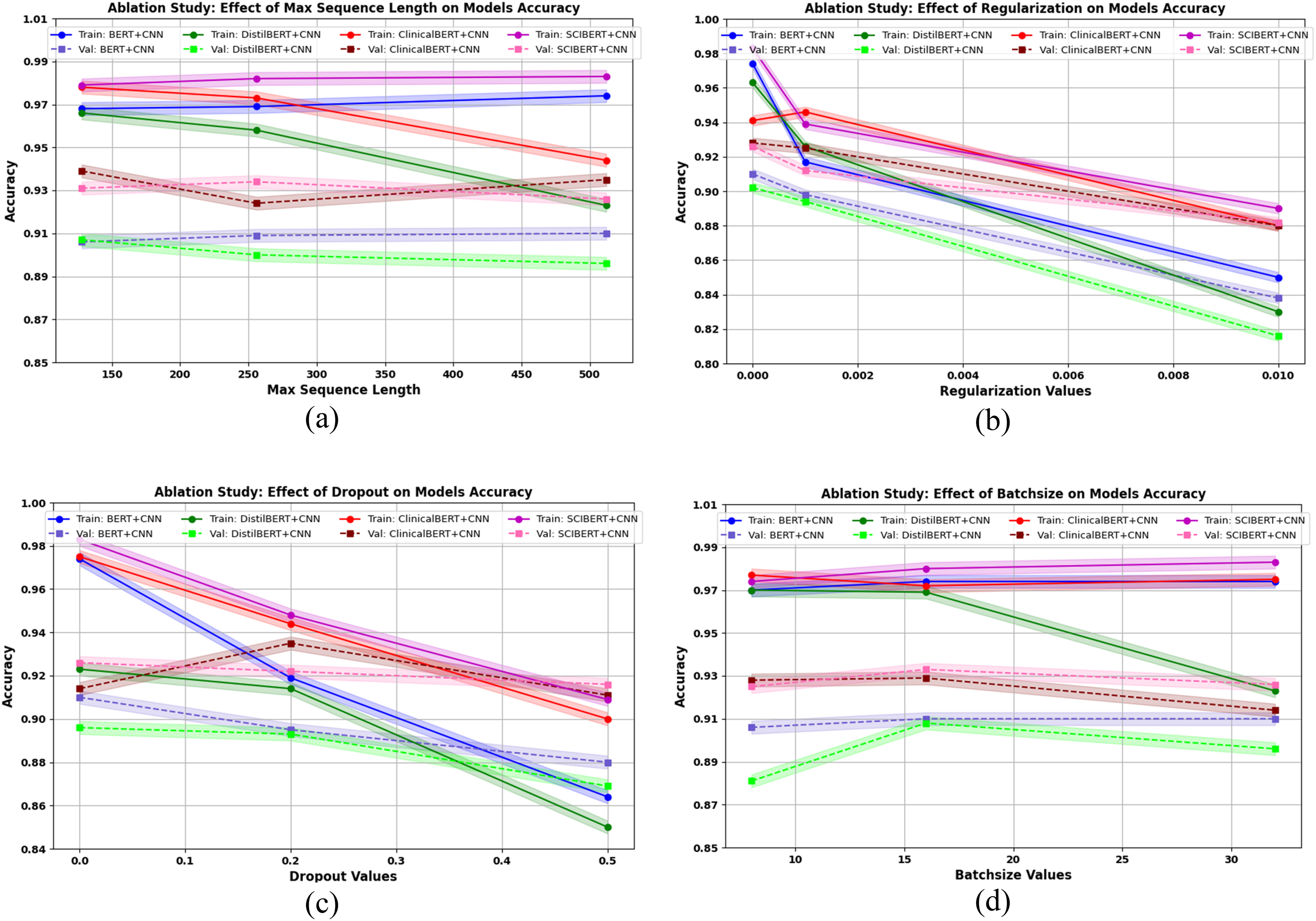
Ablation testing results for the CNN model utilizing BERT-based embeddings (BERTBase, DistilBERT, ClinicalBERT, and SCIBERT), evaluating the impact of sequence length, regularization, dropout, and batch size on model performance.
Consequently, BERTBase + CNN model shows a steady increase in training accuracy, starting at 76.36% in the first epoch and improving to 92.04% by the tenth epoch (see Figure 9(A)). Similarly, validation accuracy rises from 84.03% to 90.48%, with a slight leveling off in later epochs. The training and validation loss curves demonstrate a consistent decrease, indicating that the model is learning effectively without significant overfitting (see Figure 9(E)). The validation loss starts higher but follows a similar downward trend, ending at 0.284 in epoch 10. This suggests that BERTBase + CNN is robust and able to generalize well, maintaining relatively low validation loss.
Similarly, the DistilBERT + CNN model training accuracy starts at 74.85% and reaches 92.40% by the tenth epoch, showing a similar trajectory to BERTBase + CNN (see Figure 9(B)). However, the validation accuracy curve shows some fluctuations after the eighth epoch, peaking at 89.15% before dipping slightly. The validation loss decreases steadily but shows an upward trend in the final epochs, which could be an indication of mild overfitting (see Figure 9(F)). Despite these fluctuations, DistilBERT + CNN remains efficient and competitive, albeit less stable than the full BERT model.
Moreover, ClinicalBERT + CNN emerges as one of the most robust models. Its training accuracy rises significantly, from 78.09% to 94.36% by the final epoch, with a consistently decreasing training loss curve (see Figure 9(C and G)). The validation accuracy also steadily improves, reaching 92.42% by epoch 10, while validation loss steadily declines to 0.242. The alignment between training and validation loss suggests that ClinicalBERT + CNN effectively mitigates overfitting and achieves strong generalization, especially in clinical concept classification tasks, consequently domain-specific embeddings provide a notable advantage.
Additionally, SCIBERT + CNN demonstrates outstanding performance throughout the epochs. The training accuracy improves from 80.77% to 95.31%, with validation accuracy reaching 92.67% (see Figure 9(D)). Importantly, SCIBERT + CNN achieves the lowest validation loss of all models, starting at 0.325 and declining steadily to 0.219 by epoch 10 (see Figure 9(H)). This low validation loss, coupled with high validation accuracy, highlights the model's robustness and ability to generalize effectively in scientific and clinical text contexts. SCIBERT's specialized embeddings appear to provide a significant performance boost.
When comparing the models, SCIBERT + CNN and ClinicalBERT + CNN clearly outperform the others in terms of both training and validation accuracy, with lower validation loss, indicating better generalization capabilities and less risk of overfitting. BERT + CNN also performs well but shows slightly higher validation loss compared to the domain-specific models. DistilBERT + CNN, while efficient and fast, shows some signs of overfitting in later epochs, making it less robust than the other models, especially when working with domain-specific tasks.
Overall, Figure 9 demonstrates that domain-specific models like ClinicalBERT + CNN and SCIBERT + CNN are superior for specialized tasks in medical or scientific domain, as they achieve both high accuracy and low loss with minimal overfitting. These models are ideal for clinical concept extraction and classification task, where domain knowledge is crucial for robust model performance.
DL model ablation testing
In our comprehensive analysis of DL models for clinical concept classification, we evaluated the performance of various DL models trained over different BERTBased word embeddings, including BERTBase, DistilledBERT, SCIBERT, and ClinicalBERT. The receiver operating characteristic (ROC) curves and the corresponding area under the ROC curve (AUC) values were examined for each model. The AUC values serve as a quantitative measure of each model's ability to distinguish between positive (True Positive Rate on
Based on the results of the ablation study and the analysis of training and validation accuracy gaps, the final hyperparameters were selected to ensure the model avoids overfitting or underfitting, while promoting generalizability. In terms of sequence length, a length of 256 strikes the best balance across models, particularly for BERT + CNN, ClinicalBERT + CNN, and SCIBERT + CNN, where it minimized the gap between training and validation performance, indicating strong generalization without overfitting. For regularization, it was consistently observed that applying regularization (even at low values) increased the gap between training and validation accuracies, leading to underfitting. Therefore, no regularization (0) is recommended to maintain optimal performance and prevent underfitting. Regarding dropout, while setting dropout to 0 led to overfitting in some models, particularly BERT + CNN and DistilBERT + CNN, introducing a dropout of 0.2 improved validation accuracy and reduced the gap in models like ClinicalBERT + CNN and SCIBERT + CNN, enhancing generalizability. Finally, a batch size of 32 was found to stabilize performance, minimizing the accuracy gap and ensuring good generalization, especially for ClinicalBERT + CNN and SCIBERT + CNN. This combination of sequence length 256, no regularization, dropout of 0.2, and batch size of 32 provides the best configuration for achieving a well-generalized model across different language models, avoiding overfitting while ensuring robust validation performance.
Comparative analysis among AI based proposed and existing approaches for clinical concept classification
Table 8 provides a comprehensive comparison among several existing approaches and the proposed approaches for clinical concept classification, focusing on precision, recall, and F1-score metrics. Among the existing approaches, Zhu et al. 54 utilized a BiLSTM-CRF model, achieving precision, recall, and F1-score of 89.34%, 87.87%, and 88.60%, respectively. Wu et al. 53 experimented with both CNN and RNN architectures, achieving varying levels of performance, with CNN yielding a precision of 84.91% and an F1-score of 82.77%, while RNN achieved a higher recall of 86.56%. Tang et al. 30 explored SSVMs and CRFs, with the latter achieving a precision of 88.20% and an F1-score of 85.68%.
Performance comparison among proposed approach and existing approaches for clinical concept extraction.

TP: true positive; FN: false negative; FP: false positive; TN: true negative; A: accuracy; P: precision; R: recall; F: F1-score.
In contrast, the proposed approaches leverage various versions of BERT-based embeddings in conjunction with CNN architectures for clinical concept classification task. Consequently, our approach-1 utilized BERTBase embeddings, achieved a precision of 92.03%, a recall of 90.12%, and an F1-score of 91.01%. Similarly, our approach-2, employing DistilBERT embeddings, achieved competitive performance with a precision of 91.51% and an F1-score of 90.36%. Our approach-3, employing ClinicalBERT embeddings, achieved results similar to our approach-2, with a precision of 91.51% and an F1-score of 90.11%. Overall, our approach-4, leveraging SCIBERT embeddings, outperformed the other approaches, achieving a precision of 93.34%, a recall of 92.58%, and an F1-score of 92.68%.
Comparatively, our approaches exhibit promising performance across all metrics, with our approach-4 demonstrating the highest precision, recall, and F1-score among all approaches. This suggests that leveraging BERT-based embeddings in combination with CNN architectures enhances the model's ability to accurately classify clinical concepts. Additionally, our approaches demonstrate consistency and robustness across different variants of BERT embeddings, highlighting their effectiveness in capturing contextual information and improving concepts classification performance. Overall, our approaches offer a compelling alternative to existing methods, showcasing the potential of BERT-based models in clinical concept classification tasks.
LLM performance
Table 9 illustrates the evaluation of different LLMs on both training and testing datasets for the clinical concept classification task. Notably, ClinicalBERT stands out with the lowest training loss (0.05) and the highest training accuracy (98.4%), indicating its strong capability to learn and represent clinical concepts from the training data. Moreover, this model maintains a competitive testing performance, showcasing a testing loss of 0.15 and testing accuracy of 96.0%. These results suggest that ClinicalBERT not only excels in fitting the training data but also generalizes effectively to new, unseen clinical concepts during testing, making it a promising candidate for robust clinical concept classification. In contrast, other models like BERT, DistilBERT, and SCIBERT also exhibit commendable performance, they generally exhibit slightly higher losses and marginally lower accuracies in both training and testing phases. Overall, these four LLMs, exhibit strong performance and deliver superior results underscore their effectiveness towards clinical concept classification task as shown in Table 9.
Large language model (LLM) performance towards clinical concept extractions.

On the other hand we also calculated the time complexity of various LLMs, measured in total time taken across two epochs. The time was initially measured in seconds, but due to the large values, it was converted into minutes for easier interpretation. As a result, BERTBase exhibits the highest time complexity, requiring 121.48 minutes over two epochs, indicating a considerable computational load. In contrast, DistilBERT is the most efficient, taking only 26.84 minutes across two epochs, reflecting its design for faster performance while maintaining accuracy. SCIBERT and ClinicalBERT, which are specialized versions of BERT, show moderate time complexities, with 51.57 minutes and 52.62 minutes over two epochs, respectively. Overall, DistilBERT emerges as the most computationally efficient model in this comparison, while SCIBERT and ClinicalBERT balance specialization with moderate increases in time complexity.
Ablation study and learning rate analysis for LLMs
This section presents an ablation study that examines the impact of two key learning rates—Longest Valley (Red) and Min Numerical Gradient (Purple)—on the performance of four models: BERTBase, DistilBERT, ClinicalBERT, and SCIBERT. The learning rates were determined using the ktrain library, and the models were evaluated based on their training loss, accuracy, validation loss, and validation accuracy. Whereas, the Longest Valley represents a stable area on the learning rate plot where the loss remains low, ensuring reliable performance and reducing the risk of uncertain training behavior. This method is particularly useful for models that aim for stability and to avoid overfitting. In contrast, the Min Numerical Gradient marks the point where the loss gradient is at its lowest, facilitating faster convergence. However, this approach can introduce instability if the learning rate is too high. A comprehensive understanding of these strategies is vital for optimizing model performance, as highlighted in the analysis of the four LLM models in our study.
For BERTBase, the Longest Valley learning rate of 7.43E−06 provided better performance, with lower training loss (0.11) and higher accuracy 0.97%, compared to the Min Numerical Gradient learning rate of 3.18E−05, which resulted in slightly worse performance with higher loss (0.13) and lower accuracy 0.96% (see Figure 10(A)). Similarly, for ClinicalBERT, the Longest Valley learning rate (9.44E−06) resulted in slightly lower validation loss (0.11) and higher accuracy 0.98% compared to the Min Numerical Gradient (4.39E−06), which had a similar training loss but higher validation loss (0.12) as shown in Figure 10(C).

Presents performance comparison impact of two learning rates Longest Valley and Min Numerical Gradient on the training and validation metrics of BERTBase, DistilBERT, ClinicalBERT, and SCIBERT.
For DistilBERT, both learning rates performed exceptionally well, but the Min Numerical Gradient learning rate (2.22E−05) slightly outperformed the Longest Valley (7.29E−06) in terms of training loss (0.07 vs. 0.08) and validation accuracy (0.97% vs. 0.96%) as depicted in Figure 10(B). Similarly, SCIBERT achieved much better training loss (0.06) with the Min Numerical Gradient (3.22E−06), compared to the Longest Valley (3.89E−06), where the training loss was higher (0.11), though both learning rates produced comparable validation results (see Figure 10(D)).
Overall, this study shows that while the Longest Valley learning rate generally provides stable training and good results for BERTBase and ClinicalBERT, the Min Numerical Gradient learning rate tends to offer better convergence for DistilBERT and SCIBERT. The study emphasizes the importance of selecting an appropriate learning rate based on the specific model architecture, as different models benefit from different strategies for learning rate optimization.
Individual LLM performance statistical analysis
We performed statistical analysis of four LLMs (BERT, DistilBERT, ClinicalBERT, and SCIBERT) in classifying clinical concepts derived from the I2B2 test dataset. This dataset includes 990 clinical concepts equally distributed among three categories: Problem, Treatment, and Test, with 330 concepts in each. These models have been previously trained and fine-tuned on I2B2 datasets for accurately categorizing these unseen concepts, and the output is compared against the I2B2 gold standard to evaluate classification performance as shown in Figure 11.

Upset Analysis plot presented statistical analysis of individual LLMs toward clinical concept classification.
In Figure 11, the intersection size bars at the top of the graph depict the number of correctly classified concepts shared across different combinations of models, organized by degrees of overlap. The largest bar, marked in orange, represents the degree-4 intersection, showing 814 concepts (82.2%) that were correctly classified by all four models. This high overlap indicates strong agreement among the models for a substantial subset of clinical concepts, suggesting that these concepts may be inherently easier to classify accurately, regardless of the model used.
Moving to the right, smaller bars reflect concepts correctly classified by fewer models. For example, blue bars (degree-2 intersections) indicate concepts that were correctly classified by two models but missed by the others, while green bars (degree-3 intersections) represent concepts identified by three models. The percentages beneath each bar denote the proportion of the overall dataset for each intersection, indicating that most classifications fall within high-overlap categories, with minimal divergence across models.
On the left side, horizontal bars display the total number of correct classifications achieved by each individual model. ClinicalBERT leads with the highest correct classification count at 961 concepts (97.1%), followed by SCIBERT with 942 (95.2%), BERT with 915 (92.4%), and finally, DistilBERT with 901 (91.0%). This performance distribution suggests that ClinicalBERT, which is tailored for clinical language, outperforms the other models, with SCIBERT also demonstrating high performance due to its specialization in scientific and biomedical language. BERT and DistilBERT, while still performing well, show slightly lower accuracies in comparison, reflecting their more general-purpose language understanding capabilities.
Overall, the Upset plot reveals that ClinicalBERT and SCIBERT are particularly well-suited for clinical concept classification, with high agreement across all models on a large portion of the dataset, and relatively few cases where only one or two models accurately classified a concept. This high consistency underscores the effectiveness of domain-specific models for clinical applications.
Discussion
Clinical documents are usually available in an unstructured format, incorporating important information that assists the practitioner, patient, and hospital in terms of diagnosing disease, prescribing medication to improve patient health, and enhancing practitioner and hospital services, which is time and cost-efficient. The information available in the clinical document could be related to a patient's medical history, current health status, diagnoses, treatments, and overall healthcare management. In this study, we have focused on clinical concept identification and extraction to support biomedical information retrieval, question answering, and clinical event detection and parsing tasks. Researchers and industrialists are using NLP and AI techniques to automate the clinical concept extraction process. Utilizing NLP and AI techniques necessitates annotated or training data for model development. The process of preparing label data requires domain expertise and is time-consuming due to manual annotation. To address this challenge, we have presented an end-to-end process to automatically extract and label clinical concepts, and then evaluated various DL and LLMs to gain deep insights into these models’ performance and contribute to clinical NLP research. The proposed study comprises three modules: initial level concept labeling (M1), ATL process (M2), and DL and LLM for concept classification (M3).
In the initial concept labeling stage, we have leveraged UMLS Metathesaurus to identify clinical concepts. UMLS incorporates over 200 source vocabularies and ontologies, covering various domains of biomedicine, including anatomy, diseases, drugs, procedures, and more. Initially, after preprocessing unstructured clinical text, we can then identify the concept boundary through our proposed algorithm, which is discussed in the Proposed methodology section. Typically, clinical notes contain concepts expressed either as single words or as part of composite terms. While using just n-gram words might not accurately capture these concepts, further processing, such as applying POS tagging and regular expressions, is required to more effectively extract the valuable concepts embedded within unstructured clinical documents.
Moreover, a concept matching to UMLS approach is adapted to extract the semantic information including concept id, concept semantic type. Additionally, we have curated our own dictionaries containing explicit concept category semantic types as shown in Table 1. These dictionaries encompass relevant semantic types that play contextual role in defining explicit concept category. Following this, we establish handcrafted rules to explicitly categorize concepts into Problem, Treatment, and Test categories, upon successfully matching the concept's semantic type with the entries in the semantic dictionaries. Matching words to a lexicon is a fundamental step in generating labeled data for data-driven approaches. As a result, the proposed module (M1) has shown promising results compared to existing methods. Likewise, we assessed the effectiveness of the proposed module (M1) across distinct datasets from the I2b2 Challenge 2010, which encompassed the Beth, Partner, and Test datasets. We scrutinized each dataset meticulously, examining individual concepts to gain a thorough understanding of the proposed approach.
Applying a rule-based approach is a computationally expensive and time-consuming task. Therefore, we introduce the ATL approach to automate the labeling process while incorporating the initial level label data acquired through the previous approach. In the ATL process, we utilize LLMs that require sufficient label data to obtain promising results. To achieve our goals in the ATL process, we incorporate true label instances from the previous approach along with label instances from the gold standard. We obtained gold standard labeled data because the label data acquired from the rule-based approach was insufficient for training the LLMs.
Word embedding similarity is another widely used approach adapted by researchers for data classification based on similarity scores. Following this, we generated embeddings for labeled or known concepts (rule-based concepts and gold standard concepts) using the SCIBERT model for explicit concept categories. Similarly, we generated embeddings for the unlabeled or candidate concepts via SCIBERT, and embedding similarity was performed between known concepts and candidate concepts. As a result, the unseen concepts were ultimately classified into explicit categories based on high embedding similarity scores. Furthermore, we involved domain experts to validate the embedding similarity-based concept classification and incorporate it into the appropriate concept category. Consequently, the labeled data were automatically enhanced, and these newly classified concepts were applied during the ATL process. Therefore, we named this approach the ATL environment.
Hence, after acquiring a substantial amount of labeled data from the ATL environment, we experimented with DL and LLMs to automate the clinical concept classification task. We adapted a contextual word embedding approach for feature generation and trained various state-of-the-art DL models such as RNN, LSTM, BiLSTM, GRU, and CNN. We leveraged BERT base variant LLM models, such as BERTBase, DistilBERT, SCIBERT, and ClinicalBERT, to generate the contextual word embeddings and trained DL models over them. Throughout the experiment, the CNN model, when combined with all variant BERT base embedding models, achieved promising results for concept classification on both training and test data.
Likewise, an ablation study was conducted to evaluate the performance of a CNN model trained over various LLM embeddings, for the clinical concept classification task. The study examined the influence of key hyperparameters such as batch size, sequence length, dropout, and regularization on the model's training and validation accuracy and loss. By systematically adjusting these parameters, we fine-tuned the model to optimize performance while maintaining a balance between underfitting and overfitting. To further mitigate overfitting and ensure robust model performance, we plotted the training and testing scores, as well as training and validation loss for CNN model trained over LLM embeddings. These plots highlighted the model's ability to generalize across different embeddings, providing insights into how various LLMs can enhance the CNN's capability for clinical concept classification.
Moreover, we conducted another ablation study to determine the optimal learning rate for various LLMs in the context of the clinical concept classification task. Specifically, we examined two key learning rates: Longest Valley and Min Numerical Gradient for each LLM, as suggested by the learning rate plots generated during training. These rates varied across models, reflecting the unique learning dynamics of each LLM. Based on the results, we selected a learning rate of 2e−05 as the optimal value for all LLMs, ensuring a balance between training stability and performance. This consistent choice of learning rate allowed for improved model convergence while maintaining robust accuracy and minimizing loss on both the training and validation sets. The aim of ablation study to demonstrates the importance of selecting appropriate hyperparameters and embedding models to ensure effective and stable model performance in clinical NLP tasks. In addition, we performed a statistical analysis to measure the performance of individual LLMs on unseen clinical concepts. The results reveal that ClinicalBERT and SCIBERT are particularly well-suited for clinical concept classification, with high agreement across all models on a large portion of the dataset and relatively few cases where only one or two models accurately classified a concept. This high consistency underscores the effectiveness of domain-specific models for clinical applications.
Though there is a myth that CNN does not perform well in NLP tasks, our observation shows that CNN performs best in our study due to the concept window size, which is not greater than six words. This smaller window size allows CNN to focus on local patterns within the text, easily capturing short-range dependencies and extracting relevant features from neighboring words. Moreover, we further trained the pretrained LLMs individually, including BERTBase, DistilBERT, SCIBERT, and ClinicalBERT. As a result, ClinicalBERT outperformed the others, likely because ClinicalBERT is trained on the Medical Information Mart for Intensive Care III (MIMIC-III) dataset. The MIMIC-III dataset consists of EHRs from 58,976 unique hospital admissions of 38,597 patients in the intensive care unit of the Beth Israel Deaconess Medical Center between 2001 and 2012. Similarly, the dataset we utilized was collected from the Beth Israel Deaconess Medical Center for the I2b2 challenge 2010, on which ClinicalBERT outperformed in our study for clinical concept classification.
The motivation behind this study is to introduce an end-to-end approach to automatically prepare annotated data, advancing clinical NLP research by adopting state-of-the-art data-driven approaches for various clinical tasks. In the future, we will further extend this approach for clinical concept classification by incorporating character-level, concept-level, and UMLS semantic-level features to enhance large language model accuracy and streamline the clinical concept annotation process.
Conclusions, limitations, and future works
Labeling unstructured clinical text manually is time-consuming and requires domain expertise. To streamline this process, we proposed an automated end-to-end approach for clinical concept labeling into Problem, Treatment, and Test. The proposed approach consists of three modules: a heuristic approach (M1), an ATL approach (M2), and a DL and LLM approach (M3).
In M1, we used the UMLS dictionary and lexical semantics to categorize clinical concepts. Once a substantial amount of data was labeled, we moved to M2, where we employed BERT-based models (DistilBERT, SCIBERT, and ClinicalBERT) for automatic labeling, keeping the domain expert in loop. We generated contextual word embeddings of labeled concepts and unlabeled concepts and adopted embedding similarity approach with in some threshold scores to categorize unlabeled concepts. A domain expert is utilized to further validate the classified concepts. Finally, in M3, we explored DL models combined with various LLM embeddings. Remarkably, CNN models incorporating all variant LLM model embedding achieve promising accuracy. Notably, ClinicalBERT emerged as the leading performer among the LLM models, showcasing superior performance in concept classification. This approach not only alleviates the burden of manual labeling but also enhances the efficiency and accuracy of clinical concept classification in unstructured text.
Despite the promising results, our study has some limitations. First, the performance of the proposed model depends heavily on the availability of high-quality labeled data, as the quality and accuracy of annotations can significantly affect classification performance. Additionally, the generalizability of our model may be restricted to the specific context of the i2b2 dataset and might require adaptation for use with other datasets or in different clinical settings. Moreover, the proposed methodology is specifically designed for Problem, Treatment, and Test clinical concept classification tasks, and its effectiveness may vary when applied to other medical protocols or classification tasks.
In future research, a promising direction to explore is the incorporation of prompt engineering techniques based on LLMs. LLM-based prompt engineering has demonstrated its potential to enhance the performance of language models across various NLP tasks. Integrating these techniques into the proposed methodology for clinical text annotation and classification could lead to even greater improvements.
Furthermore, combining active learning with LLM-based prompt engineering can improve annotation quality and model performance. By utilizing the contextual knowledge and capabilities of LLMs, the model can gain a deeper understanding of the clinical context, leading to more accurate annotations during the active learning process.
Overall, incorporating LLM-based prompt engineering techniques into the proposed methodology could significantly advance clinical text annotation and classification. This approach can improve the model's efficiency, accuracy, and generalizability, making it more robust in handling variations in clinical notes and enhancing its performance in clinical practice settings.
Footnotes
Contributorship
AA conceived the idea, responsible for data curation, investigation, methodology, implementing experiments, conducting evaluations, and drafting the original manuscript. ML played a crucial role in refining the manuscripts, rewriting the sections, and supervised. NS assisted in the reviewing and writing process and participated in the project's experiment discussions. VK assisted in the writing process and manuscript refinement.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
This study did not require ethics committee review and approval.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Guarantor
AA.
Informed consent
This study used a publicly available dataset (I2b2-2010 challenge) that did not include any directly identifiable patient information. Consequently, informed patient consent was not necessary for this research.
Appendix A. Deep learning model selection computing AUC ROC
In our comprehensive analysis of deep learning models for clinical concept classification, we evaluated the performance of various DL models trained over different BERTBased word embeddings, including BERTBase, DistilledBERT, SCIBERT, and ClinicalBERT. The receiver operating characteristic (ROC) curves and the corresponding area under the ROC curve (AUC) values were examined for each model. The AUC values serve as a quantitative measure of each model's ability to distinguish between positive (True Positive Rate on
The LSTM and BiLSTM models exhibited slightly lower but still commendable AUC values of 0.96% (see Figure A1). Transitioning to DistilledBERT, we observed consistently high AUC values across all architectures, with the CNN model leading the way with an AUC of 0.97%, reinforcing the robustness of DistilledBERT embeddings (see Figure A1). Moving to SCIBERT, all models showcased exceptional discriminative power, particularly the CNN architecture with a remarkable AUC of 0.99% (see Figure A1). The RNN, GRU, LSTM, and BiLSTM models closely followed with AUC values of 0.98%, indicating the superior discriminative capability of SCIBERT embeddings for clinical concept classification (see Figure A1).
Finally, ClinicalBERT embeddings demonstrated consistently high AUC values across all deep learning architectures, reinforcing the effectiveness of this domain-specific embedding for clinical tasks (see Figure A1).
In summary, our comparative analysis underscores the nuanced interplay between LLM's based word embeddings and deep learning model, with each combination exhibiting strengths in the discriminative power for clinical concept classification. The choice of BERT-based embedding, whether BERTBase, DistilBERT, SCIBERT, or ClinicalBERT, offers distinct advantages, providing practitioners with valuable insights for optimizing model selection based on the specific requirements of their clinical applications.
Appendix B. Individual concept classification using LLMs
In the context of clinical concept classification, a comprehensive analysis of the performance metrics reveals distinctive characteristics among four state-of-the-art large language models: BERTBase, DistilBERT, SCIBERT, and ClinicalBERT as shown in Table B1.
Regarding precision (P), ClinicalBERT exhibits the highest precision score of 98%, indicating a remarkable ability to correctly identify Problem clinical concepts. DistilBERT and SCIBERT follow closely with a precision score of 97%, showcasing its robust performance towards Treatment and Test concepts classification as shown in Table B1. In terms of recall (R), DistilBERT and SCIBERT emerge as top performers, attaining an impressive score of 97% in accurately classifying the Problem concept. This achievement underscores their effectiveness in capturing a significant proportion of actual positive instances.
In the context of Treatment concepts, ClinicalBERT and SCIBERT closely boast recall scores of 97% and 98%, respectively. Notably, for Test concept categorization, BERT, DistilBERT, and ClinicalBERT exhibit consistent performance, each achieving an equal recall rate of 96%. This indicates their reliability in identifying positive instances within the Test category as shown in Table B1. The F1-score, serving as a balanced metric between precision and recall, effectively illuminates the overall performance of the models in the context of concept classification. For the categorization of Problem concepts, both BERT and SCIBERT demonstrated an impressive F1-score of 97%, surpassing the slightly lower but still commendable scores of 96% obtained by DistilBERT and ClinicalBERT. Similarly, in the realm of Treatment concept classification, SCIBERT and ClinicalBERT exhibited closely matched F1-scores of 96%, outperforming the respective scores of 95% achieved by BERT and DistilBERT.
Additionally, when assessing the classification of Test concepts, BERT consistently maintained a competitive F1-score of 96%, while DistilBERT, SCIBERT, and ClinicalBERT closely followed with F1-scores of 95% as shown in Table B1. Overall, these LLM models demonstrating a balanced performance in precision and recall, also exhibit strong F1-scores, emphasizing their suitability for clinical concept classification tasks.
