Abstract
Background
Federated Learning (FL) offers a privacy-preserving solution for multi-party data collaboration in smart healthcare. However, the data heterogeneity among hospitals and among patients often results in suboptimal performance for some hospitals when applying a global FL model. Current clustering-based FL methods struggle to adapt to complex and diverse data distributions, negatively impacting model performance.
Methods
We propose a novel framework, Federated Gaussian Mixture Clustering (FedGMC), which leverages Gaussian Mixture Clustering to train personalized FL models. FedGMC determines the optimal number of clusters prior to the FL process, reducing the time and computational cost associated with traversing multiple clustering configurations in existing approaches.
Results
The FedGMC framework was evaluated using real-world eICU datasets with various classifiers and performance metrics. Experimental results show that FedGMC outperforms other baseline methods in terms of the overall performance of combining two classifiers and two performance metrics. Moreover, it mitigates the risk of performance degraded for participating hospitals following FL.
Conclusions
The FedGMC framework effectively addresses clinical heterogeneity, enhancing predictive performance and ensuring fairness among participating medical institutions. These improvements increase the willingness of data owners to engage in the collaboration FL initiatives.
Keywords
Introduction
Applying machine learning to electronic health records (EHRs) has achieved remarkable outcomes in various medical and healthcare applications, furnishing substantial support for clinical decision-making.1,2 However, accurate machine learning modeling often depends on large-scale data samples, which are typically distributed among diverse hospitals and institutions. EHR data contains extensive patient-related information, making its use subject to stringent privacy protection requirements.3,4 Federated Learning (FL) has emerged as a promising solution to address these challenges. FL allows data owners (i.e. clients) to convey locally trained model parameters to a central server without transferring the original data. The central server aggregates these parameters and generates a global model through iterative training across multiple rounds. By leveraging FL, the diversity and scale of data available for training can be significantly enhanced, leading to models with improved performance.5–7 Given its ability to jointly develop machine learning models while preserving data privacy, FL is regarded as an ideal framework for privacy-sensitive data such as EHRs. It has been successfully applied to tasks such as disease risk prediction, clinical diagnosis, and medical image recognition, thereby demonstrating its potential to transform healthcare analytics,8,9 directly contributing to Sustainable Development Goal (SDG) 3 (Good Health and Well-being) while addressing SDG 9 (Industry, Innovation, and Infrastructure) through AI frameworks.
Significant obstacles remain in application of FL in healthcare field. A primary challenge lies in the statistical data heterogeneity among clients, termed non-independent and identically distributed (non-IID) data, which is particularly pronounced in healthcare. This heterogeneity stems from factors such as geographic location, clinical practices, patient demographics, genetic diversity, and phenotypic differences. As a result, FL algorithms often face impaired performance, with the global model showing considerable variability in effectiveness across different hospitals. In some cases, the performance of the global model may even worse than local model in specific hospitals.5,6 Furthermore, patient heterogeneity adds another layer of complexity, as a single global model may struggle to achieve desired performance across diverse patient cohorts.10,11 If these issues can be overcome, it will facilitate the collaborative training of models among medical institutions while protecting data privacy, and further support SDG 17 (Partnerships for the Goals), thus addressing the problem of data silos that hinder global health programs.
Researchers have explored clustering-based FL approaches 12 to address these challenges. The research of Stallmann 13 divided these methods into two categories. The first and more prevalent type, called as clustered federation, divides clients into multiple clusters based on their similarity. Each client participates in federated training only within its respective cluster. The second category is called as Federated Clustering or Sample Clustered FL, 14 which is concerned with identifying global clusters to which samples belong in distributed data without sharing data.
Clustered federation approach assumes that data within each cluster is identically distributed or that each client contains a single type of data.15–18 However, this assumption conflicts with the cross-silo scenarios typical in the medical domain, where heterogeneity among patients in each hospital may be significant. Consequently, these methods are better suited to cross-device scenarios, such as the Internet of Things or smart wearable devices and fail to adequately address the unique challenges posed by heterogeneity among hospitals in healthcare.17,19–22
Federated Clustering is particularly well-suited for cross-silo scenarios and effectively addresses the problem of data heterogeneity within a single data center.
13
However, research in this area remains relatively scarce. Most existing studies refined the federated clustering process by utilizing K-means and its derivative algorithms, such as C-means.22–24 In the context of smart healthcare, research has shown that the misclassification rate using K-means clustering is approximately four times higher than that of clustering methods based on probabilistic models.
25
The K-means algorithm, despite its widespread use, faces inherent limitations that reduce its effectiveness in federated clustering scenarios, particularly in handling the complex and heterogeneous data common in healthcare. Key limitations include the following:
Poor recognition of non-convex and multi-peaked clusters. K-means assumes that the clusters are circular or nearly circular, and when the clusters in a dataset were irregularly shaped, it may struggle to accurately delineate the clusters. A hard clustering method, not flexible enough. If a sample does not exactly match the characteristics of any of the clustering centers, it is still forced to be classified to a clustering center in the K-means algorithm. Sensitive to noise. Since the K-means algorithm is based on a distance metric, outliers may significantly affect the location of the clustering centers and the quality of the clustering results.
To address these challenges, we propose the Federated Gaussian Mixture Clustering (FedGMC) framework. It comprises three sequential stages: Patient Encoding, Federated Clustering, and Personalized FL. It overcomes the limitations of traditional K-means by leveraging a probabilistic clustering approach and addresses the heterogeneity issue using a personalized FL method. We evaluated the effectiveness and stability of the framework using a real-world EHR dataset of acute kidney injury (AKI) patients. The experimental results demonstrated that FedGMC not only consistently outperformed baseline methods in terms of predictive performance metrics but also maintained fairness across participating institutions by significantly reducing risk of performance degradation.
Literature review
This study follows the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) guidelines to conduct a systematic literature search on data heterogeneity solutions for federated learning based on clustering method, aiming to systematically evaluate the technical bottlenecks of existing methods. We performed a structured search in the Google Scholar databases (January 2019 to May 2025) using the keyword “federated learning clustering.” First, non-Original Research Articles such as reviews and conference abstracts were excluded; then, full-text quality evaluation was conducted on the remaining literature to exclude those lacking control groups or insufficient relevance to the theme. Finally, core research achievements were selected and are shown in Table 1.
Clustering-based FL optimization method.

The first category is clustered federation. This category of method performs clustering based on clients data distribution,16,40 model parameters, 18 or update gradients. 17 The clustering methods used include K-means, its derivative algorithm fuzzy C-means, Hierarchical clustering, and GMM. Sahinet 26 and some researchers used K-means and its derivative algorithm fuzzy C-means for clustering. Briggs et al. 15 introduced a hierarchical clustering step and used the similarity between the client's local update and the global federated model to divide the client clusters. In 2024, Malekmohammad et al. 31 uses GMM clustering to achieve client clustering by combining model update and training loss. All these methods categorized as clustered federation usually assume that the data is evenly distributed among clients or that each client contains only a single type of data. It ignores the reality that there may be significantly different sample clusters between different clients/hospitals and within the same dataset.
The second category is Federated Clustering, which clusters samples. The clustering methods used include K-means, its derivative algorithm (fuzzy C-means, fuzzy C-means), Hierarchical clustering, and GMM. In algorithm k-FED,
23
each client performs K-means clustering on the local data and sends the clustering results to the server. The server performs K-means clustering on the centroids sent by all clients. Stallmann et al.
13
proposed the FedFCM algorithm in 2022, which is an algorithm similar to k-FED that utilizes fuzzy c-means.
41
In 2024 and 2025, latest researches still focus on improving algorithms with k-means and its derivative algorithm. Bárcena
36
and Stallmann
37
used c-Means and Fuzzy c-Means clustering, respectively. In the section Introduction, we have elaborated on the limitations of this type of algorithms. Ahmed et al.
38
proposed a framework PCBFL based on spectral clustering
42
in medical and healthcare area. It has a time complexity of O (
Based on the above analysis, the clustered federation method cannot cope well with the data heterogeneity problem in the cross-silo scenario targeted by this study. Existing methods based on Federated Clustering have defects in accuracy and algorithm complexity or are not suitable for horizontal FL scenarios.
Methods
Framework overview
The FedGMC framework comprises three phases, as shown in Figure 1.
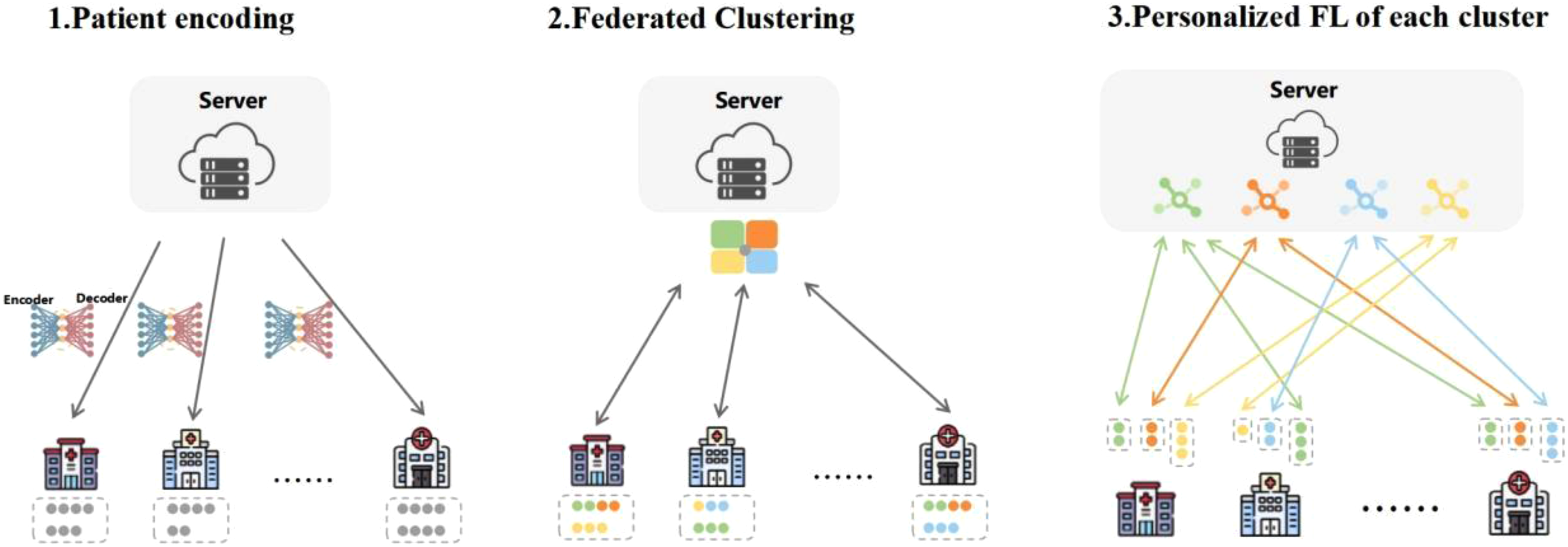
Schematic of FedGMC.
Patient encoding
This stage preprocesses patient data, creating robust feature representations while preserving data privacy. Each hospital uses the vector encoder provided by the central server to transform local patient cohorts into vector representations. Continuous data is directly encoded using its measurement values, while categorical data is processed using one-hot encoding, to maintain consistency and interpretability across the federated system.
Federated clustering
This phase involves clustering the entire patient cohort across hospitals without sharing raw data, preserving data privacy. The clustering process leverages the probabilistic Gaussian Mixture Model (GMM) to effectively capture complex data distributions and heterogeneity among patients. Detailed procedures are elaborated in section “Federated clustering.”
Personalized FL for each cluster
The framework trains personalized FL models for each cluster, and all hospitals participate only in the FL of their respective clusters, ensuring that each cluster benefits from a tailored model optimized for its unique characteristics. Comprehensive details of this phase are provided in section “Personalized federated learning.”
The FedGMC framework integrates these phases seamlessly to address the challenges of heterogeneous and privacy-sensitive data in healthcare, enabling effective and equitable predictive modeling across institutions.
Federated clustering
The GMM 44 is a probabilistic model capable of representing datasets that can be partitioned into multiple Gaussian distributions. Unlike hard clustering algorithms such as K-means, GMM assigns a likelihood to each sample belonging to a given cluster, offering a more nuanced approach to clustering. The model characterizes each cluster's position and shape using two key parameters: mean and covariance, enabling flexible, and precise clustering. In comparison to K-means and its derivative clustering algorithms, 25 GMM offers the following advantages. First, unlike K-means, which assumes spherical clusters, GMM can model clusters with diverse and irregular shapes. Second, as a soft clustering method based on probability density, it assigns each sample the probability of belonging to different clusters, which make it more adaptable to complex datasets where it is difficult to assign each sample to a hard cluster. Third, GMM's probabilistic approach makes it less sensitive to outliers and noisy data. Fourth, GMM accounts for not only the mean and centroids of clusters but also the variance, sample distribution shapes, and underlying statistical properties. Our study integrates the GMM method into federated clustering, enhancing the process by accounting not only for mean of samples and centroids but also for variance, sample distribution shapes, sample sizes, and feature differences across clusters and hospitals. This approach enables the central server to receive richer and more comprehensive distribution information about the patient cohort, resulting in significantly improved clustering performance. The improved GMM-based federated clustering phase (Phase 2) is detailed in Algorithm 1, which illustrates the step-by-step process. Compared to baseline algorithms, such as CBFL 16 and K-Fed, 23 the enhancements introduced in FedGMC focus on improving the performance of Phase 2. Table 2 summarizes the differences between these methods, highlighting how FedGMC's GMM-based approach addresses the limitations of existing federated clustering frameworks.
Comparison of phase 2 with baseline algorithms.
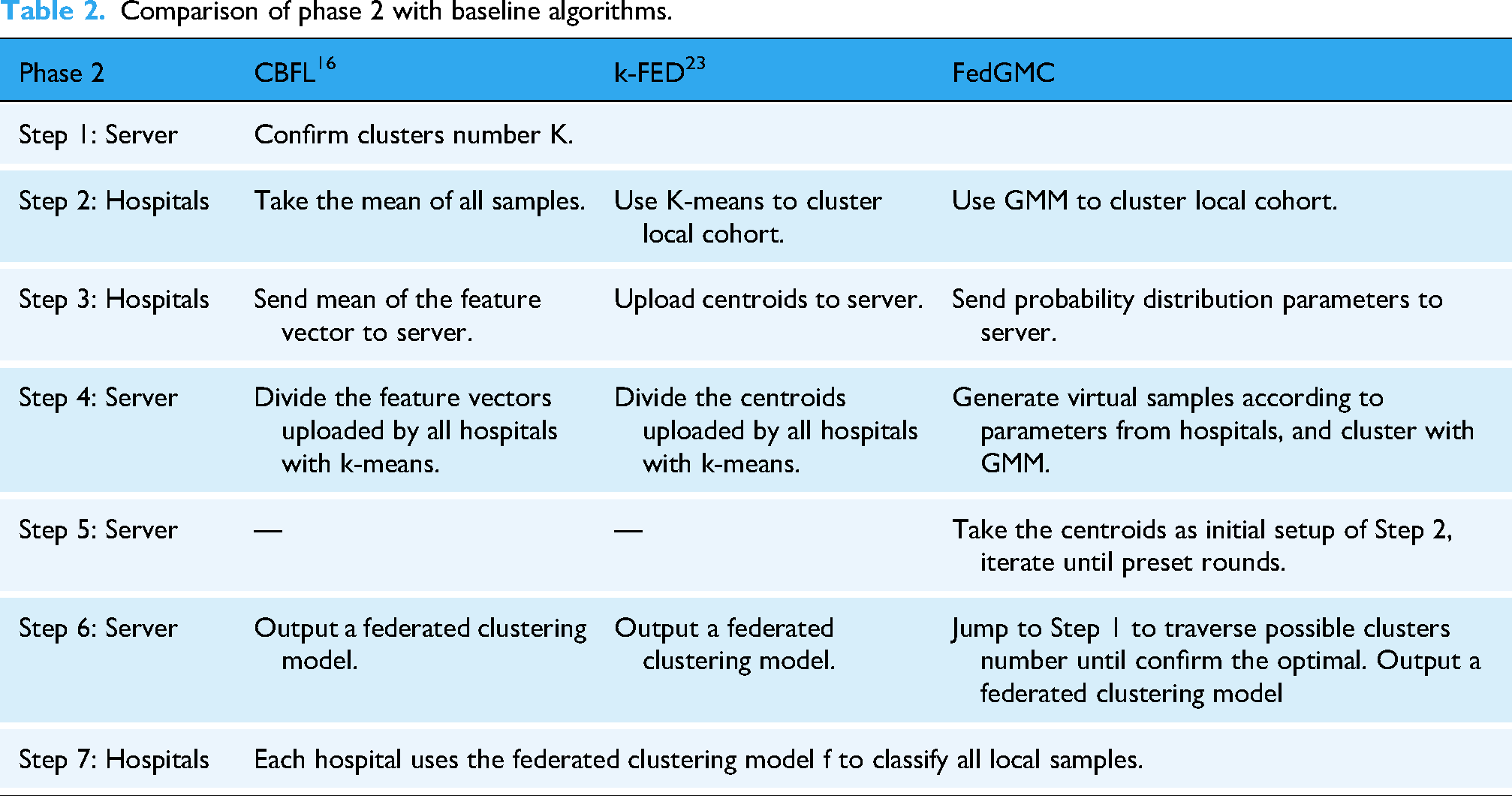

Determining the optimal number of clusters K is a critical challenge in clustering algorithms. Existing studies often require exhaustive traversal through all possible values of K, leading to significant time, communication, and computational overhead in FL settings. To address this, we introduce the silhouette coefficient in Phase 2 as an evaluation metric for clustering effectiveness. 45 The silhouette coefficient measures the difference in similarity between samples within clusters and between clusters, providing a comprehensive assessment of clustering quality. Its value range is [−1, +1]. 46 The larger the value, the better the clustering effect. The K value that provides the maximum silhouette coefficient is selected as the optimal number of clusters (see Appendix A. Algorithm 2).
Personalized federated learning
In phase 3, all hospitals engage exclusively in the training of personalized FL models for the clusters they host. The detailed training process is as follows:
The server initializes the prediction model for each cluster and distributes it to the hospitals containing patients belonging to that cluster. Each hospital conducts FL model training locally for the clusters it contains, updating the model parameters using its data. Hospitals encrypt the updated model parameters and transmit them securely to the server. The server aggregates the parameters received from all hospitals to update the cluster-specific models. If the preset convergence criterion is not reached, the server sends a new round of updated cluster model to the client containing the cluster, and the process loops back to (2) for a new round of FL training. Always criteria for convergence of FL training are generally set to one of the following, such as the change in loss value or model parameters between iterations is less than the preset value, or the preset maximum number of training rounds is reached.
47
Data and experiments design
Datasets and preprocessing
This study used a real-world eICU dataset to validate proposed method. 48 The dataset encompasses EHR data from over 200 hospitals and more than 100,000 ICU patients across the United States in 2014 and 2015. It includes diverse patient information such as demographics, medications, diagnoses, procedures, time-stamped vital signs, and laboratory test results.
The experimental prediction task was to forecast whether a patient would develop acute kidney injury (AKI) 48 h in advance. AKI is a potentially life-threatening condition that complicates treatment, impacts clinical trajectories, and can significantly worsen outcomes for a substantial number of hospitalized or ICU patients.
49
Early prediction of AKI risk can enable timely interventions, improve patient outcomes, and significantly reduce hospitalization costs and mortality rates.1,50 The definition of AKI followed the method proposed by the Kidney Disease Improving Global Outcomes (see Appendix A).
51
The study cohort excluded the following patient samples to ensure data reliability and relevance:
Patients with <2 measurements of Serum Creatinine (SCR); Patients with an estimated Glomerular Filtration Rate (eGFR) < 15 mL/min/1.73 m2 at admission; Patients aged <18 years; Patients with an ICU stay duration of <48 h from admission to discharge.
Selecting feature variables guided by expert knowledge has proven to be more effective for constructing data-driven machine learning models. 52 In this study, clinical features critical for predicting AKI were selected based on the expert recommendations from the Kidney Disease Improving Global Outcomes (KDIGO) guidelines. 51 Features with a high rate of missing data (occurrence <10%) were excluded to ensure data quality. This process resulted in the selection of 22 discrete traits and 71 continuous traits. We deleted discrete features from the clustering process for three reasons. First, as we employed the GMM algorithm, which assumes Gaussian-distributed data, only continuous features meet this requirement. Second, GMM characterizes correlations between features using covariance matrices. However, the covariance between binary and continuous variables lacks practical interpretability. Third, the discrete “0/1” transitions of binary features can distort probability density estimation, leading to ambiguous clustering boundaries. All 93 selected features were incorporated in the personalized FL module (see Table 3). Missing values were addressed using tailored imputation methods: for discrete features, a zero placeholder was used, whereas continuous features were imputed using the random forest-based filling method. We also standardized the continuous data; these preprocessing steps ensured a robust and reliable foundation for subsequent analysis.
Feature used in modeling.

In this study, we selected the 20 hospitals with the largest patient cohorts as participants, encompassing a total of 64,974 ICU stays. These hospitals exhibited variations in sample size, AKI proportion, and patient characteristics, ensuring a diverse dataset for analysis. Each hospital's cohort was randomly split into training and testing datasets using an 80:20 ratio, maintaining a consistent approach across all participants.
Compared methods
We compare our framework with the following methods.
Local: each hospital only uses its local data to train model.
Centralized: it aggregates data from all participants together to train a global model.
FedAvg 53 : Federated Averaging algorithm is the most commonly used model aggregation algorithm in federated learning. The core idea is that after multiple participants locally train models, the parameters of these models are weighted and averaged to obtain a global model.
FedProx 54 : A well-known algorithm used to solve the heterogeneity of FL data. The core idea is to introduce the proximal term in the client optimization process, constrain the difference between the local update and the global model, alleviate the divergence of the client optimization direction, and improve the generalization ability of FL under heterogeneous data.
CBFL 16 : classic clustering-based FL algorithm. It uses the k-means algorithm to cluster patients based on the average feature vector of samples in each hospital and then trains FL models for each cluster separately. This method ignores heterogeneity among patients in each hospital.
K-FED 23 : improved algorithm of CBFL. Each hospital uses the k-means algorithm to cluster local patients. The server uses the k-means to cluster patients based on all centroid's information uploaded by each hospital and then trains FL models for each cluster separately. K-Fed overlooks the weight differences between centroids.
FedGMC: algorithm proposed in this study.
Parameter settings
In this study, we employed two widely adopted classifiers: the logistic regression (LR) model, known for its interpretability, and the Multi-Layer Perceptron (MLP) model, a common neural network architecture. Consistent classifier parameters were applied across different algorithms to ensure a fair comparison.
Model performance was evaluated using Recall and Area Under the Receiver Operating Characteristic Curve (AUC). Recall measures the model's ability to identify positive cases, making it particularly relevant for risk-sensitive scenarios such as EICU. While AUC, on the other hand, provides a comprehensive assessment of overall classification performance.
To compare various FL algorithms, key parameters such as the number of communication rounds, local training iterations, and batch size were standardized. Based on expert recommendations, the range for the number of clusters was set between 3 and 8.
Results
Cluster analysis
As shown in Figure 2, the clusters present in each hospital differ significantly, with notable variations in both the proportions and sample sizes of each cluster, even among hospitals with the same cluster types. This highlights the inherent heterogeneity among hospitals and among patient populations. Additionally, hospitals with larger patient cohorts tend to contain more cluster types, indicating that cluster centroids are more heavily influenced by these hospitals.

Distribution of patients among hospitals.
Taking the LR model as an example, we calculated the top-ranked features in each clustering model based on the absolute value of the regression coefficient 55 (see Table A.1). Tables in Figure 3 show the overlap rates of the Top5, Top10, and Top20 features of the clusters generated by different algorithms. It presents the feature overlap rates for all pairwise combinations (a total of 6 pairs) of the four clusters from cluster_0 to cluster_3 and also marks the overall average overlap rate. FedGMC exhibits values of 0.2, 0.35, and 0.35 for the top 5, 10, and 20 features, respectively. This is in contrast to CBFL, which shows average values of 0.3, 0.32, and 0.44 for the corresponding feature sets, and K-fed, with average values of 0.33, 0.40, and 0.42. A smaller overlap rate indicates a more significant difference between clusters. The differences between clusters in the FedGMC algorithm are more significant, demonstrating that it effectively captures heterogeneity with a better clustering performance. Table A.1 highlights the top 10 features of FedGMC that are unique to each cluster.
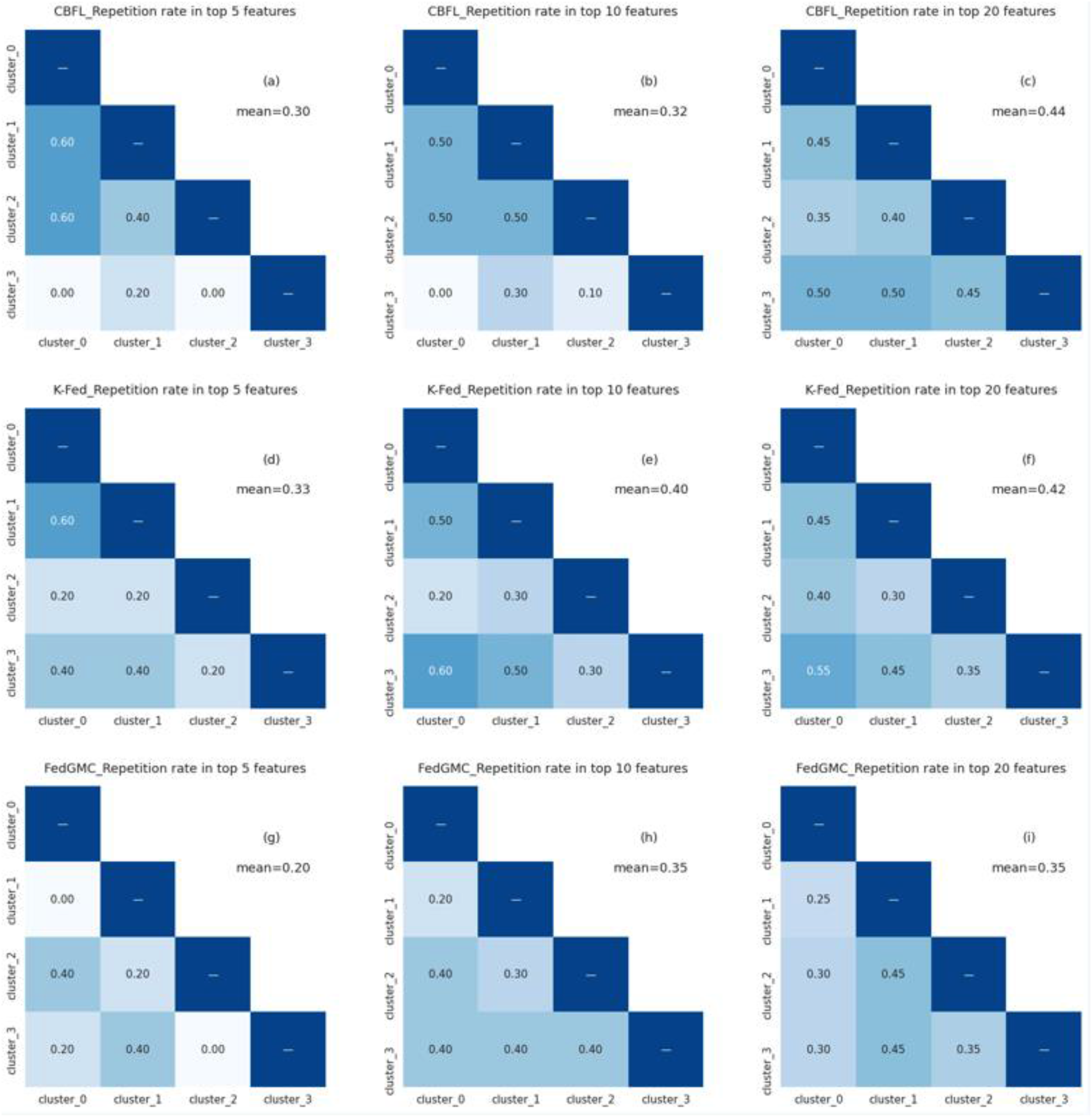
Comparison of inter-cluster repetition rates under different top features for CBFL, K-Fed, and FedGMC.
Predictive performance
We conducted 5-fold cross-validation to measure model performance. Table 4 presents the performance results of FedGMC and baseline algorithms. Considering the overall performance across the two classifiers and two performance metrics, FedGMC outperforms other baseline methods. Compared to FedAvg, CBFL, and K-Fed, FedGMC_LR exhibits improvements in mean Recall by 5.80%, 5.65%, and 2.06%, respectively, and in AUC by 2.20%, 1.03%, and 1.91%. Similarly, FedGMC_MLP demonstrates improvements in mean Recall of 4.30%, 3.77%, and 1.40%, and in AUC of 1.01%, 0.60%, and 1.63%. On the MLP classifier, although the AUC of FedProx leads FedGMC by 0.67%, it lags behind by 2.99% in terms of the Recall. On the LR classifier, the Recall and AUC metrics of FedProx lag behind those of the FedGMC by 7.17% and 2.05% even more significantly.
Predictive performance on 20 hospitals.

Figure 4 shows the number of hospitals that achieved optimal performance for each model. FedGMC_LR achieved the best performance in terms of Recall at 10 hospitals and AUC at 12 hospitals. FedGMC_MLP showed optimal performance in Recall in 12 hospitals and in AUC at 3 hospitals.

Number of hospitals achieving optimal performance for each model.
Discussion
The proposed personalized FL framework, FedGMC, which leverages a probabilistic modeling approach to overcome the limitations of existing methods. By utilizing sample distribution information, the server generates virtual samples to enhance the accuracy of federated patient clustering. Subsequently, a personalized FL model is trained for each cluster, leading to improved overall prediction performance.
Experimental results show that the FedGMC outperforms all baseline algorithms across both classifiers and performance indicators, achieving the best predictive method in most hospitals. We also observe from Table 4 that while FL improves the overall average prediction performance of participants, it does not necessarily improve that of the vast majority of participants. For example, with FedAvg_LR and CBFL_LR, only 45% of hospitals show in recall. FedGMC significantly reduces the likelihood of performance loss for participants in joint modeling. This has important implications for enhancing data owners’ willingness to participate and promoting the fairness of the algorithm.
Our study has several limitations that can be addressed in future studies. First, we used all continuous traits for clustering in our framework. Future research could explore whether selecting a subset of features would result in better clustering outcomes or be more aligned with clinical applications. Second, knowledge is not shared between clusters in our current framework. Future research could investigate whether transfer learning could be used to leverage useful information from other clusters. Third, the eICU dataset used in our study consists of hospitals exclusively from the United States, all participating in the Philips eICU program. This uniformity in data sources likely facilitated data standardization reduced the data statistical heterogeneity. 7 We suspect this is why, despite our optimization efforts, the performance of the MLP classifier still lags behind that of the LR model. Previous studies have also indicated that simpler FL algorithms, like FedAvg, may be more suitable for machine learning tasks on structured EHR data compared to more complex FL methods. We aim to validate our findings using additional datasets in the future. Fourth, the experiments show that the performance improvements of different FL algorithms for the participants vary considerably across participants, with some participants experiencing performance declines. Therefore, we recommend that further research focus more on ensuring the fairness of FL algorithms.
Conclusion
This study addresses the challenges posed by the heterogeneity of hospitals and patients in collaborative modeling. We propose a personalized FL framework FedGMC for the collaborative training of disease risk prediction models across medical institutions. This framework is designed to improve the performance of existing methods in complex datasets and scenarios. Experiments using EICU data show that FedGMC effectively captures heterogeneity. The personalized FL models generated by FedGMC not only outperform multiple baseline methods but also significantly reduce the likelihood of performance degradation among participants. The improvement is crucial for attracting more data owners to join the collaborative efforts. Beyond healthcare, the proposed is also applicable to federated clustering and joint modeling for data privacy protection in other domains, such as finance and recommendation systems. This study plays a critical role in overcoming data silos and unlocking the value of data.
Footnotes
Acknowledgements
We thank the members of the MIT Laboratory for Computational Physiology for allowing access to and use of the eICU database.
Ethical approval and consent to participate
The eICU database was accessed via the PhysioNet platform. Access to the database was approved after completing the Collaborative Institutional Training Initiative program “Data or Specimens Only Research” (certificate ID: 66813987), as well as signing the data usage agreement of the PhysioNet Review Board. The study was exempt from approval from the institutional review board of the Massachusetts Institute of Technology because of the retrospective design, lack of direct patient intervention, and the security schema, for which the re-identification risk was certified as meeting safe harbor standards by an independent privacy expert (Privacert) (Health Insurance Portability and Accountability Act Certification no. 1031219-2). The institutional review board of the Massachusetts Institute of Technology waived the need for informed consent for the same reason. The study was conducted following the Declaration of Helsinki. All methods used in this study were performed in accordance with the relevant guidelines and regulations.
Author contributions
Hong Ye: methodology, writing—original draft preparation, formal analysis, validation. Xiangzhou Zhang: project administration, writing—original draft preparation, data curation. Kang Liu: writing—reviewing and editing, methodology, formal analysis. Ziyuan Liu: validation, visualization. Weiqi Chen: software, data curation. Bo Liu: investigation. Eric W. T. Ngai: writing—reviewing and editing, conceptualization, supervision. Yong Hu: conceptualization, supervision, methodology, funding acquisition.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article. This work was supported by the Major Research Plan of the National Natural Science Foundation of China (Key Program, Grant No. 91746204), the National Natural Science Foundation of China (Grant No. 72371116), the Science and Technology Development in Guangdong Province (Major Projects of Advanced and Key Techniques Innovation, Grant No. 2017B030308008), and Guangdong Engineering Technology Research Center for Big Data Precision Healthcare (Grant No. 603141789047).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
