Abstract
Background
Infectious diseases, particularly COVID-19, continue to be a significant global health issue. Although many countries have reduced or stopped large-scale testing measures, the detection of such diseases remains a propriety.
Objective
This study aims to develop a novel, lightweight deep neural network for efficient, accurate, and cost-effective detection of COVID-19 using a nasal breathing audio data collected via smartphones.
Methodology
Nasal breathing audio from 128 patients diagnosed with the Omicron variant was collected. Mel-Frequency Cepstral Coefficients, a widely used feature in speech and sound analysis, were employed for extracting important characteristics from the audio signals. Additional feature selection was performed using random forest (RF) and principal component analysis (PCA) for dimensionality reduction. A Dense-ReLU-Dropout model was trained with K-fold cross-validation (K = 3), and performance metrics like accuracy, precision, recall, and F1-score were used to evaluate the model.
Results
The proposed model achieved 97% accuracy in detecting COVID-19 from nasal breathing sounds, outperforming state-of-the-art methods such as those by Lella and Alphonse and Abayomi-Alli et al. Our Dense-ReLU-Dropout model, using RF and PCA for feature selection, achieves high accuracy with greater computational efficiency compared to existing methods that require more complex models or larger datasets.
Conclusion
The findings suggest that the proposed method holds significant potential for clinical implementation, advancing smartphone-based diagnostics in infectious diseases. The Dense-ReLU-Dropout model, combined with innovative feature processing techniques, offers a promising approach for efficient and accurate COVID-19 detection, showcasing the capabilities of mobile device-based diagnostics
Introduction
The global outbreak of COVID-19 in 2019 has posed unprecedented challenges to public health systems, with emerging variants such as Omicron exacerbating the crisis due to heightened transmissibility and severe respiratory complications. 1 Despite advancements in diagnostic tools like RT-PCR and rapid antigen tests, widespread implementation remains hindered by cost, accessibility, and logistical barriers, particularly in resource-limited regions 2 This gap underscores the urgent need for non-invasive, scalable, and cost-effective diagnostic alternatives to curb transmission and enable timely interventions.
Recent advancements in digital health technologies have highlighted the potential of acoustic analysis for disease detection. Respiratory infections, including COVID-19, often alter vocal fold dynamics, breathing patterns, and sound production, offering a unique opportunity to leverage audio signals as diagnostic biomarkers. 3 Speech and cough analysis have been widely explored, with studies demonstrating the feasibility of machine learning models in detecting COVID-19 through vocalizations (e.g. vowel articulation, cough sounds). For instance, deep learning models analyzing cough recordings have achieved validation accuracies of 67–83%, 4 while vocal fold vibration analysis via vowel vocalization has yielded ∼80% accuracy. 5 However, these approaches often require complex speech tasks or extensive computational resources, limiting their practicality for real-world deployment. 6
A critical gap in existing research lies in the underutilization of nasal breathing sounds—a passive, non-invasive signal that directly reflects upper respiratory tract physiology. Unlike speech or cough sounds, nasal breathing is effortless, making it ideal for rapid screening in diverse populations, including asymptomatic individuals. 7 While studies have explored oral breathing and cough acoustics for COVID-19 detection,89–10 nasal breathing remains underexamined despite its clinical relevance. Existing methodologies also face challenges such as low accuracy (<85% in some studies 11 ), reliance on high-dimensional datasets, and insufficient feature optimization, which can introduce noise and overfitting. 12
To address these limitations, this study proposes a lightweight deep neural network (DNN) framework optimized for nasal breathing sound analysis. By integrating Mel-Frequency Cepstral Coefficients (MFCCs) with advanced feature selection techniques—random forest (RF) and principal component analysis (PCA)—we aim to reduce computational complexity while enhancing diagnostic accuracy.13,14 Our approach leverages smartphone-recorded nasal breathing sounds from 128 Omicron patients, focusing on key acoustic features such as fundamental frequency, sound pressure level (SPL), and MFCCs15,16 Through systematic dimensionality reduction and threefold cross-validation, we evaluate the robustness of our model against state-of-the-art methods, demonstrating its potential for scalable, real-world clinical applications.
The primary contributions of this work are threefold:
The remainder of this paper is organized as follows: Section Materials and methods details the dataset collection, preprocessing, and feature extraction methodologies. Section Statistical results describes the lightweight DNN architecture and experimental setup. Results and comparative analyses are presented in Section Experiment results, followed by a discussion of clinical implications and limitations in Section Discussion. Finally, Section Conclusion concludes the study and outlines future research directions.
Materials and methods
Experiment design
Figure 1 shows our research methodology, which is divided into four stages: data collection, feature extraction, model training, and prediction.

The general flow of the study consists of four parts: data collection, feature extraction, model training, and prediction.
Dataset
Data collection
The prospective observational study was conducted over 1 year, from March 2021 to February 2022, at Shanghai Sixth People's Hospital and Shanghai Key Laboratory of Multidimensional Information Processing, East China Normal University. Nasal breathing sounds were collected from 128 participants (67 COVID-19 positive, 61 healthy controls) using a smartphone application in a controlled clinical environment. All participants provided written informed consent, and the study protocol was approved by the institutional Ethics Committee of Shanghai Sixth People's Hospital (Approval No. 2022-KY-050(K)). These real-time respiratory samples were then analyzed for acoustic feature extraction and disease detection.
Participants were included if they met the following criteria:
Aged 18–65 years; Diagnosed with COVID-19 (Omicron variant), confirmed by PCR of rapid antigen test; Able to provide informed consent and participate in this study; Presenting symptoms of COVID-19, such as fever, cough, or difficulty breathing, at the time of data collection.
Participants were excluded if they met any of the following criteria:
History of severe respiratory conditions such as asthma, chronic obstructive pulmonary disease (COPD), or pneumonia; Pregnancy of breastfeeding; Any known neurological disorders or hearing impairments that could affect the ability to participate in the study; Use of medications that could affect respiratory function (e.g. sedatives or narcotics)
The dataset comprises standard audio recordings of nasal breathing sounds from 67 patients diagnosed with neoplastic disease, collected from the Shanghai Sixth People's Hospital. Each recording has an average duration of approximately six seconds. For the control group, nasal breath sounds were collected from 61 adults who tested negative for neoplastic pneumonia using the same recording procedure. Written consent was obtained from all participants, and the study was reviewed and approved by an ethics committee. Unlike other nasal breathing datasets, such as the COVID-19 dataset from Sonde Health (2020), no category in our dataset contains more than 25 individuals. This dataset provides crucial insights and supports research into the association between neoplastic disease and COVID-19 through nasal breathing sound characteristics. Additionally, it aids in the development of methods for early COVID-19 detection (see Table 1 for detail).
Number of patients enrolled in our study.

Data preprocessing
The raw nasal breathing audio signals collected via smartphones are inherently one-dimensional (1D) time-series data, representing variations in SPLs over time. However, convolutional neural networks (CNNs) are traditionally designed to process two-dimensional (2D) data, such as images, where spatial hierarchies and local patterns are critical for feature extraction. To leverage the powerful pattern recognition capabilities of CNNs, we transformed the 1D audio signals into a 2D format.
This approach allows the CNN to capture local features in the data, which are essential for distinguishing COVID-19-related acoustic features from healthy controls. By reshaping the 1D audio data into 2D formats, we enable the CNN to apply convolutional filters across both time and frequency dimensions, enhancing its ability to detect subtle associated with respiratory abnormalities.
Statistical analysis
Feature selection
This study employed nine key acoustic features to provide a comprehensive analysis of the audio signal to achieve accurate results. These characteristics include voiced and unvoiced sounds, effective speech segments, fundamental frequency (F0), log energy, short-term energy, zero crossing rate, SPL and MFCCs (see Figure 2), with detailed explanations below:17,18

Mel-scale frequency cepstral coefficients map for nasal breath sound in tested adults for COVID-19 (left: positive; right: negative).
Each acoustic feature provides a unique and reliable perspective on voice signal analysis. In this study, these features were used for audio data analysis, offering comprehensive acoustic insights crucial for subsequent model training. The analysis revealed significant variations in the acoustic properties of nasal breathing between individuals with different COVID-19 test results.
Figure 2 demonstrates that the MFCC plots of COVID-19 positive patients exhibit significantly darker tones, indicating a lower frequency component in their nasal breathing sounds compared to healthy individuals. This variation may be linked to the impact of COVID-19 infection on the sound produced.
Figure 3 shows that the fundamental frequency (f0) curves of COVID-19 patients are significantly lower than those of healthy individuals, suggesting that their nasal breathing sounds are relatively low pitched. The stable frequency observed on the right-hand side, with no lower frequency in that period, supports the notion that 2k can be the fundamental frequency.
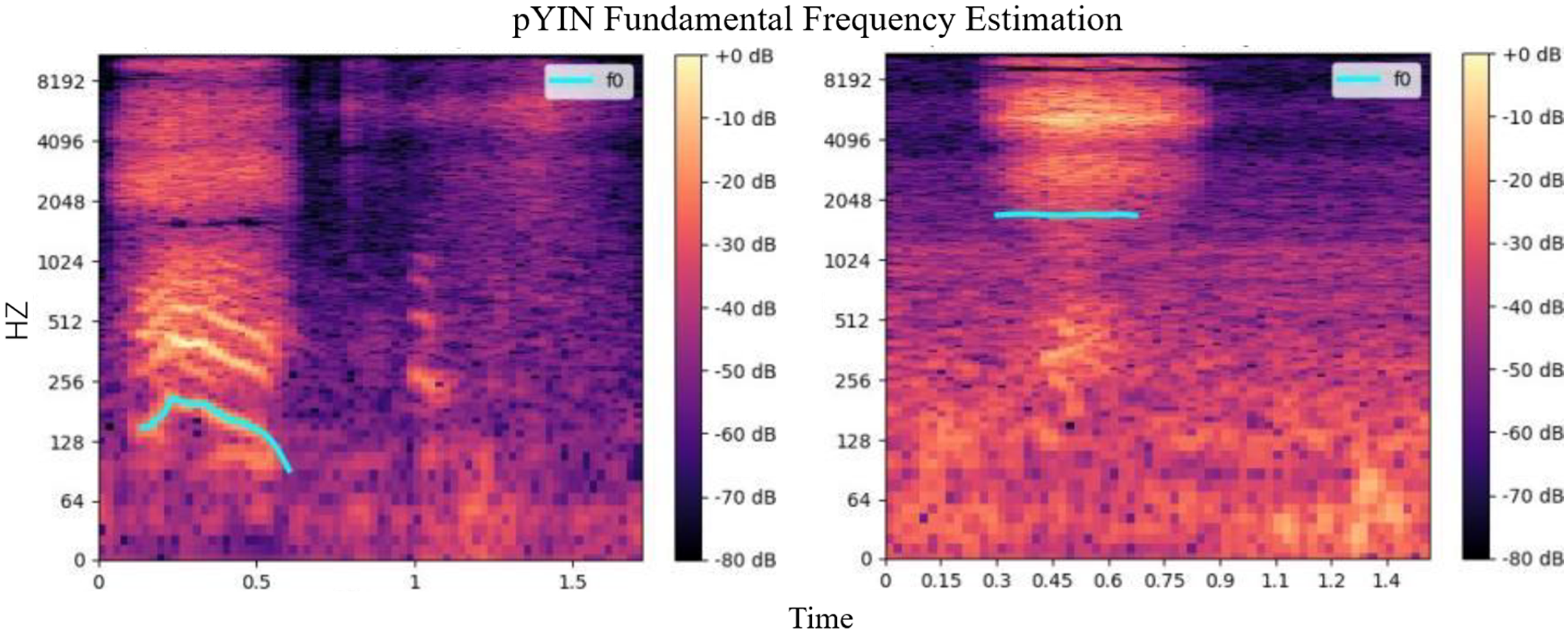
Fundamental frequency curve for nasal breath sound in adults who tested for COVID-19 (left: positive; right: negative).
We extract statistical information such as extreme values, mean values, standard deviations, peak values, and skewness from a range of acoustic features. These metrics collectively provide a comprehensive quantification of the key characteristics of each feature. This process offers an analysis that is more profound and informative than that provided by a 2D image. Detailed statistical metrics selected can be seen in the Table 2:
Selected statistical metrics in this study.

We extract statistical information such as extreme values, mean values, standard variances, and skewness from a range of acoustic features. These metrics collectively provide a comprehensive quantification of the key characteristics of each feature. This process offers an analysis that is more profound and informative than that provided by a 2D image. The extreme value outlines the sound feature's upper and lower boundaries. The mean value illustrates its central tendency. The standard variance is indicative of the feature's variability. The peak value points to the maximum sound intensity. The skewness measures the skewness of the distribution. The process of condensing data from 2D images into these vital statistical metrics helps minimizes redundant information. This step contributes significantly to making the model more streamlined.
Feature dimension reduction
In this subsection, we will introduce the methods we applied for feature dimension reduction. The model input the more redundant information involved.192021–22
In traditional studies of COVID-19 detection utilizing acoustic features, the challenge of low accuracy often arises. To counter this, we explore critical comparisons of these acoustic features and employ a strategy designed to reduce model complexity, enhancing operational efficiency. Consider data from 2D images into these vital statistical metrics helps minimize redundant information and significantly contributes to making the model more streamlined.
In recent years, dimension reduction techniques have evolved significantly, with several new methods offering promising improvements over traditional approaches. In this study, we employed PCA and RF for the feature selection and dimensionality reduction. While newer methods such as t-Distributed Stochastic Neighbor Embedding (t-SNE) and autoencoders have gained popularity in the field, recent studies still demonstrated the effectiveness of PCA in acoustic and sound-based analysis. For example, Wang et al. 23 show that PDCA remains a reliable method for extracting relevant features from high-dimensional audio data, especially when computational resources are constrained. Similarly, Chen et al. 24 found that RF-based feature selection, when combined with PCA, continues to yield high performance in diagnostic tasks using sound data, making these traditional techniques a strong choice for our study.
To maintain model efficiency, we adopted strategies including RF25,26 and PCA2728–29 for ranking and prioritizing the extracted features.
RF ranks and prioritizes features based on their importance scores, where the importance score for a feature 
where
PCA reduces dimensionality by transforming the original features into a new set of uncorrelated features (principal components) that maximize variance. The principal components are given by:
Beyond the implementation of RF and PCA, we explored the correlation analysis between input features and labels. We selected the top eight features exhibiting the highest correlation coefficients to form a new subset of features. The correlation coefficient 
This approach ensures that the chosen attributes have a significant relationship with the target label, enhancing our model's diagnostic accuracy.
Network structure
To achieve a high-accuracy lightweight model suitable for rapid disease diagnosis on mobile devices, we developed architectures based on CNNs and DNNs. The input data specifications for both architectures are determined by the chosen feature extraction method, with all features used as input, as illustrated in Figure 4.
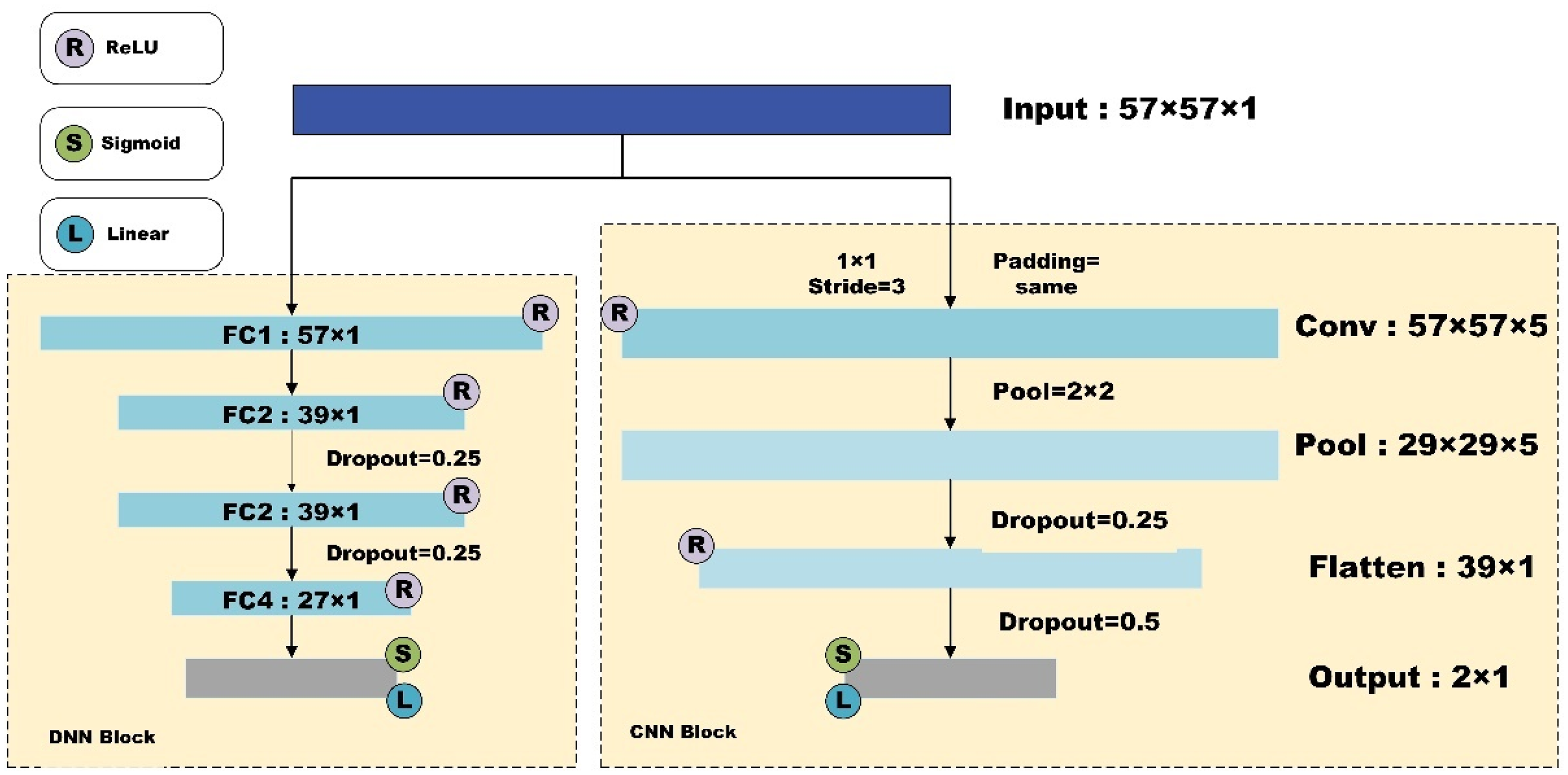
Network architecture of our study.
The CNN architecture consists of a single convolutional layer with a kernel size of
The DNN architecture includes four fully connected layers. Nonlinear transformations between each layer are performed using the ReLU activation function, and dropout rates of 0.25 are applied between the second, third, and fourth fully connected layers to exclude some data. The final classification results are obtained using a sigmoid activation function in the linear layer.
Evaluation metrics
During this process, we did not use an additional split to create a separate validation set. Instead, one of the three parts of the dataset was designated as the validation set in each fold. This ensured that every data point was used both for training and validation, and helped provide a more robust evaluation of the model's performance while avoiding any data leakage.
Before each fold in the threefold cross-validation process, we first randomly shuffle the dataset to prevent potential distribution bias. For each fold, we ensure that there is no overlap between the training and test sets at the patient ID level to avoid data leakage. Additionally, PCA dimensionality reduction and RF feature selection is performed separately within each training set of each fold, preventing test set information from leaking into the feature engineering process and ensuring the independence of model evaluation.
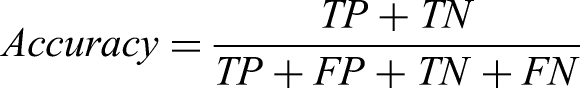



Statistical results
Feature selection results
Using the selected statistical metrics, and employing the feature selection techniques, Table 3 demonstrates the feature extracted by RF (23 features) and PCA (27 features), respectively.
Results of feature extraction of random forest and principal component analysis.

MFCC: Mel-Frequency Cepstral Coefficient; PCA; principal component analysis.
The results of top 8 correlated features and corresponding correlation coefficients can be seen in the Figure 5. In the following study, we will utilize these to train lightweight deep learning-based model. The reduced dimension of the features can be seen in Table 4.
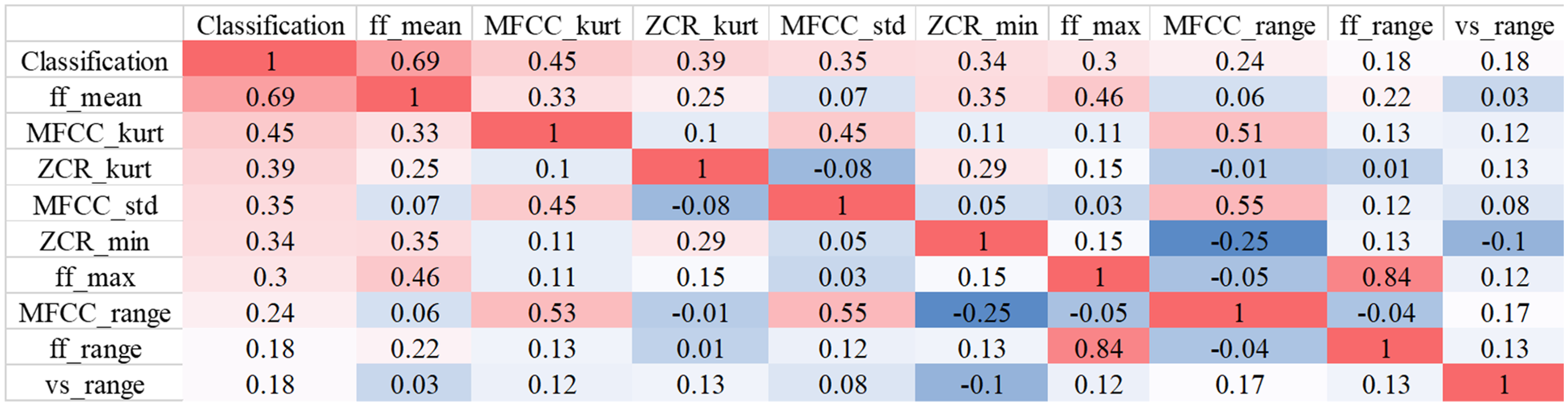
Heatmap of top eight correlated coefficients from correlation analysis.
Feature dimension reduction results. The values represent the dimension of features after reduction.

MFCC: Mel-Frequency Cepstral Coefficient.
By leveraging these highly correlated features, we construct more robust models for effective COVID-19 detection. This strategy underscores the importance of the feature selection process in building precise and efficient diagnostic models.
Despite the rise of newer dimension reduction methods, we opted for PCA and RF because of their simplicity, efficiency, and strong performance in our dataset, which consists of nasal breath sound recordings. Moreover, the trade-off between newer techniques and the computational cost was a critical factor in our choice. As demonstrated in recent literature, PCA remains effective in preserving the most important features while significantly reducing dimensionality, thereby improving model training efficiency without sacrificing accuracy.
Experiment results
The experimental results (Tables 5 and 6) demonstrated that the DNNs model achieved the highest accuracy of 97.67% on the full feature dataset. In comparison, the initial accuracy of the CNN model on the same dataset was 76%. Notably, the accuracy of the CNN model improved by nearly 10% after applying feature filtering techniques using RF and PCA. However, when the CNN model was applied to a dataset refined by correlation selection, its performance dropped by approximately 16% compared to the full feature dataset.
ROC accuracy for different models with different feature selection methods and results.

RF: random forest; PCA: principal component analysis; DNN: deep neural network; CNN: convolutional neural network.
F1 score for different models with different feature selection methods and results.

RF: random forest; PCA: principal component analysis; DNN: deep neural network; CNN: convolutional neural network.
In contrast, the DNN model's accuracy only slightly decreased by about 1% despite the reduction of nearly 30 features through RF and PCA feature filtering. The corresponding F1 score reduction was about 5%. Even with nearly 50 features eliminated from the relevance-selected dataset, the DNN model still achieved an accuracy of 88% and an F1 score of 86%. This indicates that the DNN model maintained high performance across different feature subsets, demonstrating greater robustness and adaptability in managing acoustic features for COVID-19 diagnosis.
It is important to note that training models on the full feature dataset can introduce noise. This noise increases the likelihood of overfitting, where the model performs well on the training data but poorly on unseen data. The feature filtering techniques, such as RF and PCA, help mitigate this issue by removing irrelevant features, thus enhancing the model's generalizability and reducing overfitting risks. The experimental results confirm that the DNN model is more resilient to feature subset variations, maintaining high effectiveness and reliability for COVID-19 diagnosis.
Note: Due to computational constraints and dataset limitations, visualizations of the cross-validation and performance metrics were not included in this manuscript. However, we recognize the importance of such visualizations for enhancing the clarity of model performance and plan to incorporate them in future versions of this work.
Discussion
In this study, we introduced a lightweight classification model for COVID-19 detection using nasal breathing sounds, aiming to contribute to the growing field of non-invasive diagnostics. The key accomplishments of our study, the novel contributions, and the limitations are discussed in the following texts.
Our research successfully developed a highly accurate and efficient method for detecting COVID-19 through nasal breathing sounds recorded on smartphones. The classification model achieved an impressive accuracy rate of 97% and an F1 score of 98% when utilizing the full feature set. By employing DNN, our model consistently outperformed CNN, particularly with datasets containing comprehensive audio features. These results suggest the robustness and effectiveness of DNN in handling complex audio signal features, making it a viable tool for disease detection in real world.
The inclusion of patients with neoplastic disease in this study introduces potential biases, as their nasal breathing sound characteristics may differ from those of the general population, particularly in individuals without underlying health conditions. This could affect the model's ability to generalize across diverse populations. In future work, we plan to explore this potential bias more thoroughly and consider methods to mitigate its impact, such as testing the model on separate datasets or applying stratified sampling techniques.
A key innovation of our study lies in the use and processing of nasal breathing sounds as input data. Unlike general breath sounds, nasal breathing sounds are more reflective of the nasal cavity, nasopharynx, and upper airway characteristics, which are crucial in detecting COVID-19-related respiratory symptoms. This makes nasal breathing sounds a more targeted and informative source of data compared to broader, less specific breath sounds. Furthermore, our approach to feature selection focused on using nine key acoustic features, with a particular emphasis on voiced sounds, which are essential for capturing the dynamics of respiratory health and disease-related changes.
The method of dimensionality reduction using RF and PCA enhanced the representativeness of the audio features, thus improving model performance. Although we employed PCA and RF for feature dimensionality reduction and selection, dataset bias may still exist, such as the potential omission of certain high-dimensional features. Future research could further optimize this process by incorporating additional data augmentation strategies or integrating multiple dimensionality reduction techniques, such as t-SNE or Autoencoder.
The reduction in feature set from 57 to 23 and ultimately to 8 features resulted in the maintenance of high accuracy and F1 scores, demonstrating the effectiveness of both feature selection and dimensionality reduction strategies. These results align with recent studies on acoustic signal analysis, which highlight the importance of selecting relevant features and reducing dimensionality for improving the efficiency of machine learning models in health diagnostics.3031–32
We observed that dimensionality reduction improved the performance of the CNN, enhancing its ability to classify COVID-19 positive and negative samples. However, for the DNN, performance was actually best when the full feature set was used without any dimensionality reduction. This suggests that while dimensionality reduction is useful for some models, it may not be beneficial for others, such as the DNN, where reducing the number of features resulted in a slight drop in accuracy.
Despite the promising results, our study has certain limitations that warrant further investigation. Firstly, while our study incorporated basic noise reduction techniques, more sophisticated methods are needed to address the noise interference commonly present in acoustic recordings, particularly in uncontrolled environments. Secondly, although our dataset contains 128 samples, expanding it to include more diverse and temporally varied samples would significantly improve the model's generalizability and robustness across different populations, environments, and COVID-19 variants. Thirdly, while we focused primarily on feature extraction and classification, future research should explore other machine learning models, ensemble methods, and hybrid architectures to enhance reliability and accuracy further. Finally, the real-world clinical application of our model requires validation through prospective studies to ensure its feasibility and utility in front-line healthcare settings, where the model could aid in rapid COVID-19 detection and assist healthcare professionals in decision-making.
Beyond COVID-19, the potential of our methodology extends to diagnosing other respiratory diseases such as influenza, bronchitis, and COPD. Similar to COVID-19, these diseases can cause characteristic changes in nasal breathing sounds and vocal fold vibrations, which can be captured and analyzed using similar acoustic features and machine learning models. Notably, studies such as 24 have demonstrated the utility of sound-based diagnostic methods in identifying anomalies related to respiratory diseases, showing how such models can provide early diagnosis and continuous monitoring in real-world applications. Future research should investigate the applicability of our approach across a broader spectrum of respiratory conditions, enhancing early detection, patient management, and the use of non-invasive, cost-effective methods in diverse healthcare settings. Integrating our approach with wearable devices or smartphone applications can enable continuous, real-time monitoring, significantly improving healthcare accessibility, especially in low-resource settings.
Conclusion
Our study highlights the potential of nasal breathing sounds as a reliable, non-invasive diagnostic tool for detecting diseases like COVID-19 using a novel lightweight Dense-ReLU-Dropout model. By integrating advanced feature selection techniques such as RF and PCA, we achieved 97% accuracy and a 98% F1 score, demonstrating the feasibility of smartphone-based rapid disease detection. This approach has broader applicability to other respiratory illnesses and mobile health technologies, offering a cost-effective solution for real-time diagnostics. Future work should focus on expanding the dataset through multi-center collaborations to enhance generalizability, validating the model across diverse clinical settings, and addressing challenges such as noise interference in real-world environments to ensure robust performance and seamless clinical integration.
Footnotes
Acknowledgements
Thanks to Hailin Ma, Han Lu, Jiayi Guo, and Rui Su for their support during this study. The authors wish to thank the anonymous referees for their thoughtful comments, which helped in the improvement of the presentation.
ORCID iDs
Ethical considerations
This study was approved by the Ethics Committee of Shanghai Sixth People's Hospital Affiliated to Shanghai Jiaotong University School of Medicine (Approval Number: 2022-KY-050(K)) with the informed consent of all participants.
Author contributions
JS contributed to methodology, experiment, software, and writing—review and editing. PL and RL contributed to software, visualization, writing—original draft, review and editing. SL contributed to software and writing—review. LS contributed to software, validation, data curation, and project management. LG contributed to conceptualization, formal analysis, data curation, and visualization. ZT, RS, and ZY contributed to project administration and writing—review and editing. JC, LF, and YJ contributed to inspiration, conceptualization, writing—review and editing, and supervision.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Clinical Research Plan of SHDC, Scientific Development funds for Local Region from the Chinese Government in 2023, Shanghai Acupuncture and Moxibustion Clinical Medical Research Center, Science and Technology Commission of Shanghai Municipality, the Key Research & Development Project of Zhejiang Province, Jilin Province science and technology development plan project, National Natural Science Foundation of China, Shanghai Association of Traditional Chinese Medicine Program, Shanghai Committee of Science and Technology, China, 2022 ”Chunhui Plan” cooperative scientific research project of the Ministry of Education, (grant number SHDC2020CR4057, 16CR2026B, XZ202301YD0032C, 20MC1920500, 22DZ2229004, 22JC1403603, 21Y11902500, 2024C03240, 20230204094YY, 82151318,82004446, 2023-HPZY-04, (23Y11920900D, 22Y11923000, 20Y21901100).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The code and data used in this study are available upon reasonable request. Interested researchers may contact the corresponding author to obtain access.
