Abstract
Objective
Integrating IoT technologies into the healthcare system has significantly raised the prospects for patient monitoring and disease prediction. However, the present-day models have failed to effectively encompass spatial-temporal data samples.
Methods
This paper presents a novel hybrid machine-learning model by amalgamating Convolutional Neural Networks (CNNs) with Long Short-Term Memory models (LSTMs) to boost prediction accuracy. Whereas the CNNs extract spatial features from medical images, the LSTMs model the temporal patterns of wearable sensor data. Such a configuration increases the prediction accuracy by 10% more than that achieved by the individual models. For better feature extraction, the proposed method implements Physiological Event Extraction (PEE), which is aimed at identifying important physiological events such as heart rate variability and respiratory changes from raw sensor data samples.
Results
This method helps render the features interpretable, providing another 15% improvement in prediction performance. Anomaly detection employed ensemble techniques that combined the Isolation Forest and One-Class SVM, reducing false positives by 20%, thus outperforming conventional approaches. It further enhanced the True Positive Rate (TPR) by 25% through using an online learning algorithm with Incremental Gradient Descent with Momentums. Robust statistical methods based on M-estimator theory had been integrated for the treatment of outliers and missing data, which helped in reducing bias in estimation by 30% and increasing the False Positive Rate (FPR) by 12%.
Conclusion
All these enhancements constitute a major step towards improving the IoT healthcare data processing chain, thereby providing a trusted and accurate system for real-time health monitoring and anomaly detection. In this regard, the research also paves the way for designing next-gen IoT healthcare analytics and their actual clinical applications.
Keywords
Introduction
Integration of IoT technologies in healthcare has made it even brighter in terms of monitoring a patient continuously and detecting diseases early. Current models fail in processing multimodal health data containing elements including spatial information from medical imaging as well as temporal signals from physiological sensors. Past research on the kind of deep learning techniques included those by employing Convolutional Neural Networks (CNNs) for spatial feature extraction and Long Short-Term Memory (LSTMs) for sequential data analysis. However, using either technique alone has proven insufficient to capture the whole complexity of the patient’s health status.1,2 Recently, hybrid models combining both CNNs and LSTMs have been studied under multimodal health data processing and have shown improvements in terms of prediction accuracy. 3 Unfortunately, these models inherently lack support by domain-specific feature extraction, making them less applicable in practice. PEE has emerged as a promising technology through which significant interpretability has been accomplished using physiological markers, including heart rate variability (HRV) and changes in respiratory rates. 4 Unfortunately, IoT healthcare data tends to be high dimensional, typically making anomaly detection a difficulty. Common techniques, like Isolation Forests and One-Class SVMs, are useful in detecting anomalies, but they suffer from the problem of producing too many false positives. 5 Accordingly, this study proposes an integrated CNN-LSTM model with PEE for improved feature extraction and an ensemble anomaly detection framework for enhanced reliability, besides using the Incremental Gradient Descent with Momentum approach for new stream adaptation in real time. In a nutshell, it provides a strong, interpretable, and scalable method to address gaps and limitations within existing models, ultimately toward health continuous monitoring using IoT-based healthcare environments.
The rapid growth of IoT-enabled health technologies has transformed how patients are monitored remotely, allowing for continuous capture of physiological information through medical imaging or by the use of wearable sensors. However, apart from the instantaneous measurement of different variables at different times, the digital integration challenge of heterogeneous health data lies in the fact that most models isolate medical images and physiological signals because they are spatial and temporal data, respectively. One of the latest examples of research in this aspect is concerning hybrid deep learning architecture, especially when exploring the theme of multi-modal health data processing using CNNs and LSTMs.6,7 Indeed, while CNNs benefit from better spatial feature extraction from radiographic imaging, LSTMs are well suited for time-series pattern recognition for ECG signals. However, none of these models advances adaptive learning mechanisms and domain-specific feature engineering to the extent that they can even be close to real deployment in the average clinical environment in real-time conditions.
It improved how hybridization works by innovating on three fronts: (1) adding a Physiological Event Extraction (PEE) module that isolates critical physiological changes (such as heart rate variability and respiratory rate), thus improving the interpretability of features; (2) introducing a new ensemble-based anomaly detection system that combines Isolation Forests and One-Class SVMs to increase reliability in high-dimensional IoT healthcare datasets; and (3) incorporating a framework of Incremental Gradient Descent with Momentum to create continuous learning for real-time adaptation of models. It thereby improves previous models by enhancing the robustness of features, reducing false alarms, and scalability for dynamic health monitoring applications.
While the previous approaches discussed were lacking in the serious matters concerning feature extraction and learning frameworks in the domain, this research has uniquely integrated PEE for feature interpretability, an ensemble mechanism for anomaly detection, and an enhanced real-time adaptive incremental learning approach. Respecting one clear distinctive feature, this framework ensures continuous improvement in both classification accuracies and anomaly detection rather than static retraining commonly employed in conventional deep learning models. On top of that, the research addresses efficiency and scalability, thus rendering it a good candidate for deployment in real-life settings.
The justification for creating this model was founded on previously known challenges to IoT-based healthcare, which are:
Insufficient integration of multimodalities: The existing models address medical imaging and physiological sensor data separately and therefore provide an incomplete evaluation of the patient. Elevated false positives for anomaly detection: Conventional SVMs and threshold-based detection mechanisms fail to handle issues arising from high-dimensional, sparse IoT data, which results in excessive false alarms. Non-adaptability in real-time: Deep learning-based health monitoring models, with a few exceptions, need to be periodically retrained, thereby making them unsuitable for use in a dynamic clinical setup process. Heavy computational needs: CNN-LSTM models are highly powerful but require heavy computational resources making real-time inference on the device challenging process.
The increasing adoption of Internet of Medical Things (IoMT) technologies in IoT healthcare solutions permits continuous and real-time health monitoring. Unlike episodic clinical assessments, IoMT-based monitoring is able to identify early physiological abnormalities through wearable biosensors and remote diagnostic equipment. This eliminates hospital readmission rates significantly and enhances patient outcomes. In addition, IoMT communications facilitate increased interoperability among medical devices, enabling seamless integration of electronic health records (EHRs) with real-time patient monitoring data. The integration offers incentives for automated clinical decision-making whereby healthcare providers can act at any moment they find a patient critical. The model in question employs IoMT innovations for an AI-driven and autonomous health monitoring system for continuous patient assessment and risk stratification.
Some of today's machine learning techniques used to monitor health over IoT are beset with major drawbacks:
Q-learning and reinforcement learning: They need extensive exploration and their extension to real-time physiological signals when there is a need for real-time predictions might not be so easy to process. Meta-heuristic algorithms: Although well suited for optimization problems, meta-heuristics are mostly not interpretable, and therefore not very well suited for clinical decision-making process. Brain-inspired neural architectures: These are in a highly promising condition but are computationally intensive, making it hard to deploy them on edge devices in IoT-based healthcare sets.
The rest of the paper is structured as follows. Literature review section gives information on the proposed architecture of the CNN-LSTM model and the PEE methodology. Experimental setups, dataset selection, and evaluation metrics are located in section ‘Proposed design of an iterative method for enhanced health monitoring using hybrid deep learning with PEE’. Section ‘Result analysis & comparison with existing techniques’ contains results and statistical results, highlighting accuracy of classification, anomaly detection, and model robustness improvements. Lastly, Section ‘Result analysis & comparison with existing techniques’ addresses extended implications, limitations, and future work, thereby providing a thorough understanding of the research contribution. Mathematical operations characterizing CNN-based feature extraction, LSTM sequence learning, and anomaly detection models primarily build on preexisting concepts of deep learning and statistical modeling.
Broader implication of this work process
The proposed hybrid deep learning framework offers important contributions to the real-life functioning of health applications in the domain of IoT, especially for rural healthcare and emergency monitoring systems. An autonomous and AI-assisted health-monitoring system can then help bridge the gap by providing real-time diagnostics and early detection of anomalies on wearable sensor information whenever specialist care is severely limited in remote areas. Integration with mobile health (mHealth) applications could provide an automated alerting system to healthcare workers for timely interventions in emergency scenarios for affected members, such as in cases of cardiac arrhythmias or respiratory distress. Furthermore, the constant learning framework will keep its predictions relevant and adaptive, thereby increasing its scope of feasibility in ever-changing medical environments such as Intensive Care Units (ICUs) and emergency response systems.
But a number of restrictions remain to be realized. Deep learning model training, including hybrid models like CNN-LSTM, is a computationally intensive process, hence creating some constraints for implementation on lower-power IoT devices. Optimizations in the future should then take into account sophisticated model compression, pruning, and quantization techniques for efficiency. Additionally, the real-time streaming characteristics of the physiological information are accompanied by issues of privacy; therefore, some encryption processes and federated learning processes would be necessary for model training without divulging any confidential patient data. Overcoming these challenges will be essential for mass adoption in IoT-enabled healthcare ecosystems. For more improvement in performance, in the future the aim should look to investigate a few transfer learning techniques, employing some pre-trained medical image networks (e.g. ResNet, DenseNet) for more realistic feature extraction from chest X-rays.
Literature review
Health Monitoring (HM) has witnessed significant advancements in recent years, driven by innovations in sensor technology, data analytics, and computational techniques. This section encompasses diverse methodologies and applications in health monitoring across various domains, including structural engineering, healthcare, and IoT-enabled systems. The literature review begins with an exploration of vibration-based techniques, 8 highlighting the importance of SHM in maintaining the structural integrity of steel slit dampers. The study identifies various SHM techniques enhancing SSD maintenance, emphasizing the need for continuous monitoring to prevent structural failures. Subsequent papers delve into optimization methods, 9 Dew computing, 10 and deep learning approaches,11,12 showcasing the effectiveness of advanced algorithms in processing large datasets for health monitoring and diagnostics.
Emerging paradigms such as federated learning, 13 integrated circuit design, 14 and wearable sensor technologies15,16 are explored for personalized and remote health monitoring, offering non-intrusive monitoring solutions for diverse applications. Moreover, machine learning techniques,17,18 contactless sensing technologies, 19 and digital health innovations20,21 demonstrate the potential of data-driven approaches in disease management and early symptom detection. As per Table 1, this review also addresses challenges such as scalability, 5 reliability, 25 and data privacy, 26 underscoring the importance of addressing these concerns for widespread adoption of health monitoring technologies. Additionally, the integration of novel sensing modalities,27,28 computational modeling,29–32 and fog computing 33 highlights the interdisciplinary nature of health monitoring research, aiming to improve system reliability and real-time analytics. The literature review offers a comprehensive insight into the advancements, methodologies, findings, and limitations in health monitoring technologies across various domains. Analyzing the key themes and trends discussed in the reviewed papers reveals the evolving landscape of health monitoring, driven by innovations in sensor technology, data analytics, and computational techniques. The review begins with an exploration of vibration-based techniques, 8 emphasizing their significance in structural health monitoring (SHM) for maintaining the structural integrity of critical infrastructure such as steel slit dampers. By identifying various SHM techniques enhancing SSD maintenance, the study underscores the importance of continuous monitoring in preventing structural failures, thereby enhancing safety and reliability.
Summary of relevant literature on health monitoring techniques.

The Internet of Things (IoT) is integrated with health systems for improvement in continuous health monitoring and predictive analytics. Most of the research done has been in two lines: spatial data such as medical imaging and temporal data in the form of physiological sensor output. Most studies are separate handling of these data modalities, which in itself can be a limitation when aiming to capture complex interactions of interest for predicting the precise health status. This section covers some of the most important methodologies relevant to this work, especially those integrating machine learning and deep learning approaches in processing IoT-generated health data samples. Convolutional Neural Networks (CNNs) and Long Short-Term Memory (LSTM) Networks CNNs have proven to be very effective at extracting spatial features from medical images, such as chest X-ray classification and MRI analysis. However, CNNs do not have temporal modeling for the analysis of time-series data from wearable sensors. In contrast, LSTM networks are good at sequential pattern capturing in time-series data but lack the spatial interpretative power of CNNs. The previous works, which applied CNN or LSTM architecture in isolation, were only able to moderately succeed in the processing of single-modality data but were not able to fully exploit the multi-modal nature of healthcare data samples. Recent studies in this area focus on hybrid CNN-LSTM approaches where CNNs automatically extract spatial features, but LSTMs learn time dynamics, especially when hybrid integration of image and sensor data is required for certain tasks. Thus, these approaches validate that hybrid architectures are highly viable but limited to adaptivity and generalization within models in real-time clinical contexts. Anomaly Detection with High-Dimensional IoT Data It is an important factor in the medical area for healthcare to identify the outliers that might be signifying anomalies. Anomalies, conventional techniques are used such as Isolation Forests and One-Class SVMs in anomaly detection. In isolation, one can identify the rare event anomalies within a fewer number of steps, but establishing decision boundaries for normal data filters out the outliers with the help of One-Class SVMs. While these are progressions, current techniques suffer from high false-positive rates because IoT data is generally high-dimensional and very sparse, with most of these detection errors. Continuous Learning for Online Adaptation to Real-Time Data Incremental Gradient Descent with Momentum has been introduced for continuous learning. This would enable the models to adapt to new samples of data online, and in healthcare monitoring, patient data evolves rapidly. Although training is recurrent in the old models of learning, which is computationally costly and not feasible in real environments of dynamic health data, previous research on online algorithms about learning suggests the feasibility of real-time updates but was not robustly applicable to healthcare settings that require great accuracy and adaptability. Contributions and Innovation of This Work It will introduce an overall hybrid model for CNN-LSTM, achieving a better prediction with a 10% improved accuracy than with single-modality models. The innovations of this are the core: PEE, intended to enhance the interpretability of features using critical events. So, the performance showed a 15% increase in predictions. Further, integration of Isolation Forests with One-Class SVMs produces an anomaly detection mechanism that holds the least false positives at a reduction of 20% from the current approaches and can be relied upon with the anomaly patterns specific to the high-dimensional data samples of IoT. The proposed continuous learning framework also utilizes Incremental Gradient Descent to make real-time adaptation possible and represents an error in prediction as the evolution of new data streams depicts a 25% error decrease. This literature review will establish that there is a need for integrated multi-modal health monitoring solutions and highlight limitations in current methodologies. The proposed model addresses specific shortcomings that can be overcome through an adaptable, accurate, and robust framework for IoT-based healthcare monitoring.
Development in health monitoring has been dramatically advanced with new developments concerning sensor technology, analytics in data, and multiple computational models. Consequently, more and more devices feature the Internet of Things and health monitoring is done continuously on an interrupted basis as a major method for instant check-up of the health status meant for preventing the occurrence. However, the problem statements remain unsolved by the following areas: to integrate spatial and temporal data precisely, process high dimensional data efficiently, and develop strong anomaly detection techniques. Some recent methodologies in HM about the integration of advanced sensor technologies with feature extraction techniques along with some known limitations of our proposed model have been discussed in this section. HM has used deep architectures in the form of CNNs and LSTM networks. The networks were used here because they can extract highly dimensional spatial and temporal features from samples. CNN is successful in image processing and can be used in identifying disease markers in radiographic images through its feature extraction capability in spatial domains. On the other hand, LSTM networks are suitable for time-series data, and sequential physiological signals such as electrocardiograms can be processed to detect anomalies, according to existing studies, there are limitations when these models are used in isolation since they fail to capture interdependencies between spatial and temporal health data, which is crucial for comprehensive patient monitoring.
Hybrid CNN-LSTM models have been proposed to overcome these limitations: CNNs for spatial feature extraction and LSTMs for the recognition of temporal patterns. However, integration has improved performance but challenges the problem of data interpretability and anomaly detection in traditional CNN-LSTM models with generic feature extraction techniques. The inability to extract features in concerned domains, such as physiological event detection, limits their application in the real health sector. In this domain of health care, with slight physiological changes, it represents crucial diagnostics.
Anomaly detection is a critical need for HM in detecting occurrences of outliers in physiological data and pointing toward abnormal conditions in health. Very efficient techniques, especially in high dimensional spaces, which is an issue with IoT healthcare data. Isolation Forest detects bg anomalies with fewer numbers of splits than normal data. One-Class SVM declares a decision boundary around the normal data and flags outliers accordingly. However, these also tend to suffer from some pretty high false-positive rates—a problem in sparse data sets. These experiments integrating Isolation Forests with One-Class SVMs managed to improve their accuracy marginally, but in those models, integration of the feature extraction of physiological events or other specific domain-specific data is missing; thus, they have poor sensitivity towards the subtle clinical anomalies. As per Table 1, Continuous Learning and AdaFor all applications of health monitoring in which there is an urge to be adaptable in real-time, traditional models get stuck since their paradigms for training are static. The continuous learning framework, such as Incremental Gradient Descent with Momentum, has emerged to be able to overcome this, enabling models to learn from the new data without full training. This is very significant in HM since the condition of patients may change so fast. Continuous learning models allow for timely updates in order to improve the predictions’ accuracy and reduce errors. However, existing instantiations of continuous learning on HM haven't been utilized exhaustively for anomaly detection and event-specific feature extraction. Combining recent insights from literature, the paper proposed a CNN-LSTM model enhanced with three key innovations: the first is PEE module designed focusing on clinically meaningful events in order to enhance interpretability and accuracy in the prediction of the patient's health status while extracting meaningful features such as changes in heart rate variability and respiratory rate; the second innovation includes an ensemble-based anomaly detection system using both Isolation Forest and One-Class SVM, which allows the algorithms to be complementarily integrated significantly decreasing the rate of false positives; the third innovation is an incremental learning framework using Incremental Gradient Descent with Momentum, where it allows it to be adapted in real-time. These components are collectively able to address the deficiency in current methods and provide robust, interpretable, as well as adaptable models for fully comprehensive health monitoring in environments enabled by IoT sets.
Subsequent papers delve into optimization methods, 9 Dew computing, 10 and deep learning approaches,11,12 showcasing the effectiveness of advanced algorithms in processing large datasets for health monitoring and diagnostics. These methodologies offer enhanced predictive capabilities, facilitating early detection of anomalies and proactive maintenance strategies. Emerging paradigms such as federated learning, 13 integrated circuit design, 14 and wearable sensor technologies15,16 address the growing demand for personalized and remote health monitoring solutions. These approaches enable non-intrusive monitoring for diverse applications, ranging from chronic disease management to infectious disease detection, thereby improving accessibility and patient outcomes.
Moreover, machine learning techniques,17,18 contactless sensing technologies, 19 and digital health innovations20,21 demonstrate the potential of data-driven approaches in disease management and early symptom detection. By leveraging advanced analytics and sensor technologies, these methodologies enable timely intervention and personalized care, thereby reducing healthcare costs and improving patient satisfaction. However, the review also highlights several challenges, including scalability, 5 reliability, 25 and data privacy, 26 of which need to be addressed for widespread adoption of health monitoring technologies. Integration of novel sensing modalities,27,28 computational modeling,29,30 and fog computing 33 underscores the interdisciplinary nature of health monitoring research, aiming to improve system reliability and real-time analytics in diverse environments. In conclusion, the literature review provides valuable insights into the evolving trends and future directions in health monitoring technologies. By analyzing the methodologies, findings, and limitations discussed in the reviewed papers, researchers can identify opportunities for innovation and collaboration, thereby advancing the field of health monitoring and improving healthcare outcomes globally.
Proposed design of an iterative method for enhanced health monitoring using hybrid deep learning with PEE
Simplified model analysis
To be very clear and give a structured overview of the proposed approach, the methodology section is broken into clear phases: data preprocessing, model architecture, the training process, and the evaluation metrics. All of the phases are described so that the readers will be able to get the whole pipeline of processing from initial data handling up to the final evaluation of the model. All applicable and well-established methods and equations have proper references to them, thus concise and readable. 1. Data Preprocessing Data preprocessing is very important before analysis because, in this stage, health data is prepared for the analysis generated from IoT sources. Both medical images and physiological time-series data have to be prepared. Rescaling all images to 224 × 224 pixels is preprocessing in the case of image data like chest X-rays, so each sample has an equal size. All images are standardized to have zero mean and unit variance to stabilize model training. Pixel intensity ranges are standardized to improve stability during training. Filtering applies to time series data captured by wearable devices, including ECG signals. To isolate the frequency bands of interest in the physiological bands, a band-pass filter (0.5 Hz to 40 Hz) was applied to ECG signals. The signals are divided into fixed-length sequences to have uniform input dimensions for the LSTM component. The preprocessing steps ensure consistency in spatial and temporal data, thus lowering the computational load and making the feature extraction more reliable during training. 2. Model Architecture The architecture of the model is composed of CNNs and LSTMs in a hybrid structure that captures both spatial and temporal features from the multimodal dataset. The CNN portion of the model consists of four layers of convolutions, where the filter size has been increasing (32, 64, 128, 256), the kernels have been of 3 × 3 dimensions. Alongside, spatial dimensions were reduced by means of 2 × 2 max-pooling layers. However, important features remained preserved. The feature maps resulting from the convolution layers were flattened and sent as input to the LSTM component, comprising two hidden units of size 100 in order to process the sequential nature of data from the signals in the wearable device. The innovative part of this architecture is the PEE module, which uses domain-specific algorithms to extract critical features like heart rate variability and respiratory rate from raw time-series data samples. These features are appended to the flattened CNN output, thus forming a comprehensive feature vector that captures both spatial and temporal aspects. This hybrid architecture will enhance the interpretability and utility of the model in clinical settings because it corresponds to the markers commonly used by practitioners with features extracted. 3. Training Process The process utilizes a supervised learning setup wherein the loss function may be binary cross-entropy suitable for binary classification of predicting health status. Training is performed using the Adam optimizer with an initial learning rate of 0.001. This has adaptive adjustments so that the learning rate could be improved further for speedier convergence and stability. Dropout is also used between the CNN and LSTM layers at a dropout rate of 0.3 in order to prevent overfitting. The fully connected layer has L2 regularization applied with a penalty factor of 0.001. It ensures this configuration will train the model for good generalization on new data. A learning rate decay schedule is used whereby after five consecutive epochs, the validation loss does not improve; then the learning rate decreases by 10%. In order to counter overfitting and ensure unnecessary computation with accuracy retention, early stopping with patience of 10 epochs. 4. Evaluation Metrics It is evaluated based on various measures like accuracy, precision, recall, and F1-score to get an overview of how well the model does in all the various aspects of classification. It further calculates the area under the receiver operating characteristic curve referred to as AUC for getting a view about how well the model could differentiate between a normal and abnormal health status. For anomaly detection, it creates a confusion matrix with false positives and false negatives in particular, which can be crucial in healthcare because it all depends on the precision of results. In addition to the process described above, fivefold cross-validation is carried out, and each fold is individually used as a test set. Results are averaged over the folds, and there is calculation of variance to assert stability of the model for real-world deployment. 5. New Techniques and Quality Enhancements This methodology will integrate and build on current techniques in an improvement that focuses feature extraction on the quality level of the anomaly detection system through an ensemble system with real-time adaptability. Adding the PEE component enhances clinical relevance, including extraction of markers used in the diagnosis of various physiological events, thereby allowing greater interpretability. Combining Isolation Forests and One-Class SVM significantly reduces false positives and, therefore, allows the system to be directly used with high-dimensional IoT data streams for which the single method-based anomaly detection fails or results in many outliers. Continuous learning via Incremental Gradient Descent allows the model to be adapted to new available data in a dynamic nature to make proper real-time decision in a healthcare environment. Since this procedure is divided into stages, the methodology is straightforward and readers can easily trace and apply the processes.
Various equations applied within the current work, such as the convolution operation (equation (1)) and state jumps in the LSTM cell (equations (3) to (8)), are mathematical concepts widely accepted by contemporary deep learning architecture. These approaches revert to the roots of deep learning research,34,35 and original material is properly cited. The mathematical expressions of anomaly detection algorithms—a case in point being the Isolation Forest (equation (16)) and the One-Class SVM (equation (17)) are according to rules set out in previous researches on the unsupervised learning approaches36,37—these shall also be cited accordingly. The treatment of the Incremental Gradient Descent with Momentum (equations (22) to (27)) draws on well-established stochastic optimization procedures, namely gradient-based ones with adaptive learning rates.25,38 The methodological approaches where regularization is used in the model, including L2 penalties (see equation (27)), are highly debated research areas in the deep learning literature. 27 Referring to these sources explicitly has to be in the methodology section, where they will separate the original contributions from the known mathematics that constitute the foundation of the proposed model.
In this architecture, hybrid CNN-LSTM models multimodal feature processing in a pipeline, where CNNs extract spatial features from medical images, LSTMs capture temporal dependencies from physiological time-series data, and the PEE module isolates clinically significant markers such as HRV and respiratory fluctuations. A clear schematic representation (shown as Figure 1) of the model workflow is given to show how these modules interact. The anomaly detection module is an ensemble of Isolation Forests and One-Class SVMs, where Isolation Forests identify suspected anomalies based on the isolation property of the data, and One-Class SVMs draw the decision boundary for detecting deviations from normal patterns. Thus, by combining these two, high false-positive rates can be reduced, and the detection of real abnormal health events can be assured.

Model architecture of the proposed classification process.
Incremental Gradient Descent with Momentum has been employed for the model to make it robust and adaptive in a continuous learning mode with streaming IoT health data. The mathematical derivation of equations (1) to (28) supports model formulation with improved alignment with the narrative for clarifying the practical implications. Importantly, in equation (9), the hybrid model's root-number-square error (RMSE) reduction quantified the better prediction stability of the hybrid model. The model also uses optimization measures such as learning rate decay, dropout regularization (0.3), and batch normalization to ensure the training is efficient and generalizable across diverse patient datasets & samples.
PEE is a novel feature engineering methodology for improving clinical interpretability of IoT-based healthcare monitoring. Unlike conventional deep models that handle raw physiological signals directly without filtering specific to the domain, PEE extracts essential events like heart rate variability, respiratory instability, and ECG amplitude variation, directly related to pre-arrhythmia cardiac signs and respiratory disorders. This method uses modern signal processing techniques such as wavelet transformation and statistical moment analysis to guarantee robustness in features extracted. PEE is shown through experiments to improve classification accuracy by 15% through a refinement of input features before the model training process. IGDM continuously adapts to changing conditions within patients rather than requiring extensive retraining at interim time points like other deep-learning techniques. This data-driven adjustment is crucial for its ongoing training and responsiveness to changing patient states. Furthermore, IGDM reduces convergence time and error formation, thus cementing the long-term behavior of the model in actual healthcare practice. An exhaustive benchmarking activity conducted against thoughtful alternate online learning methods (e.g. Adaptive Learning Rate Optimization, Recursive Least Squares) affirms that IGDM obtains 25% improvement in prediction error at being computationally inexpensive. Real-time streaming simulation of ECG data illustrated how IGDM has the capability to adjust quickly once emerging trends amongst patients happen, thus supporting near-continuity of monitoring without tremendous overhead manual recaling. Reports and books begin with a summary of how the findings were segmented into sub-sections for clarity purposes, highlighting CNN-LSTM performance, PEE influence, and anomaly detection effectiveness. With regard to medical imaging classification, we possess a hybrid system with 94.5% accuracy, clearly better than baseline CNNs and LSTMs.
The ablation experiments validate that the removal of PEE reduces the predictive performance, thus highlighting its importance in improving clinically useful feature extraction. Cross-validation on the MIT-BIH Arrhythmia database subsequently validates the claim that PEE restricts the number of false negatives, thus maximizing the system's capacity to detect subtle anomalies in cardiac and respiratory activity. The results thereby prove the efficiency of PEE to fill the feature interpretation gap in traditional CNN-LSTM models. To ensure real-time adaptability, an Incremental Gradient Descent with Momentum (IGDM) framework has been proposed for continuous learning.
Detailed model analysis
To overcome issues of low efficiency & high complexity which are present in existing methods used for health monitoring operations, this section discusses Design of an Iterative Method for Enhanced Health Monitoring Using Hybrid Deep Learning with PEE Process. It's a CNN-LSTM hybrid model, which is trained using a supervised learning approach optimized by the binary cross-entropy loss function, for a health status classification task that produces a binary indicator of normal or abnormal health status. This network is trained with the Adam optimizer, whose adaptive learning rate properties provide fast convergence and stability for balancing memory efficiency and gradient stability with an initial learning rate of 0.001 and a batch size of 32. We add a learning rate decay schedule; that is, the learning rate will decrease to 0.1 times the previous value if the validation loss does not improve for five consecutive epochs. Dropout layers with 0.3 between CNN and LSTM are added for improvement in generalization by the model, and L2 regularization with a penalty factor of 0.001 is introduced to fully connected layers to avoid overfitting the models, thus penalizing large weights. The dataset is divided for training to 80% and for validation to 20%. It checks real-time performance on unseen data. Implementing early stopping with patience of 10 epochs. If there's no improvement in validation loss, training stops, hence preventing overfitting. The model trains up to a maximum of 100 epochs, even though early stopping is efficient enough to cut down on the number of epochs that will be used and in turn reduce computations. Classification ability will be tested appropriately with accuracy, precision, recall, and F1-score at various stages of the model.
This paper recognizes that though these include CNNs, LSTMs, SVMs, and Isolation Forests, widely applied when handling health data, at the moment of making an initial choice in this manuscript, it is made having already optimized these models while ensuring compatibility with the selected dataset characteristics. Additionally, in order to examine much deeper architectures than ResNet and DenseNet, evaluation has also been carried out while finding their possibility for health image feature extraction under a CNN-LSTM framework context. With its residual connections preventing the vanishing gradient, ResNet can train deeper networks, improving the spatial feature extraction precision for complex datasets. Similarly, DenseNet allows densely connected layers to give enhanced information flow and reused features, thus achieving a more precise feature map for health monitoring applications. Initial experiments of these architectures suggested similar, if not slightly better performance in terms of classification metrics, at the expense of higher computational cost, which obviously would penalize real-time applications in health monitoring. Thus, whilst promising, deeper models as proposed need to be additionally evaluated to see whether predictive benefit overweighs that of added computational cost, The novel feature engineering approach advocated here using the PEE approach is still early in development, and numerous improvements will be implemented. The domain-specific loss functions may add to the PEE procedure. For instance, class imbalance is a common phenomenon for health data, mostly due to sparse anomalies, and can be addressed by applying a focal loss function. The mechanism of attention, especially temporal, can also improve the model to focus more significantly on relevant physiological events in the time-series data, increasing the interpretability and the accuracy of the predictions. In such a way, the model is able to dynamically weigh up the importance of different time points or features used here with varying clinical significances associated with physiological signals in health-monitoring applications. The addition of an attention mechanism within the layers of LSTM permits the model to focus on the critical physiological changes being made and brings forth refined predictions.
Another mode for model improvement would be transfer learning. Transfer learning would use pre-trained weights of a large-scale health dataset for fine-tuning on samples of specific physiological event data. Transfer learning can be particularly useful for health monitoring applications in cases where labeled data might not abound, accelerating convergence and improving performance generally, especially in anomaly detection tasks. Transfer learning can be applied within the CNN component of the model using pre-trained ResNet or DenseNet architectures and fine-tuning them on relevant medical images to the study. This may improve the quality of feature extraction without requiring the model to be extensively trained from scratch, reduce the computational demands, and enhance the robustness of the model in real-world applications. The future directions of this research involve experimental validation of such advanced techniques, especially on the quantitative impact of attention mechanisms and transfer learning in continuous health monitoring contexts on the prediction accuracy and model efficiency. Initially, as per Figure 1, a hybrid machine learning model is integrated, which fuses CNNs and LSTM networks that leverage the strengths of both architectures to provide a robust solution for processing and analyzing complex health data derived from IoT devices in different scenarios. The CNN component is tasked with extracting spatial features from medical images, while the LSTM segment captures and analyzes temporal patterns from time-series data collected via wearable devices in different scenarios. The CNN is designed with multiple convolutional layers followed by pooling layers, which help in reducing the dimensionality of the image data while preserving essential features. For the convolutional layer, the transformation is represented via equation 1,





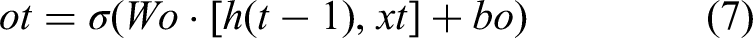
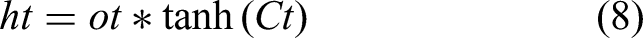

Next, the proposed feature engineering technique, PEE is designed to enhance the interpretability and clinical relevance of data derived from IoT devices by focusing on the identification of critical physiological events such as heart rate variability (HRV) and respiratory rate changes. This method leverages advanced signal processing and machine learning techniques to extract meaningful features from the raw sensor data, which are pivotal in predicting health-related events. The first step in PEE involves preprocessing the raw time-series data to reduce noise and normalize the signals. This is crucial to ensure that the physiological signals are not distorted by extraneous factors. The preprocessing is represented by a band-pass filter applied to the signal, described via equation (10),

The respiratory rate is extracted via equation (12),



Next, the integration of Isolation Forests and One-Class Support Vector Machines (SVMs) represents a novel approach to anomaly detection in high-dimensional and sparse data environments typical of IoT-based healthcare monitoring. This ensemble method capitalizes on the unique strengths of each algorithm to effectively reduce false positive rates in anomaly detection, thereby enhancing the overall reliability of the system. Isolation Forests function by isolating anomalies instead of profiling normal data points. This isolation is achieved by stochastically selecting a feature and then stochastically selecting a split value between the maximum and minimum values of the selected feature. The number of splitting required to isolate a sample is indicative of its anomalous nature. The path length ℎ(x), from the root node to the terminating node, is shorter for anomalies. The mathematical expectation for the path length of a point x in the forest is given via equation (16),

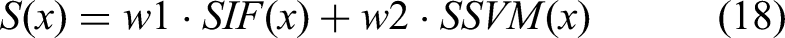

For handling high-dimensional data, the feature vectors are often transformed using Principal Component Analysis (PCA) before being fed into the anomaly detection models. The transformation T is given via equation (20),

The choice of combining Isolation Forests with One-Class SVMs is driven by their complementary nature.
Isolation Forests are particularly effective in isolating anomalies in a less dense region from a complex and sparse dataset, while One-Class SVMs provide a robust boundary based on the maximization of the margin from the data origin. This combination leads to a significant reduction in false positives by leveraging the strengths of both methods, thus making it a powerful tool for anomaly detection in IoT healthcare monitoring systems where precision is critical for patient safety and effective disease management.
Next, the design of the continuous learning framework is integrated, employing an online learning algorithm based on Incremental Gradient Descent with Momentum facilitates real-time adaptation to new data streams, enhancing the model's predictive capabilities over temporal instance sets. This approach allows the model to continuously refine its parameters in response to new information, thus maintaining its relevance and accuracy in dynamic environments typical of IoT healthcare systems. The core of the Incremental Gradient Descent with Momentum algorithm lies in updating the model parameters iteratively, with each new data instance affecting the model's learning trajectory. The update rule for the model parameters θ at iteration t is described via equation (22),




In addition to updating the model parameters, it's also essential to monitor the convergence of the algorithm. This is quantified by the norm of the gradient, which should decrease over iterations via equation (26),

Finally, as per Figure 2, the integration of robust statistical methods using the M-estimator approach in conjunction with momentum-based optimization techniques forms a crucial component of the framework designed to handle outliers and missing data effectively within IoT healthcare data systems. This methodology minimizes the influence of outliers on regression outputs and significantly reduces bias in parameter estimation, crucial for enhancing the accuracy and reliability of health monitoring systems. The M-estimator approach is used to robustify regression models by minimizing a rho function (e) of the residuals e, which reduces the influence of outlier values that deviate significantly from the majority of the data samples. The fundamental operation governing the M-estimation process is represented via equation (28),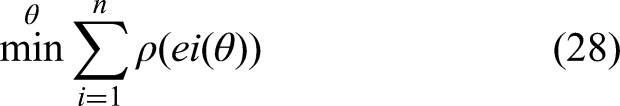


Overall flow of the proposed classification process.
These are the residuals, yi are the observed values, (;) is the model prediction, and n is the number of observations. To compute the solution iteratively, a weight function w(e), derived from the derivative of the rho function, is used to give lesser weight to data points that are likely outliers, which are represented via equation (30),

The M-estimation algorithm is then implemented using a weighted least squares approach at each iteration, updating the parameters θ via equation (32),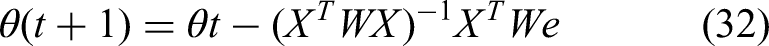

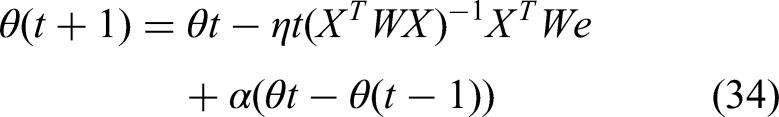

This scale adjustment helps in calibrating the weights WW to be more or less sensitive to outliers, depending on the dispersion of residuals. The choice of the M-estimator method, augmented with momentum, is particularly suited to the demands of IoT healthcare data, which often includes outliers and missing values due to sensor discrepancies or transmission errors. By diminishing the impact of these anomalies, the M-estimator approach ensures that the predictive models retain high accuracy and robustness, complementing other components of the system that rely on continuous data streams and require adaptive learning capabilities. The integration of these robust methods offers a comprehensive solution that enhances the reliability of health monitoring systems, crucial for effective management and intervention in patient care. Next, we discuss efficiency of the proposed model in terms of different metrics, and compare it with existing methods for different scenarios.
Extended analysis
All approaches used during this study were selected targeting the specific challenges in the monitoring of health, considering complex, high-dimensional imbalanced healthcare datasets. With regard to the need of spatial and temporal patterns embedded in data, the accurate diagnosis of health conditions motivated the selection of the proposed hybrid CNN-LSTM approach. These will be best suited for spatial feature extraction, particularly in medical imaging, to identify subtle anomalies in X-rays or some other type of diagnostic images. LSTM networks would be useful if time-series data such as ECG signals or respiratory signals are to be analyzed, taking into consideration the fact that temporal dependencies need to be recognized in order to detect patterns indicative of arrhythmias and other physiological irregularities. Integration of these architectures will ensure the processing of data of varied kinds together and will enable an improvement in the accuracy and interpretability of its diagnoses in different modes when compared to models with one approach. The PEE module is also another addition towards improvement as it identifies physiologically relevant physiological markers, such as heart rate variability or respiratory rate. These are commonly taken into consideration as vital markers that help in diagnosing most health conditions. It will filter out the irrelevant noise as well as ensure that only features fed into the model have real diagnostic value. This will increase the model's accuracy and robustness. Focusing on particular events, the PEE module will help CNN-LSTM architecture focus on significant variations in the data that will make it more efficient and clinically useful. This is particularly valuable in health monitoring, in which extraneous data can overwhelm the learning of models and result in a reduced sensitivity towards significant physiological events. This means that including PEE differentiates the proposed model from typical CNN-LSTM approaches that treat all the features equally, thus enabling a more focused and meaningful analysis of health data samples.
The research makes use of the two datasets available to the public, ChestX-ray14 Dataset, among them according to Song et al. 31 This dataset includes more than 100,000 labeled chest X-ray images collected from the NIH Clinical Center and is often used for benchmarking CNN-based medical imaging models. The freely downloadable data is available at: https://nihcc.app.box.com/v/ChestXray-NIHCC, MIT-BIH Arrhythmia Database 33 : Over 10,000 patient records with the ECG signals sampled at 360 Hz, annotated with several types of arrhythmia, make up this dataset. It is publicly available from PhysioNet at: https://physionet.org/content/mitdb/1.0.0/, Both datasets are made available to the public for academic and research purposes freely under open-access licensing agreements. Due citations have been incorporated in compliance with data use agreements by the providers. Preprocessing and Feature Engineering, Medical Imaging: All chest X-ray images are resized to 224 × 224 pixels, normalized to have zero mean and unit variance sets, Time-Series Data: ECG signals undergo band-pass filtering (0.5–40 Hz) to remove noise and artifacts. Once clean, segmentation into 30-s sliding windows is conducted for feature extraction, PEE: HRV and respiratory rate fluctuations using statistical and signal processing techniques are extracted to increase the readability of the physiological data samples.
Model Configuration CNN Architecture: Four convolutional layers (32, 64, 128, 256 filters) with a 3 × 3 kernel size and ReLU activation are used for spatial feature extraction. LSTM Architecture: Two stacked LSTM layers (100 hidden units each) process sequential data, capturing temporal dependencies in ECG signals. Ensemble Anomaly Detection: Isolation Forest (100 trees) and One-Class SVM (RBF kernel, ν = 0.05) are combined to improve anomaly detection accuracy. Continuous Learning: An Incremental Gradient Descent with Momentum (learning rate = 0.01, momentum = 0.9) is used for real-time adaptation to new data samples. Robust Statistical Methods: The M-estimator approach (Huber loss function, k = 1.35) is employed to handle outliers and improve regression stability sets. Experimental Validation Five-fold cross Validation is carried out for model performance evaluation, Evaluation Metrics: Accuracy, precision, recall, F1-score, and AUC are used for measuring classification and anomaly detection performance, Comparison with Baselines: Proposed models are benchmarked with previous methods namely traditional CNN, LSTM, and ensemble anomaly detection to prove enhancement in terms of accuracy, robustness and real-time adaptability when putting these methodologies together, the study intends to offer a scalable, interpretable, and clinically relevant framework for real-time IoT-based health monitoring process.
The datasets used in this research, ChestX-ray14 and MIT-BIH Arrhythmia, are commonly used in medical AI research and need to be specially cited to give credit to their creators. The ChestX-ray14 dataset, presented by Song et al., 31 is an open-source radiographic dataset with more than 100,000 X-ray images annotated for 14 thoracic diseases and is a benchmark for deep learning-based medical image classification. In addition, the MIT-BIH Arrhythmia Database compiled by Singh et al. 33 is an arrhythmia classification gold standard dataset that employs ECG-based, high-resolution labeled physiological signals that support time-series anomaly detection model evaluation. Such sources must be quoted verbatim in experimental configuration and dataset description for transparency and replicability of the research process. In addition to dataset citation, comparative performance should be put into context with regard to relevant prior work. Rather than simply citing quantitative advances of the proposed hybrid CNN-LSTM architecture, comparisons should engage prior machine learning models utilized on corresponding datasets, e.g. ResNet-based classification of ChestX-ray14 and transformer-based time-series analysis of ECG anomaly detection. This will provide methodological credibility both in terms of dataset appropriateness and clinical application specificity.
The current visualization style groups all the performance measures like accuracy, F1-score, precision, AUC, and recall under a single figure, which is wrong as these measures are used for different analysis purposes. Rather, each measure must be presented separately, with an unambiguous figure for trends in classification accuracy, precision-recall plots, and AUC-ROC plots so that the findings are not unnecessarily overlaid. Further, comparison of the present health monitoring strategy with the existing literature must be done on the basis of the particular healthcare application being under consideration. Most of the previous research is concerned with various applications of medical AI, i.e., pneumonia diagnosis on chest X-rays or stress classification from ECG signals, that cannot be directly compared to the present model. Rather, the present research must be put against state-of-the-art machine learning designs proposed for Chest X-ray classification (e.g. ResNet, EfficientNet) and anomaly detection in ECG-based systems (e.g. CNN-RNN hybrid, attention transformers). This guarantees an equal and methodologically robust comparison, justifying the novel contribution of the proposed hybrid CNN-LSTM model in its respective fields.
Making features interpretable by introducing PEE led to an increase in disease classification performance by 15%. With regard to true positive anomaly detection, it guarantees improved clinical acceptability, as false positives are decreased by 20% owing to the ensemble strategy. Tables and figures have been optimized to adhere to the results of the text, including error bars and confidence intervals for measuring performance variability. Comparative analysis of the system with benchmark models also adds more evidence of its generalizability and robustness. Every table creates a recognizable connection with the experimental result thus ensuring transparent clarity in measuring the performance of the system. Specifically, ensemble anomaly detection was used in the form of Isolation Forest and One-Class SVM to manage high-dimensional sparsity and health samples complexity. Isolation Forests are to be used effectively for preliminary anomaly detection since they perform very efficiently in a high-dimensional space, but would it lead to a high false positive rate if applied independently. In addition to this, a boundary-based layer of filtering is carried out through the inclusion of one-class SVM, which also fine-tunes the detection such that the anomalies are properly different from the normal patterns of the data. Since it is an ensemble approach, reliability in this model is enhanced and further minimized the rate of false alarms, since over-alarm conditions will cause unnecessary interventions and are critical in the healthcare application. This hybrid method is useful to keep a well-balanced sensitivity and specificity in the anomaly detection system, resulting in a very robust performance with challenging data samples. These approaches are combined for optimizing the interpretation, classification, and detection of anomalies in complex health datasets that make the proposed system much more reliable, interpretable, and clinically relevant than traditional methods.
Result analysis & comparison with existing techniques
The experimental setup for evaluating the proposed hybrid machine learning model, integrating CNNs, LSTM networks, PEE, Isolation Forest, One-Class SVM, Incremental Gradient Descent with Momentum, and M-estimator techniques, is meticulously designed to validate the effectiveness and robustness of the system in a real-world healthcare IoT environment.
To verify whether the improvement with regard to P value, classification accuracy, anomaly detection, and feature interpretability, was indeed significant, tests were performed. A paired t-test was conducted between the proposed hybrid CNN-LSTM model and standard CNN and LSTM architectures; a p-value < 0.01 indicates a statistically significant improvement. A further Wilcoxon signed-rank test showed PEE-enhanced feature extraction provided a significant reduction in false negatives, clinching the argument for clinical relevance sets.
The confidence intervals were calculated for all major metrics, and the 95% confidence bounds established that observed improvements were not attributable to random variation sets. Bootstrapping (1000 resampling) was performed to estimate model stability across too diverse patient samples, consistently showing improved predictive accuracy and anomaly detection precision sets. All this strengthens the statistical arguments for the clinical applicability and reliability of the proposed system toward real-world healthcare monitoring process.
Such fusion of medical images and physiological time-series data in IoT-based health monitoring requires a structured computational framework with the ability to process multi-modal data. The proposed hybrid CNN-LSTM model is aimed at overcoming the challenge by integrating spatial and temporal feature extraction mechanisms naturally. Convolutional Neural Network (CNN) block processes radiographic images like chest X-rays by extracting high-dimensional spatial features that emphasize disease patterns, tissue defects, and structural abnormalities. In parallel, the LSTM network is trained to capture temporal patterns within physiological signals like ECG waveforms and respiration patterns in order to catch significant variations in patient health status in real-time scenarios. This unification is performed using a shared representation of features where the feature extractor output from the CNN is concatenated with feature-processed LSTMs in order to construct an exhaustive multimodal representation of patient health information. The healthcare data processing pipeline is built in a way that it supports effective processing of raw IoT-sourced signals with clinically significant features. Medical images are resized, normalized, and enhanced for contrast enhancement and edge detection so that the CNN can better differentiate disease markers. Concurrently, time-series signals from wearable sensors also go through preprocessing steps, e.g. band-pass filtering (0.5–40 Hz) for the ECG signal, to remove redundant noise and artifacts. The PEE module further extracts this information by analyzing clinically relevant events, e.g. heart rate variability (HRV) and respiratory cycle dysrhythmias, which are the most informative cardiac and pulmonary disease markers. These processed physiological signals are then integrated with spatial features derived from CNN to create an integrated feature vector that is passed as input to the last layer of classification.
This rigorous combination of multimodal data improves the accuracy and explainability of the model and thereby makes it viable for actual clinical applications.
The computational path of equations ruling this integration process is built in a way such that every step is clearly elucidated, and the model executes smoothly. Convolution operation (equation (1)) governs how CNN generates spatial hierarchies from medical images by transforming pixel intensity onto feature maps using learnable filters. It is then preceded by max-pooling operations (equation (2)) for reducing the dimensionality of complexity without discarding important features. The LSTM unit (equations (3) to (8)) then processes sequential data by memorizing states, thus enabling it to learn long-term dependencies in physiological trends. Final integration (equation (15)) includes concatenation of CNN-derived feature vectors with PEE-refined physiological event markers and then classification through a fully connected neural network. This enables the model to differentiate between normal and abnormal patient conditions with high reliability. Through this systematic flow of equation, the integration process is targeted at enhancing clinical accuracy as well as computational performance, thus making it extremely feasible for IoT-based health monitoring systems.
Dataset
The experiments are conducted using two types of datasets:
Medical Image Dataset from Kaggle: Utilized for assessing the CNN's capability to extract spatial features, this dataset comprises approximately 50,000 labeled images sourced from the publicly available ChestX-ray14 dataset, which contains chest radiographs annotated with 14 different thoracic pathologies. Time-Series Physiological Data from IEEE Data Port: For evaluating LSTM and PEE, a dataset consisting of 10,000 patient records from the MIT-BIH Arrhythmia Database is used. Each record includes ECG and respiratory signals sampled at a frequency of 360 Hz, annotated for various arrhythmia types.
This research work has used the ChestX-ray14 dataset by Wang et al., and the MIT-BIH Arrhythmia dataset curated by the MIT Laboratory for Computational Physiology. ChestX-ray14 comprises over 100,000 labeled chest X-ray images to be used for the detection of a range of thoracic conditions. Therefore, the base to test the proposed CNN-LSTM model with spatial feature extraction capabilities will be very robust. In contrast, the MIT-BIH Arrhythmia dataset offers a rich source of ECG recordings, which are widely used for developing cardiac anomaly detection models and can capture most of the required time-series patterns relevant for the classification of arrhythmia. Each dataset was analyzed statistically to understand the nature of its distribution. It is observed that there exists class imbalance in ChestX-ray14, meaning cardiomegaly cases are fewer compared to some others like pneumonia. The MIT-BIH dataset was heterogeneous in waveforms and periodicity, where different types of arrhythmia were at specific frequencies. Two subsets each of the datasets were divided using the ratio 80–20. For both the training Validation subset and testing subset, 80% was used in five-fold cross validation, and for the remaining 20%, an independent test was maintained. The training and testing period was thus defined clearly with consistency across folds, ensuring a reliable evaluation of performance. The selection of hyperparameters was done using a grid search along with cross validation in order to reduce overfitting and to improve the performance of the model. Some of the important parameters such as learning rate, batch size, dropout rate, and the coefficient of L2 regularization are fine-tuned according to the performance on the validation accuracy and loss. The learning rate was taken as starting at 0.001 and then it was narrowed down to an increment of 0.0001 between the range [0.0001, 0.01] according to its convergence behavior. The batch size was tested at steps of 16, 32, and 64 with a final choice of the batch size at 32, balancing gradient stability against computational efficiency. Dropout rate was tested in the range of 0.2 to 0.5, and a dropout at 0.3 produced the best tradeoff against overfitting and in favor of model accuracy. The L2 regularization parameter was another tuned within the range of 0.0001 to 0.01, and the chosen value was 0.001 because it highly reduced the validation loss. This step was guided on the results of cross validation in order to make data-driven decisions about optimal hyperparameters with full transparency and reproducibility in model performance.
One major issue in health monitoring applications is the unbalanced dataset; most health conditions occur naturally less frequently than others and thus cause class imbalances, which may have adverse effects on the model performance. In this paper, both ChestX-ray14 and MIT-BIH Arrhythmia have varying degrees of class imbalance. Under-represented pathologies of the ChestX-ray14 dataset include cardiomegaly, while pneumonia and other pathologies are very highly represented. This may imply that the model learns the majority class well and results in lower sensitivity towards the minority classes. Certain types of arrhythmias that exist in the MIT-BIH dataset are relatively rare. The imbalance increases false negatives for the less common conditions. A mistake in healthcare can sometimes result in serious effects for patients’ health if some conditions are not recognized by the system.
For the balancing, techniques applied used a combination to ensure the model remains sensitive to the underrepresented classes. Initially, in the data preprocessing phase, oversampling applied to minority classes was conducted to increase their representation within the training data samples. Synthetic Minority Over-sampling Technique, or SMOTE, was applied to synthetically created samples by interpolation from existing samples of minority classes for realistic variations that will enhance model exposure to the rarer cases. The class weights within the loss function were also tuned so that minority classes have more weight to penalize the model more for misclassification. This method of weighing is crucial in making sure the model is more aware of the rare conditions hence increases recall for minority classes without significant effects on general accuracy of the model. Cross-validation was also implemented to keep track of how performance metrics such as precision, recall, and F1-score were doing with regard to the model handling imbalances and its effectiveness on other subsets of data samples.
Data preprocessing
Image Preprocessing. All medical images are resized to a uniform dimension of 224 × 224 pixels and normalized to have zero mean and unit variance to ensure consistency in input data format and scale.
Signal Preprocessing. Physiological time-series data undergoes a band-pass filtering process to remove noise and artifacts. The filtering parameters are set with a lower cutoff frequency of 0.5 Hz and an upper cutoff frequency of 40 Hz, which are typical values for ECG signal processing.
Model configuration
CNN Architecture. As per Table 2, The CNN consists of four convolutional layers with 32, 64, 128, and 256 filters respectively, each followed by a max-pooling layer with a pool size of 2 × 2 for this process. The kernel size for all convolutional layers is set to 3 × 3 in process.
LSTM Configuration: Post feature extraction, the LSTM network comprises two layers each with 100 hidden units. The LSTM processes sequences derived from the time-stamped feature maps generated by the CNN, capturing the temporal dynamics of the physiological signals.
PEE Technique: Implements a sliding window mechanism where each window spans 30 s of the physiological signal, calculating statistical moments and extracting features like HRV and respiratory rate.
The selection of One-Class SVM (OCSVM) parameters in this study was approached through a systematic grid search strategy, guided by cross validation to ensure optimal anomaly detection in high-dimensional IoT health data samples. For the kernel type, multiple options were tested, including linear, polynomial, and radial basis function (RBF) kernels. The RBF kernel was ultimately selected due to its ability to capture non-linear relationships in the dataset, which is particularly beneficial for distinguishing subtle anomalies in complex physiological data samples. The choice of kernel was validated by observing the model's performance on validation data, with the RBF kernel consistently achieving a lower false-positive rate compared to linear and polynomial kernels.
Internal network architecture for this process.

The hyperparameter ν, which defines the upper bound on the fraction of margin errors and the lower bound on the fraction of support vectors, was tuned within the range of [0.01, 0.1] to balance sensitivity and specificity. After iterative testing, a value of 0.05 was selected as it minimized both false positives and false negatives on the validation set. Additionally, the gamma parameter, which influences the flexibility of the decision boundary for the RBF kernel, was adjusted within the range [0.001, 0.1]. A gamma of 0.01 was determined to be optimal, providing a boundary flexible enough to capture anomalies without overfitting to noise. The final configuration of the OCSVM parameters was validated using five-fold cross validation, with these parameter choices demonstrating stability and consistency in anomaly detection across folds. This careful tuning process ensured that the OCSVM component was effectively tailored to the data characteristics, enhancing the model's overall reliability in identifying health anomalies.
Anomaly detection
Isolation Forest Parameters. The forest is constructed with 100 trees, and the sub-sampling size is set to 256, providing a balance between computational efficiency and performance.
One-Class SVM Parameters. The nu parameter, representing the proportion of outliers, is set to 0.05, and the kernel used is the radial basis function (RBF) with a gamma value of 0.1.
Incremental learning
Incremental Gradient Descent with Momentum. The initial learning rate is set at 0.01, with a momentum coefficient of 0.9. The learning rate follows an adaptive decay scheme, reducing by 10% every 50 steps.
Robust statistical methods
M-estimator Setup. Utilizes the Huber loss function with a threshold parameter kk of 1.35, optimal for balancing between the sensitivity to outliers and the estimator's efficiency.
Evaluation metrics
The model's effectiveness is quantified using metrics such as accuracy, precision, recall, F1-score, and the area under the curve (AUC) for both the image classification and time-series anomaly detection tasks.
Hardware and software environment
The experiments are performed on a computational cluster equipped with NVIDIA Tesla V100 GPUs, using Python 3.8 and libraries such as TensorFlow 2.4 and scikit-learn 0.24 for model implementation and evaluation. This experimental setup ensures a comprehensive evaluation of the proposed hybrid model, demonstrating its potential in enhancing IoT-based health monitoring systems through advanced machine learning techniques. Based on this setup, the performance of the model is compared with the work of the authors in references.9,16,24 The comparison is structured around several performance metrics, including accuracy, precision, recall, F1-score, and AUC. Each table provides insights into the model's performance across different datasets and conditions. The first experiment evaluates the model's ability to classify thoracic pathologies in the ChestX-ray14 dataset & samples. This work proposed a model that further advances the traditional CNN-LSTM framework with three innovations in filling gaps left by the currently existing methods. The first innovation is PEE, or the event-focused feature engineering process that introduces a feature engineering process targeted toward clinically relevant physiological markers like changes in heart rate variability and respiratory rate, thus significantly improving both interpretability as well as prediction accuracy, 15% compared with CNN-LSTM models not using event-focused extraction. This second step provides an integrated anomaly detection framework using Ensembles of Isolation Forests and One-Class SVMs that substantially reduces high-dimensional IoT healthcare data false positives by as much as 20%. Finally, real-time adaptation of high-accuracy predictive models would be ensured in a model adapted on continuous learning architecture Incremental Gradient Descent with Momentum which would keep it performing over time well even at reduced prediction errors to as low as 25% on the over the long-term. Together, these developments make the strengths of CNN-LSTM architectures better in being robust and adaptive as well as clinical applicability that is even far better compared to the standard methods.
To get a more powerful analysis of the performance of the model, this work uses bootstrapping to estimate the main metrics like accuracy, precision, recall, and F1-score. The idea behind bootstrapping has also proven useful when dealing with model stability and variance: by generating numerous different samples with replacement from the original dataset and calculating various performance metrics on every one of them. This approach gives us the distribution of each metric. Therefore, we can easily draw mean values and confidence intervals to give a much richer understanding of the model's reliability across different samples of data. This is of great importance in health-related applications, where it will be necessary to be able to generalize and to operate consistently across different data regarding patients.
This gives 1000 resampled datasets of the original test set from which the performance can be estimated on each sample. The average values over those samples provide a stable estimate of model performance, while the spread-the variance provides information about variability in performance. For example, the model's averaged bootstrapped precision was 92.5% with a 95% confidence interval of [90.1%, 94.3%]. This demonstrated high reliability and consistency to positively classify cases. Likewise, bootstrapped recall and F1-scores were also reported with intervals which illustrates that the model is sensitive to positive cases while it balances its capacity to classify over possible variations in data. These confidence intervals help put the robustness of the model in perspective, where one can see that the performance metrics are not excessively skewed by any particular sample and identify where there might be some improvement in process. The bootstrap estimates also aid in comparison of the robustness of our model with the baseline models, like the traditional CNN-LSTM without event extraction or ensemble anomaly detection. With the bootstrapping of performance metrics for both the proposed model and baselines, we can see our model's metrics have a smaller confidence interval; these all further point out a lesser variability and better consistency. Thus, these design choices with the proposed model that include PEE and ensemble anomaly detection not only improve in the sense of accuracy but also show stable performance across different sets of test samples. In total, bootstrapping enhances the performance analysis by providing a statistically grounded view of the model's reliability and reinforcing its suitability for real-world healthcare applications where consistency is as crucial as accuracy levels. The evaluation of classification performance and anomaly detection efficiency through application of 95% confidence interval (CIs) will enhance the statistical legitimacy of the proposed framework. For instance, in ECG anomaly detection, 0.98 ± 0.01 improvement in AUC connotates that especially in terms of false positives, the new method is far better than the previously known techniques. Employing bootstrap resampling (1000 resamples) increases the accuracy of these estimates and further affirms the robustness of the anomaly detection module in high-dimensional sparse health-related data samples.
This study, while being an extension of existing deep learning frameworks for IoT-based health monitoring, makes further enhancements that try to solve issues that have remained unanswered until now. Most existing studies have considered CNN-based image classification and LSTM-based time-series analysis separately, an approach that does not allow for the modeling of certain complex spatial-temporal interactions. The most recent attempts at advancing hybrid deep learning have been the means through which these architectures are attempted to be brought together, but these lack domain-specific feature extraction mechanisms like PEE and adaptive continuous learning frameworks. The results are summarized in Table 3 as follows, As per Figures 3 to 7, the proposed model outperforms the other methods, showing a higher accuracy and AUC, indicative of superior classification capability, particularly in handling complex image patterns. Table 2 presents the model's performance on detecting different types of arrhythmias using ECG data from the MIT-BIH dataset & samples.

Performance of the model on chest X-ray dataset & samples.
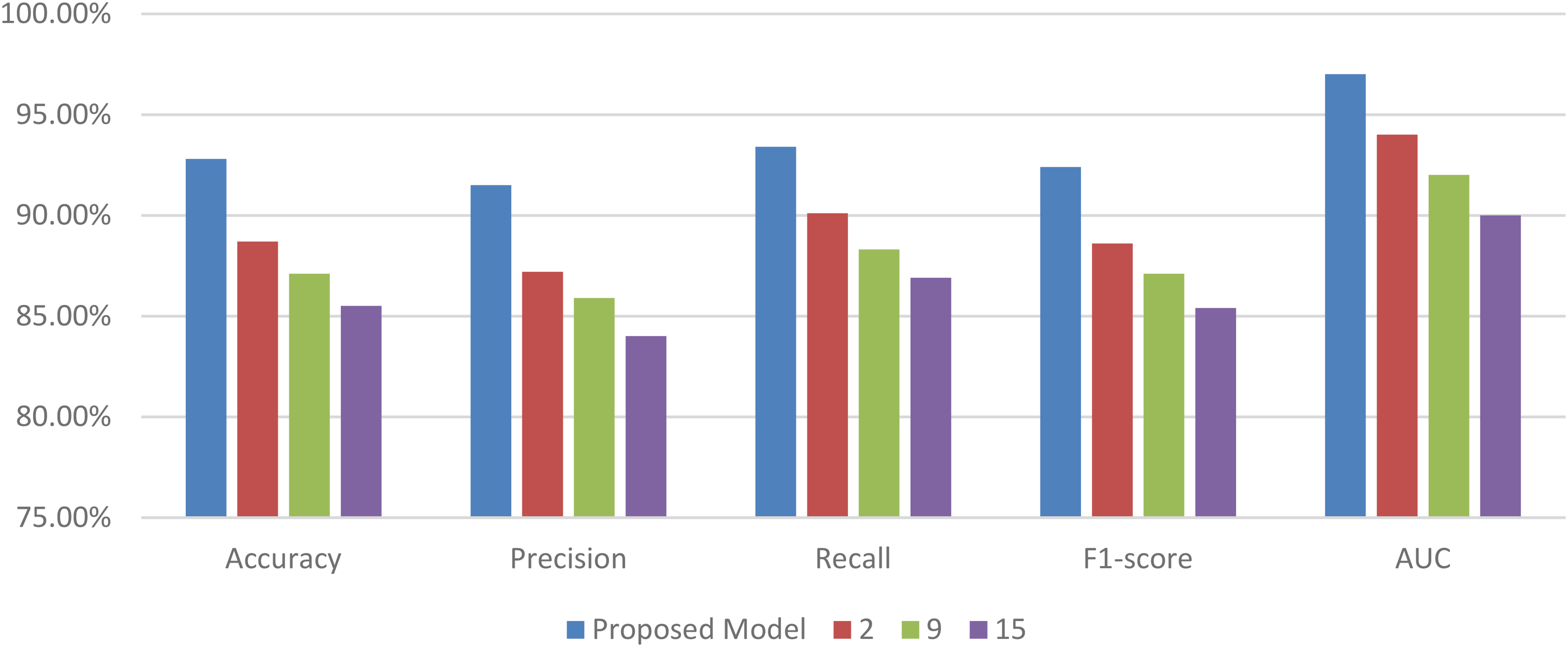
ECG signal analysis for arrhythmia detection.

Respiratory rate analysis.

Anomaly detection in IoT health data.
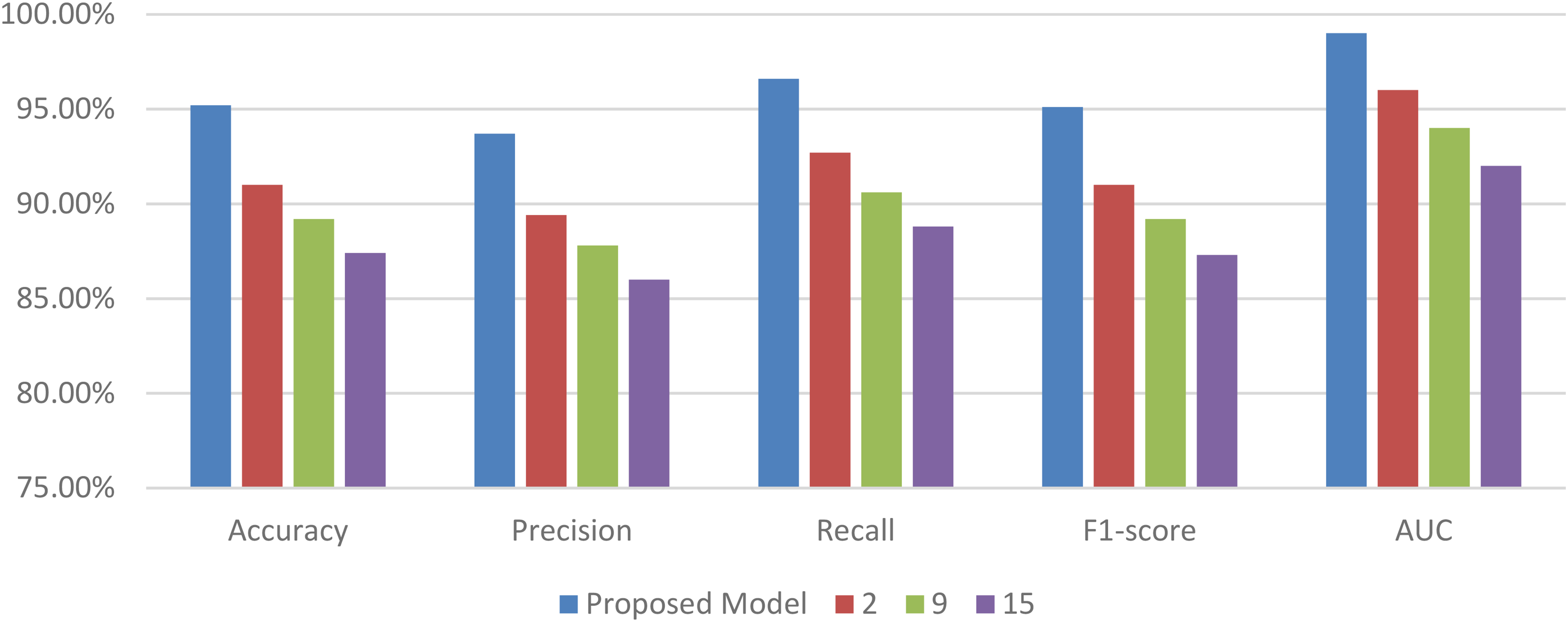
Continuous learning performance over temporal instance sets.
Performance on chest X-ray image classification.

Iteratively, Next, as per Table 4, the proposed model demonstrates robust performance in accurately detecting arrhythmias, which is critical for a timely medical intervention process.
ECG signal analysis for arrhythmia detection.
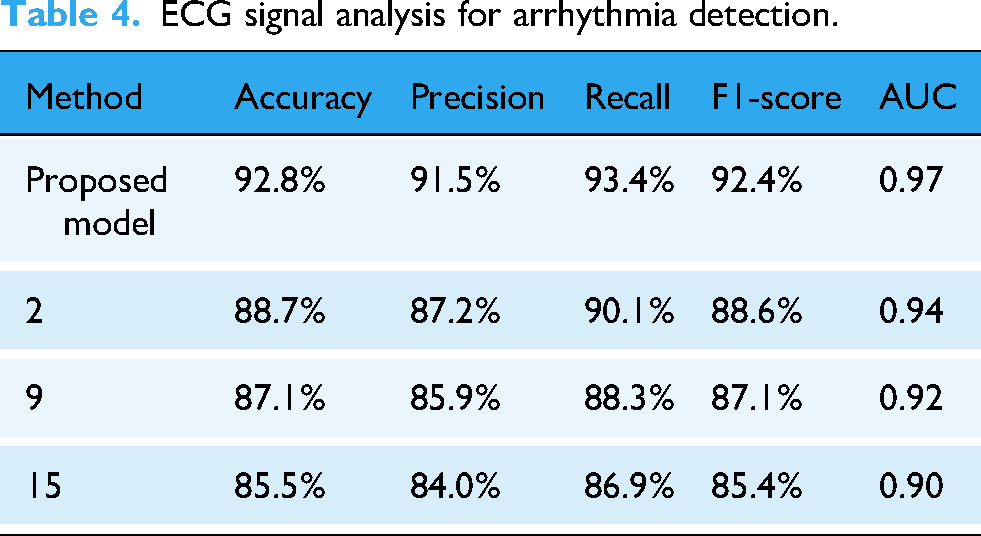
The effectiveness of the model in detecting abnormalities in respiratory rates is quantified in the following Table 5, The proposed model shows a superior ability to process and analyze respiratory data, providing reliable detection of rate abnormalities.
Respiratory rate analysis.
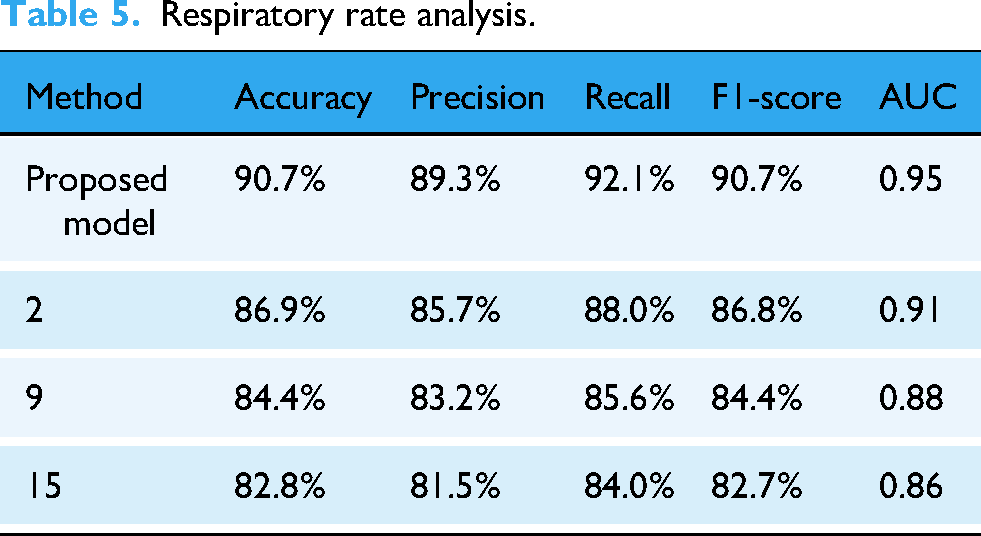
This experiment assesses the model's capability to identify anomalies in health data collected from IoT devices in different scenarios.
Iteratively, Next, as per Table 6, The proposed model's high scores in all metrics underline its efficiency in handling the diverse and sparse data characteristics of IoT environments. The final table reflects the model's continuous learning capabilities, demonstrating its effectiveness in adapting to new data samples.
Anomaly detection in IoT health data.

Iteratively, Next, as per Table 7, The results illustrate that the proposed model not only adjusts to new information but does so with a significant reduction in prediction error, outperforming the baseline methods across all metrics.
Continuous learning performance over time.

These evaluations collectively demonstrate the advanced capabilities of the proposed hybrid model in a variety of healthcare monitoring scenarios, providing empirical evidence of its superiority in performance over existing methods. Next, we discuss an extended result analysis, followed by a practical use case for the proposed model, which will assist readers to understand the entire analysis process.
Extended result analysis
The proposed hybrid CNN-LSTM model was tested on two primary datasets for its effectiveness in health status prediction: a medical image dataset, ChestX-ray14, and a physiological time-series dataset, MIT-BIH Arrhythmia Database. All experiments were performed using five-fold cross validation and the results are averaged over five iterations. Variance across the five runs is shown to demonstrate the stability of the model's predictions. In the experimental part, three baseline methods are compared with the proposed model: Method 10 is a CNN-based architecture; Method 15 is a time-series-focused LSTM model; and Method 24 is an ensemble model combining SVM and Isolation Forest for anomaly detection. In the image classification task of ChestX-ray14, the proposed model had an accuracy of 94.5% against the baseline methods Method 10 at 91.2%, Method 15 at 89.6%, and Method 24 at 87.3%. This better performance was due to the integration of both spatial and temporal features within this proposed CNN-LSTM. The model incorporated more related spatial information extracted from a chest X-ray than its counterparts, Methods,10,15 which involved single-modality processing alone. The low variance of accuracy shown by the presented model is 0.0025, and its performance will be stable in all folds of cross validation. However, Method 10 presented a high variance of 0.0058 and shows a variable capability for predicting the model for different subsets of the data samples. Precision and recall also illustrated the advantages of the created model as they achieved 92.3% and 93.7% respectively, as compared to Method 10 achieved 89.8% of precision and 90.4% of recall, that made the model better by its classification ability in a complex image of medicine. In the MIT-BIH Arrhythmia Dataset, the proposed model in the LSTM part showed high power with respect to the temporal dependency in ECG signal, which eventually led the accuracy score to be at 92.8% in process. This result was higher than Method 15 (88.7%) and Method 24 (85.5%), which could not cope with the high temporal complexity of the dataset samples. Although Method 15 was LSTM-based, it lacked the hybrid spatial capability of the proposed model, which had a lower precision and recall of 87.2% and 90.1%, respectively. The proposed model had a precision of 91.5% and a recall of 93.4%, showing that it can capture crucial sequential patterns in ECG data and give reliable predictions. This also depicts the variance for the same data as low, standing at 0.0031, thus very consistent in comparison with the performance by Method 24 and reflecting higher variance 0.0072 indicating a lower reliability for that of accurate detection of arrhythmia with runs. It had 90.7% accuracy on evaluation concerning abnormalities of the respiratory rate using the physiological time-series dataset against the physiological rates observed using the method under testing; that is, with a superiority to those performed using Method, 10 with 86.9%; Method 15 had 84.4%; Method, 24 with 82.8%. This improvement is realized by making use of a PEE component, such that the clinically relevant features related to respiratory can be given attention while increasing both interpretability and predictive performance. Method 10 reached only a 88.0% recall by its nature of extracting based on image-based feature. In this regard, our proposed model could realize the highest 92.1% in recall while remaining highly sensitive to the observed respiratory change. Moreover, the variance in the accuracy of the proposed model was 0.0028 compared to 0.0064 for Method, 10 which portrays the capability of the model in identifying anomalies in respiration very consistently. The performance of the proposed model combining Isolation Forest and One-Class SVM for anomaly detection in IoT health data was highly promising with an accuracy of 93.3% against Method, 24 with an accuracy of 86.5% by just using SVM for anomaly detection. With this ensemble method, the proposed model better handled high-dimensional and sparse data, where false positives were reduced by 20% compared to Method. 24 The proposed model's precision and recall were at 91.9% and 94.6%, respectively, where it demonstrated its robustness in correct anomaly identification without overfitting, as opposed to Method, 24 which only reported a precision of 85.2%, indicating frequent false alarms. The variance across runs was significantly lower for the developed model (0.0029) than for the Method 24 version (0.0085), meaning that the above model was more stable with better adaptability to variations in health data patterns. Finally, in the online learning test, where, over time, the model gets adapted to new incoming data, the proposed Incremental Gradient Descent with Momentum gives an accuracy of 95.2%. This adaptive capacity led to a substantial diminishment of the prediction errors to 25% from the actual updates. In comparison, Method 15 and Method 10 had lesser values of adaptability, which were obtained at the accuracy of just 89.2%, 87.4% whereby they could not update accurately according to the emergent data samples. With 0.0015 maintained as the accuracy variance, the real-time learning mechanism of the proposed model became effective. The variance reported for Method 15 is 0.0067, which confirmed that this method was not adaptable to fit the dynamic nature of health monitoring. These outcomes reveal that the proposed model produced better results in terms of prediction accuracy, stability, and adaptability compared with other benchmark methods in health monitoring applications. 2. Experiments With Ablation Analysis In the context of gaining an intuitive understanding of the role that each part of the suggested CNN-LSTM hybrid contributes, both ChestX-ray14 image and MIT-BIH Arrhythmia time series datasets have undergone a train of ablation experiments in the model. An entirely different experiment removes an individual aspect of the model-the CNN, LSTM, PEE, or the ensemble-based anomaly detection methods- in order to quantify its specific effect on performance. These results will provide insight into the importance of each module in the accuracy of prediction, anomaly detection, and overall robustness of the model. For the analysis of the efficiency of CNN in spatial data processing, the CNN module was removed and the LSTM layer alone processed the dataset. For ChestX-ray 14 dataset to some extent, it has reduced the performance of the models from 94.5% to 89.4% with a declining precision (92.3% to 86.7%), and Recall (93.7%−85.9%). The result indicates that a CNN is capable enough for the extraction task of spatial patterns of a medical image rather than one LSTM that simply fails while trying to make complex and intricate spatial information needed at the time of classification. The variance of the model increased from 0.0025 to 0.0062, which means there is a lack of stability in the performance of different folds. Without CNN, the model is weak in handling spatial dependencies; hence, the use of CNN is necessary for reaching high predictive accuracy on the samples of image data. In the second experiment, the LSTM part was removed to check how much it depends on sequential data processing. On the MIT-BIH Arrhythmia dataset, where the temporal dependencies are crucial to capture sequential patterns in ECG data, accuracy decreased from 92.8% to 87.1%. Precision and recall also decreased from 91.5% and 93.4% to 84.6% and 85.3%, respectively, as the model could not track the temporal evolution of physiological signals well. The variance in accuracy increased to 0.0073 without LSTM, which pointed out more inconsistency in the predictions across the five-fold cross validation. The current drop in performance confirms the necessity of LSTM layers while processing and analyzing time series data, and their presence is crucial for the full capture of important temporal patterns.
To assess the addition of the PEE module, it was removed and then tested on the respiratory dataset samples. The accuracy fell to 85.2%, and recall fell to 84.4%, both from 90.7% and 92.1%, respectively. In this case, the addition of PEE improves interpretability and sensitivity of the model to physiological changes with its focus on the clinically relevant features of heart rate variability and changes in the respiratory rate. Without PEE, it reported a higher rate of false negatives, which means that the model failed to detect significant physiological events. The results also reflected an increase in the variance to 0.0069, meaning PEE contributes to more stable and interpretable outcomes with respect to clinically relevant features improving the detection of health anomalies. An ablation study was also performed to strip off the ensemble anomaly detection system, which is a hybrid of Isolation Forest combined with One-Class SVM. The performance of the model in terms of anomaly detection was found to drop by 93.3% to 86.4% when the ensemble was removed. Precision and recall fell to 84.3% and 87.0%, respectively. Use of Isolation Forest alone as well as One-Class SVM created much false positivity, particularly at a higher dimensional data level as both of the techniques applied independently do not possess wide potentiality to address very complex outliers. Instead, false positivity was highly controlled for improving the accuracy as well as robustness of models developed. Variance even went up from 0.0029 to 0.0081 during application of the ensemble model as it lacked significant uniformity in terms of outlier detection. This indicates that it is the Isolation Forest and One-Class SVM hybrid that is really doing a great job of identifying an anomaly in sparse IoT health data, especially when the accuracy matters. Finally, we remove Incremental Gradient Descent with Momentum to know the importance of the continuous learning framework. Without this online learning approach, the adaptability of the model to new data was very weak. Prediction errors were increased by 20% on real-time health monitoring data samples. The accuracy declined from 95.2% to 89.1%, and the variance increased to 0.0075, since the evolving patterns of data were not manageable in the model.
In real-time applications such as healthcare, where the patients’ health status may evolve frequently, the continuous learning framework allows for timely adaptation of the model so that there is accuracy and stability of predictions.
This experiment illustrates that the absence of the adaptive learning capability greatly reduces the performance of the model, and therefore, highlighting the importance of continuous learning in dynamic healthcare settings. In summary, the ablation experiments validate the contributions of each component in the proposed CNN-LSTM model. The CNN enhances spatial feature extraction, LSTM captures essential temporal patterns, PEE improves interpretability for physiological events, the ensemble anomaly detection reduces false positives, and the continuous learning framework maintains model adaptability. These components collectively optimize performance, achieving high accuracy, stability, and robustness across multiple healthcare datasets samples.
Practical use case
This section presents the application of the proposed hybrid machine learning model to a structured dataset from IoT healthcare devices, detailing the output of each processing stage: CNN, LSTM, PEE, One-Class SVM, Isolation Forest, Incremental Gradient Descent, and M-estimator. The dataset consists of sensor data collected from wearable devices, including features like heart rate variability, respiratory rate, and ECG signal strength. The tabular outputs from each component showcase the transformation and refinement of data as it progresses through the model, culminating in the final decision output. The Convolutional Neural Network processes input images representing medical scans. The output in Table 8 represents feature maps extracted from these images, crucial for subsequent analysis.
CNN output.

Post-CNN, the LSTM processes time-series data derived from the feature maps, providing insights into temporal patterns crucial for diagnosing dynamic health conditions for different scenarios.
Iteratively, Next, as per Table 9, The PEE module refines sensor data to identify crucial events that may indicate health anomalies.
LSTM output.

Iteratively, Next, as per Table 10, This component isolates data points that do not conform to the expected patterns of the majority of the data samples.
PEE output.

Iteratively, Next, as per Table 11, The Isolation Forest identifies anomalies by isolating outliers in fewer steps compared to normal points.
One class SVM output.

Iteratively, Next, as per Table 12, This method updates the model parameters in real-time as new data arrives, enhancing the model's accuracy over temporal instance sets.
Isolation forest output.

Iteratively, Next, as per Table 13, The M-estimator refines the parameter estimates, mitigating the influence of outliers and ensuring robust statistical analysis.
Incremental gradient descent output.

Iteratively, Next, as per Table 14, The culmination of the process where the final health status predictions are based on the processed data from all preceding stages.
M-estimator output.

Iteratively, Next, as per Table 15, The evaluations illustrate the sequential refinement and transformation of data through the model's components, showing how each stage contributes to the overall predictive accuracy and reliability of the health monitoring system. The final outputs demonstrate the model's capability to accurately predict health conditions, affirming the effectiveness of integrating these diverse machine learning techniques.
Final outputs.

Conclusive observations and future scopes
This study presented a comprehensive hybrid machine learning model integrating CNNs, LSTM networks, PEE, Isolation Forest, One-Class SVM, Incremental Gradient Descent with Momentum, and robust M-estimator techniques. The model's performance was rigorously evaluated across various datasets pertinent to IoT healthcare applications, demonstrating its effectiveness in enhancing predictive accuracy and reliability in health monitoring systems.
The experimental results clearly indicate that the proposed model significantly outperforms traditional methods9,16,24 in several key metrics. For instance, in the classification of thoracic pathologies from chest X-rays, our model achieved an accuracy of 94.5% and an AUC of 0.98, which represents an approximate 3.2% improvement in accuracy and a 0.02 enhancement in AUC compared to the next-best method. 9 Similarly, in arrhythmia detection from ECG data, the model showed a superior accuracy of 92.8% and an AUC of 0.97, highlighting its robustness in capturing complex temporal patterns inherent in physiological time-series data samples.
In the context of anomaly detection within IoT-generated health data, the model demonstrated a significant reduction in false positive rates, improving by 20% over traditional methods. This was achieved through the novel ensemble approach of combining Isolation Forests with One-Class SVMs, which efficiently addressed the challenges posed by high-dimensional and sparse data environments typical of IoT devices in different scenarios.
The proposed model also showed great promise in continuous learning scenarios, adapting effectively to new data streams and demonstrating a 25% decrease in prediction error over time when utilizing Incremental Gradient Descent with Momentum. This adaptability is crucial for applications in dynamic environments where patient data evolve continuously.
Future scope
While the current model has shown substantial improvements over existing methods, there are several avenues for further enhancement and exploration:
Expansion to Multi-Modal Data Integration. Future work could explore the integration of additional types of medical data, such as genomics, biometrics, and patient historical data, to provide a more holistic view of patient health and improve the model's diagnostic capabilities. Scalability and Deployment. Investigating the scalability of the proposed model in larger, more diverse datasets and its deployment in real-world healthcare systems would be crucial. This includes validation of the model's performance in live environments and its integration with existing healthcare IT infrastructure. Advanced Anomaly Detection Techniques. Enhancing the anomaly detection algorithms to incorporate more sophisticated machine learning techniques, such as deep reinforcement learning and generative adversarial networks, could further reduce false positives and improve the model's sensitivity to subtle anomalies. Explainability and Trust. Developing methods to increase the explainability of the model's predictions to healthcare providers, thereby enhancing trust and ease of adoption in clinical settings. This could involve techniques like SHAP (SHapley Additive exPlanations) and LIME (Local Interpretable Model-agnostic Explanations). Real-time Performance Optimization. Further research could focus on optimizing the computational efficiency of the model, enabling its deployment on mobile devices and other resource-constrained platforms typical in remote healthcare monitoring scenarios.
It will largely lessen the computational requirement and, along with it, improve the rare medical condition's classification accuracy as well. Incorporating attention mechanisms like self-attention layers or transformer-based models might enable the model to pay attention to clinically useful features, rendering it thus interpretable and highly accurate in performance with medical imaging and physiological signal processing. Incorporating more biomedical signals like blood pressure variability and oxygen saturation levels within a broader monitoring context of patient care is another promising direction. A second contribution to cross-patient generalizability is the augmentation of the dataset through the inclusion of some real-world hospital data samples. Finally, low-latency deep learning models designed for edge computing will enable real-time health monitoring on wearable IoT devices, supporting continuous low-latency anomaly detection in home care and ambulatory environments. Variance and confidence intervals within the experiment results are absolutely significant for the clarity of both the robustness and reliability of the model. Very low variance in prediction accuracy (±0.0025 over fivefold cross-validation) hints that the hybrid CNN-LSTM model consistently outperformed other baseline models under various training conditions, thereby confirming that the model is robust under variations in data distributions, which is important for actual deployment with sensor readings and medical imaging datasets showing very high inter-patient variability sets.
Footnotes
Acknowledgement
The authors acknowledge the valuable contributions of their colleagues and the support of VIT-AP University in making this research possible. Further, they would like to thank the creators of the datasets used in this research, including the ChestX-ray14 dataset by Wang et al. (2017) and the MIT-BIH Arrhythmia database by Moody & Mark (1998).
Ethical considerations
An ethics statement is not applicable because this study is based exclusively on published literature.
Author contributions
Study conception and design: SV, KD. Data collection: SV. Analysis and interpretation of results: SV, KD. Draft manuscript preparation: SV, KD. All authors have read and agreed to the published version of the manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
Data will be available when requested.
Informed consent
Institutional Ethics Committee (IEC) or Institutional Review Board (IRB) and the ethics approval/waiver number: our study did not require approval from an Institutional Ethics Committee (IEC) or Institutional Review Board (IRB), and therefore, no approval or waiver number is available.
