Abstract
Objective
The objective of this study is to develop a machine learning (ML)-based predictive model for bone metastasis (BM) in esophageal cancer (EC) patients.
Methods
This study utilized data from the Surveillance, Epidemiology, and End Results database spanning 2010 to 2020 to analyze EC patients. A total of 21,032 confirmed cases of EC were included in the study. Through univariate and multivariate logistic regression (LR) analysis, 10 indicators associated with the risk of BM were identified. These factors were incorporated into seven different ML classifiers to establish predictive models. The performance of these models was assessed and compared using various metrics including the area under the receiver operating characteristic curve (AUC), accuracy, sensitivity, specificity, F-score, precision, and decision curve analysis.
Results
Factors such as age, gender, histological type, T stage, N stage, surgical intervention, chemotherapy, and the presence of brain, lung, and liver metastases were identified as independent risk factors for BM in EC patients. Among the seven models developed, the ML model based on LR algorithm demonstrated excellent performance in the internal validation set. The AUC, accuracy, sensitivity, and specificity of this model were 0.831, 0.721, 0.787, and 0.717, respectively.
Conclusion
We have successfully developed an online calculator utilizing a LR model to assist clinicians in accurately assessing the risk of BM in patients with EC. This tool demonstrates high accuracy and specificity, thereby enhancing the development of personalized treatment plans.
Introduction
Esophageal cancer (EC) ranks seventh among the most prevalent malignant tumors worldwide, resulting in an annual mortality rate of 500,000 and accounting for 5.5% of all cancer-related deaths. 1 The 5-year survival rate for patients with distant metastasis is less than 5%, 2 while approximately 50% of patients already exhibit distant metastasis at the time of diagnosis. 3 Furthermore, nearly one-third of patients develop distant metastasis following surgical intervention or radiotherapy/chemotherapy. 4 Early detection and accurate diagnosis play a pivotal role in enhancing prognosis. 5
The development of EC can be influenced by various factors, such as biological characteristics, 6 dietary choices, 7 lifestyle patterns and behaviors, 8 psychological well-being, 9 as well as environmental conditions. 10 Bone metastasis (BM) is one of the most common forms of metastasis in malignant tumors and is the third most common site of metastasis in patients with EC, 11 after the liver and lungs.12,13 Currently, there are few treatment options that can effectively improve the survival rate of patients once BM occurs. Patients may experience a series of bone-related symptoms, including pathological fractures, hypercalcemia, pain, and neurological compression syndrome. 14 Prevention and early screening are particularly important for patients with EC as timely detection and intervention can improve their quality of life and prolong survival time. 15
Traditional bone marrow puncture biopsy, although considered the gold standard for diagnosing bone marrow metastasis, is limited in clinical application due to its invasiveness and discomfort. Additionally, imaging studies such as CT and MRI have limited sensitivity for early microscopic metastatic lesions, often leading to missed diagnoses. 16 In recent years, with the rapid development of artificial intelligence (AI) technology, its application in medical imaging analysis, data mining, and clinical decision support has become increasingly widespread.17,18 Particularly in the field of tumor diagnosis, AI, through deep learning algorithms, can automatically identify and extract features from vast amounts of medical imaging data, significantly improving the accuracy and efficiency of disease diagnosis.
As a branch of artificial intelligence, machine learning (ML) has become a powerful tool in the field of healthcare, completely changing disease diagnosis, prognosis, and treatment methods.19,20 Compared to traditional logical models, ML technology can reveal more information and achieve result prediction with higher accuracy when dealing with large-scale datasets. 21 This promotes personalized medical approaches and enhances clinical decision-making ability. 22 Currently, research on AI applications for detecting bone marrow metastasis in EC remains relatively limited. This study aims to develop a predictive model for BM in EC using ML methods. By applying ML technology, we aim to construct a robust predictive model that can assess the risk of BM in patients in advance and improve clinical outcomes.
Methods
Study population
This study utilized Surveillance, Epidemiology, and End Results (SEER)*stat 8.4.1 software to retrieve clinical data of patients diagnosed with EC from the SEER database. The study included patients diagnosed with EC between 2010 and 2020 in the SEER database. Exclusion criteria were as follows: (a) unknown race and grade; (b) unknown primary site; (c) histological types other than adenocarcinoma and squamous cell carcinoma; (d) unknown AJCC T, N stage; (e) unknown bone, brain, liver, and lung metastatic status. A total of 21,032 cases of EC were identified. The complete selection process is illustrated in Figure 1.

The study flow chart of case screening.
Data selection
In this study, we selected 15 variables related to patients’ clinical pathology and demographic characteristics for analysis. Demographic variables included age, gender, race, etc., while clinical pathology indicators included primary site, tumor histology, tumor grade, T stage, N stage, surgery, radiotherapy, chemotherapy, brain metastasis, BM, lung metastasis, liver metastasis, etc. According to the ICD-O-3 coding, the histological types of EC are divided into adenocarcinoma (8140–8573) and squamous cell carcinoma (8050–8082). All EC patients were staged according to the AJCC 8th edition guidelines and SEER staging information. Chi-square analysis and variance analysis were used to reveal the differences between factors in the training and validation sets.
Data preprocessing
We processed the data collected from the SEER database using Python 3.12 and SPSS 28 software with the aim of identifying variables suitable for ML models. Firstly, we conducted single-factor logistic regression (LR) analysis to determine variables that were significant in patients with bone metastases (p < 0.05). Subsequently, these variables were included in a multiple LR analysis, and ML models were used to further analyze the variables with p < 0.05 in the multiple LR analysis. To examine the correlation between the selected features, we employed a correlation analysis method. In addressing the imbalance within our dataset, we employed the synthetic minority oversampling technique (SMOTE) as an oversampling method to balance the distribution of minority and majority classes. 23 SMOTE is a widely used technique that synthesizes new samples for the minority class by creating synthetic instances between existing minority class samples and their nearest neighbors. The crucial aspect of this approach lies in augmenting the number of samples from the minority class data through oversampling, which ultimately enhances the model's precision without altering its accuracy. 24
Model establishment and evaluation
We randomly assigned the data in the SEER database to a training set and an internal test set at a ratio of 7:3. We selected seven common classifier algorithms, including random forest, extreme gradient boosting, LR, K-nearest neighbors, Naive Bayes classifier (NBC), decision tree and support vector machine. We used Python software to train the ML models and divided all SEER data into 10 parts for 10-fold cross-validation in the training set. In the internal test set, we directly imported the built models for validation. Performance metrics for evaluating ML algorithms included the receiver operating characteristic curve (AUC), sensitivity, specificity, accuracy, F-Score, and precision. We also used decision curve analysis (DCA) to compare the actual application effects of various models in clinical decision-making. Finally, based on the best-performing model, we built a web-based online calculator to facilitate more convenient predictions and decision-making.
Results
Patient clinical characteristics
A total of 21,032 patients diagnosed with EC were retrieved from the SEER database (Supplementary Table 1), and then divided into training set (n = 14,722) and test set (n = 6310) at a ratio of 7:3. Fifteen factors were collected, including age, gender, race, primary site, histological type, tumor grade, T stage, N stage, surgery, radiotherapy, chemotherapy, brain metastasis, BM, lung metastasis, and liver metastasis. Among them, there were 1157 cases of BM (5.5%) and 19,875 cases without BM (94.5%). Chi-square and variance analyses within the two groups (training and validation groups) showed no statistically significant differences between the factors (p > 0.05, Table 1).
Characteristics of train set and internal test set.

Univariate and multivariate logistic regression analysis
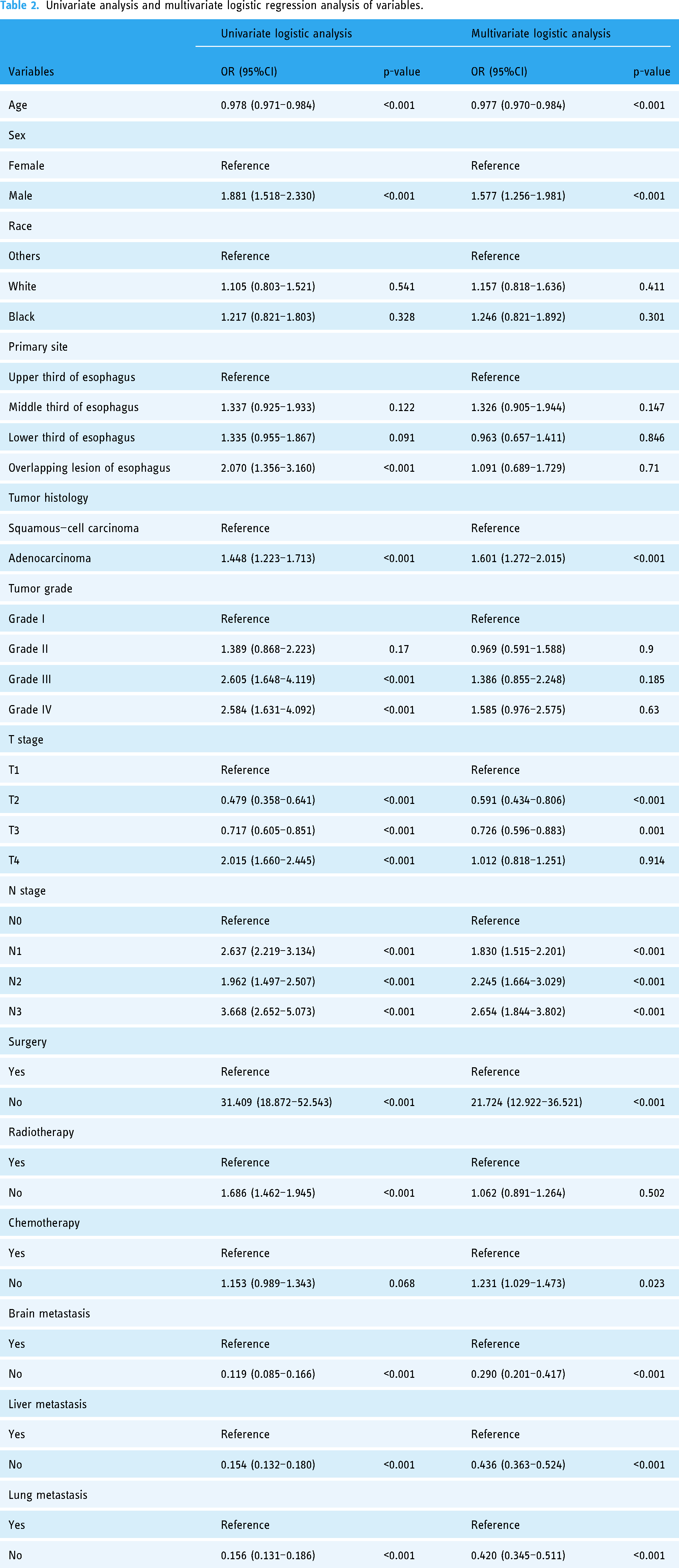
Through univariate and multivariate LR analyses, 10 risk factors related to BM were identified, including age, gender, tumor histology, T stage, N stage, surgery, chemotherapy, brain metastasis, liver metastasis, and lung metastasis (p < 0.05, Table 2). Based on these independent prognostic factors, seven different models were developed using ML algorithms.
Univariate analysis and multivariate logistic regression analysis of variables.
Correlation analysis
The Spearman correlation analysis was used to evaluate the relationships among the 14 factors (Supplementary Table 2). The results (Figure 2) showed that there was no strong linear correlation among these 14 features.

Heat map of the correlation of features.
Model performance
Ten-fold cross-validation on the train set revealed that among the seven models, the LR model demonstrated superior performance with an average AUC of 0.834, closely followed by the NBC model (AUC = 0.832) Figure 3(a). In the test set, the results of ROC curve analysis showed that the AUC score of the LR model was 0.831, which was superior to other models (Figure 3(b)). To further evaluate the performance of the models, we compared the AUC, accuracy, sensitivity, specificity, F-Score, and precision of the seven ML models. Overall, the LR model performed the best, and its indicators were as follows: AUC 0.831, accuracy 0.725, sensitivity 0.791, specificity 0.721, F-Score 0.241, and precision 0.142 (Table 3). Subsequently, the importance of the predictive variables of the LR model was ranked through permutation tests (Figure 3(c)), and the results indicated that surgery, brain metastasis, lung metastasis, and liver metastasis had significant effects on the prediction results. In addition, DCA (Figure 3(d)) showed that the LR model had better clinical utility compared to other models in this study. Finally, the confusion matrices of the training set and the test set (Figure 3(e)) further verified the high accuracy of the LR model. We used SHAP and LIME analysis methods to conduct global and local interpretations of the LR model. The results showed that the feature “Whether surgery” had the greatest impact on the model's prediction results (Supplementary Figures 1 and 2).
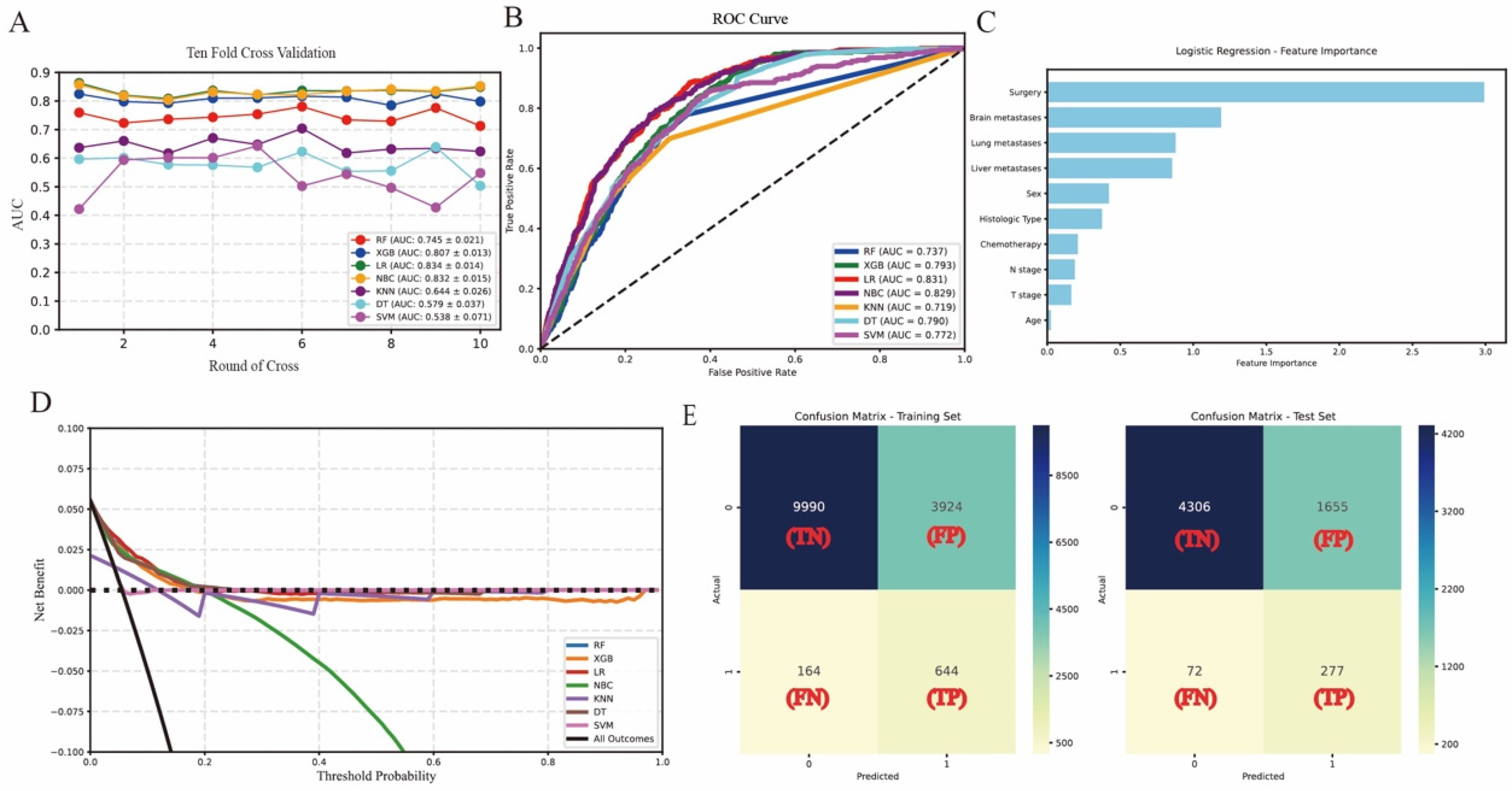
(a) Ten-fold cross-validation results of different machine learning models. (b) The roc curves of different machine learning models in internal test set. (c) Feature importance ranks in logistic regression. (d) Decision clinical analysis curves of algorithms in the internal test set. (e) The confusion matrix of the LR model in the train set and the internal test set. TP: true positive; TN: true negative; FP: false positive; FN: false negative; LR: logistic regression.
Prediction performance of different models.
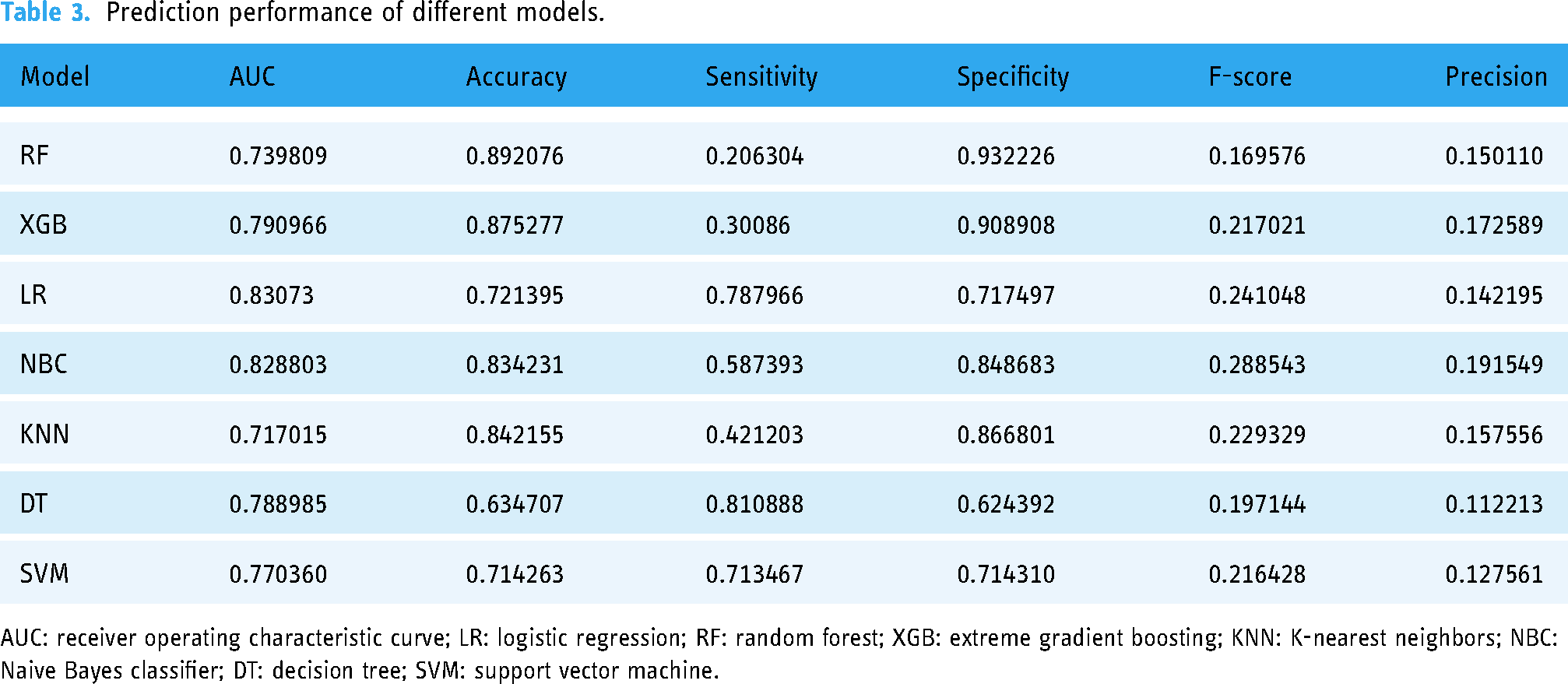
AUC: receiver operating characteristic curve; LR: logistic regression; RF: random forest; XGB: extreme gradient boosting; KNN: K-nearest neighbors; NBC: Naive Bayes classifier; DT: decision tree; SVM: support vector machine.
Network prediction
The LR model was utilized to develop a network predictor, which demonstrated superior predictive performance in the identification of BM among patients with EC. This network predictor serves as a valuable tool for healthcare professionals, enabling them to make more precise clinical decisions. By inputting relevant variables associated with BM into the network predictor, physicians can easily calculate the probability of BM occurrence in patients with EC. To access the network predictor, please refer to the following link: (https://project-1-sz2zqgdwbewjz4pmjiluya.streamlit.app/) (Figure 4)

A web predictor for predicting BM in EC. EC: esophageal cancer; BM: bone metastasis.
Discussion
The application of AI in the medical field is becoming increasingly widespread, particularly in areas such as precision diagnosis, personalized treatment, and disease prediction. For instance, the use of AI in COVID-19 detection and diagnosis has demonstrated its critical role in responding to public health emergencies. 25 In the field of nuclear medicine, the integration of AI has not only enhanced diagnostic accuracy but also opened up new possibilities for personalized treatment. 26 These studies indicate that AI technology is driving rapid advancements in the medical field.
Recent studies have used this innovative approach to explore clinical issues related to early cancer metastasis27,28; however, there hasn't been any research using ML methods to assess the risk of BM in diagnosed EC patients. The primary objective of this study is to develop a ML-based model for predicting the risk of bone marrow metastasis in EC patients. Previous studies, such as,29,30 Bei Yuan and Liming Jiang constructed traditional nomogram prediction models that not only used clinical data from 2010 to 2015 as patient samples but also included fewer variables. This study not only improved the sample size but also added variables compared with previous studies. In our study, we identified 10 independent risk factors influencing BM in EC through univariate and multivariate LR analysis and incorporated them into seven ML algorithms for development and validation to predict BM in EC patients. Among them, the LR model showed excellent performance in internal validation with AUC values of 0.831 on internal testing sets. The DCA also demonstrated good applicability.
In this study, we have for the first time utilized ML techniques to establish a statistical model that accurately predicts the risk of BM in EC patients. Compared to traditional nomogram prediction models (such as those constructed by Bei Yuan and Liming Jiang),29,30 our model demonstrates higher AUC values. The improved predictive ability of this model holds significant importance for clinical practice, as it can help doctors identify the risk of bone metastases earlier during the initial diagnosis of patients. Early identification facilitates the development of more personalized treatment plans, enhancing patient prognosis and quality of life. For instance, based on the model's predictions, doctors can monitor high-risk patients more frequently or adopt more aggressive treatment measures, potentially delaying the onset of bone metastases or mitigating their impact. However, there are limitations to this study. Firstly, the training and validation data for the model primarily originate from the SEER database, which encompasses a relatively restricted range of variables. Notably, detailed information on chemotherapy regimens and dosages is lacking in the SEER database, potentially impacting the real-world accuracy of our model. For instance, certain chemotherapy drugs may influence the risk of bone metastases; however, these factors have not been incorporated into our current model. Secondly, internal validation alone has been conducted for this study without external validation. Although internal validation results demonstrate favorable predictive ability, it is essential to confirm the generalizability of our model across diverse populations through future research involving external validation in different medical institutions and populations to ensure reliability and effectiveness. Additionally, selection bias exists within the SEER database concerning patients’ races, age groups, or disease stages that might limit the predictive ability due to insufficient sample diversity. Therefore, future research should consider incorporating sample diversity during model development and validation processes to enhance generalizability. Despite these limitations acknowledged hereinabove, our developed predictive model exhibits significant advantages in forecasting BM risk in EC patients; nevertheless further refinement and extensive verification are warranted for improved accuracy and generalizability of our proposed approach towards providing more effective support for prevention and treatment strategies among EC patients.
Conclusion
This study developed seven predictive models based on ML, among which the LR algorithm demonstrated the best overall performance among all candidate algorithms. Based on this algorithm, we constructed a web-based predictor specifically designed for BM in EC. This predictor is capable of effectively identifying the risk of BM in EC patients, thereby assisting clinicians in making more precise decisions in actual clinical practice. By providing early warnings and personalized treatment recommendations, this tool has the potential to significantly improve the treatment outcomes and prognosis management of EC patients.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251325960 - Supplemental material for Use machine learning to predict bone metastasis of esophageal cancer: A population-based study
Supplemental material, sj-docx-1-dhj-10.1177_20552076251325960 for Use machine learning to predict bone metastasis of esophageal cancer: A population-based study by Jun Wan and Jia Zhou in DIGITAL HEALTH
Supplemental Material
sj-docx-2-dhj-10.1177_20552076251325960 - Supplemental material for Use machine learning to predict bone metastasis of esophageal cancer: A population-based study
Supplemental material, sj-docx-2-dhj-10.1177_20552076251325960 for Use machine learning to predict bone metastasis of esophageal cancer: A population-based study by Jun Wan and Jia Zhou in DIGITAL HEALTH
Supplemental Material
sj-jpg-3-dhj-10.1177_20552076251325960 - Supplemental material for Use machine learning to predict bone metastasis of esophageal cancer: A population-based study
Supplemental material, sj-jpg-3-dhj-10.1177_20552076251325960 for Use machine learning to predict bone metastasis of esophageal cancer: A population-based study by Jun Wan and Jia Zhou in DIGITAL HEALTH
Supplemental Material
sj-jpg-4-dhj-10.1177_20552076251325960 - Supplemental material for Use machine learning to predict bone metastasis of esophageal cancer: A population-based study
Supplemental material, sj-jpg-4-dhj-10.1177_20552076251325960 for Use machine learning to predict bone metastasis of esophageal cancer: A population-based study by Jun Wan and Jia Zhou in DIGITAL HEALTH
Footnotes
Acknowledgements
I would like to express my gratitude to all the patients, investigators, and institutions involved in this study, with special thanks to the SEER database.
Availability of data and materials
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
Authors’ contributions
Wan Jun and Zhou Jia are responsible for the entire work and bear the responsibility for all aspects.
Consent for publication
The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Competing interests
The authors declare that they have no competing interests
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical disclosure statement
The authors stated that no experiments involving humans or animals were conducted in this study.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
