Abstract
Background
Personalized medicine has gained more attention for cancer precision treatment due to patient genetic heterogeneity in recent years. However, predicting the efficacy of antitumor drugs in advance remains a significant challenge to achieve this task.
Objective
This study aims to predict the efficacy of antitumor drugs in individual cancer patients based on clinical data.
Methods
This paper proposes to predict personalized antitumor drug efficacy based on clinical data. Specifically, we encode the clinical text of cancer patients as a probability distribution vector in hidden topics space using the Latent Dirichlet Allocation (LDA) model, named LDA representation. Then, a neural network is designed, and the LDA representation is input into the neural network to predict drug response in cancer patients treated with platinum drugs. To evaluate the effectiveness of the proposed method, we gathered and organized clinical records of lung and bowel cancer patients who underwent platinum-based treatment. The prediction performance is assessed using the following metrics: Precision, Recall, F1-score, Accuracy, and Area Under the ROC Curve (AUC).
Results
The study analyzed a dataset of 958 patients with non-small cell cancer treated with antitumor drugs. The proposed method achieved a stratified 5-fold cross-validation average Precision of 0.81, Recall of 0.89, F1-score of 0.85, Accuracy of 0.77, and AUC of 0.81 for cisplatin efficacy prediction on the data, which most are better than those of previous methods. Of these, the AUC value is at least 4% higher than those of the previous. At the same time, the superior result over the previous method persisted on an independent dataset of 266 bowel cancer patients, showing the generalizability of the proposed method. These results demonstrate the potential value of precise tumor treatment in clinical practice.
Conclusions
Combining LDA and neural networks can help predict the efficacy of antitumor drugs based on clinical text. Our approach outperforms previous methods in predicting drug clinical efficacy.
Introduction
Conventional cancer treatment strategies typically treat the same disease with the same treatment. Due to the heterogeneity of individuals, cancer patients with the same cancer often exhibit different responses to the same anticancer drugs in cancer treatment. 1 Drug efficacy prediction makes precision medicine possible, which can improve patient's treatment outcomes by recommending the most proper drugs to individual patients of cancer. 2
Machine learning3,4 has grown substantially in recent years in the field of precision medicine. Different from traditional approaches, they can uncover latent patterns that may elude even the most experienced clinicians in large amounts of data. Doaa et al. 5 developed a machine learning model to predict pelvic tilt and lumbar angle, which may contribute to improving the quality of life for women with urinary incontinence and sexual dysfunction. Doaa et al. 6 also explored machine and deep learning techniques to predict changes in core muscles during female sexual dysfunction (FSD) effectively, which can help women with FSD achieve optimal sexual health. Entesar et al. 7 proposed an approach using Convolutional Neural Networks (NNs) to classify monkeypox skin lesions. Results demonstrated that it can improve the accuracy and efficiency of monkeypox diagnosis and surveillance.
Currently, most methods are developed based on the genomic data of patients for predicting drug efficacy.8–11 For example, Peng et al. 12 proposed to predict drug response based on multiple omics data and a Graph Convolution Network. Tarek et al. 13 presented a machine learning framework based on omics data to aid physicians in choosing the best cancer drug combinations. The genomic data, however, are expensive to measure for clinical use. Compared with genomic data, clinical data, including patient's demographics, symptoms and signs, laboratory test results, radiology examination, etc., are inexpensive and very large in amount. In particular, imaging technology, such as computed tomography (CT), magnetic resonance imaging (MRI), and B-mode ultrasound (BU), can visually view the shape and volume of the pathological tissue, helping doctors better judge the disease's progress and prognosis. 14 Generally, the imaging-based examination results are recorded in electronic medical records (EMRs) in an unstructured text format. The information in clinical texts is crucial for understanding the underlying disease processes and predicting the most appropriate drug therapy. By analyzing unstructured texts, researchers and physicians can gain insights into the specific characteristics of a patient's condition, enabling more personalized drug predictions. 15 As EMR technology develops rapidly, the amount of the resulting radiomic examination report data constantly increases, which allows for the large-scale analysis of patient records. Recently, we have argued that the radiometric examination reports could provide an alternative low-cost way to predict personalized antitumor drug efficacy.
In this paper, we present a new method for predicting the efficacy of antitumor drugs. This method is based on clinical text big data. It uses the Latent Dirichlet Allocation (LDA) model 16 to extract semantic features related to drug efficacy in text data, obtain low-dimensional feature vectors for drug efficacy prediction, and predict the efficacy of antitumor drugs combined with a NN 17 classifier. To test the effectiveness of this method, we collaborated with a tertiary-class hospital in China to apply it to the clinical efficacy prediction management of patients with non-small cell lung cancer (NSCLC). The experimental results show that our method has the potential value of adjuvant clinical diagnosis and treatment. The contributions of our work can be listed as follows: (1) The clinical text of radiomic examination can be used to predict personalized antitumor drug efficacy. (2) We design a novel method for predicting drug efficacy by combining NNs with topic model analysis of clinical text. (3) Our method provides an effective and low-cost way to drug efficacy prediction for cancer precision medicine.
Materials and methods
Figure 1 shows the flowchart of the proposed drug efficacy prediction method based on clinical text big data. The specific algorithm steps are as follows: First, the clinical text data of the patients are preprocessed by deleting numbers and letters, segmenting words, and removing stop words. Second, we obtain the optimal number of topics by minimizing perplexity and use the LDA model for each patient's text to extract the underlying topic and encode the text data for each patient. Finally, the personalized drug efficacy prediction model represented by text topic was established using the NNs model.

The flowchart of our drug efficacy prediction method. Data preprocessing: preprocessing patient texts by deleting numbers and letters, segmenting words, and removing stop words. Word embedding: getting the optimal number of topics by modeling perplexity and encoding patient texts using the LDA model. Drug efficacy prediction: predicting outcome results using neural networks.
Latent Dirichlet Allocation model
The LDA model 18 is an unsupervised machine learning model that can be used to mine valuable information hidden in massive document sets or corpora. It assumes that documents are composed of multiple topics, where each topic is characterized by a word distribution. The model has been widely used in many fields, such as natural language processing, biomedicine, personalized recommendation, and so on. 19
Latent Dirichlet Allocation is a document topic generation model proposed by David Blei et al.
20
It is a three-layer Bayesian probability model,
21
including word, topic, and document. Its basic idea is that each document represents a probability distribution of several issues, and each case represents a probability distribution of many words. It has four main parameters: the document-topic distribution
Encoding patient clinical text data based on the LDA model
We encode the imageology examination text for each patient before treatment. Assuming 

The architecture of LDA model.
Then, from equation (1), eliminate hidden variables and obtain the generation probability of each document W:

The optimal parameters and potential variables are solved through EM algorithm.
22
Following equation (3), we use 


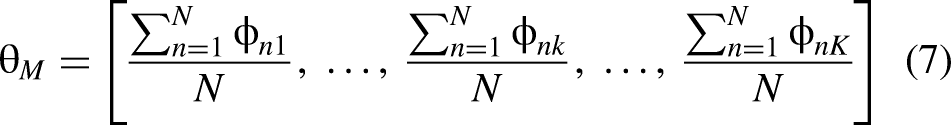

Neural network-drug efficacy prediction model
Neural networks, also referred to as artificial NNs, are computational models that mimic the behavior of biological NNs. They are composed of interconnected nodes, called neurons, that process information using a weighted sum of inputs and pass the result through an activation function to produce an output. Neural networks are typically organized into layers, including an input layer, hidden layers, and an output layer. Each layer consists of a set of neurons. Neurons apply a nonlinear transformation to their inputs using activation functions such as sigmoid, ReLU, or tanh. This allows the network to learn and represent complex patterns. The advantages of NN include the fact that they can model complex nonlinear relationships between input and output variables, making them suitable for a wide range of tasks. They can scale to large datasets and complex problems, as they can be trained on massive amounts of data and contain millions of parameters. They provide a powerful and flexible framework for modeling complex real-world problems, enabling researchers to achieve state-of-the-art results in various domains.
After extracting features based on the LDA model, we construct binary NN classifiers for evaluation. We consider the prediction task a binary classification problem: responsive (1) or nonresponsive (0). During the training process, we use the grid search algorithm 23 to determine the optimal hyperparameters for the models. In NNs, the hyperparameters include the number of layers in the network, the number of neurons in each layer, the number of learning rounds, the selection of cost functions, the dropout rate, and so on. 24 We attempted different combinations of hyperparameters and chose the optimal value as the experimental parameter. The architecture of our model is shown in Figure 3.
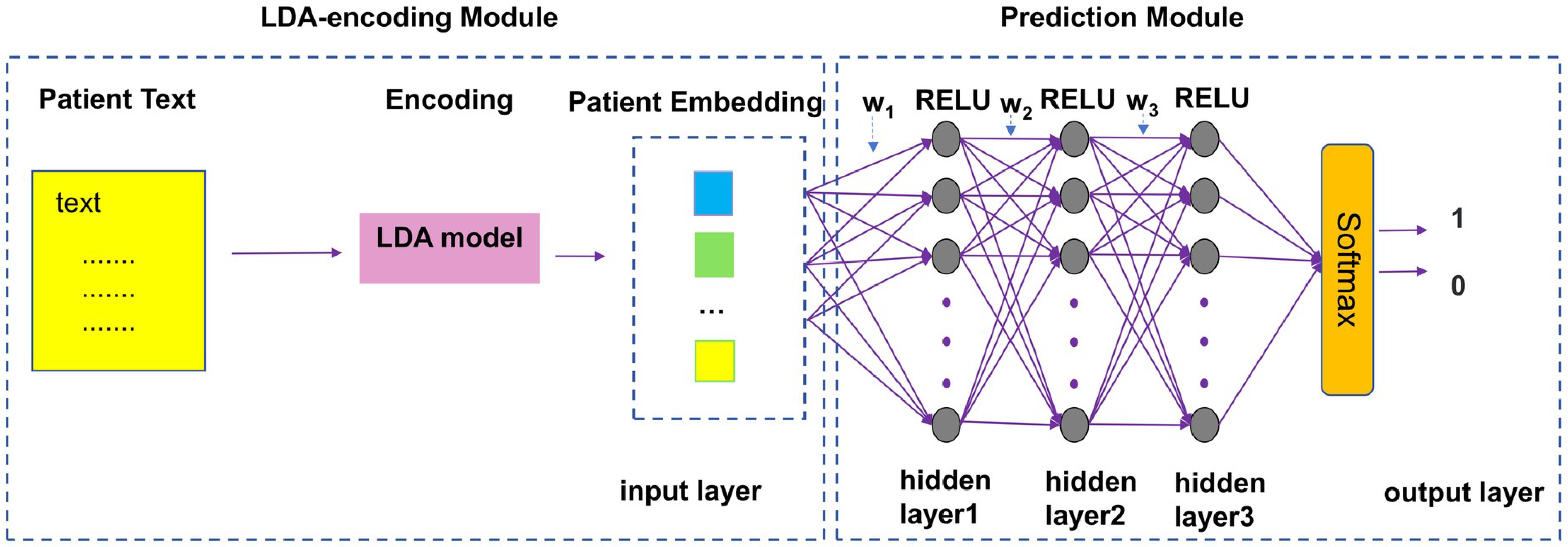
The architecture of our model. This model consists of an LDA-encoding module and a prediction module. The LDA-encoding module encodes patient texts using the LDA model, while the prediction module predicts outcome results using the deep neural network model.
Evaluation metrics
We use five metrics to measure the performance of the models, namely precision (Prec), recall (Rec), F1 value, accuracy (Acc), and area under the receiver operating curve (AUC).
25
In particular, the larger the AUC value, the better the model's classification performance, and AUC is defined as the area under the receiver operating characteristic (ROC) curve. The other metrics can be calculated as follows: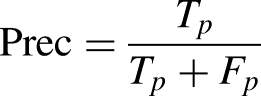


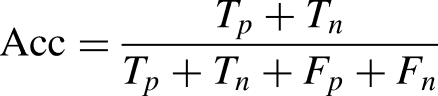
where
Datasets
We screened patients using the following criteria: (1) patients with NSCLC and who had received first-line platinum-based chemotherapy; (2) patients with treatment outcomes available after one or two courses of treatment. For the screened patients, we collected the radiomic examination text data before treatment, including CT, MRI, and BU. We labeled patients with a binary efficacy label (responsive or nonresponsive) for platinum drug. Table 1 provides a summary of the patients used in our experiment. The experimental data set was derived from actual clinical data from a hospital in China. The dataset included 958 patients with NSCLC 26 receiving first-line treatment with platinum-based drugs. Data for each patient consisted of text data from pre-chemotherapy imaging (CT, MRI, BU) examination reports and drug efficacy ratings. Efficacy rating was referred to RECIST criteria (Response Evaluation Criteria in Solid Tumors). The RECIST criteria classify tumor treatment effects as complete response (CR), partial response (PR), stable disease (SD), or disease progression (PD). Only the binary responses of efficacy, responsive (1) and nonresponsive (0), were considered in our experiment. Efficacy grade CR, PR, and SD were classified as reactivity, and efficacy grade PD was classified as nonreactivity; 958 patient samples were classified, of which 691 were responsive and 267 were nonresponsive. The numbers of patients in the training set and the test set are 764 and 194, respectively. The clinical data of the patients are provided in Supplementary Table S1 for reference.
Summary of lung cancer clinical dataset.

Results
Word embedding
We preprocessed the text of the patient's imaging examination before word embedding. 27 First, we removed unrelated characters, including numbers and letters, participles, and stop words consisting of prepositions, quantifiers, and nonmedical nouns. Each patient's imaging examination text was then cut into individual words using the Jieba word segmentation technique. 28 Finally, all the patient text data in the dataset were divided into 2372 words.
We represented each patient's text as a vector using word embeddings. Then, we ran LDA on a dataset containing 958 patients and obtained the optimal number of hidden topics by varying the perplexity. 29 After a series of practices, we chose 16 as the number of hidden topics based on the best performance. Figure 4 shows the variation curve of the degree of confusion with the number of topics. As shown in Figure 4, the lowest perplexity is the optimal topic number. Finally, the word embedding vector of each patient's text data was obtained using the LDA topic model.
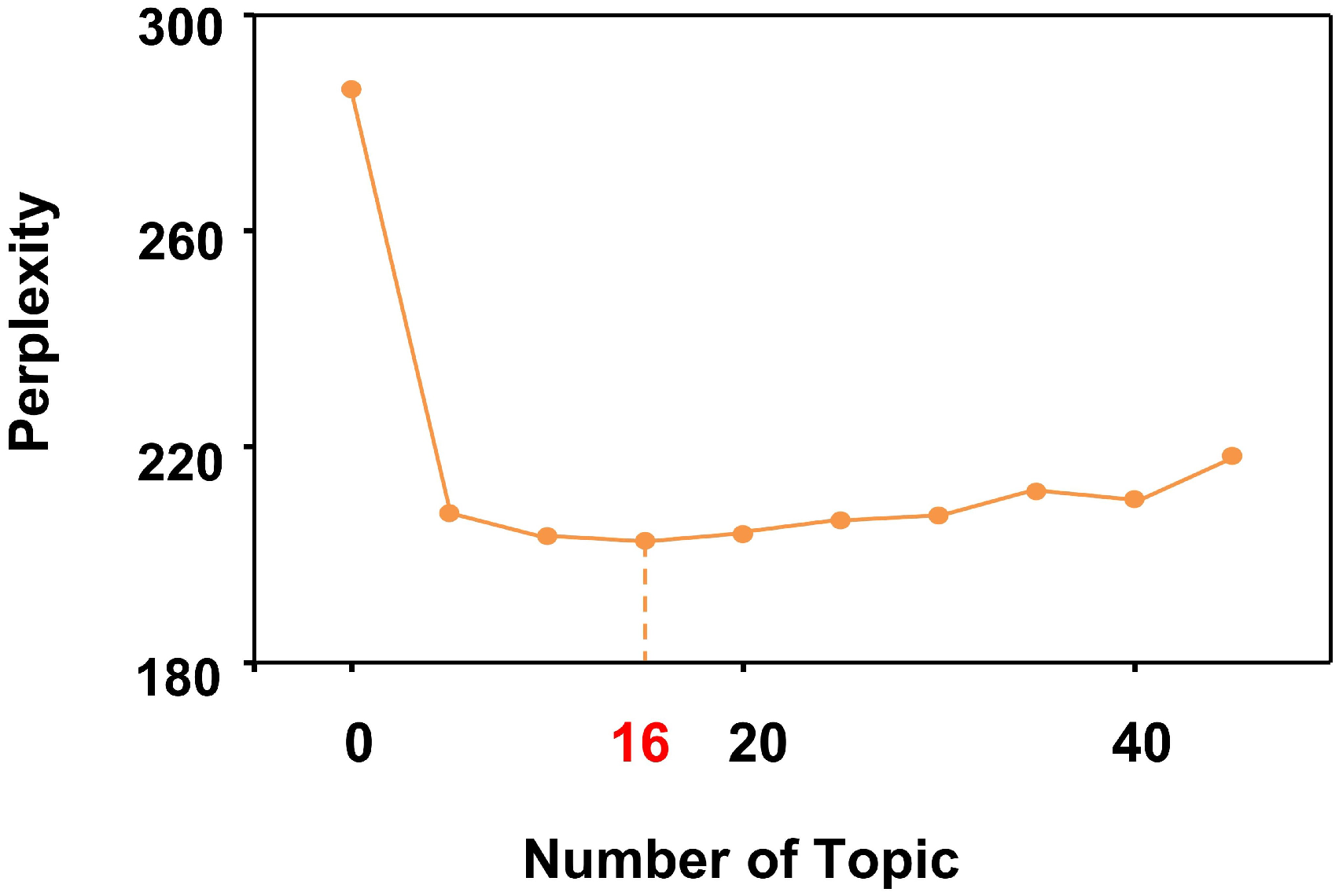
Perplexity-topic number curve.
Parameter setting
For the LDA model, the number of topics was set to 16 by the number optimization result, and α and β were set to 0.0625. At the same time, for the NN, the parameters were optimized as follows: We have tried various network structures, including 300-400-500, 500-400-300, 500-300-500, 500-500-500, 500-500-500-500-500. As a result, the best performance was obtained when the network structure was set to 500-500-500. We used AdaGard to optimize the model's parameters and the nonlinear rectification (ReLU) activation function. Considering the overfitting issue, we added dropout regularization with a dropout rate of 0.5. We utilize binary cross entropy loss function to train our model. The main parameters used in the model are listed in Table 2.
Parameters used in our model.

Prediction performance
Considering the imbalance of positive and negative samples in the dataset, we used stratified 5-fold cross-validation to evaluate model performance. 30 Specifically, the entire dataset was divided into five equally sized folds, with the proportion of each category in each fold being the same as that in the whole dataset.
To compare with our methods, we also applied previous classifiers, namely Logistic Regression, K-Nearest Neighbor, Decision Tree, and Support Vector Machines to the dataset. In the validation, we use a grid search algorithm and an internal stratified 5-fold cross-validation on the training set to find the optimal model parameters for each classifier. Table 3 shows the results of the experiment. From this table, we can see that all five classifiers resulted in AUCs of higher than 70%, of which our method achieved the best drug efficacy prediction performance: a recall of 0.89, an F1 value of 0.85, an accuracy of 0.77 and an AUC value of 0.81, The evaluation experiment on real clinical text datasets demonstrated that our approach is excellent at predicting the efficacy of personalized drugs in individual cancer patients, it achieves absolute improvements of 4% in AUC upon state of the art, suggesting the effectiveness of our method. Figure 5 illustrates the ROC curves of these methods and the training loss curves of the proposed model in the stratified 5-fold cross-validation. As the number of training epochs increases, the training loss gradually decreases and eventually reaches stability, indicating the convergence of the model.

The ROC curves of five methods: (a) LR (b) KNN (c) DT (d) SVM (e) our method, and (f) the train loss curves of our method in the stratified 5-fold cross-validation.
Performance comparison (mean ± SD) with previous methods on lung cancer patients dataset.

The computation cost is important in clinical applications. Our method's average training time is 1.53 s, and the prediction time for a sample takes less than 0.005 s. This suggests that it can be potentially useful in clinical practice.
Comparative experiments of different representation models
We used the Term Frequency-Inverse Document Frequency (TF-IDF) 31 and Graph Convolutional Neural Network (GCN) 32 models to verify the validity of LDA characterization. The TF-IDF model is a widely used statistical method in natural language processing and information retrieval. It provides a numerical score that reflects the importance of a word in a document or a set of documents. One typical application of GCN in natural language processing is modeling syntactic and semantic dependencies in text. It enables learning representations of nodes, edges, and entire graphs by capturing both their feature information and structural information. For TF-IDF, we selected the 16 terms with the highest TF-IDF value. In the GCN model experiment, we initialized the word using the bag of words model, 33 and then learned the word's embedding. The bag of words model counts the number of times each word appears in the text. In the experiment, the Count Vectorizer function of the scikit-learn library 34 was used to convert text into word frequency vectors. Its working principle is based on two key steps: tokenization and frequency counting. In the tokenization step, the text in each document is segmented or broken down into smaller tokens. After tokenization, Count Vectorizer counts the frequency of each token in each document. It generates a vocabulary, which is a set of all unique tokens found in the collection of documents. Each token in the vocabulary is assigned a unique index or feature ID. The experiment trained a two-layer Graph Convolutional NN. All the tokenized words were treated as nodes. The edges between nodes were defined as if two words appeared in the same document, there was an edge between them with a weight of 1. The parameters for the graph convolutional filters were set as follows: dropout rate: 0.5; optimizer: Adam; loss function: cross-entropy loss function; regularization technique: L2 regularization; learning rate: 0.01; the number of convolutional layers: 2. The experimental results are shown in Table 4. The results showed that the LDA model was superior to the TF-IDF and GCN models in all indexes, indicating that LDA performs well in extracting information related to drug efficacy from clinical text data.
The experiment used three different text embedding models (LDA, TF-IDF, and GCN) and the same classifier (NN). The result shows that LDA model achieved the best results on all metrics.

Statistical significance analysis
The permutation test
35
further verified the statistical significance of our method. In the permutation test, it is assumed that there is no difference between the actual result and the random result. The label of the sample of the stratified 5-fold cross-validation training set is kept unchanged, while the sample label of the test set is randomly scrambled, and then the model is retrained and predicted on this basis. This process is repeated 1000 times, with the proportion of AUC values more significant than the actual situation (0.81) as the

Distribution statistics of AUC values for random results.
Evaluation on an independent test set
Further, we used an independent data set from another type of cancer to verify the proposed approach's validity. 36 The dataset consists of 266 patients with bowel cancer whose first-line treatment was platinum-based chemotherapy. Data for each patient was composed of pre-chemotherapy imaging reports. Of the 266 patient samples, 225 were responsive, and 41 were nonresponsive. To compare with previous methods, four previous methods were also applied to the independent dataset. Table 5 shows the results of the experiment. From this table, it can be clearly seen that our method achieved the best drug efficacy prediction performance: a precision of 0.85, a recall rate of 0.93, an F1 value of 0.89, an accuracy of 0.8, and an AUC value of 0.6, showing that our proposed method is effective in predicting the therapeutic effect of bowel cancer.
Performance comparison (mean ± SD) with previous methods on bowel cancer patients dataset.

Discussion
The current study has successfully demonstrated the potential of combining LDA and NNs to predict the efficacy of antitumor drugs based on clinical data. The results achieved on both NSCLC and bowel cancer datasets are significantly effective, indicating the potential of clinical text data in drug efficacy prediction. The LDA model has proven to be more effective in mining the hidden information behind clinical text data compared to TF-IDF and GCN. At the same time, the NN classifier has surpassed the performance of traditional classifiers.
One of the key strengths of our approach lies in the use of LDA to encode clinical text into probability distribution vectors with hidden topics. This approach allows us to capture the latent semantic structure of the clinical text, which is often complex and contains valuable information relevant to drug efficacy. By transforming the text into a numerical representation, we are able to feed it into the NN model and utilize its ability to learn complex patterns and relationships hidden behind data.
The NN model used in this study accurately predicted drug efficacy. However, the model's performance could be further improved by incorporating additional sources of information, such as genomic or proteomic data, which could provide a more comprehensive picture of the patient's disease and response to treatment.
Limitations
There are several limitations in this study. First, the performance of the model depends heavily on the quality and quantity of the clinical data available. Incomplete or noisy data could impact the model's ability to make accurate predictions. Second, the model was trained and evaluated on a relatively small dataset, which could limit its generalization to a wider range of cancer types and patients. Lastly, the performance of the method can be further improved by introducing more advanced word segmentation techniques and optimizing the NN architecture.
Conclusions
We have proposed a method for predicting drug efficacy by combining LDA and NNs based on clinic text data. The evaluation experiment on real clinical text datasets demonstrated that our approach is excellent at predicting the efficacy of personalized drugs in individual cancer patients, compared with previous methods. Specifically, our method achieved a stratified 5-fold cross-validation average Precision of 0.81, Recall of 0.89, F1-score of 0.85, Accuracy of 0.77, and AUC of 0.81 for cisplatin efficacy prediction on the data, which most are better than those of previous methods. Of these, the AUC value is at least 4% higher than those of the previous. The advantage should lie in better mining hidden patterns associated with treatment outcomes from the clinical text by combining topic representation and NNs. In addition, the performance on the independent bowel data set further confirms that clinical text data has potential clinical application value in drug efficacy prediction. Future work will be focused on introducing large-scale clinic data for better drug efficacy prediction, as well as developing a clinical application system to assist doctors in personalized medicine decision-making.
Supplemental Material
sj-xlsx-1-dhj-10.1177_20552076241280103 - Supplemental material for A method combining LDA and neural networks for antitumor drug efficacy prediction
Supplemental material, sj-xlsx-1-dhj-10.1177_20552076241280103 for A method combining LDA and neural networks for antitumor drug efficacy prediction by Weiwei Zhu, Lei Zhang, Xiaodong Jiang, Peng Zhou, Xinping Xie and Hongqiang Wang in DIGITAL HEALTH
Footnotes
Acknowledgements
The authors thank the anonymous Reviewers for their careful reading of our manuscript and their many insightful comments.
Contributorship
Hongqiang Wang, Xinping Xie, and Weiwei Zhu conceived the idea of the study and wrote the main manuscript text; Xiaodong Jiang, Lei Zhang, and Peng Zhou performed data collection, data analysis, and system programming. Xinping Xie and Hongqiang Wang reviewed the manuscript.
Ethical approval
This study was reviewed and approved by the Ethics Committee of The First Affiliated Hospital of University of Science and Technology of China, with the approval number: 2021-RE-85. Informed consent was not required for this study because the data are anonymized.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the University Science Research Project of the Education Department of Anhui Province, Anhui Province’s key Research and Development Project, National Natural Science Foundation of China, Laboratory of Operations Research and Data Science of Anhui Jianzhu University, National Natural Science Foundation of China, Introduction of high-level talent research funding projects of Hefei Normal University, (grant number KJ2021A0633, 201904a07020092, 81872276, YCSJ2024ZR02, 61973295, 60423018).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
