Abstract
Medical text classification, as a fundamental medical natural language processing task, aims to identify the categories to which a short medical text belongs. Current research has focused on performing the medical text classification task using a pre-training language model through fine-tuning. However, this paradigm introduces additional parameters when training extra classifiers. Recent studies have shown that the “prompt-tuning” paradigm induces better performance in many natural language processing tasks because it bridges the gap between pre-training goals and downstream tasks. The main idea of prompt-tuning is to transform binary or multi-classification tasks into mask prediction tasks by fully exploiting the features learned by pre-training language models. This study explores, for the first time, how to classify medical texts using a discriminative pre-training language model called ERNIE-Health through prompt-tuning. Specifically, we attempt to perform prompt-tuning based on the multi-token selection task, which is a pre-training task of ERNIE-Health. The raw text is wrapped into a new sequence with a template in which the category label is replaced by a [UNK] token. The model is then trained to calculate the probability distribution of the candidate categories. Our method is tested on the KUAKE-Question Intention Classification and CHiP-Clinical Trial Criterion datasets and obtains the accuracy values of 0.866 and 0.861. In addition, the loss values of our model decrease faster throughout the training period compared to the fine-tuning. The experimental results provide valuable insights to the community and suggest that prompt-tuning can be a promising approach to improve the performance of pre-training models in domain-specific tasks.
Keywords
Introduction
Medical texts such as medical literature and electronic medical records, contain valuable medical knowledge, including the symptoms, diagnosis, and medications of a particular disease. 1 Since learning such knowledge by human experts can be labor-intensive, natural language processing (NLP) has increasingly influenced the medical information research.2,3 Medical text classification (MTC), as a fundamental medical NLP task, aims to identify the categories to which a short medical text belongs, such as disease stage, allergy intolerance, and organ status. The classification results affect the performance of downstream tasks, such as the detection of adverse medical events 4 or the construction of a clinical decision support system (CDSS). 5
For the MTC task, previous research has established that the performance of machine learning 6 and statistical approaches 7 is highly dependent on the quality of feature engineering. In contrast, deep learning models, such as the convolutional neural network (CNN) 8 or the recurrent neural network (RNN), 9 can provide improved performance without additional manual feature selection. However, regardless of the approach being used, its essence is still to generate representations with abstract semantics and predict the categories based on them. Thus, considering that such models can only capture the semantic knowledge within training sets, they are unsatisfactory since the degree of improvement is still limited by the accessibility of labeled training sets. Therefore, how to enrich the semantic knowledge contained in the representations is the key to improving the performance of MTC.
Much of the current literature in NLP pays special attention to pre-training language models (PLMs), such as bidirectional encoder representations from transformers (BERT), enhanced language representation with informative entities (ERNIE), and ELECTRA.10–12 Researchers have conducted extensive studies and suggested that PLMs have acquired rich prior semantic knowledge during the pre-training process, such as the masked language model (MLM) and next sentence prediction (NSP).13,14 In the early stages of exploiting such knowledge for the MTC, researchers added additional classifiers on the top of PLMs and fine-tuned both of them using task-specific objective functions.15,16 However, one major problem in such a “fine-tuning” paradigm is that additional parameters are introduced when tuning the extra classifiers. Therefore, how to bridge the gap between pre-training objectives and classification tasks has been considered as a critical factor in making a PLM more suitable for MTC. 17
Recently, a number of studies have demonstrated that the “prompt-tuning” paradigm of PLMs induces better performance on a variety of NLP tasks.
18
The core concept of prompt-tuning is to use a pre-training task from the pre-training phase for a particular downstream task. For example, given that BERT uses the masked language model (MLM) during its pre-training phase, prompt-tuning methods based on the BERT would be built around the MLM task, which helps to reduce the difference between the pre-training task and the downstream task. A typical way to achieve such a paradigm is to wrap the input sentence in a natural language template and have a PLM perform the MLM. For instance, the sentence
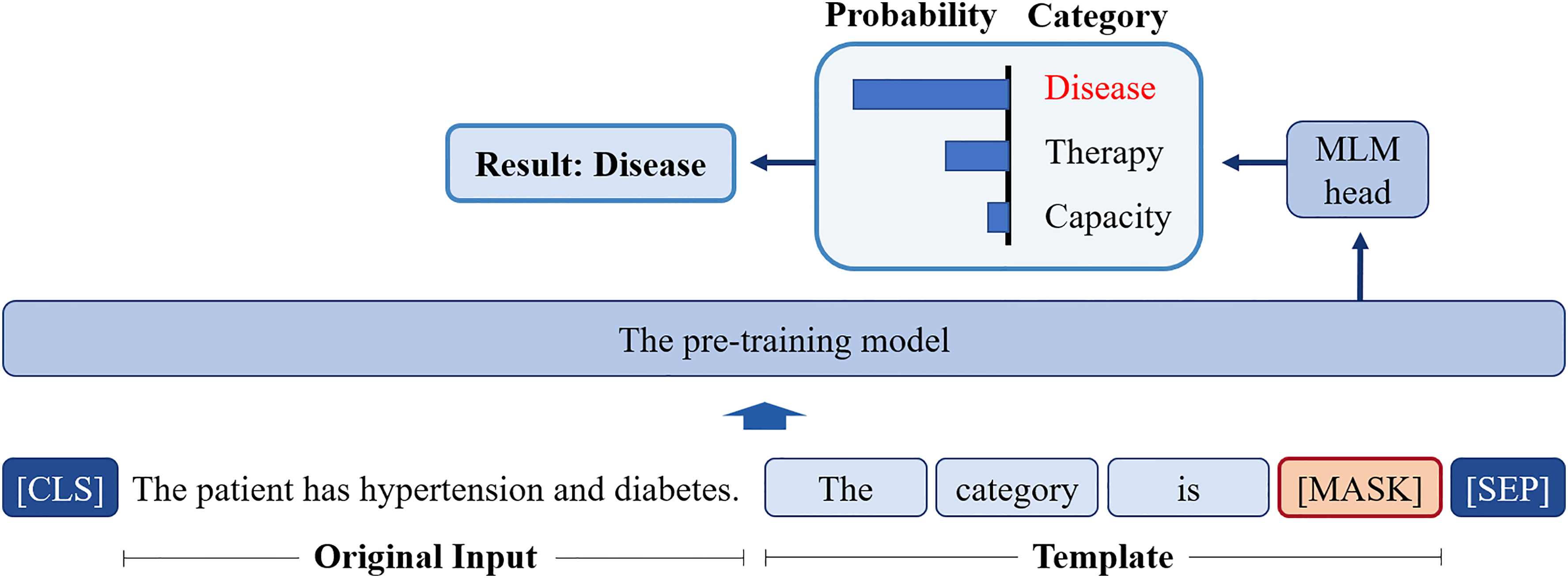
The illustration of prompt-tuning. The category “Disease” can be obtained by predicting the candidate word through masked language model (MLM).
In addition, current research on prompt-tuning has focused more on performing the downstream tasks based on the BERT series PLMs, ignoring that the discriminative PLMs, such as ELECTRA and ERNIE-Health 19 are strong alternatives. Inspired by the application of generative adversarial networks (GANs) in the field of computer vision (CV), 20 the discriminative PLMs aim to design a discriminator to distinguish whether a generator replaces a single token. It has been previously observed that the discriminative PLMs perform better in a variety of NLP tasks, 21 but their properties in the MTC remain unexplored.
The present research explores, for the first time, how to classify medical texts leveraging a discriminative PLM in a prompt-tuning paradigm. This study provides valuable insights to the community and suggests that prompt-tuning, can be a promising approach to improve the performance of pre-training models in domain-specific tasks. Moreover, this study mainly focuses on the fixed or hard prompt-tuning, as these methods are more concise. The main contributions of this study can be summarized as follows:
Firstly, this study provides new insights into MTC using the ERNIE-Health, a discriminative PLM consisting of a generator and a discriminator. ERNIE-Health is specifically designed for the medical domain, which means that it may have a better understanding of medical concepts and generate the representations with more abstract prior semantic knowledge than BERT, a more general-purpose model. This could lead to better performance on tasks that require domain-specific knowledge, such as the MTC task. Secondly, in contrast to the existing prompt-tuning approaches based on MLM, this study attempts to perform prompt-tuning based on the multi-token selection (MTS) task, which is a pre-training task of ERNIE-Health. The rationale for choosing the MTS task for prompt-tuning in our study is based on the core concept of prompt-tuning, which is to use a pre-training task from the pre-training phase to reduce the discrepancy between the pre-training and downstream tasks. Unlike other discriminative PLMs, the discriminator of ERNIE-Health performs the MTS task in addition to the replaced token detection (RTD) task, that is, when the generator replaces a token, the MTS aims to predict the original token from a candidate word set. Specifically, we wrap the raw text into the new input sequence with a template, and use the [UNK] to replace the [MASK], forcing ERNIE-Health to predict the word replaced by [UNK] and achieving the prompt-tuning. In this way, the gap between the pre-training goals and classification tasks is bridged, and the category of a medical text can be inferred from the predicted word without additional parameters or classifiers. Because prompt-tuning is a flexible method that allows for fine-grained control over the input to the model, the proposed method could be beneficial for the tasks that require specific or complex input formats, such as natural language inference or question answering. Thirdly, we test our method on the KUAKE-Question Intention Classification (KUAKE-QIC) and CHiP-Clinical Trial Criterion (CHIP-CTC) datasets, which are sub-tasks in the Chinese Biomedical Language Understanding Evaluation (CBLUE).
22
The experimental results show that the accuracy values of our approach are between 0.866 and 0.861, which outperform the benchmark and previous approaches. In addition, the loss values of our model decrease faster throughout the training period compared to the method based on the fine-tuning paradigm.
In summary, our proposed prompt-tuning approach bridges the gap between pre-training and downstream tasks, leverages prior semantic knowledge, and leads to improved performance on specific tasks. We believe that this approach represents a promising direction for future research on MTC or other NLP tasks using PLMs in the medical domain.
The overall structure of this study takes the form of six sections. A brief review of the related work is presented in the “Related work” section. The “Methods” section deals with the methodology used in this study. The experimental results are presented in the “Experiments and results” section, while the discussion is provided in the “Discussion” section. Finally, the “Conclusions” section concludes this study with a summary.
Related work
Given that the method we propose falls under the theory of supervised learning, this section provides an overview of related work on MTC in supervised learning. Research on machine learning methods for MTC has a long history, with early examples including support vector machine (SVM) and hidden Markov model (HMM). For instance, Sarker and Gonzalez 6 used an SVM-based classifier to automatically detect adverse drug reactions from medical text, while Koopman et al. 23 trained SVM classifiers with detailed features to identify cancer-related causes. Kocbek et al. 24 presented a text-mining system to detect positive cancer admissions, while Yi and Beheshti 7 tested the application of HMM in medical text classification. However, the manual selection of appropriate features remains a key challenge, as it is time-consuming and labor-intensive. Therefore, the existing literature on MTC mainly focuses on identifying the categories based on deep learning and pre-training models.
Deep learning
Deep learning methods have become popular since 2012 with the improvement of computer hardware, especially the graphics processing unit (GPU). Over the past decade, most research in the field of MTC has emphasized the use of two standard deep learning models: the CNNs 8 and RNNs. 9
Convolutional neural networks
The CNN, originally used in CV, 25 consists of convolutional kernels and pooling layers. In NLP, researchers have applied CNNs to generate word embeddings, capturing local features through pooling. Rios and Kavuluru 26 showed that CNNs outperformed previous approaches in biomedical text classification for assigning medical subject headings. Hughes et al. 27 applied CNNs to sentence classification in the Merck Manual dataset, the first study of its kind in medical texts. Nii et al. 28 used CNNs to classify nursing-care texts and extract essential parts for classification. Yao et al. 29 combined rule-based features with a knowledge-guided CNN model trained with word and entity embeddings from Unified Medical Language System.
Recurrent neural networks
RNNs are more suitable than CNNs for sequence modeling, which is crucial for text classification with sequential data. In medical text classification, the long short-term memory (LSTM) and gated recurrent unit (GRU), both RNN variants with gated units, are more widely used due to their better handling of gradient vanishing and explosion. Oleynik et al. 30 evaluated machine learning and LSTM for multi-label binary text classification. Venkataraman et al. 31 trained an RNN to automate clinical record assignment. Liang et al. 32 improved LSTM with a dual-channel mechanism for Chinese medical text classification. Bangyal et al. 33 applied RNN and LSTM to COVID-19 fake news detection, a medical text classification task.
Hybrid methods and attention
A number of studies have also proposed methods that take advantage of both CNNs and RNNs. Zhou et al. 34 used an integrated CNN–RNN framework to extract semantic and sequential features from patient queries. Li et al. 35 proposed a three-stage hybrid method using BiLSTM and a regular expression-based classifier for medical text classification. Ibrahim et al. 36 used a CNN and LSTM in a deep generic learning-based hybrid multi-label classification method. Shin et al. 37 combined the attention mechanism and CNN to classify radiology head CT reports and generated a heat map of attended terms.
As can be seen, the core of these deep learning approaches is to generate representations with semantic knowledge using CNNs or RNNs. However, the semantic knowledge can only be abstracted from the training sets. In other words, the performance of these methods is still limited by the inaccessible labeled training sets.
Pre-training language models
Much of the current literature on medical text classification pays special attention to pre-training models. Inspired by the ImageNet 38 in the field of Computer Version, researchers have proposed pre-training language models, such as BERT, ERNIE, ELECTRA, and ERNIE-Health for NLP. All PLMs adopt a hierarchical architecture consisting of several layers of transformer blocks as shown in Figure 2. Each transformer block contains a self-attention mechanism 39 that allows the model to pay attention to the different parts of the input sequence and to capture long-range dependencies. These models are pre-trained on large-scale text corpora using various pre-training tasks to learn contextual representations of words and sentences. For example, BERT uses the MLM and the NSP task, while discriminative PLMs such as ELECTRA and ERNIE-Health have an additional discriminator to determine whether a token is “Original” or “Replaced,” that is, the Replaced Token Detection (RTD) task, which allows them to generate representations with more abstract semantics. In addition, ERNIE-Health uses a multi-task learning approach with tasks such as RTD and MTS to capture domain-specific knowledge and semantics relevant to the medical domain. As shown in Figure 3, ERNIE-Health consists of a generator and a discriminator. The generator is used to perform an MLM task that infers what the masked word is in the original sentence. The discriminator determines sequentially whether each token has been replaced, that is, the same RTD task as in ELECTRA. However, based on the RTD results, ERNIE-Health also selects the original word of the replaced word from the candidate word set, that is, the MTS task.

The difference between BERT series PLMs and discriminative PLMs. BERT: bidirectional encoder representations from transformers; PLMs: pre-training language models.

The overview of enhanced language representation with informative entities (ERNIE)-Health.
There are two paradigms for transferring prior knowledge to a specific downstream task: fine-tuning and prompt-tuning. For the first one, extra classifiers are added on the top of PLMs, and both are tuned using task-specific objective functions. Almost all current work on MTC based on the PLMs follows this paradigm. As an example, Qasim et al. 15 detected the fake news of COVID-19 by tuning BERT, RoBERTa, and XLNet. Guo et al. 16 evaluated six types of PLMs on social media health-related text classification tasks. An elaborate experiment conducted by Lu et al. 40 verified that BERT performed best in all scenarios for classifying the presence or the absence of 16 diseases from patient discharge summaries. Peng et al. 41 found that the BERT model pre-trained on PubMed abstracts and MIMIC-III clinical notes performed best when tested on the biomedical language understanding evaluation (BLUE) benchmark. However, Gao et al. 42 showed that BERT often fails to outperform these simpler benchmarks when classifying MIMIC-III discharge summaries and cancer pathology reports, limiting the application of transformers for document classification on long clinical texts.
The disadvantage of fine-tuning is that it introduces additional parameters and tunes extra classifiers. Recently, a number of studies have demonstrated that prompt-tuning induces better performance for NLP tasks. In such a paradigm, the input sentence is wrapped into a natural language template in order to bridge the gap between pre-training goals and downstream tasks. For example, Jiang et al. 43 proposed a knowledgeable prompt-tuning method for the fake news detection task. Li et al. 44 proposed a knowledge-injected prompt-tuning model to detect events from the text by identifying and classifying event triggers. To address the problem that the number of labeled texts is small due to the lack of specialized expertise, a prompt-tuning method for multi-label text classification was introduced by Wei et al. 45 However, a literature search revealed that no previous study has attempted to prove that prompt-tuning is also effective in classifying the medical text.
Methods
This section first defines the MTC task, and then the proposed model is introduced in detail.
Problem definition
For the MTC task, the input is a medical related sentence, while the output is the category to which the sentence belongs. Give a dataset
Model architecture
The proposed model is shown in Figure 4, and the selected pre-training model is the discriminator of ERNIE-Health, as it is pre-trained on medical corpora. In contrast to the most common paradigm fine-tuning, we utilize prompt-tuning to formalize the classification task into a multi-token selection problem, which is identical to the pre-training process.
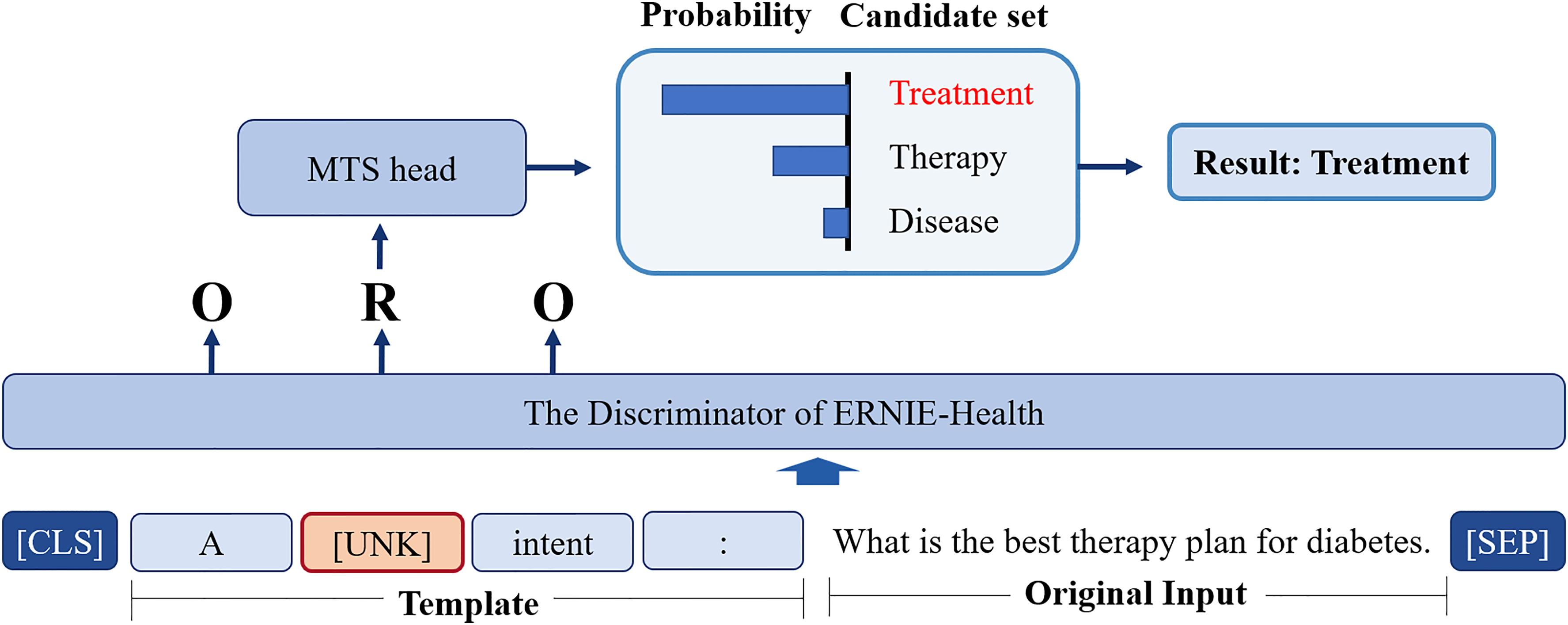
The architecture of the proposed model.
Give a text sequence 
Then, 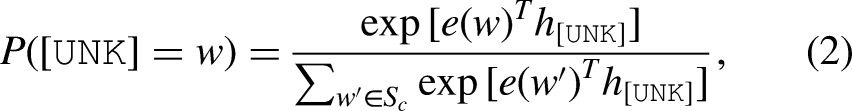

Experiments and results
This section introduces the dataset used in the experiment and shows the performance of the proposed model. The goal of this experiment is to validate the effectiveness of the proposed method by evaluating the classification accuracy of all models listed in Tables 1 to 4 for the MTC task, using the “KUAKE-QIC” and “CHIP-CTC” datasets. Since previous models were based on fine-tuning, we first tested the performance of fine-tuning with ERNIE-Health, and then evaluated its performance with prompt-tuning. The software environment for this experiment is the Paddlepaddle 1 , an end-to-end open-source deep learning platform developed by Baidu. In our experiments, we used a server equipped with an 8-core CPU and a NVIDIA Tesla V100 GPU, and 16 GB of RAM.
The distribution of labels in the CHiP-clinical trial criterion (CHIP-CTC) dataset.

The distribution of labels in the KUAKE-question intention classification (KUAKE-QIC) dataset.

Hyper-parameters.
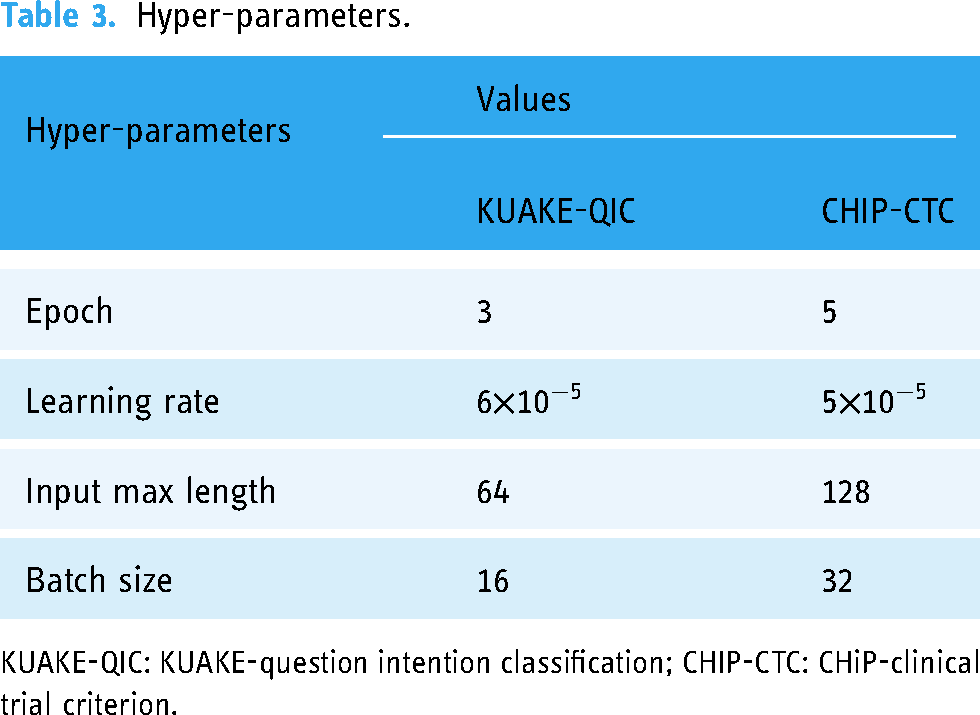
KUAKE-QIC: KUAKE-question intention classification; CHIP-CTC: CHiP-clinical trial criterion.
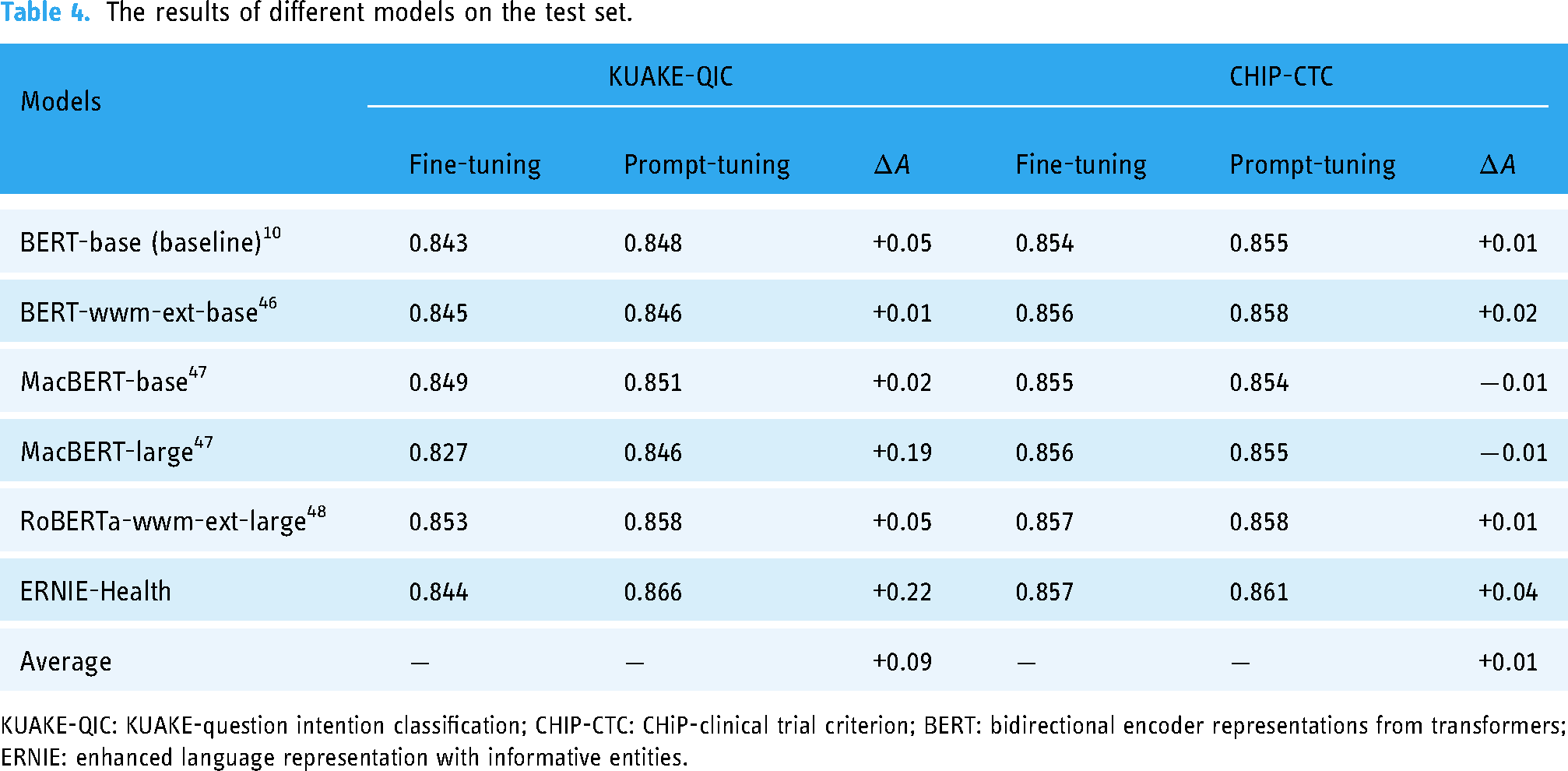
The results of different models on the test set.
KUAKE-QIC: KUAKE-question intention classification; CHIP-CTC: CHiP-clinical trial criterion; BERT: bidirectional encoder representations from transformers; ERNIE: enhanced language representation with informative entities.
Dataset
The datasets used for our experiment are the “KUAKE-QIC” and “CHIP-CTC,” which are sub-tasks of the CBLUE 22 dataset. 2 The KUAKE-QIC contains short Chinese texts related to patient inquiries and has proven to be a valuable resource for the development and evaluation of natural language processing techniques. Each sample in the dataset is assigned a label indicating one of the following intentions: “Diagnosis,” “Cause,” “Method,” “Advice,” “Metric explain,” “Disease express,” “Result,” “Attention,” “Effect,” or “Price.” If the intent of a sample does not fall into one of these categories, it is labeled as“Other.” The dataset provides a diverse range of intentions that a patient may have when seeking medical advice. This diversity allows for a more comprehensive evaluation of the performance about the proposed method on various patient inquiries. The train set contains a total of 6931 samples, and the distribution of the text labels is presented in Table 2. The distribution of labels in the dataset indicates that the most common intention is “Query for method,” which accounts for 25% of the total samples.
The CHIP-CTC dataset contains 44 pre-defined semantic categories for filtering criteria and a set of descriptive sentences for Chinese clinical trial screening criteria. The goal of this task is to determine the specific category for each screening criterion. It can be seen from Table 1 that the training sample size of CHIP-CTC is three times larger than that of KUAKE-QIC, and the number of categories is four times larger.
Hyper-parameters
The hyper-parameters involved in this experiment are listed in Table 3. We used Adam as the optimizer and trained our model for three epochs with a batch size of 16 with KUAKE-QIC, and for five epochs with a batch size of 32 with CHIP-CTC.
Evaluation metrics
We introduce the accuracy to evaluate the performance of the models listed in Table 4. This metric is calculated according to the following formulation where 
Results of different models
First, we tested the performance of some popular PLMs on the KUAKE-QIC dataset, and the experimental results are shown in Table 4, where the BERT-base is the baseline model. As mentioned before, BERT 10 is a PLM that encodes the representations with prior semantic and syntactic knowledge through pre-training tasks MLM and NSP. It is selected as the baseline model in our experiment, and the baseline accuracy is 0.843 with fine-tuning. BERT-wwm-ext-base is an extended version of BERT-base. This PLM is retrained based on the initial checkpoint of the BERT-base and performs the pre-training task on an extended corpus with Whole Word Mask (WWM). Given that the WWM is more suitable for a Chinese MTC because there is no separator between Chinese words, BERT-wwm-ext-base improves the accuracy to 0.845 with fine-tuning. Furthermore, the base and large versions of MacBERT 47 are all extended PLMs of BERT-base. The difference between these two kinds of versions lies in the number of whole parameters and the dimensional size of the hidden layers. The MacBERT optimized the WWM of BERT-wwm-ext-base with N-gram masking strategies, and the two types of versions of MacBERT gets the accuracy of 0.849 and 0.827 with fine-tuning, respectively. Moreover, the RoBERT-large 48 utilizes a random masking strategy and obtains the representations with richer semantics compared to the static masking strategy of BERT. The RoBERTa-wwm-ext-large improved RoBERT-large using WWM, and this PLM obtained the highest accuracy of 0.853 among the PLMs listed in Table 4 except ERNIE-Health.
Given that the ERNIE-Health is pre-trained on medical corpora, it gets an accuracy of 0.844 when fine-tuning, slightly higher than the baseline. The approach we proposed is to classify the medical texts with prompt-tuning using ERNIE-Health, and this model obtains an accuracy of 0.866, which is higher than the result of RoBERTa-wwm-ext-large. Moreover, the architecture of the latter one is more complex than it of our model. The number of transformer block layers and hidden layer sizes are 12 and 768 in our model, while these two indices are 16 and 1024 in RoBERTa-wwm-ext-large, respectively. That means the RoBERTa-wwm-ext-large may need more training time but obtain a lower accuracy value than our model.
Second, we also tested the performance of the PLMs on the CHIP-CTC dataset, and the experimental results are shown in Table 4, where the BERT-base is also the baseline model. The baseline accuracy is 0.854 with fine-tuning. BERT-wwm-ext-base, as an extended version of BERT-base, improves the accuracy to 0.856 with fine-tuning. Furthermore, for the base-version and large-version of MacBERT, 47 the two types of versions of MacBERT get the accuracy of 0.854 and 0.855 with fine-tuning, respectively. Furthermore, the RoBERTa-wwm-ext-large, which is the improved version of the RoBERT-large 48 obtains the highest accuracy of 0.858 among the PLMs listed in Table 4 except ERNIE-Health.
The ERNIE-Health obtains an accuracy of 0.857 when fine-tuning, which is slightly higher than the baseline. The approach we proposed achieves an accuracy of 0.861, which is higher than the result of RoBERTa-wwm-ext-large, but the improvement is modest. Similarly, given that the architecture of the latter one is more complex than that of our model, the RoBERTa-wwm-ext-large may need more training time but obtain a lower accuracy value than our model on the CHIP-CTC dataset, too.
Results of fine-tuning and prompt-tuning
Moreover, the performance of two types of paradigms: fine-tuning and prompt-tuning based on the PLMs listed in Table 4 are tested on both the KUAKE-QIC and CHIP-CTC datasets. The experimental results are shown in Table 4, where all the models based on the prompt-tuning outperform those based on the fine-tuning when testing on KUAKE-QIC, increasing the accuracy with an average value of 0.09. However, the performance of the proposed method on CHIP-CTC is not significant when compared with KUAKE-QIC. Although the prompt-tuning based on ERNIE-Health can still improve the accuracy by 0.04, the average accuracy is only increased by 0.01, and even the accuracy of the MacBERT series models is decreased.
Based on the findings presented in Table 4, it is evident that the performance improvement of prompt-tuning is relatively small for some PLMs. Consequently, we conduct additional experiments to validate the statistical significance of this accuracy improvement. Specifically, we conduct random experiments based on fine-tuning and prompt-tuning, respectively, and the seeds are randomly assigned. Subsequently, a paired
The results of

KUAKE-QIC: KUAKE-question intention classification; CHIP-CTC: CHiP-clinical trial criterion; BERT: bidirectional encoder representations from transformers; ERNIE: enhanced language representation with informative entities.
In addition, we also record the loss-values at each training batch for fine-tuning and prompt-tuning based on the ERNIE-Health. As shown in Figure 5(a), the red line represents the loss-values of fine-tuning and the blue line represents those of prompt-tuning. It is clear from this figure that the prompt-tuning curve decreases more rapidly than the fine-tuning curve, especially in the early stages of training. However, since the loss-values are oscillating and decreasing, we also provide the smooth version of the loss curves to compare the two types of paradigms more clearly, which can be found in Figure 5(b) and (c). The loss curves, as shown in Figure 5(b), show more obvious that the loss-values of prompt-tuning fall faster in the early stages. Furthermore, Figure 5(c) illustrates the results when the loss-values are taken logarithmically and shows that the loss-values of prompt-tuning also decrease faster at the later stage.

(a) The loss curves of fine-tuning and prompt-tuning, (b) the smoothing curves of loss curves, and (c) the smoothing curves of loss curves under logarithm.
Results of different templates
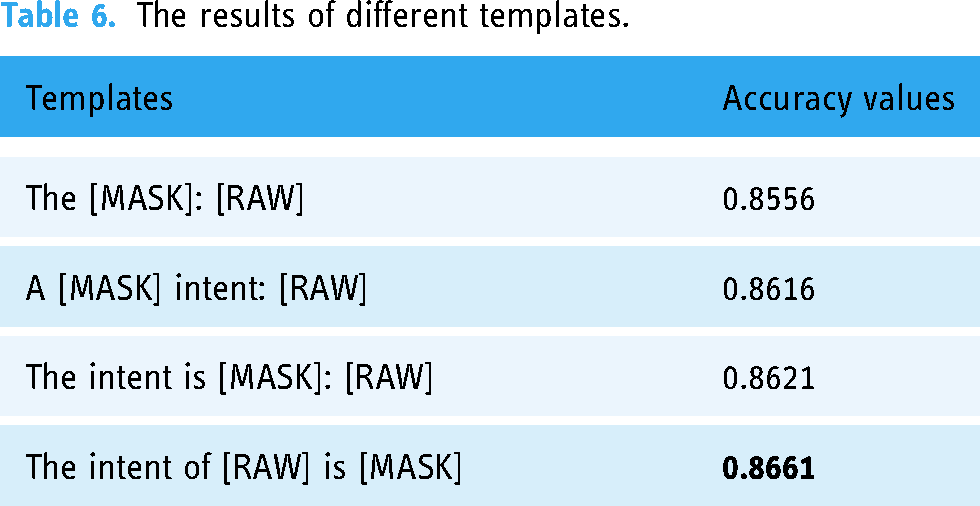
Considering that the templates used in prompt-tuning may affect the performance of text classification, we conducted experiments to test the impact of different templates on the accuracy of text classification, as shown in Table 6. We differentiate these templates based on whether they are closer to the natural language used by humans. From Table 6, we can see that the closer the template is to the natural language used by humans, the higher the accuracy obtained by prompt-tuning.
The results of different templates.
Discussion
We tested our method on the KUAKE-QIC and CHIP-CTC datasets and compared the results with those obtained by some popular PLMs. It can be seen from Table 4 that our method, for example, ERNIE-Health with prompt-tuning, achieves the highest accuracy on both datasets. It can increase the accuracy by 0.08 and 0.03 compared to the RoBERTa-wwm-ext-large. In addition, the total parameters and the hidden layers of our method are smaller than those of RoBERTa-wwm-ext-large. Therefore, our model spends less time on training and is less likely to overfit, which makes it more competitive.
The results presented in Table 4 provide strong evidence that prompt-tuning is an effective method for improving the performance of PLMs. Essentially, the prompt-tuning paradigm still performs a classification task, that is, calculating the probability of the output word corresponding to the [MASK] token in the template. The process still involves computing each word in the entire word list and updating the weights of the PLM which may still be downstream task oriented. However, unlike the fine-tuning paradigm that uses the [CLS] token, prompt-tuning constructs a template with the [MASK] token based on the raw input and then allows the PLM to predict the output. The prompt-tuning paradigm helps bridge the gap between pre-training and downstream tasks, resulting in improved performance on specific tasks. In contrast to the fine-tuning approach adopted by other PLMs listed in Table 4, where additional classifiers are added to the PLMs and both are fine-tuned using task-specific objective functions, our proposed model performs prompt-tuning leveraging the pre-training task: MTS. This approach can be thought of as a kind of MLM, where we wrap the raw text into a new input sequence with a template and train the model to predict the correct token to be replaced by [UNK] in the template. The reason we use [UNK] to replace [MASK] is to force the ERNIE-Health discriminator to select a word from the candidate word set to replace [UNK], thus achieving prompt-tuning.
Importantly, our model is trained based on the initial parameters that are the same as those of the pre-trained ERNIE-Health, allowing us to fully exploit the prior semantic knowledge encoded in ERNIE-Health. Moreover, ERNIE-Health is pre-trained on medical corpora, and its representations contain semantic knowledge relevant to the medical domain. Therefore, our proposed method is particularly well-suited for tasks in the medical domain. We note that the fine-tuning paradigm introduces additional parameters, and the training process starts from scratch, making it computationally expensive and time-consuming. In contrast, our prompt-tuning approach is more efficient and does not require additional parameters to be added to the model. However, we found that the performance improvement of our proposed method on the CHIP-CTC dataset is limited, and in some cases, it is even inferior to fine-tuning. We believe that the reason for this is related to the size of the training set. The number of training samples of CHIP-CTC is four times higher than that of KUAKE-QIC. At this point, the advantage of prompt-tuning in bridging the gap between pre-training tasks and specific downstream tasks may no longer be significant.
Furthermore, the experimental results presented in Table 4 show that the accuracy of ERNIE-Health is slightly higher than that of BERT-base when performing fine-tuning. This observation highlights the importance of pre-training on domain-specific corpora and leveraging prior semantic knowledge when training PLMs for specific downstream tasks.
In addition, we also show the loss curves for ERNIE-Health based on fine-tuning and prompt-tuning. As shown in Figure 5, the loss-values of prompt-tuning decrease faster than those of fine-tuning, especially in the early stages of training. Moreover, when plotting the smooth version of the loss curve and illustrating the logarithmical loss value, another important finding is that the loss-values of prompt-tuning are lower than those of fine-tuning not only in the early stage but also in the later stage. This result can be explained by the fact that our model (ERNIE-Health with prompt-tuning) is trained based on the initial parameters that are the same as those of a trained ERNIE-Health. Therefore, the model can converge faster considering that the fine-tuning paradigm requires additional classifiers to be trained from scratch.
We also conduct random experiments to validate the statistical significance of the accuracy improvement presented in Table 4. The
Finally, we also tested the performance of different templates on prompt-tuning. As shown in Table 6, the closer the template is to natural language, the higher the accuracy obtained. This is again due to the nature of prompt-tuning. The closer the template is to natural language, the more likely the PLM has “seen” it during pre-training. Therefore, the probability of predicting the correct replacement word is higher.
Conclusions
This study provides new insights into classifying medical text using a discriminative PLM with prompt-tuning. The discriminative PLM selected for this study is ERNIE-Health, which is pre-trained on medical corpora. The MTC task is performed following a pre-training task MTS instead of adding additional classifiers as previous methods did. Specifically, we wrap the raw text into a new input sequence with a template and calculate the probability distribution of candidate words corresponding to the [UNK]. The category of a medical text can be inferred by the predicted word without using extra parameters or classifiers. Finally, the experimental results show that our method outperforms the benchmark and previous approaches on both KUAKE-QIC and CHIP-CTC datasets.
However, there are still potential challenges and opportunities for future research. Firstly, our method still relies on the quality and quantity of training data. Insufficient or noisy training data may negatively affect the performance. Secondly, our method may not be robust to noise and errors in the input. In the real world, the input text may contain errors or noise, which may affect the performance. Lastly, the generalizability of our model to other languages and domains is an open research question. In our experiments, we focused on the Chinese language and the medical domain. However, the effectiveness of our model in other languages and domains remains to be explored. For the future work, we can further explore the importance of domain-specific pre-training and prompt engineering for other NLP tasks based on the proposed method, or explore the generalizability of our method to other languages and domains.
Footnotes
Acknowledgements
This research was supported by the Medical Big Data Supercomputing Center System of Anhui Medical University.
Contributorship
YW (Yu Wang) was the lead in conceptualization, methodology, and initial manuscript draft writing. YW (Yuan Wang) conducted the literature review. ZP and FZ prepared the dataset. ZY contributed to code implementation. FY was the lead in funding acquisition and supervision. The manuscript was reviewed and edited by all authors, and the final version was approved.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
This study not involve primary data collection, and formal ethics approval will therefore not be required.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Natural Science Foundation of Anhui Province of China (Nos. 2108085MH303 and 2108085QF274) and the Initiation Fund of Anhui Medical University (No. 1401039201).
Guarantor
FY.
Informed consent
Patient consent is not required because the research does not involve primary data collection and we are using publicly available medical text datasets that do not contain private information.
Notes
Appendix
The appendix provides the details of the random experiments mentioned in the “Results of fine-tuning and prompt-tuning” section. Table 7 displays the results of fine-tuning and prompt-tuning using BERT and ERNIE-Health on the KUAKE-QIC dataset, including the accuracy values,
Based on the experimental findings, it can be observed that the results obtained from eight random experiments satisfy the normality assumption, as evidenced by the
