Abstract
Although databases have been well-defined and thoroughly discussed in the computer science literature, the actual users of databases often have varying definitions and expectations of this essential computational infrastructure. Systems administrators and computer science textbooks may expect databases to be instantiated in a small number of technologies (e.g., relational or graph-based database management systems), but there are numerous examples of databases in non-conventional or unexpected technologies, such as spreadsheets or other assemblages of files linked through code. Consequently, we ask: How do the materialities of non-conventional databases differ from or align with the materialities of conventional relational systems? What properties of the database do the creators of these artifacts invoke in their rhetoric describing these systems—or in the data models underlying these digital objects? To answer these questions, we conducted a close analysis of four non-conventional scientific databases. By examining the materialities of information representation in each case, we show how scholarly communication regimes shape database materialities—
Introduction
What is a database? What do we talk about when we talk about databases? In our work as information scientists who study scholarly data practices and teach information modeling, we have found that these seemingly straightforward questions can be surprisingly hard to answer. Computer scientists and systems administrators often have very different ideas of what constitutes a database than the end-users of these systems. Although computer scientists typically define databases in terms of the information models supporting the system (e.g., Edgar “Ted” Codd’s relational model, or the graph databases of Neo4j or NoSQL systems), users tend to more generously define “database” as any sort of system of data in a computer that is organized for storage and retrieval. For instance, Amy Voida et al. (2011) have described “homebrew” databases built through
Why does it matter what form a database takes? Because “databases make the world”—in that both they are omnipresent and we increasingly treat the world as “something that can be encoded in a database” (Dourish, 2014). Databases are fundamental to the organization and access of data big and small. The specific technology stacks and information models deployed by a database have major implications for its use and maintenance over time. Changes in these materialities of databases both reflect and cause changes to work arrangements and collaborations. Understanding the impacts of specific materialities is crucial to supporting work with and preservation of databases. Furthermore, “examining the materialities of the database form uncovers a range of historical specificities that help contextualize broader arguments about digital media, and also illuminate the significance of contemporary shifts, dislocations, disruptions and evolutions” of media use (Dourish, 2017: 107).
For this article, we conducted a close examination of four exemplar scientific “databases” that do not use traditional database technology but rather rely on spreadsheets, plain text files, and programming languages such as R. We analyzed these databases with careful consideration of the materialities of information at play in their creation and use, and contrasted these with best practices for database construction typically taught in classrooms. We asked: How do the materialities of non-conventional databases differ from or align with the materialities of conventional relational systems? What properties of databases do the creators of these artifacts invoke in their rhetoric describing these systems—or in their underlying data models?
We found that though our database creators have abandoned certain norms of relational database construction, they nevertheless maintain some relationally aligned database practices. By examining the materialities of information representation in each case, we uncovered how scholarly communication regimes shape database materialities—
Our goal in this project was fairly practical in some ways: we wanted to support work with data as it is actually done rather than uncritically advocate for normative best practices. However, we also wished to contribute to broader studies of data materiality—and the role that different materialities play in society and scholarship. We believe that understanding the implications and persistence of relationally aligned data modeling is important in understanding the legacy of relational databases on modern data practices.
Background: The materiality of databases
As originally conceptualized by Codd, a relational database organizes data into tables, which are operated on with programming languages that use mathematical set theory to query across tables (Codd, 1970; see also Elmasri and Navathe, 2010). Codd’s approach has been deeply influential in how database systems are built. Importantly from our point of view, it has also been influential in how databases are explained in textbooks and taught to students.
However, a database can be more broadly defined as “a usually large collection of data organized especially for rapid search and retrieval (as by a computer)” (Merriam-Webster, 2019). Further, many other methods of information organization and retrieval have been developed since Codd’s day. Thus, despite the relational database’s continued dominance in many contexts, modern databases’ specific material forms can vary dramatically. For instance, though all relational databases organize data into sets of interlinked tables, the specific file types and querying languages vary depending on the software platform being used. Emergent “NoSQL” or “schema-less” database platforms store data variously as key-value pairs, collections of tagged documents, or “graphs” of related values—again, depending on the specific software platform being used, and the decisions or needs of the people using them. Several scholars have even argued for the existence of the “database before the computer,” such as the paper-based “compendia” of late 19th century paleontology (Sepkoski, 2017); “slip-boxes” of bibliographic annotations (Krajewski, 2013); collections of 18th century newspaper clippings consulted and re-arranged for the production of the abolitionist book
Lev Manovich, a scholar of digital media and computing, has famously argued that it is the database’s unordered nature at the heart of these disparate forms, and that this sets databases apart from other types of media regardless of the specific technology used to enact the database. Whether a media object uses a relational schema or an object-oriented infrastructure, From the point of view of the user’s experience a large proportion of [new media objects] are databases in a more basic sense. They appear as collections of items on which the user can perform various operations: view, navigate, search. The user experience of such computerised collections is therefore quite distinct from reading a narrative or watching a film or navigating an architectural site. (Manovich, 1999: 81)
While Manovich’s point is taken, we echo prior work on data materiality (Bowker, 2008; Dourish, 2017; Gitelman, 2013; Ribes and Polk, 2015) to note that the specific forms and formats of media objects have major impacts on their use, interpretation, dissemination—as well as the structuring of work and society that goes on around the objects. Manovich’s databases are the new media objects of the late 1990s—“multimedia encyclopedias” and similar compendia made possible by the random access afforded by hypertext, CD-ROMs, and the ability to “browse” a unique path through a digital collection. We do not dispute that these are databases; rather, we note that the experience of using a multimedia CD-ROM differs considerably from that of working with a relational database to query scientific data. Close examination of the materiality of different information systems is crucial to understanding the implications of design choices in said systems.
We draw our definition of materiality from Dourish and Mazmanian (2013), though we note that theirs built on prior work by Manovich (2001), Michael Castelle (2013), and others. In a digital context,
Understanding these cultural practices of information is fundamental to understanding the role that data and data systems play in society more broadly. Data practices are the “ways in which people collect, make sense of and engage” with data and data systems, including the types of “data materializations” that are generated (Lupton, 2015; see also Cragin et al., 2010). The artifacts produced by different communities of practice are embedded with and shaped by the cultural norms of the groups that created them (Gitelman, 2013). Even raw data bear the imprint of some sort of selection process of lens on to the world (Bowker, 2008; see also Coombs, 1964). Researchers in science, technology and society and in the information sciences have repeatedly shown that diverse data practices have profound impacts on the way in which data are collected, shared, and interpreted (e.g., Edwards et al., 2011; Karasti et al., 2007; Stvilia et al., 2013; Vertesi and Dourish, 2011; Zimmerman, 2008), and which persist over time (Ribes and Finholt, 2009; Ribes and Jackson, 2013). Understanding and supporting pluralism in data practices is crucial to supporting the needs of those traditionally marginalized by information technologies—whether in their personal or disciplinary identity (Bardzell, 2010; D’Ignazio and Klein, 2020; Loukissas, 2016; Rawson and Muñoz, 2016).
Studying databases’ material forms is additionally important for understanding the role these systems play in the social settings in which they are used. Christine Hine (2006), for instance, via ethnography of the development of a genomic database, showed that databases are as much “communication regimes” as “scientific instruments”; though used to store and analyze data, they structure data sharing and information transfer workflows over the course of their use. Sabina Leonelli (2014) similarly explored the fundamental impact of large shared model organism databases on local data practices and norms. Taking a “data journeys” approach to trace the lifecycle of one particular scientific observation, Jo Bates et al. (2016) showed that the form and properties of data are as much shaped by the social, organizational, and infrastructural contexts in which they are created and transmitted as they are by the phenomena from which they are derived. And Bietz and Lee (2009) described databases as “boundary negotiation artifacts” that facilitate the organization and distribution of work among different groups; they noted that users have a notion of an ideal database that is often in tension with the realities of the specific material instantiation of the realized database.
We take inspiration from these studies of the role of databases in scholarly communication as well as Dourish’s call to take the materiality of information seriously and aim to contribute to the analysis of the historical and material contingencies shaping database practices. Here, though, we are interested in a subtler shift in technology use than relational-to-NoSQL databases: databases instantiated in other-than-strictly-database-technology. As briefly described in the Introduction, we identified this phenomenon through ongoing work studying database practices in memory institutions, beginning with natural history museums. We have found multiple cases in which databases are instantiated in, or augmented by, not-quite-database technology—primarily spreadsheets. Many of these were once relational databases but migrated into spreadsheets for a host of practical and organizational reasons. Further work is needed to understand the specific affordances of these databases, as well as to understand the impact of these shifting materialities on scientific work (Thomer et al., 2018).
Methods
A number of approaches might be taken to study scientific data practices, including interview-based methods, ethnography, or artifact-focused approaches of content analysis. Here we blended the latter two approaches: we conducted a close examination of data products, informed by methods of trace ethnography.
Data communicate as much through their structure (the way in which values are organized for human and machine readability) and invocation of standards (controlled vocabularies, best practices) as they do through their associated articles and metadata records. Close consideration of materialities information representation is important in understanding the cultures of the communities in which they are shared. For instance, the differing ways in which a standard is enacted can reveal rifts in communities (Millerand and Bowker, 2009). By centering our analysis on the artifacts themselves, we mirror the experience of the re-users of these artifacts, who must often infer meaning from a data object’s structure in addition to “metadata” and contextualizing documentation. An artifact-centric analytical approach was therefore important in our goal of understanding and supporting diverse data practices.
As in similar prior work from the information sciences, we used qualitative content analysis to examine “significant aspects of texts that are not amenable to quantitative techniques”—such as themes, structure, or metaphor (Pickering, 2004; see also Mayring, 2000). While data objects do not necessarily have themes to their structure, they do exhibit different styles of construction, and different methods of expressing data semantics and structure. Given our interest in exploring ways in which databases are being made, and the consequences of these decisions, qualitative content analysis was appropriate.
Our analysis was informed by methods of trace ethnography (Geiger and Ribes, 2011). Trace ethnography extends forms of documentary ethnography to environments that are not amenable to simple observation: for instance, online environments in which collaboration is facilitated through digital “traces” of action like logs and records, which may seem like incomprehensible markup to outsiders but are deeply meaningful to community members. Prior applications of trace ethnography have focused on collaborative work in sites like Wikipedia (Geiger and Ribes, 2011), the Zooniverse citizen science platform (Mugar et al., 2014), and in open-source communities (Howison and Crowston, 2014). In each of these examples, a traditional ethnography would largely involve watching people as they sat at a computer, whereas a trace ethnography places the ethnographer in the middle of the “site” of a community’s collaboration in the form of traces and documents. Here, our site is the system of scholarly communication and data sharing, as facilitated through repositories like PubMed and Dryad; the traces we studied are those created in the process of collecting and processing data for sharing and reuse.
Purposive selection of data objects for analysis
In prior work, we identified several instances of technologies such as spreadsheet software being referred to and treated as databases (Thomer et al., 2018). These data objects could be split in two categories: (1) a single spreadsheet being referred to as a database, and (2) multiple spreadsheets or plain text files “linked” through identifiers or code being used as databases. We used a purposive sampling strategy to identify further examples of “databases” exhibiting these features. Purposive sampling is “the deliberate seeking out of participants with particular characteristics, according to the needs of the developing analysis and emerging theory” (Morse, 2004: 884). It is not meant to be a statistically representative sample; rather, it is paradigmatically representative of trends identified in earlier work. We identified potential cases through keyword searches of PubMed Central and the Dryad Data Repository. In PubMed, we looked for articles describing “databases;” in Dryad, we looked for published database files. In both cases, our searches returned results spanning disciplines and size. Our search of Dryad (in January 2019) retrieved 280 results, and our search of PubMed retrieved more than 7000 (though we note that the PubMed results included databases using both traditional and nontraditional formats; the Dryad results entirely consisted of spreadsheets or plain text files being used as databases). We reviewed a subset of the results by hand until we found two paradigmatic cases of single-spreadsheet-as-database and two paradigmatic cases of multiple-spreadsheets-as-database use.
Analytical workflow
For each case, we began by gathering and reading the articles and documentation associated with each database. We then:
Read publications associated with the database to identify stated motivations for construction, techniques used in the construction, and anticipated uses of the database. Qualitatively assessed format, character, content, and queriability of the database to determine how data is structured and how that structure impacted the meaning and interpretation of the data. Evaluated each database for its use of best practices in relational database design, as articulated through a popular textbook by Elmasri and Navathe (2010). Drafted UML diagrams of the databases’ underlying information models. Categorized each database according to degree of normalization, clarity and consistency of data semantics, and support for querying or running complex analyses over the data.
Throughout, we wrote memos about our work and discussed the cases in weekly meetings. The cases described below were derived from these memos and discussions. Because each of these databases is publicly available, our study did not require oversight from our universities’ institutional review boards, and there are no required privacy restrictions on our presentation of the data. However, we chose to lightly anonymize them to avoid unintended reputational damage to their creators via our critique. Links to the specific resources are available by request of the authors.
Results: Four scientific “databases”
We grouped our analysis according to the two primary types of databases identified in our previous work: highly interlinked relational data objects, and “flat file” relational data objects. Throughout this article, we refer to these files as data objects (following Open Archival Information System terminology; Consultive Committee for Space Data Systems, 2012) to better distinguish them from the relational database paradigms we wish to contrast them to. We describe the highly interlinked relational data objects first.
Highly interlinked relational data objects
In this category of databases, we include data objects with multiple files of data that are linked through a combination of identifiers and scripts yet do not make use of traditional relational database technology. This group of objects exhibits a high level of alignment to relational data paradigms.
Case 1: Carbon cycling database
Our first case, the Carbon Cycling Database, was published as part of a broader project aggregating and curating previously published data measuring ecosystem cycling of certain key nutrients, such as carbon. We found this data object in the Dryad Data Repository, where it was archived as supplementary material for a journal article presenting an ecological analysis of the chemical and elemental dynamics its data describe. The supplementary material included “sample R code for joining the tables,” strongly suggesting that the authors envisioned use of the resource in conjunction with other computational workflows.
On Dryad, the data object is described as a “series of cross-referenced data tables”. It is stored as a zip archive containing 10 individual comma-separated value files (CSVs), along with a README (a simple human-readable text document often included as metadata for a dataset or code) stored separately from the zip archive. The README contains a summary table that describes each of the data tables in terms of column names, descriptions, and units (where applicable; Figure 1).

UML diagram shows table structure of Carbon Cycling data resource, created by the authors. The Measurements table might be considered the core data table; it references six of the seven other tables. For a given plot at a given site, the Measurements table records the data used to construct the analysis of carbon cycling presented in the article.
In some ways, this data object quite clearly follows typical practices in relational database design. Most significantly, rather than keeping data in one large file, it decomposes data into multiple
That said, this data object also departs significantly from conventional database practices. A key best practice for database normalization is data cell atomicity: each cell should only contain one value. However, we noted several instances of non-atomic values. We also noted several violations of referential integrity constraints, in which a column referencing another table contains values other than the primary key of the referenced table. Most of the violations of integrity constraints arise from indicators of “missing data” instead of references to primary keys. This information is important in understanding the provenance of each observation aggregated in this data object. However, it is not stored in a way that satisfies the constraints of a database management system (DBMS).
More seriously from a relational database management perspective, there is a partial conditional dependency between two columns in the Measurements table (shown with dashed connections in the structure diagram shown in Figure 1), where some, but not all, values in the column are references to another table. In a scripting language like R one could simply write a conditional “if/then” statement to retrieve the necessary values for processing. But this approach would not work in a conventional relational DBMS, where retrieval languages are implemented with an assumption of consistent semantics for a given column in a table. This data object thus blends relational database approaches to data management with programmatic approaches common in scientific computing.
Case 2: Organismal Growth Curves Database
Similar to the Carbon Cycling Database, the Organismal Growth Curves Database aggregates published scientific data—in this case, data about the growth rate of a particular species of earthworm—to support visualization and analysis of the impact of environmental factors on the development of individual worms. We found this resource via an article describing its creation in PubMed Central. The data object itself is archived in Zenodo but is also available as a dynamic website that can generate visualizations of query results. The article associated with this data object describes a high level of data curation that went into compiling and organizing data from 162 published sources into a form that supports the dynamic visualization tool hosted online by the researcher.
Despite the complexity of the analyses, the data in this object are simple and relatively “small”—limited columns, limited rows. The files archived on Zenodo include three CSVs: one containing the growth curve data, one containing environmental information, and one containing bibliographic information for the sources of the data. As in the Carbon Cycling resource, each CSV is treated as a separate table (Figure 2). The article presenting this resource includes instructions and code for accessing the data through “the R software console”; again, as with the Carbon Cycling resource, these files are intended to be accessed through a programming language like R rather than database management software.
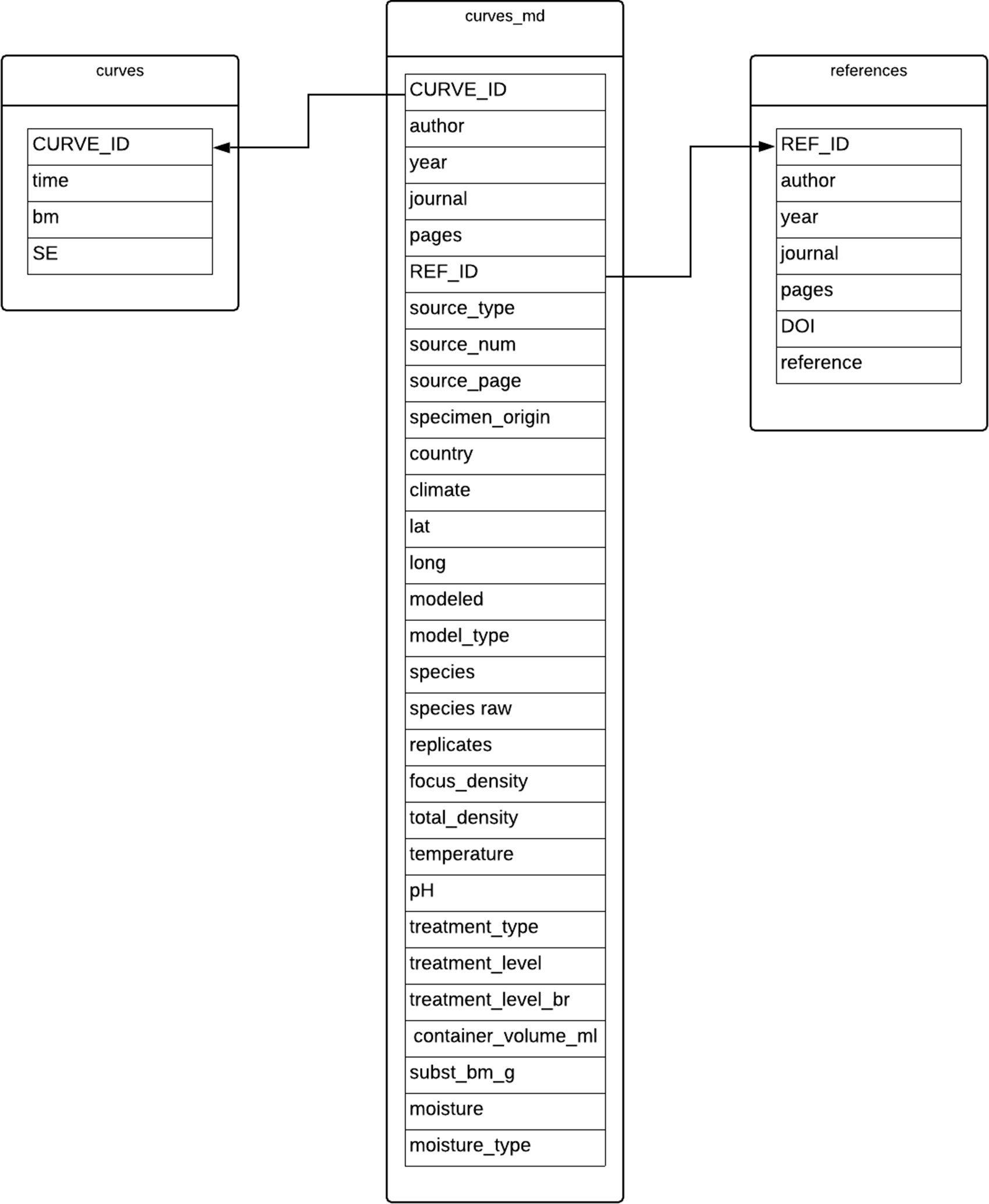
UML diagram shows table structure of Organismal Growth Curves Database, created by the authors. We note that the article describing the data object includes a similar UML-like diagram depicting attribute and reference information, though it diverges from standard UML modeling techniques because it does not identify classes or entity types in the diagram—rather, it identifies the filenames hosted in Zenodo, and illustrates how those might be merged and queried with R.
Both of our highly interlinked data objects allow end-users a high level of direct interaction with each so-called database. Both CSVs and R code are stored as plain text files and are easily viewable and alterable in typical operating systems. Notably, this is a level of access that traditional DBMSs were designed to prevent. In the late 1970s, Codd’s relational data model was codified into the ANSI/SPARC three-level architecture for DBMSs (Tsichritzis and Klug, 1978), which defines three levels of information organization that occur in a DBMS: the internal, external, and conceptual levels. This creates a separation between conceptual organization and physical storage of information referred to as
“Flat file” relational data objects
In this category of “databases” we include data objects that are presented as a single spreadsheet (typically, Excel) file. Although these data objects may appear to be simply spreadsheets on first glance, they are nevertheless referred to as databases by their creators. We find that they also exhibit select alignment to relational data paradigms—albeit in a more limited way than the highly interlinked data objects discussed previously.
Case 3: Sediment Geology Database
The Sediment Geology Database is a compilation of data about “paleocurrents”—patterns in sedimentary rocks that show evidence and direction of ancient ocean movement—from previously published journal articles, theses, and dissertations. We found the data object in Dryad, and it is supplemented by a data description article published in a popular multidisciplinary open-access journal. This data object consists of an Excel workbook containing 17 sheets, and a README file with text explaining how the resource was created. Despite the many sheets, we consider this data object a “flat file” because 16 of the sheets simply duplicate data from the first sheet (titled “Main”; Figure 3). Thus, the “main” sheet of the data resource can be viewed as one data object; the subsequent 16 sheets might be thought of as filtered results from this first sheet.

Diagram shows repeated table structure of Sediment Geology Database, created by the authors. The first sheet in the Excel file, titled “Main,” contains all of the data from the resource; the next 16 sheets repeat subsets of the Main tab, separated out by geographic region (“Continental Area,” seen in Figure 3), with no data in addition to what are listed in the Main tab.
This data object has a few features that complicate this flatness, however. Each row in the spreadsheet represents a paleocurrent data point and includes the name of an originating geologic formation and a citation to the source publication from which the data was aggregated. Because each article contains many data points, there are many data points with the same citation. There are also many data points from the same geologic formation. Rather than repeating these duplicate citations, the authors chose to list the details of each bibliographic reference only once. Subsequent data points from the same article have the other details of the data point listed but leave the cells blank for bibliographic details. Similarly, each geologic formation is assigned a period number that appears in every data point, and the formation name is recorded once for each period (Figure 4).
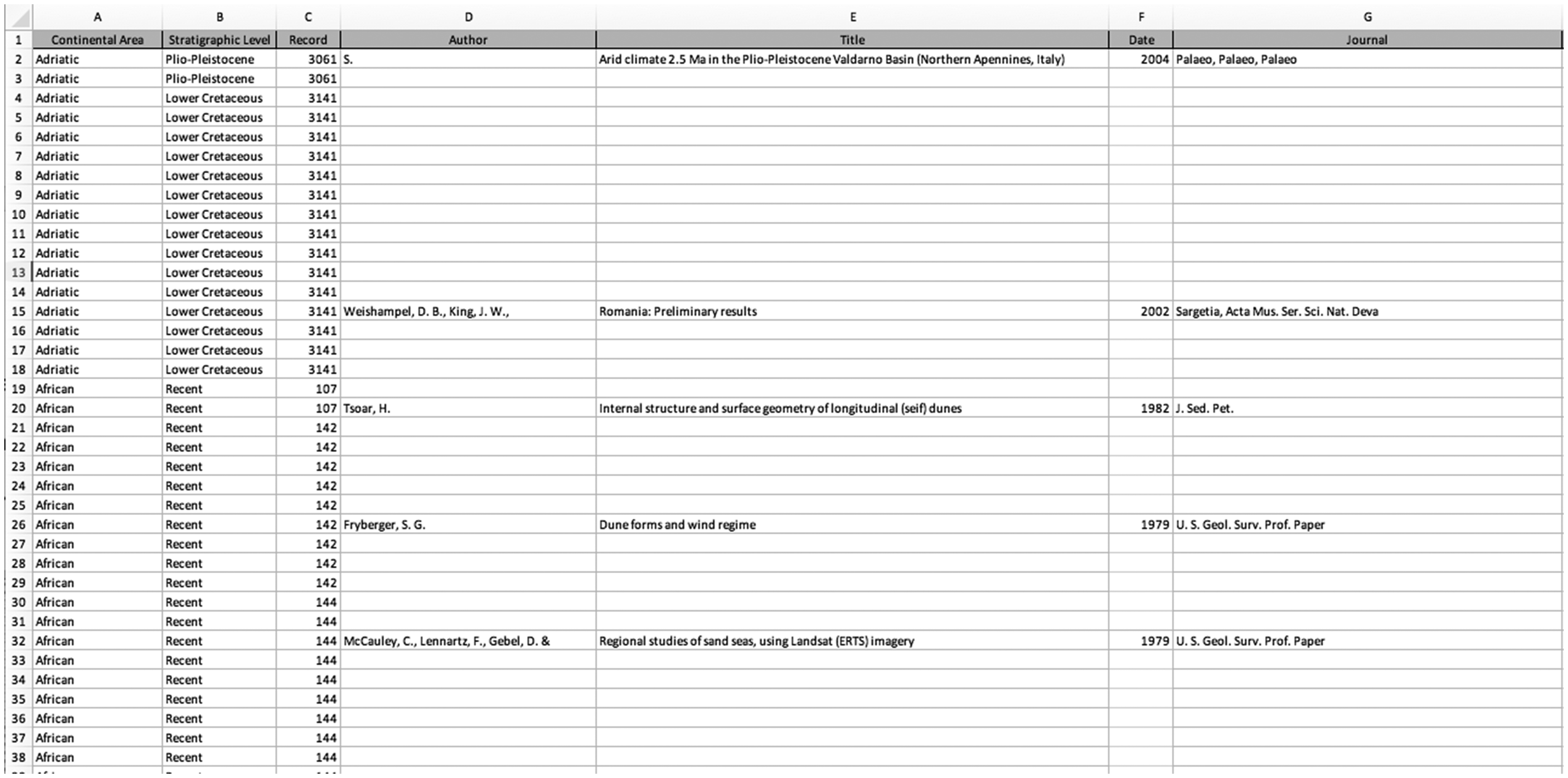
Screenshot of spreadsheet containing Sediment Geology Database. Each blank cell is meant to indicate a duplication of the most recent non-blank cell above.
UML diagram shows table structure of the Agricultural Nutrient Database, created by the authors. Each sheet of the spreadsheet is represented as a table.
On one hand, this is a clear violation of spreadsheet best practices (Teal et al., 2019). The creators of this data object seem instead to be relying on the spreadsheet display and assuming that future readers can infer the information from the placement of record in the display. This strategy departs from spreadsheet best practices because there is a significant risk that information will be lost or confused, especially if the spreadsheet is sorted or divided up. On the other hand, the authors’ choice to not duplicate information in the table does, in a sense, align with the spirit of the relational model’s focus on eliminating the need to store duplicate data, thereby reducing storage costs and potential errors. The typical strategy for handling this kind of redundancy in relational databases is to create additional tables for the repeated information and to reference those tables. In this case, one could normalize by creating a table recording the bibliographic details of each data source, and a table recording details of geologic formations, and then link to those tables with foreign key references in the primary data table. The authors adopted the first step of reducing redundancy but did not follow the database practice of splitting the duplicated information into a separate table that can be consistently referenced.
The Sediment Geology Database contains over 30,000 rows of data, which were aggregated over 30 years, and contains data from around the globe. The article associated with the data resource describes a purpose-built program “written in CPP [that] allows the data to be graphically represented on a map of the world,” although it is unclear whether this program would use the data as recorded in the spreadsheet or in some other form. Nevertheless, the article clearly presents the Excel spreadsheet as a canonical version of the resource and includes a level of documentation of the processes and concepts used in aggregating data that would be of use to those re-using the Excel file.
Case 4: Agricultural Nutrient Database
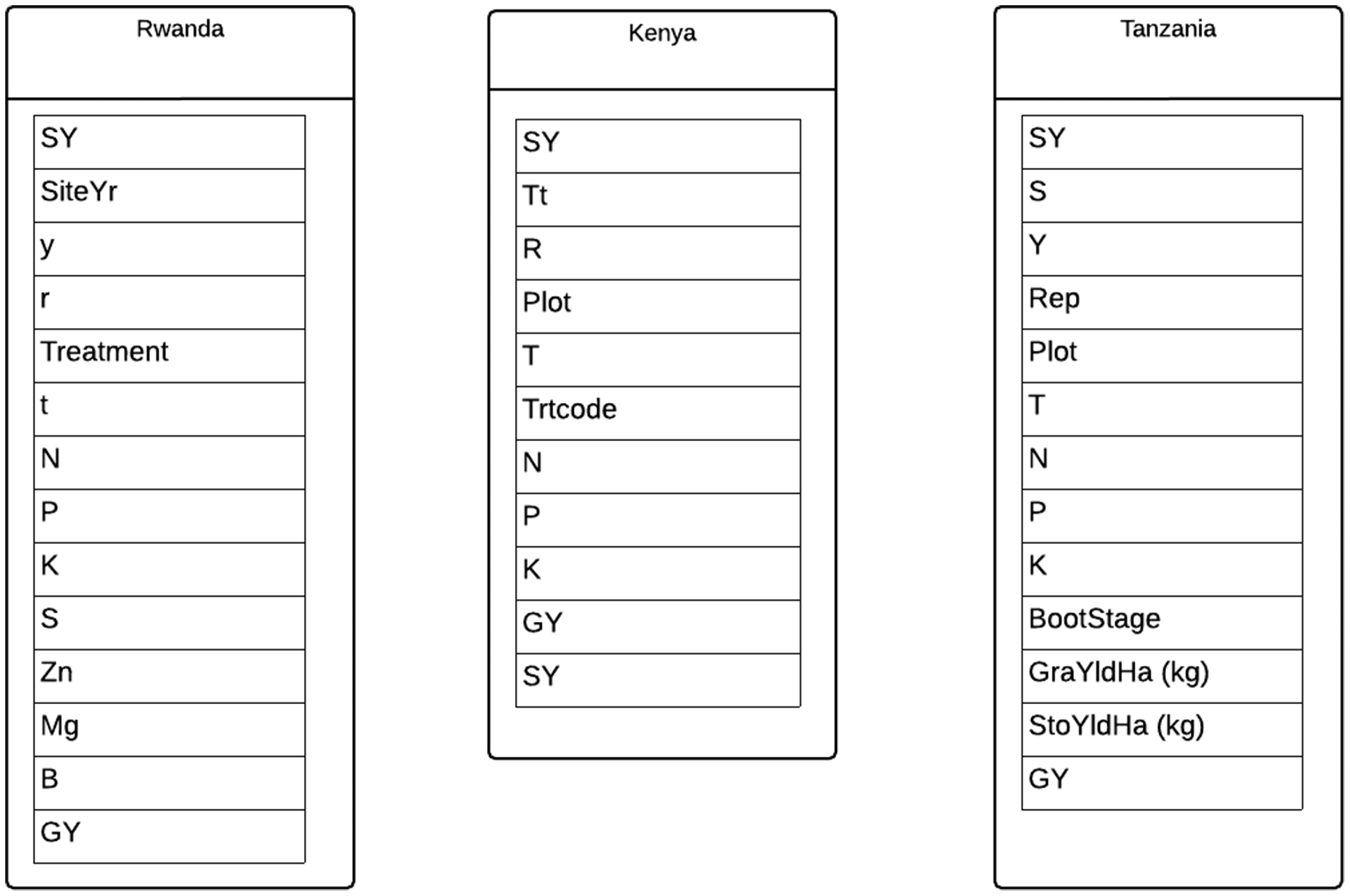
Our final data object for analysis, the Agricultural Nutrient Database, was also found on Dryad, where it was deposited in support of a journal article published in a disciplinary research journal. Like the Sediment Geology Database, this resource can be considered a “flat” data object, instantiated via four sheets of an Excel workbook. However, unlike the Sediment Geology Database, the Agricultural Nutrient Database contains novel data, collected by the authors from field sites in eastern Africa—not aggregated data from previously published literature. That said, it does act as an aggregated data resource in that the authors compiled it from 18 years of research across wheat-growing areas of Kenya, Rwanda and Tanzania, with the goal of determining “wheat grain yield response functions”. The first sheet of the Excel workbook is labeled “Readme” and contains metadata (e.g., definitions of parameters in the following sheets, and some description of data collection methods) as well as summary charts and graphs, which also appear in the associated article. The next three sheets contain subsets of the data, divided by site. In the Sediment Geology Database, the fields were consistent across each sheet; here, they vary because of changes in data collection methods at each study site (Figure 5).
The data could likely be aggregated into one sheet with some minimal cleaning and curation work. As it stands, the inconsistency between the column headers makes issuing meaningful queries across the three locations impossible to do in a straightforward way. The ability to do that kind of querying, and subsequent analysis in relational data systems, relies on the use of consistently structured data, including the assignment of regularized names for attributes.
Where the previous three cases of relational data objects have relatively clear database-like attributes, this case has fewer. The spreadsheet is not organized with any relational logic beyond a basic table structure (Figure 5), and it lacks consistent names for columns of data. The article associated with the resource does not mention the development of any kind of reusable programmatic access or analytic tools for interaction with the data. However, the creators of this resource
Comparison of cases
In tandem with our content analysis, we examined each database to determine its level of Primary key: Does each row of a table have a “primary key” (a unique identifier)? Is each row of the table unique? No repeating groups: Are fields repeated in a given table? Atomicity: Does each cell contain a single value? Referential integrity: Does each field pointing to a key in another table (or file) only consist of referenced keys, or a NULL value?
We also assessed them for two key characteristics that we identified during our analysis: the description of programmatic access to the resource, and whether the resource aggregates previously published data (Table 1).
Comparison of the degree of normalization across cases.
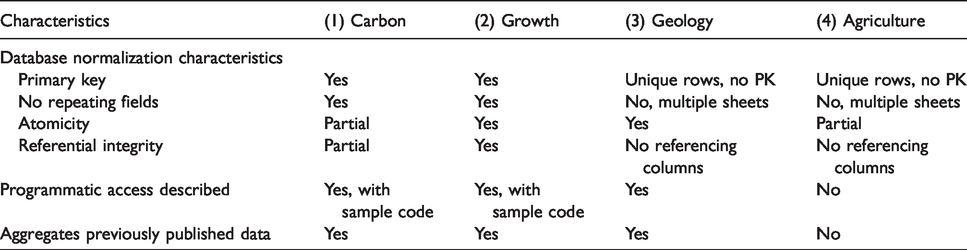
Generally, the data objects we examined were only partially normalized. For instance, the tables in the Carbon Cycling data object have primary keys, but the fields could be further atomized and have some unique issues with their referential integrity. The Organismal Growth data object features well-atomized fields, but the tables feature some redundancy and could be further decomposed. The Sediment Geology and Agricultural Nutrient data objects have unique data for every row in their tables, but they do not have any single attribute that can function as a primary key.
We conducted the normal form analysis because we see normalization as a materiality of information representation (Dourish, 2017) inherent in the use of a relational DBMS. By applying the degree to which normal forms have been adopted in these objects, we get a picture of the degree to which these scientists have adopted database materialities. The presence of these materialities, which are integral to relational database systems, speaks to the continued influence of
The aggregation of previously published data into these data objects, which appears in three of the cases, suggests a reading of “databases” as a kind of curated, enhanced dataset. The programmatic access discussed in articles in the three of the cases we examined, and even more strongly demonstrated by the inclusion of sample code in two of the cases, are evidence of a shift in scholarly communication regimes. We discuss relational data paradigms, curation and aggregation, and scholarly communication regimes in depth in the next section.
Discussion: What do we learn by taking the materiality of databases seriously?
Relational data paradigms
Part of our motivation for this study is the observation that many “best practices” for data management are in tension with the realities and priorities of scientific data production. Our aim in this work was to contribute to a picture of scientific communication that recognizes the legitimacy and function of a
We argue that the tension between “best” and actual database practices, as well as changes to scientific communication norms (described later) have resulted in some abandonment of traditional database systems by scientists in favor of the more one or more tables with unique identifiers, or at least unique rows; a minimization of redundancy by decomposing data into separate class-based tables that can be cross-referenced via identifiers and perhaps even code to support aggregation or compilation over time.
Each case discussed in this article shows the influence of these relational paradigms: the Carbon Cycling data object is almost formally normalized, despite not being manifested in a database system; the Growth Curves data object has similarly mimicked relational structure via R; and the Sediment Geology data object reflects an attempt to minimize redundancy despite its flat format. Even the simple and spreadsheet-like Agricultural Nutrient data object features unique tuples.
It is beyond the scope of this study to concretely pinpoint the reasons for the persistence of relational data paradigms. However, we note that the four cases we examined—and indeed, many of the other cases we considered—are the products of a modern scientific “communication regime” (Hine, 2006) of open science and a push toward data sharing (Borgman, 2012; Lyon, 2016; McKiernan et al., 2016; Roure, 2014). More and more journals and funding agencies require that scientists share the data underlying their studies (see, for example, Holdren, 2013; Nature News, 2016; Stodden et al., 2018). Additionally, growing international movements toward “open science” have created social incentives for sharing data and making resources FAIR: that is, findable, accessible, interoperable, and reusable (Wilkinson et al., 2016). The authors of the data objects studied here might have chosen spreadsheets and text files as their file format because they are commonly used and easy to open using common software. They may also have chosen text files and spreadsheets simply because the journals and repositories storing these files would not accept database system files. The Dryad Data Repository, for instance, does not strictly prohibit database files but does express a preference for open files and formats, which many database files (such as an Access.mdb) are not (Dryad, 2018). Thus, where Hine observed that databases can shape and structure the modes and rhythms of communication, here we find that they are, in turn, shaped by the external workflows, practices, and, indeed, materialities of the systems of scholarly communication in which their creators participate.
The regime change toward open science has been accompanied by a shift in user roles around databases, which could additionally impact database materialities. In short, databases need administrators. In our review of database textbooks, we noted that every definition of a database is paired with a definition of a database administrator; this role is fundamentally assumed in the design of the system. And as we have observed in prior work (Thomer et al., 2018), Codd’s relational model assumes a division of labor between administrators and users: users do data entry and retrieval, and administrators manage the mappings among the physical, logical, and conceptual levels of the database. However, in many modern contexts, user and administrator are one and the same (for examples see Dourish and Edwards, 2000; Jagadish et al., 2007; Li and Lochovsky, 1996; Olson, 2009; Voida et al., 2011). This collapse between users and administrators forces users to adapt databases to user-friendlier formats. Thus, changes in the user roles around databases have led to a shift in the materialities of databases, particularly when it comes to the use of databases by scientists. Scientists are creating data objects to meet the needs of their projects, using platforms and languages to achieve database goals in systems not necessarily designed for database management. Beyond the working lives of scientists, large-scale data infrastructures are playing an increasing part in our everyday lives. It is therefore essential for scholars to develop methods and frameworks that let us examine the shifting roles of the sources, producers, administrators, and users of those data systems.
Practical implications
Curating ‘curated databases’
In addition to informing design of data curation tools and workflows, this work informs our understanding of what computer scientist Peter Buneman and co-authors previously called “curated databases”—“those databases whose content (often about a specialized topic) has been collected by a great deal of human effort through the consultation, verification, and aggregation of existing sources, and the interpretation of new (often experimentally obtained) raw data” (Buneman et al., 2008: 1). Each of the data objects we selected shows some degree of curation and aggregation. The Sediment Geology and Growth Curves data resources each aggregate and curate data points from previously published literature; the Carbon Cycling data resource is designed to funnel data into a larger aggregation of similar data; and the Agricultural Nutrient data resource compiles data from 18 years of work. We argue that this “compilation” and “aggregation” is central to the relational paradigm, and to the “purposeful work” of data curation that goes into creating and maintaining a database (Palmer et al., 2013). Datasets are often framed as novel observations created from a singular experiment or field trip, whereas databases are holding places for multiple observations collated over time. Databases are for storage, whereas datasets are for independent, short-term analysis. Further examination of the scope of curated databases might provide important insights into their maintenance needs over time.
What is in a name?
Understanding why users might call one spreadsheet a database and another a dataset has implications for how we treat, manage, curate, and preserve these objects. It also has implications for how we design tools and infrastructure. Defining information entities is not a merely semantic exercise; rather, these definitions are translated into system rules and requirements, which have long-reaching (and often unexpected) impacts (Renear et al., 2010). Even developers at influential organizations like Google have stated that defining and constraining information entities is a key challenge in supporting data search and use. In a 2017 blog post detailing a new dataset search initiative, Google researchers Natasha Noy and David Brickley listed “Defining more consistently what constitutes a dataset” as a fundamental challenge: For example, is a single table a dataset? What about a collection of related tables? What about a protein sequence? A set of images? An API that provides access to data? We hope that a better understanding of what a dataset is will emerge as we gain more experience with how data providers define, describe, and use data. (Noy and Brickley, 2017)
There is a clear need to better support the curation and management of databases and database-like spreadsheets in repositories and scholarly publications. Our work here and our prior work in natural history museums have suggested a number of motivations underlying users’ choices to archive and share databases in formats such as spreadsheets, which we offer here as promising avenues for research:
The unique uses and affordances of spreadsheets will be important to account for in the design of future data platforms, archives, and tools. How might we design database systems that mimic the familiarity, openness, and flexibility that spreadsheets afford, while preserving the stability and integrability offered by database systems? Conversely, how might we better support the creation and curation of spreadsheet databases, and thereby avoid the “coerciveness” of database technology in forcing users to work “on the infrastructure’s terms” rather their own (Bopp et al., 2019: 1)? There are clear opportunities for innovation in this space, which we are excited to pursue in future work.
Conclusion
In this article, we followed Dourish’s call to take the materiality of databases seriously by conducting a close examination of four paradigmatic instances of non-conventional databases. In doing so, we have shown that a tension between scientific data practices and the material contingencies of relational data systems has resulted in the abandonment of traditional database systems by scientists, along with the adoption of relational data organization strategies in data modeling and file format choices. We show that scientists still draw on relational data paradigms when working with file formats not specifically designed for databases. We have also shown that the materialities of common communication regimes in the sciences have potentially shaped the materialities of these non-conventional databases. In addition to the specific analysis of four cases, we contribute a definition of relational data paradigms, as well as some insights into the nature of databases that are important for future innovations and developments in data curation and sharing infrastructure.
The artifact-centric methods deployed in our present work mean that we cannot be certain about the particular ways the authors of the resources we examined thought about their data. However, we argue that the features we identified are strong evidence of information representation strategies that center on data organization and modeling techniques fundamental to the relational database model. By examining the technical artifacts shared by these scientists, we have developed an account that is grounded in what has actually been done to organize and structure data. Approaches that center analysis on the actual instantiation of information objects into our communication practices and infrastructures, rather than the stated intentions of data producers or system builders, are essential as large-scale data aggregations and infrastructures take on greater roles in our society.
Footnotes
Acknowledgements
Many thanks for the thoughtful feedback from the reviewers and editors of this manuscript, whose comments improved our work immeasurably. Thanks also to Beth Yakel, Margaret Hedstrom, Paul Conway, Tre Tomaszewski, Mike Twidale, Jodi Schneider, Allen Renear, Elizabeth Wickes, and Nic Weber for feedback on various iterations of this project.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded in part by IMLS grant #RE-07-18-0118-18.
