Abstract
Objectives:
Several vaccine adjuvants comprise complex nano- or micro-particle formulations, such as oil-in-water emulsions. In order to characterize interactions and compatibility of oil-in-water emulsion adjuvants with protein antigens in vaccines, effective protein characterization methods that can accommodate potential interference from high concentrations of lipid-based particles are needed.
Methods:
Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) is a standard protein characterization technique which is affected by the presence of adjuvants such as oil-in-water emulsions. In this article, we investigate variations in SDS-PAGE methods that result in a reduction of adjuvant-induced staining artifacts. We have investigated whether the SDS method or the adjuvant composition were the reason for these artifacts and succeeded in reducing the artifacts with a modified sample preparation and different staining procedures.
Results:
The best results were obtained by using gold staining or silver staining instead of a Coomassie Blue staining procedure. Moreover, the replacement of the dilution buffer (20% SDS to disrupt emulsion) by alternative detergents such as Tween® 80 and Triton® X-100 removed adjuvant-induced streaking artifacts at the top of the gel.
Conclusions:
These methods may be useful for improving characterization approaches of antigen–adjuvant mixtures by SDS-PAGE.
Keywords
Introduction
Adjuvants are a critical component of numerous vaccines. 1 Effective adjuvants can enhance the antigen-specific immunogenicity as well as the quality of the immune response, 2 which may lead to a higher efficacy of the vaccine or the possibility of reducing the antigen content per vaccine dose (dose sparing). After aluminum salts, the most widely used adjuvants in human vaccines are oil-in-water emulsions based on squalene oil. 3 Squalene oil-in-water emulsions generally consist of nano-droplets of squalene oil (< 200 nm in diameter) emulsified by phospholipids or nonionic surfactants in an aqueous bulk phase. Licensed vaccines for seasonal and pandemic influenza (e.g. Fluad®) have demonstrated that inclusion of squalene emulsion enhances antigen-specific immune responses, facilitating antigen dose sparing and higher vaccine efficacy including among high-risk populations such as the elderly or infants.3,4
Characterization of the integrity of protein antigens is an essential component of any vaccine development program. Such characterization is even more important when adjuvants are added within the vaccine preparation since antigen integrity or stability could potentially be affected by the presence of an adjuvant (and vice versa). One of the easiest and fastest characterization methods for protein antigens is the sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). This method allows the determination of the molecular weight of the protein of interest; thus, degradation of the protein primary structure can be monitored.
With well-defined and highly pure protein antigens, the SDS-PAGE technique results in clearly visible protein bands. Useful variations of the SDS-PAGE technique include reducing or nonreducing sample conditions and different staining techniques for the visualization of protein bands. However, we and others have already observed that some vaccine formulations containing squalene-in-water emulsions or other particulates cause artifacts (e.g. streaking) on polyacrylamide gels.5–9 These artifacts may mask, completely or partially, the protein antigen bands and therefore have the potential to impair the analysis of the assay results.
In order to reduce or potentially remove the artifacts induced by squalene-in-water emulsions on polyacrylamide gels, we attempted to identify which specific component was responsible for inducing the artifacts. We also explored modifications of the SDS-PAGE methods, including the sample heating time, the presence of a reducing agent, and the introduction of a washing step. Moreover, we also investigated the use of different detergents and stains, expanding on related work from our laboratory recently reported elsewhere. 9 All of these evaluations were first performed using bovine serum albumin (BSA) as a model protein. The optimized conditions were then applied to ID93, a recombinant tuberculosis vaccine antigen currently being evaluated in the clinic, with a squalene emulsion-based adjuvant. Moreover, we tested the general applicability of the methods by employing another squalene-based emulsion with different excipient composition. Our results highlight effective approaches to reducing or removing adjuvant-induced artifacts on SDS-PAGE gels.
Materials and methods
SE, a stable oil-in-water emulsion based on squalene oil (composed of squalene, dimyristoylphosphatidylcholine [DMPC], Pluronic F68, glycerol, and phosphate buffer) and recombinant protein antigen ID93 were produced at the Infectious Disease Research Institute as described previously.10,11 A squalene-in-water emulsion (SWE) (composed of 3.9% w/v squalene, 0.5% w/v Tween 80, and 0.5% w/v Span 85 in citrate buffer) was manufactured by the Vaccine Formulation Laboratory at the University of Lausanne. BSA was obtained from Sigma (Saint Louis, CA, USA), Tween 80 from JT Baker (Center Valley, PA, USA), Triton X-100 from Thermo Fisher Scientific (Waltham, MA), Zwittergent 3–14 from Calbiochem (Merck, Darmstadt, Germany), and saline solution from Teknova (Hollister, CA, USA). The SimplyBlue SafeStain solution was obtained from Invitrogen (Carlsbad, CA), the gold stain solution from Bio-Rad (Hercules, CA, USA), and the ProteoSilver Plus Silver stain kit from Sigma. Reagents for the SDS-PAGE were purchased from Invitrogen. The molecular weight marker was SeeBlue®Plus2 Prestained Protein Standard (Invitrogen).
The general SDS-PAGE method consisted of mixing the protein antigen with adjuvant or adjuvant components and then diluting the mixture with a solution composed of 20% SDS (to disrupt the emulsion) and a loading buffer composed of 4× lithium dodecyl sulfate (LDS), at a ratio of 1:2:1 (vaccine formulation/diluent/loading buffer). Samples were then heated at 90°C for 5 min without reducing agent (unless indicated otherwise) and then 20 µl of each sample were loaded per well on a 4–20% tris-glycine gel (Invitrogen), containing 12 wells. The 20 µl contained either 500 ng of BSA or 1 µl of ID93. The first well was loaded with Prestained Protein Standard. The gels were run for 1 h at 200 V and then stained by incubating in SimplyBlue SafeStain solution overnight. The following day, the gels were destained in demineralized water for 3 h, with the water changed every hour. Alternative staining preparations were conducted as described below. Other modifications to the above protocol are described below.
The silver stain was performed using the ProteoSilver Plus Silver stain kit and its corresponding protocol from Sigma. The gels were fixed overnight in the fix solution (consisting of ethanol:acetic acid:water at 5:1:4 volume ratio). The following day, they were washed for 10 min in a solution of ethanol 30% and then a further 10 min in demineralized water. The gels were then incubated with the sensitizer solution from the kit for 10 min and washed twice in demineralized water. The gels were then incubated in the silver solution for 10 min, rinsed for 1 min in demineralized water, developed in the develop solution for 3 min, and stopped for 5 min using the stop solution.
The gold stain procedure consisted of a transfer of the unstained gel to a polyvinylidene fluoride (PVDF) membrane and then coloration of the membrane. The protein bands were transferred on to PVDF membranes using the X-Cell II Blot module (Invitrogen). The transfer was performed at 35 V for 1 h with the NuPAGE Transfer Buffer (Invitrogen). After transfer, the membrane was rinsed in phosphate-buffered saline (PBS)-Tween for 15 min and washed three times in PBS. The colloidal gold total protein stain solution was then added to the membrane. After 1 h, the membrane was rinsed with demineralized water and dried.
Results
Initial analysis of BSA in the presence of SE adjuvant using nonreducing SDS-PAGE resulted in obvious streaking artifacts at the top and bottom of the gel and strong diminution of the intensity of the expected BSA bands (Figure 1a). First, to determine if any residual adjuvant after the migration was preventing the staining of the protein, a 20-min prestain rinsing step with demineralized water was added prior to the staining procedure. This resulted in a clearer protein band but no attenuation of the streaking artifacts (Figure 1b). All subsequent gels described below included this 20-min prestain rinsing step.

Rinsing time optimization in nonreducing sodium dodecyl sulfate polyacrylamide gel electrophoresis. A prestain rinsing step affected protein band clarity in the presence of oil-in-water emulsion (SE) adjuvant. (a) No rinse: molecular weight (MW) marker (lane 1), bovine serum albumin (BSA) (lane 2), BSA + SE (lane 3). (b) A 20-min rinse: MW marker (lane 1), BSA (lane 2), BSA + SE (lane 3). A total of 500 ng of BSA was loaded per well.
Next, in order to determine which component of the adjuvant was responsible for the streaking, each raw component of the SE adjuvant was prepared alone at the same concentration as in the formulated SE. Insoluble components DMPC and squalene were sonicated in a water bath sonicator for 5 min to obtain suspensions. Each component was then mixed with BSA or with a saline solution. The samples were then prepared, run, and stained as described above, including the addition of the prestain rinsing step (Figure 2). Only the formulated SE resulted in the full streaking pattern, whereas the raw component DMPC seemed to be accountable for the staining observed near the bottom of the gel. Other raw components did not result in staining artifacts, although in the case of insoluble squalene oil it is unclear if the oil was able to enter the gel as indicated by the stained material remaining in the well of lanes 7 and 12 (Figure 2). Subsequent method development focused on the complete formulated SE adjuvant.
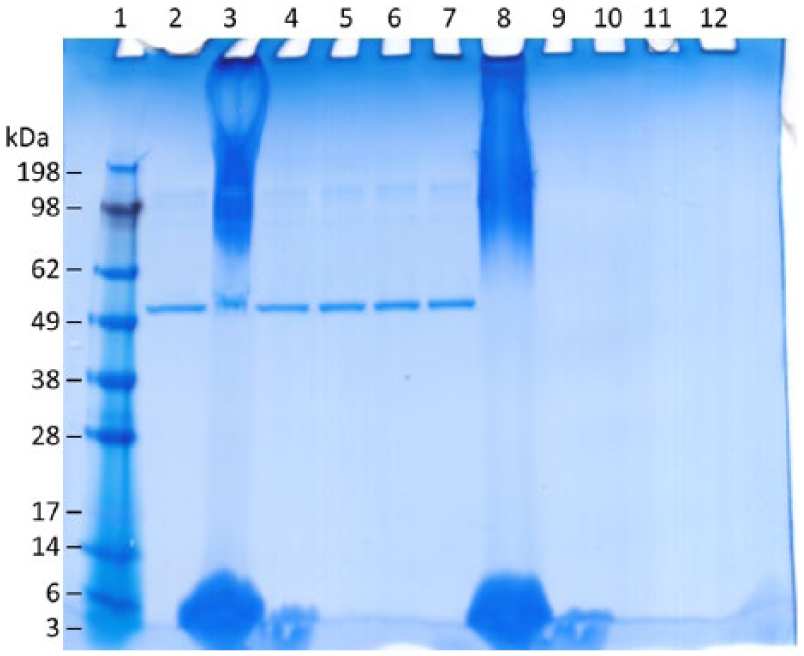
Impact of individual components on adjuvant gel interference in nonreducing sodium dodecyl sulfate polyacrylamide gel electrophoresis. Molecular weight marker (lane 1), bovine serum albumin (BSA) (lane 2), BSA + oil-in-water emulsion (SE) (lane 3), BSA + dimyristoylphosphatidylcholine (DMPC) (lane 4), BSA + Pluronic F68 (lane 5), BSA + glycerol (lane 6), BSA + squalene (lane 7), SE (lane 8), DMPC (lane 9), Pluronic F68 (lane 10), glycerol (lane 11), and squalene (lane 12). A total of 500 ng of BSA was loaded per well.
Another basic modification to the initial protocol was to change the amount of heating time of the prepared samples prior to loading them on to the gel. The samples, with the exception of the molecular weight marker, were mixed with loading buffer and 20% SDS and heated for 5 min or 15 min at 90°C before being loaded on to the gel. The longer heating time appeared to slightly reduce the severity of the adjuvant-induced streaking at the top and bottom of the gel (Figure 3). However, the BSA bands were lighter and less well defined. For subsequent experiments, a 5-min heating time was employed.

Effect of time at 90°C on adjuvant gel streaking in nonreducing sodium dodecyl sulfate polyacrylamide gel electrophoresis. (a) A 5-min heating time, (b) a 15-min heating time. Molecular weight marker (lane 1), bovine serum albumin (BSA) (lane 2), BSA + oil-in-water emulsion (SE) (lane 3), SE (lane 4). A total of 500 ng of BSA was loaded per well.
Next we evaluated the effect of a reducing agent. In this case, the samples (BSA, BSA + SE adjuvant, SE adjuvant) were prepared by adding the NuPage® Sample Reducing Agent 10×. The ratio of sample to reducing agent to SDS diluent to LDS 4× sample buffer was 5:2:8:5. A second set of samples was prepared by substituting the reducing agent with water. The gel with the reducing agent showed only a moderate reduction in the streaking artifacts caused by the adjuvant (Figure 4), therefore we decided not to include the reducing agent for subsequent experiments.

Effect of reducing agent on adjuvant gel streaking. (a) No reducing agent, (b) reducing agent. Molecular weight marker (lane 1), bovine serum albumin (BSA) (lane 2), BSA + oil-in-water emulsion (SE) (lane 3), SE (lane 4). A total of 500 ng of BSA was loaded per well.
To determine whether the artifacts were influenced by the type of stain employed, we compared alternative staining approaches to the Coomassie Blue stain, including silver stain and gold stain following PVDF membrane blot as described above and employed in previously published work from our laboratory. 9 Both the silver stain and the gold stain approach appeared to completely remove the adjuvant-induced artifacts compared with the Coomassie Blue stain (Figure 5). As expected, the silver stain and the gold stain approaches appeared to be much more sensitive than the Coomassie stain in the detection of minor protein bands, which may indicate protein aggregates, degradation products, and/or impurities present in small amounts. It is interesting to note that the higher molecular weight protein bands in the silver and gold stain gels did not appear in the samples containing adjuvant, even when the adjuvant-induced artifacts were not present. Also, the lanes containing adjuvant with protein resulted in protein bands with decreased width compared to lanes with protein alone, regardless of stain technique.
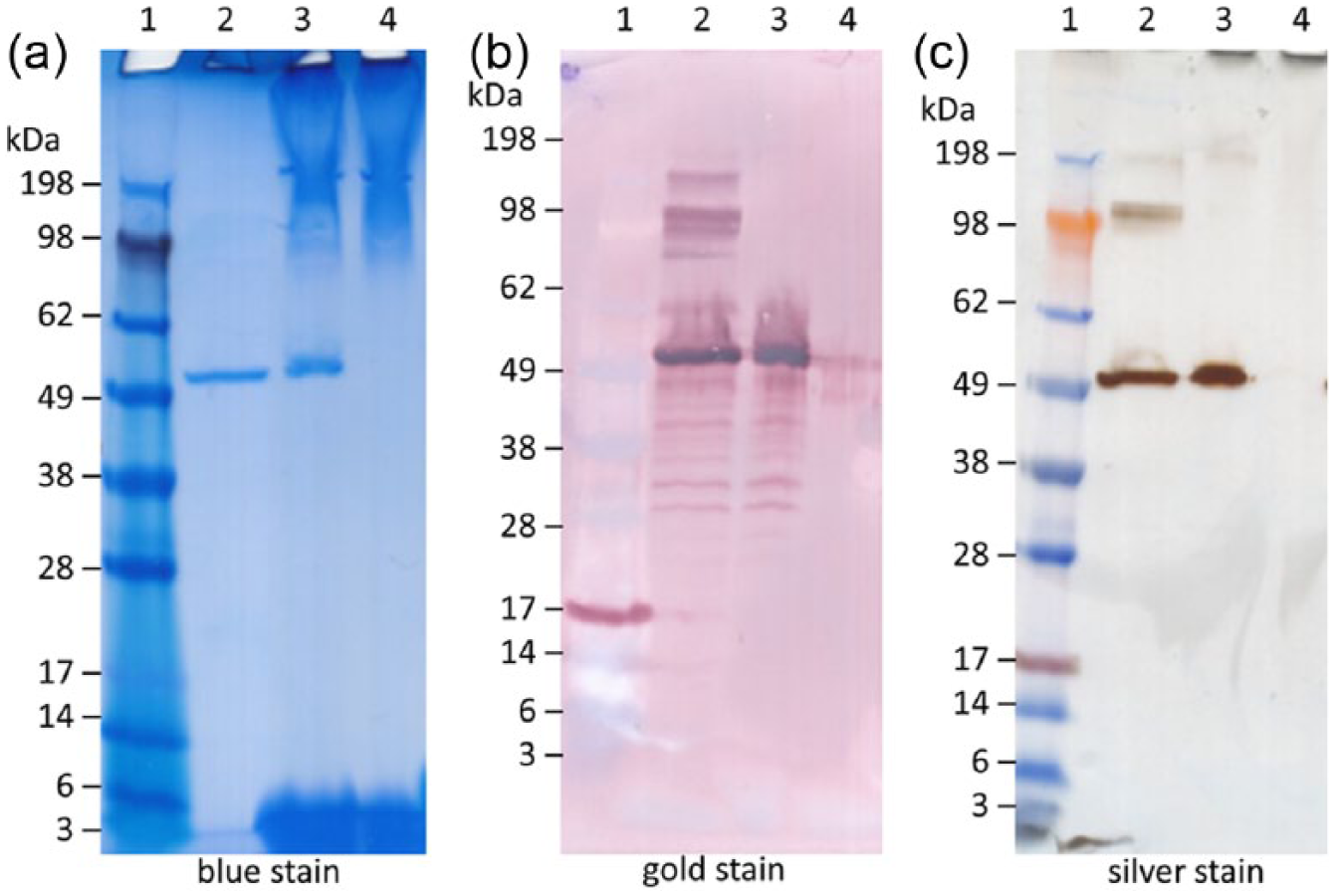
Adjuvant artifact appearance in nonreducing sodium dodecyl sulfate polyacrylamide gel electrophoresis gels stained with Coomassie Blue or silver stain, or a blotted polyvinylidene fluoride (PVDF) membrane stained with colloidal gold. (a) Blue-stained gel, (b) gold stain of PVDF membrane, (c) silver-stained gel. Molecular weight (MW) marker (lane 1), bovine serum albumin (BSA) (lane 2), BSA + oil-in-water emulsion (SE) (lane 3), SE (lane 4). A total of 500 ng of BSA was loaded per well.
Since the presence of adequate detergent may promote disruption of the squalene emulsion, different surfactants were tested to replace SDS. Thus, Tween 80 or Triton X-100 solutions were used to dilute samples instead of 20% SDS. The samples were diluted in Tween 80 or Triton X-100 solutions at different concentrations from 0.25% to 4%. The samples were then processed as described above, including the prestain rinse. Both gels were then stained using the Coomassie Blue stain protocol (Figure 6). The 4% Tween 80 solution resulted in the absence of the SE streaks at the top of the gel but not the streaks at the bottom of the gel. The samples diluted in the 2% or 4% Triton X-100 solution also resulted in the absence of the adjuvant-induced artifacts at the top of the gel. However, with the 4% Triton X-100 concentration, the streaking at the bottom of the gel was noticeably increased. The solutions of Tween 80 at 4% and Triton X-100 at 2% were selected as optimized diluent solutions. A Zwitterionic detergent, Zwittergent 3–14, was also tested but was found to contribute additional interference in gel appearance and was therefore not investigated further (data not shown).

Dilution of sample in different detergents affects the appearance of artifacts in nonreducing sodium dodecyl sulfate polyacrylamide gel electrophoresis gels. (a) Sample diluted in Tween 80, (b) sample diluted in Triton X-100. Molecular weight (MW) marker (lane 1), bovine serum albumin (BSA) diluted with 4% detergent (lane 2), BSA + oil-in-water emulsion (SE) diluted with 4% detergent (lane 3), SE diluted with 4% detergent (lane 4), BSA + SE diluted with 2% detergent (lane 5), SE diluted with 2% detergent (lane 6), BSA + SE diluted with 1% detergent (lane 7), SE diluted with 1% detergent (lane 8), BSA + SE diluted with 0.5% detergent (lane 9), SE diluted with 0.5% detergent (lane 10), BSA + SE diluted with 0.25% detergent (lane 11), SE diluted with 0.25% detergent (lane 12). A total of 500 ng of BSA was loaded per well.
In light of the evaluations performed above, we considered as optimized conditions the nonreducing sample preparation containing 4% Tween 80 or 2% Triton X-100 with a 5-min heating time and a 20-min prestain rinse. We note here that for proteins that contain disulfide bridges, reducing sample preparation may be preferred. These optimized conditions were then applied with the recombinant ID93 protein to be compared with BSA. Protein alone (ID93 or BSA), protein in the presence of SE, and SE alone were each diluted in either Tween 80 4% or Triton X-100 2%, as well as SDS 20% as a control. The samples were then heated for 5 min and 20 µl of each sample were loaded on to three different gels that were prepared according to the three different staining procedures (blue, silver, and membrane transfer follow by gold stain) (see Figure 7). The reduction in the adjuvant-induced streaking in the blue-stained gels by diluting with Tween 80 or Triton X-100 is apparent for both proteins. Overall, gel appearance for all three optimized diluent preparations appeared similar in the silver-stained gels and the gold-stained PVDF membranes, with the notable exception that the 4% Tween 80 preparation of BSA with adjuvant maintained the upper protein bands in the gold-stained membrane that were not visible with other detergent preparations and/or stains.

Comparison of modified sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) methods with two different proteins. (a) Bovine serum albumin (BSA) (500 ng per well), (b) ID93 (1 µg per well). Staining procedures were (left) Coomassie Blue stain, (middle) gold-stain polyvinylidene fluoride (PVDF), or (right) silver stain. Molecular weight (MW) marker (lane 1), protein diluted with 20% SDS (lane 2), protein + oil-in-water emulsion (SE) diluted with 20% SDS (lane 3), SE diluted with 20% SDS (lane 4), protein diluted with 4% Tween 80 (lane 5), protein + SE diluted with 4% Tween 80 (lane 6), SE diluted with 4% Tween 80 (lane 7), protein diluted with 2% Triton X-100 (lane 8), protein + SE diluted with 2% Triton X-100 (lane 9), SE diluted with 2% Triton X-100 (lane 10).
Finally, the optimized conditions were applied to BSA samples formulated with another squalene-based emulsion (SWE from the University of Lausanne). Protein alone (BSA), protein in the presence of SWE, and SWE alone were each diluted in either Tween 80 4% or Triton X-100 2%, as well as SDS 20% as a control. The samples were then heated for 5 min and 20 µl of each sample were loaded on to two different gels that were prepared according to different staining procedures (blue or silver) (see Figure 8). No artifacts were observed at the top of the gels with the SWE adjuvant; however, on the Coomassie gel some streaking was visible at the bottom of the gel.
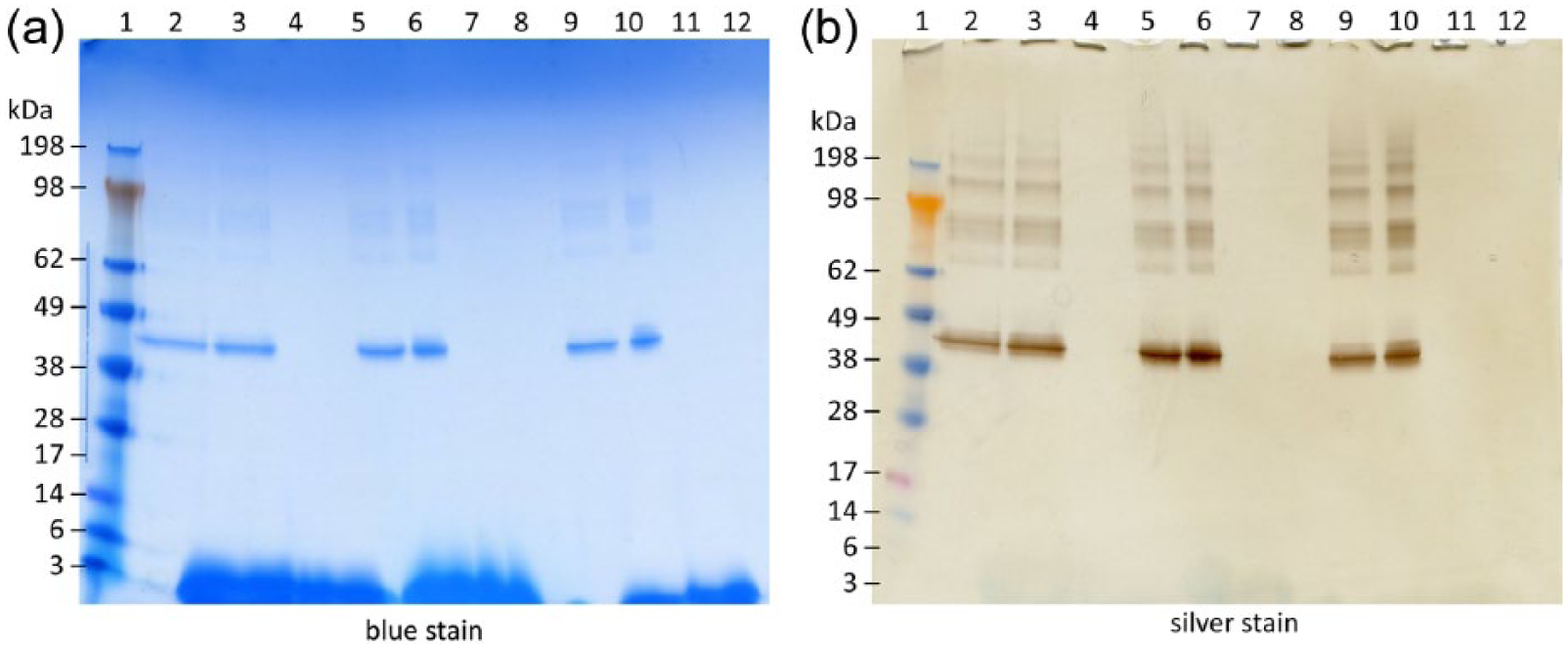
Squalene oil-in-water emulsion (SWE) with bovine serum albumin (BSA) with modified sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) methods. BSA (500 ng per well) (a) Coomassie Blue stain, (b) silver stain. Molecular weight (MW) marker (lane 1), BSA diluted with 20% SDS (lane 2), BSA + SWE diluted with 20% SDS (lane 3), SWE diluted with 20% SDS (lane 4), BSA diluted with 4% Tween 80 (lane 5), BSA + SWE diluted with 4% Tween 80 (lane 6), SWE diluted with 4% Tween 80 (lane 7), 4% Tween (lane 8), BSA diluted with 2% Triton X-100 (lane 9), BSA + SWE diluted with 2% Triton X-100 (lane 10), SWE diluted with 2% Triton X-100 (lane 11), 2% Triton X-100 (lane 12).
Discussion
This study expands on recently reported work from our laboratory 9 focused on reducing adjuvant-induced artifacts with SDS-PAGE, which is a useful assay for characterizing protein antigens within vaccine formulations. We have studied the following parameters in order to reduce artifacts associated with protein formulations containing oil-in-water emulsion: the contribution from each emulsion component, prestain rinsing, sample heating time, presence of a reducing agent, different staining procedures and membrane transfer, and the effect of different detergents. As speculated previously, 5 smearing in emulsion-containing gels could be due to incomplete separation of aqueous and oil phases of the emulsion. However, when the SE is prepared in a detergent such as Tween 80 or Triton X-100, the upper smears are reduced and even removed according to the surfactant concentration, which might be explained by the fact that the oily phase is better dissolved by these detergents and the separation of all components of the SE is thereby facilitated. However, the Tween 80 and Triton X-100 preparations did not remove the streaking artifacts at the bottom of the Coomassie Blue-stained gels, in fact these detergents may have exacerbated it compared with 20% SDS (Figure 7). Interestingly, another SWE with different emulsifier composition did not show any artifacts at the top of the gel and reduced artifacts at the bottom of the gel when Coomassie Blue staining was used. A decreased or lack of interaction of the blue staining with the different surfactants used in the SWE emulsion may explain why SWE showed fewer adjuvant-induced artifacts. Nevertheless, as with SE, the streaking due to SWE was not apparent on the silver-stained gel. A limitation of the current study is that only one type of gel system (4–20% tris-glycine) was employed.
Coomassie is used in chemistry to stain lipids on silica gel thin-layer chromatography plates, 12 and it is known to stain a wide array of lipids such as cholesterol and phospholipids. It was demonstrated that at least part of the contribution to the spot in the bottom of the gel is due to the DMPC component (Figure 2). Nevertheless, compared with the Coomassie Blue-stained gels, all streaking artifacts were greatly reduced on the silver-stained gels and the gold-stained PVDF membranes. Since Coomassie Blue complexes proteins through a variety of mechanisms, including electrostatic, Van der Waals, and hydrophobic interactions, similar phenomena are likely at work in the complexation of the stain to DMPC-coated squalene droplets (DMPC is zwitterionic and squalene is highly hydrophobic), thus resulting in the streaking artifacts. 13 In contrast, silver and gold stains presumably bind through electrostatic interactions14,15 due to the highly charged nature of the metals, which may help explain a reduced tendency to bind hydrophobic SE components. In the case of the gold-stained PVDF membrane, it is also possible that the SE components do not transfer efficiently from the gel to the PVDF membrane.
Conclusion
This study focused on optimizing SDS-PAGE protocols in order to reduce the streaking artifacts in the presence of an SE adjuvant. While no single component of the emulsion was entirely responsible for the artifacts, we identified that the emulsifier DMPC was at least partially involved in these artifacts. Implementation of variations in the SDS-PAGE protocols resulted in some reduction in the magnitude of the streaking artifacts. However, the greatest effects in the Coomassie-stained gels were achieved with the use of different detergents, such as Tween 80 and Triton X-100, to dilute vaccine samples formulated with the SE, resulting in reduced adjuvant SDS-PAGE artifacts. Other effective methods to reduce adjuvant-induced artifacts included using different staining and/or membrane transfer methods; that is, silver-stained gels or gold-stained PVDF membranes are more sensitive alternatives to Coomassie SDS-PAGE and less susceptible to adjuvant-induced artifacts. The optimized methods were compatible with another squalene-based emulsion although the artifact pattern differed, indicating the importance of emulsion composition and the importance of researchers optimizing characterization assays based on the properties of each unique formulation. This study highlights the importance of developing improved protocols for the characterization of adjuvanted vaccines.
Footnotes
Funding
This work was supported in part with federal funds from the National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Services, under Contract No. HHSN272201400041C, and by the Bill and Melinda Gates Foundation under Grant No. OPP1130379.
Conflict of interest statement
The authors declare that there is no conflict of interest.
