Abstract
Background
Polymorphisms of genes involved in the regulation of the immune response are risk factors for achalasia, but their contribution to disease pathogenesis is unknown. Nitric oxide is involved both in immune function and inhibitory neurotransmission.
Objective
The objective of this article is to assess the association and the functional relevance of the CCTTT-inducible nitric oxide synthase (NOS2) gene promoter polymorphism in achalasia.
Methods
Genomic DNA was isolated from 181 achalasia patients and 220 controls. Genotyping of the (CCTTT)n repeats was performed by PCR and capillary electrophoresis, and data analyzed by considering the frequency of the different alleles. HT29 cells were transfected with iNOS luciferase promoter-reporter plasmids containing different (CCTTT)n.
Results
The alleles’ distribution ranged from 7 to 18, with a peak frequency at 12 repeats. Analysis of the allele frequencies revealed that individuals carrying 10 and 13 CCTTT repeats were respectively less and more frequent in achalasia (OR 0.5, 95% CI 0.3–0.5 and OR 1.6, 95% CI 1–2.4, all p < 0.05). Long repeats were also significantly associated with an earlier onset of the disease (OR 1.69, 95% CI 1.13–2.53, p = 0.01). Transfection experiments revealed a similar allele-specific iNOS transcriptional activity.
Conclusion
The functional polymorphism (CCTTT) of NOS2 promoter is associated with achalasia, likely by an allele-specific modulation of nitric oxide production.
Introduction
Idiopathic achalasia is a rare esophageal motor disorder characterized by aperistalsis and defective relaxation of the lower esophageal sphincter (LES), leading to bolus impaction and symptoms of dysphagia and regurgitation.1-3 Although a wealth of evidence points toward the loss of the nitrergic innervation as the underlying pathophysiological abnormality of achalasia; 4 the mechanism leading to this selective neurodegeneration remains to be elucidated.
Hereditary, neurodegenerative, infectious and autoimmune mechanisms have all been forwarded as putative pathogenetic hypotheses and to date achalasia is widely considered a multifactorial disorder. In the wake of the strong pathogenetic role of T. Cruzii infection in Chagas disease, it has indeed been suggested that sporadic achalasia, as well, may be the result of a self-sustained inflammatory process secondary to acute gastrointestinal infections and that individual susceptibility of developing achalasia following such an initial trigger may be genetically determined.5,6 Achalasia, albeit rarely inherited, has indeed been associated with several polymorphisms in genes involved in the regulation of the immune response7-12 and the control of esophageal motility.13-20 In this complex scenario, nitric oxide (NO) represents a unique molecule since, depending on its concentration, it is involved in either inhibitory neurotransmission, or defense against infections.21-23
NO is constitutively produced by endothelial (eNOS or NOS3) or neuronal (nNOS or NOS1) NO synthases and, at higher concentrations, by the inducible form of NO synthase (iNOS or NOS2), 24 under stimulation of a variety of proinflammatory cytokines.20-22 Despite its antitumoral and antimicrobial activities,25,26 aberrant iNOS expression may have detrimental consequences as excessive NO production has been proved to exert neurotoxic effects, particularly for nitrergic neurons. NO release mediated by iNOS isoform may, indeed, induce transcriptional downregulation of nNOS, thus eventually leading to impaired nitrergic innervation.27-29
iNOS-dependent NO release is genetically determined and different iNOS gene promoter polymorphisms have been involved in individual responses to infection-induced immune activation. 30 The highly polymorphic pentanucleotide (CCTTT)n repeat located in the iNOS gene promoter region may be functionally relevant for the regulation of iNOS gene transcription. 31 The distribution of pentanucleotide microsatellite (CCTTT)n alleles has been studied in different ethnic groups and it has been associated with predisposition to infectious and autoimmune diseases.32-35
Based on this background, we aimed to examine whether the polymorphic pentanucleotide (CCTTT)n of the iNOS gene promoter is involved in the susceptibility to suffer from idiopathic achalasia and to investigate the functional role of this genetic polymorphism.
Materials and methods
Study participants
A total of 181 consecutive adult unrelated Caucasian Italian achalasia patients (male 97, mean age 56 ± 18 years) were recruited from October 2008 until November 2010. Diagnosis of achalasia was based on standard clinical, radiological, endoscopic tests and confirmed by esophageal manometry according to international criteria. 36 None of the patients had a family history of achalasia so all were considered as sporadic cases; furthermore 12 patients with comorbid autoimmune disorders (five patients with diabetes mellitus type I, six with rheumatoid arthritis, one with primary biliary cirrhosis) were excluded from the study.
A group of 220 healthy white, unrelated individuals (130 males, mean age 50 ± 13 years) without symptoms of or a history of gastrointestinal disease were included as ethnically matched controls. The control group consisted mainly of blood donors and ethnically matched hospital employees. All individuals gave their consent to participate in the protocol and the study was approved by the University Ethics Commitee.
Genotyping
Total DNA was extracted from peripheral blood leukocytes using the Nucleon BACC Genomic DNA Extraction Kit (GE Healthcare Europe GmbH, 79111 Freiburg, Germany). The iNOS pentanucleotide alleles were analyzed after polymerase chain reaction (PCR) amplification with the following set of primers: forward 5′-FAM ACCCCTGGAAGCCTACAACTGCAT-3′ and reverse 5′-CCACTGCACCCTAGCCTGTCTCA-3′. The size of the labeled PCR products was analyzed by capillary electrophoresis on an ABI PRISM 3130 sequencer with a GeneScan 500LIZ size standard.
Constructions of iNOS luciferase promoter-reporter plasmids containing different numbers of (CCTTT)n repeats
PCR was used to obtain a 1.2 Kb fragment immediately upstream of the transcription start site of the human iNOS gene (pINOS). The forward primer 5′-CAAAGTGTTGGTACCGTGAGATCA-3′ is located –1183 bp from the transcription start site and the reverse primer 5′-CTTCGGGACTCTCGAGAACTGCCCAG-3′ is located + 122 bp.
The PCR product was cloned into a pGL4 vector (Promega Madison, WI, USA), which contains the promoter without firefly luciferase reporter.
The (CCTTT)n pentanucleotide repeat region was cloned into the pGL4 construct using a pair of primers, 5′-ATGGAGGTACCATGGCATCCTGATTATCTCCA-3′ (forward) and 5′-TTCCAAGATCTAAGCAGGAATGAGGCTGAGT-3′ (reverse), by directional PCR from human genomic DNA obtained from individuals with different repeats. We obtained constructs with 9, 10, 11, 12, 13, 14, 15 and 16 repeats. All the constructs were also sequenced to confirm the authenticity of the PCR products.
Cell cultures, transient transfections, cell induction and luciferase assays
Human colon adenocarcinoma grade II cell line, HT29 (Sigma, Milan, Italy), was maintained in Dulbecco’s Modified Eagle’s Medium (Sigma) supplemented with 10% (v/v) heat-inactivated fetal bovine serum at 37℃ in a humidified 5% CO2-containing atmosphere. Cell cultures were kept sub-confluent and transiently transfected for luciferase assays.
Transfection of HT29 was performed with Lipofectamine 2000 (Invitrogen Inc, USA). In brief, the day before transfection, cells were plated into Falcon 12 well plates at a density of 1.5 × 105/ml in Optimem medium (Invitrogen Inc, USA). Cells were transiently transfected with 0.25 µg of each construct. To normalize the luciferase assay, 0.025 µg of the pRL-CMV vector (Promega) coding for the Renilla luciferase was transiently co-transfected. The pGL4-null was used as a negative control, whereas the pCMVluc (0.05 µg) was the positive control for the assay. After transfection, cells were treated for four hours with a mixture containing bacterial lipopolysaccharide (LPS) (10 µg/ml) (Sigma), interferon gamma (IFNγ) (100 units/ml) (R&D Systems, Minneapolis, MN, USA) and tumor necrosis factor alpha (TNFα) (2 ng/ml) (Sigma) for iNOS induction. Cell extracts were prepared 24 hours after induction, and 40 µl of lysate was used for the determination of luciferase activity using the Dual-Luciferase Reporter Assay System (Promega) on a 20/20 n luminometer (Turner Biosystems, Sunnyvale, CA, USA), according to the manufacturers’ protocols.
Statistical analysis
Results are given as number of cases and percentages for categorical data, and as mean ± standard deviation for quantitative variables. Data were analyzed by use of t-test for independent samples in case of quantitative variables and with the Fisher exact test in case of categorical variables. Association among iNOS CCTTT polymorphism and the presence of achalasia was quantified through the use of crude and stratified odds ratio (OR).
The statistical significance level was set at 5% (α = 0.05), and two-tailed tests were used throughout. Confidence intervals (CIs) are based on 95% CI. All the statistical analyses have been realized using R version 3.01.
Results
Patients’ demographic and clinical features
Demographics and clinical features of achalasia (181) and control participants (220)
Association between iNOS CCTTT gene polymorphisms and achalasia
The distribution of alleles having various repeat numbers ranged from 7 to 18 CCTTT and is shown in Figure 1. In patients as well as controls the number of (CCTTT)n repeats showed a central distribution and a peak frequency at 12 repeats. This pattern was similar to that reported previously for the white Caucasian population.
32
Allelic distribution of (CCTTT)n in achalasia and control participants. The allele distribution ranged from 7 to 18 with a peak frequency at 12 repeats in patients as well as controls. Individuals carrying 10 (CCTTT) repeats showed a reduced risk of developing achalasia, while no significant differences were observed in the allelic distribution of the other (CCTTT) repeats.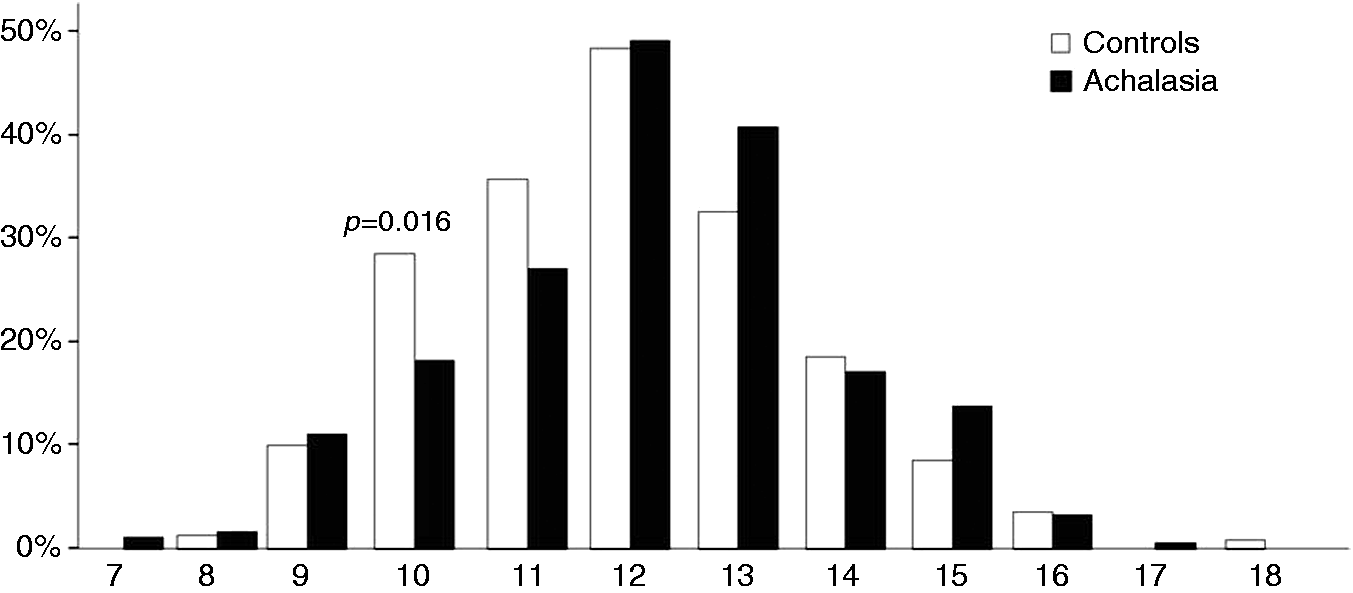
Frequencies of (CCTTT)n alleles in achalasia and healthy individuals
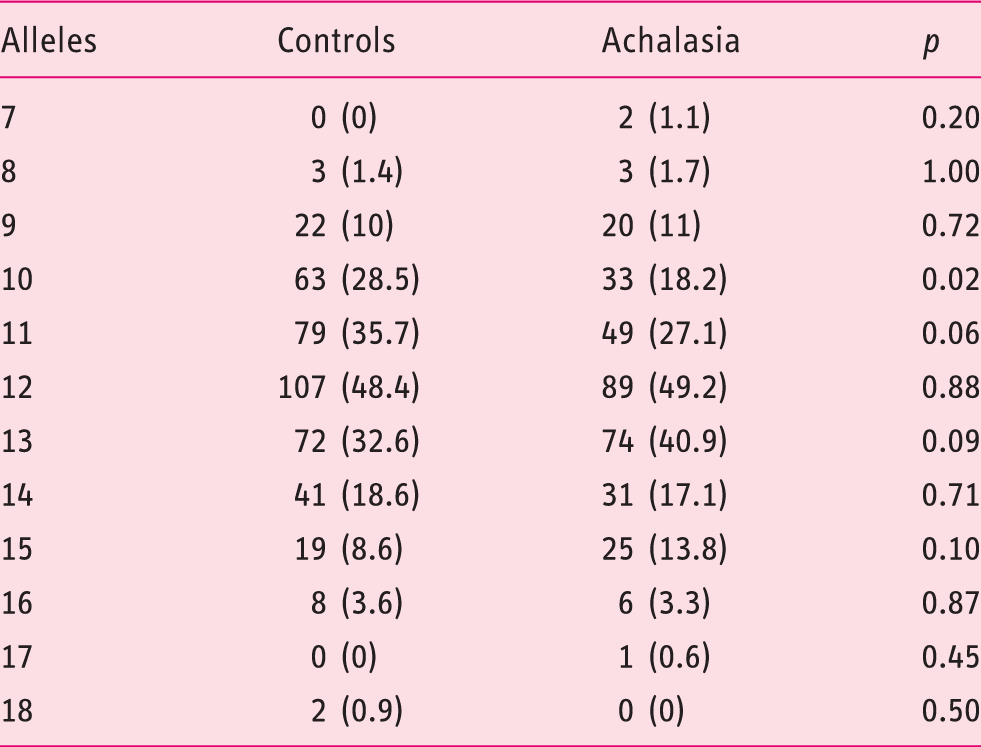
When data were stratified by gender, it emerged that among the females, those with 10 and 13 CCTTT repeats had a reduced and increased risk of achalasia, respectively (OR 0.39, 95% CI 0.19–0.80, p = 0.009 and OR 2.14, CI 95% 1.11–4.12, p = 0.022). In males, only those with 11 CCTTT repeats had a significant reduction in achalasia susceptibility (OR 0.52, 95% CI 0.29–0.93, p = 0.026).
Previous studies classified CCTTT alleles into short (7–11) and long (12–18) forms, according to the number of repeats.37-39 Thus, by sorting our cohort into long and short alleles, we found that individuals with CCTTT>11 had a significantly increased cumulative risk for achalasia (OR 1.69, 95% CI 1.13–2.53, p = 0.01); Figure 2 shows that gender likely influences such an effect, as this association was significant in males (OR 2.01, 95% CI 1.16–3.46, p = 0.012), but not in females (OR 1.42, 95% CI 0.77–2.62, p = 0.261).
Allelic distribution of long (7–11) and short (12–18) (CCTTT) repeats by gender in achalasia and healthy individuals. Males carrying the long alleles form had an increased risk of having achalasia (odds ratio (OR) 2.01, 95% confidence interval (CI) 1.16–3.46, p = 0.012).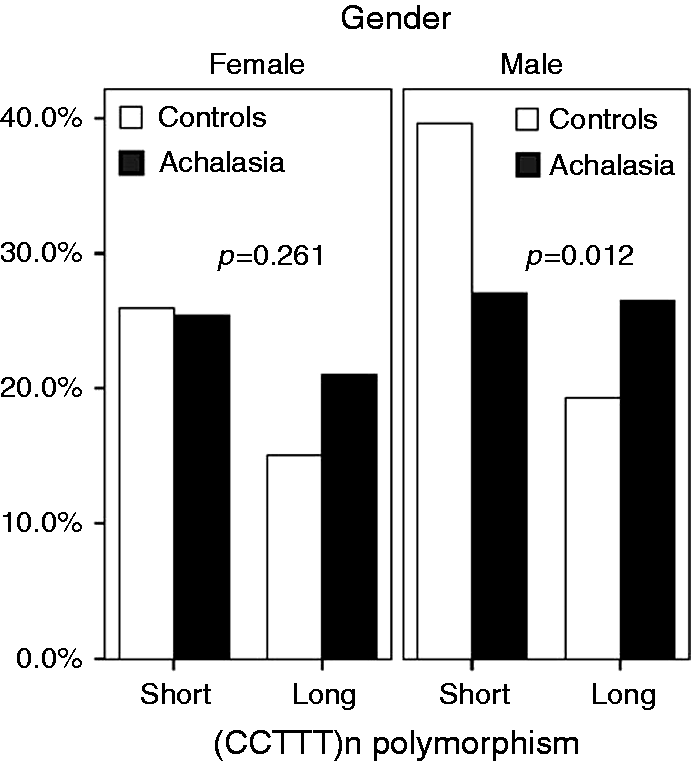
Effect of iNOS CCTTT polymorphisms and age of onset of achalasia
To evaluate whether the iNOS CCTTT polymorphism could represent a risk factor making some individuals more susceptible to the development of achalasia, we evaluated the effect of (CCTTT)n on achalasia onset. We failed to find any significant association between single different CCTTT repeats and age or the duration of the disease (data not shown). However, when we computed the analysis by considering the short and long CCTTT alleles forms, we found that patients carrying a longer number of CCTTT repeats (i.e. >11) develop the disease significantly earlier than those with short alleles (42 ± 18 vs. 51 ± 17 years, p = 0.01).
Effects of different CCTTT polymorphisms on transcriptional activity of the iNOS gene
To determine whether variable numbers of the CCTTT repeats were associated with the modulation of gene expression, we evaluated promoter activities using a variety of iNOS promoter-luciferase constructs in transient transfection experiments.
As shown in Figure 3, luciferase activity was almost silent in cells transfected with the empty vector; in contrast, in cells transfected with vectors containing the promoter region of the iNOS gene, luciferase activity was significantly increased. More interestingly, stimulation with LPS, IFNγ and TNFα resulted in a gradual and significant increase of luciferase activity, along with the increasing number of CCTTT repeats, with a peak occurring for constructs bearing 12 and 13 (CCTTT) repeats, and the lowest transcriptional activity for 10 (CCTTT).
Effects of varying numbers of (CCTTT)n repeats on NOS2 gene transcription. The CCTTT repeat sequences enhance the minimal NOS2 promoter induction in response to LPS, IFNγ and TNFα stimulation, with a significant increase in luciferase activity (*p < 0.0001 vs. unstimulated). Analysis of the difference among the different (CCTTT)n within the stimulated group revealed that constructs containing 12 repeats produced a significantly greater induction of luciferase as compared to all the other constructs (°p < 0.01), whereas the 13-repeat construct produced significantly greater luciferase activity than the 9, 10, 14, 15 and 16-repeat constructs (#p < 0.01 and ## p < 0.05, respectively). Conversely, the 10-repeat construct was associated with a significantly lower transcriptional activity than 11, 12 and 13 constructs (all p < 0.01). Data are the mean of six determinations and expressed as mean ± SEM. Data analysis was performed by ANOVA with Bonferroni post-test. NOS2: nitric oxide synthase; LPS: lipopolysaccharide; IFNγ: interferon gamma; TNFα: tumor necrosis factor alpha; ANOVA: analysis of variance.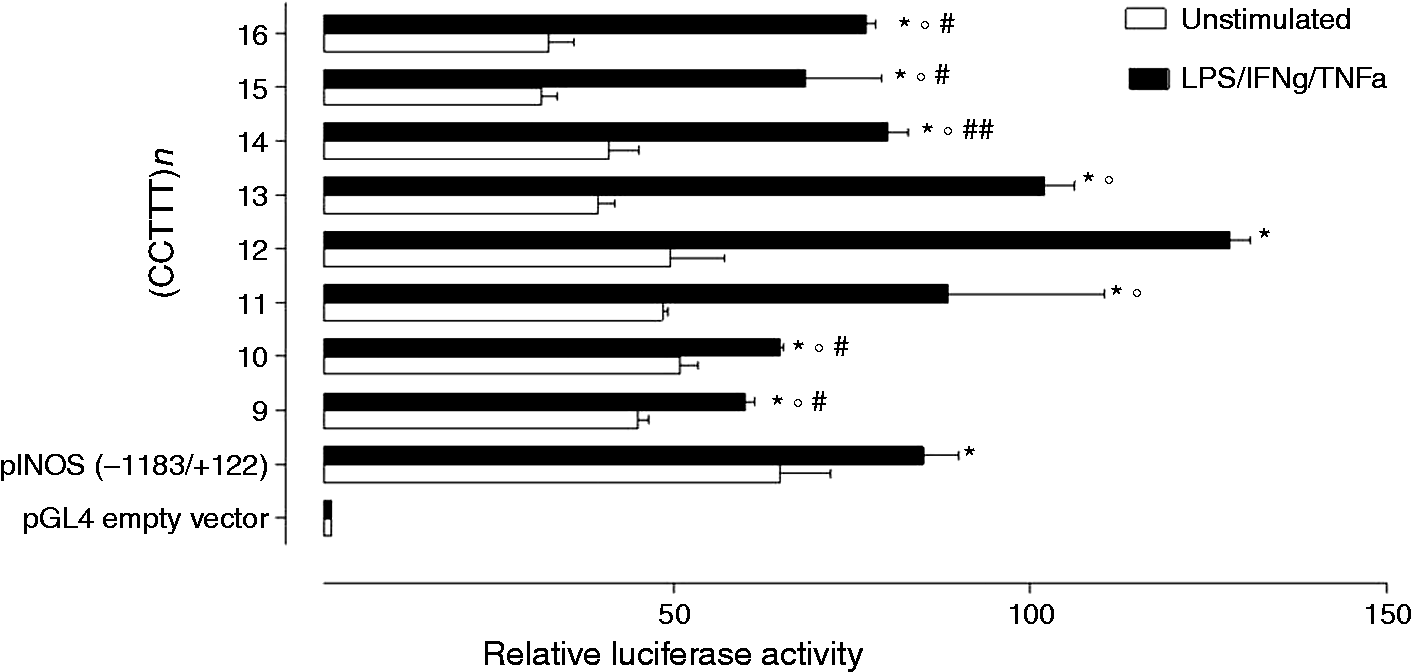
Conclusions
Idiopathic achalasia is the best characterized esophageal motor disorder, nonetheless its pathogenesis is not yet resolved. The occurrence of familial achalasia and its association with well-defined genetic syndromes suggests the involvement of genetic factors.40-44
To date, several genetic association studies have been reported in achalasia and some of these studies focused on candidate genes possibly linked to achalasia through their involvement in particular cell pathways, above all the regulation of immune response and the inhibitory neurotransmission.6-11,16
Although the genetic contribution to achalasia has been strongly supported,10,11 the clinical relevance of the reports are hampered either by the low number of studied patients or by the weak pathophysiological translation of the studied genes. NO may represent an ideal candidate to explain inhibitory nerve degeneration occurring in achalasia patients because it is involved both in defense against infections and inhibitory neurotransmission, and excessive concentrations of NO have been demonstrated in vitro to be neurotoxic, particularly for NOS-expressing neurons.29,45,46
Several single-nucleotide (SNP) or microsatellite (STR) polymorphisms have been described in the iNOS promoter region and many attempts have been made so far to investigate their possible functional significance in modulation of iNOS expression.27-29 Although iNOS activity can be regulated by post-transcriptional mechanisms, the human iNOS gene is regulated predominantly at the transcriptional level by a complex cytokine combination including IFNγ, interleukin (IL)-1β and TNFα.
In the present study we investigated the role of a longer polymorphic pentanucleotide repeat in the iNOS gene promoter region that has already been linked to predisposition to different immune-mediated conditions like infectious or degenerative diseases.46,47 The distribution of (CCTTT)n pentanucleotide in our population reproduced that observed in the Caucasian population in previous studies with a peak frequency at 12 repeats. 35 As reported from others association studies,40,48 in the subgroup of female participants, a significantly reduced or increased risk of developing achalasia was observed in individuals carrying 10 and 13 (CCTTT) repeats, respectively. Sorting into long (12–18) and short (7–11) alleles in our cohort, we found that long alleles were more frequent in achalasia patients than in the control group, and, although this association is significant in male patients but not in females, it is likely that this may be a result of the small analyzed sample size. In addition we also provide evidence that among achalasia patients, those with longer (CCTTT)n have a significant risk of developing an earlier onset achalasia as compared to those with the shorter alleles form, further indicating that such a genetic background, if present, may account for a more premature disease onset.
Few studies have tried to assess whether polymorphisms in the nNOS, iNOS or eNOS genes were associated with achalasia yielding contrasting results.14-16 Some of these studies failed to produce any conclusive association because no significant differences in genotypes and allele distribution were found, likely excluding a causative role of these polymorphisms.14,15 In a more recent study from India the iNOS22GA and nNOS29TT genotypes were identified as risk factors for achalasia, respectively. 16 However, it is of note that though the SNP of iNOS gene explored by Mearin et al.14 is an exonic polymorphism, it does not determine a change in the amino acid sequence, and its functionality in iNOS expression is still unclear. On the contrary, the (CCTTT)n microsatellite is a polymorphism that has already been linked to several autoimmune and degenerative disorders and has a well-established functional significance in the regulation of iNOS gene expression. Longer repeat numbers of this polymorphism had indeed been related with a higher iNOS transcriptional rate induced by IL-1β. 33 In our research, the luciferase activities of cells transfected with vectors containing the promoter region of the iNOS gene gradually increased along with the increasing number of CCTTT repeats, until the constructs with 12 and 13 (CCTTT) repeats, thus pointing out the functional relevance of this polymorphism in regulating the expression of the iNOS gene, and likely reflecting an increased production of NO. Similarly, the observation that constructs with 10 repeats was associated with a lower transcriptional activity seems to suggest that the reduced NO production is protective and is in line with the reduced risk of achalasia in individuals carrying such an allele.
Since there is evidence suggesting that iNOS-dependent NO production could induce downregulation of nNOS expression, 28 one can speculate that, under proinflammatory stimuli, high levels of NO may contribute to impair the normal functioning of nitrergic neurons thus leading to the highly selective neurodegeneration observed in achalasia patients. In this context, our study is limited because we did not study the nNOS in our patients and thus our pathogenetic hypothesis remains speculative. However, a more detailed knowledge of cellular responses in vivo would be warranted to identify the complex interplay between iNOS and nNOS isoforms and since several post-transcriptional modifications in NOS genes have been described so far, this cannot be obtained by studying isolated genetic polymorphisms on peripheral blood mononuclear cells (PBMCs). 49
Here we provide evidence that genetic variations in the promoter region of the iNOS gene are associated with the susceptibility to achalasia. Furthermore, we demonstrated that patients carrying longer alleles have a significant risk of developing an earlier onset of achalasia, possibly as a result of increased iNOS gene expression. Although limited by the low number of the studied population, our data provide a plausible pathophysiological mechanism to explain the selective neurodegeneration and the reduced nNOS expression occurring in the myenteric plexus of achalasia patients. Therefore, larger multicentric studies aimed at understanding molecular mechanisms regulating iNOS gene expression could help to pave the way to novel therapeutic tools able to control excessive NO production and also to identify genetic factors determining the susceptibility to achalasia.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was partially funded by Regione Campania (L.R. 5 2008) to G.S.
