Abstract
Background
Clinical use of hepatitis B viral (HBV) quantitative seromarker\s remains questionable since it is not precisely known whether they represent intrahepatic viral replication. Covalently closed circular DNA (cccDNA), relaxed circular DNA (rcDNA), and pregenomic RNA (pgRNA) are more likely to represent active HBV replication and their measurement can be used to derive virion productivity (VP; rcDNA/cccDNA), subviral particle (SVP) productivity (quantitative HBsAg/cccDNA), and replicative activity (RA; pgRNA/cccDNA). These can be used to compare relative HBV replication between HBeAg-negative and -positive patients.
Objective
To study the clinical significance of intrahepatic HBV replication phenomenon between HBeAg-negative and -positive patients and its correlation with quantitative HBV seromarkers.
Method
This was a prospective study between January 2010 and December 2011. Study subjects were naive chronic hepatitis B patients from Cipto Mangunkusumo and Medistra Hospitals. All patient samples underwent liver biochemistry and HBV seromarkers testing (HBeAg, quantitative HBsAg and HBV DNA levels), and patients underwent liver biopsy. Stored liver specimens were analysed for intrahepatic rcDNA, cccDNA, and pgRNA with quantification performed by real-time PCR. Comparison of HBV markers between HBsAg-positive and -negative patients was carried out using the Mann–Whitney U-test. Pearson’s correlation test was performed among HBV intrahepatic and seromarkers using their log-transformed values.
Results
A total of 104 patients were enrolled in this study; 54 (51.9%) were male. Patients’ mean age was 41.9 ± 11.63 years (range 19–70 years). Sixty-one patients (58.7%) were HBeAg-negative. All HBV markers were significantly higher in HBeAg-positive than HBeAg-negative patients, except for SVP productivity and RA. Serum HBV DNA was strongly correlated with intrahepatic total HBV DNA (r = 0.771), cccDNA (r = 0.774), and rcDNA (r = 0.780) while serum quantitative HBsAg showed only moderate correlation with intrahepatic total DNA (r = 0.671), cccDNA (r = 0.632), rcDNA (r = 0.675), and SVP productivity (r = 0.557).
Conclusions
Serum HBV DNA concentration and quantitative HBsAg might not accurately predict intrahepatic viral activity. Virion and SVP production do not occur in parallel with replicative activity.
Keywords
Introduction
Treatment of chronic hepatitis B (CHB) infection relies upon monitoring serological markers (HBsAg, HBeAg) and HBV DNA levels. 1 –3 Decreases in HBV DNA levels are used as an indicator of efficacious antiviral therapy, 4 whereas HBsAg loss is considered the ultimate goal of serological viral response. In patients with positive HBeAg, HBeAg seroconversion is a desirable treatment endpoint.
Complete eradication of HBV by antiviral therapy is still difficult to achieve because of the persistence of viral covalently closed circular DNA (cccDNA) in the hepatocyte nucleus. 5,6 This HBV replicative form is produced from the partially double-stranded genomic DNA (relaxed circular DNA, rcDNA) in the virion, which after infection, is converted by host cell enzymes in the nucleus into a fully double-stranded form. From this, HBV cccDNA molecule is generated as the transcriptional template for HBV RNA production. The hepatocyte RNA polymerase II transcribes all the viral mRNA from the HBV cccDNA, including the HBV pregenomic RNA (pgRNA). 7
The pgRNA functions as the mRNA not only for the polymerase and core protein production, but it can also be reverse transcribed after encapsidation into new genomic rcDNA. Capsids containing the newly formed genomic DNA can be enveloped by surface proteins and secreted into the blood or redirected back to the nucleus to maintain a pool of HBV cccDNA. 6 Therefore, the measurement of pgRNA provides more accurate information on viral replication and the ratio of pgRNA per cccDNA molecule reflect the true viral capacity to replicate its genomic material. This ratio has been termed as replicative activity (RA). Presuming in established CHB infection that most of the intrahepatic rcDNA is aimed for virion production, it has been proposed also that the ratio of rcDNA per cccDNA molecule reflects virion productivity (VP). 8 Since not all new rcDNA will be part of the new intact virion, neither rcDNA nor RA can be used to reflect the capacity of viral particle production. Virion productivity may thus be a measure of the actual production of viral particles.
Recently, quantitative HBsAg has been introduced to help predict treatment response in CHB patients. This marker is believed to better reflect intrahepatic viral replication than serum HBV DNA levels. 9,10 Indeed, HBsAg titres have been proposed as a surrogate marker for transcriptionally active HBV cccDNA, and in a recent study of treatment-naive patients with CHB, an association between HBsAg titres, cccDNA, and total intrahepatic HBV DNA was shown but only in HBeAg-positive patients. 11 However, commercial assays to quantify HBsAg detect all forms of surface antigen in the blood, either as an intact viral particle (virion) or as spherical and filamentous subviral particles (SVP). 12,13 Conflicting results have been published among studies conducted in different countries on the use of quantitative HBsAg in clinical practice and its exact role is yet to be determined. 14,15 The lack of association in HBeAg-negative patients indicates that the association of HBsAg titres with intrahepatic HBV DNA is not straightforward and is likely affected by both virus and host factors.
The different natural history of CHB infection and HBV genotypes between Western and Asian countries, and even between Asian countries themselves might be one of the reasons for the variation in results in clinical studies. 16,17 Molecular studies on intrahepatic HBV DNA kinetics may give a new perspective in understanding HBV replication and CHB, distinct from the biomarkers that are usually monitored in clinical practice. This study was aimed at evaluating the clinical significance of virion and SVP productivity by measuring intrahepatic HBV DNA replicative forms and serological markers in HBeAg-positive and HBeAg-negative patients.
Methods
Study design and subjects
This was a prospective cross-sectional study conducted in the Department of Internal Medicine, University of Indonesia from January 2010 to December 2011. All consecutive, treatment-naïve, newly diagnosed CHB patients from two big referral hospitals, Cipto Mangunkusumo and Medistra Hospitals, Jakarta who presented to the outpatient clinics and agreed to participate in the study after signing informed consent were enrolled. This study was approved by the Ethical Committee for Medical Research, Faculty of Medicine, University of Indonesia.
Chronic hepatitis B infection was diagnosed by the presence of HBsAg in the patient’s serum at least for 6 months based on a chemiluminescent microparticle immunoassay (CMIA; ARCHITECT HBsAg Reagent Kit; Abbott Diagnostics, Abbott Park, IL, USA). Inclusion criteria were newly diagnosed, treatment-naive CHB patients, who were willing to undergo liver biopsy before and after antiviral treatment. Patients were excluded if there was a coinfection with hepatitis C virus or human immunodeficiency virus, decompensated liver cirrhosis, or hepatocellular carcinoma. Quantitative serum HBsAg measurement, HBV DNA levels, and genotyping were done in a private laboratory in Jakarta, whereas measurement of intrahepatic markers (cccDNA and pgRNA assay) and identification of the PC/BCP variants were done in Victorian Infectious Disease Reference Laboratory, Melbourne, Australia.
Quantitative HBV seromarker measurement
Quantitative HBsAg was measured using CMIA. The results were expressed as signal sample/cutoff ratios (S/CO). The values were then calibrated (ARCHITECT HBsAg Calibrators) and verified (ARCHITECT HBsAg Controls) to obtain quantitative levels (IU/ml). Manual dilution was performed if the HBsAg value exceeding 250 IU/ml. The suggested dilution was 1:500, which was done by adding 25 µl specimens to 475 μl ARCHITECT HBsAg Manual Diluent for 1:20 dilution, followed by adding 20 µl of the first solution to 480 μl ARCHITECT HBsAg Manual Diluent for a 1:500 dilution. According to the manufacturer, the standard error was 0.018 and the coefficient of variation was 7.8%. Specimens were not analysed in duplicate. Serum HBV DNA level was measured with COBAS TaqMan HBV Test (Roche Diagnostics, Mannheim, Germany). The detection range was ∼30 IU/ml to 1 × 108 IU/ml. Samples were diluted and reanalysed if the titre was above the upper limit of quantification. Serum samples were obtained in the same day of liver biopsy.
Identification of HBV genotype and precore and basal core promoter variants
Hepatitis B virus genotyping was initially determined using line probe assay (INNO-LiPA HBV; Innogenetics, Belgium). HBV genotype was confirmed by sequence analysis of the HBV surface/polymerase gene as previously described. 18 Variants with substitutions in the precore (PC) region and basal core promoter (BCP) region were identified by sequencing after PCR amplification, as described by Ayres et al. 18 The PC variant was defined by the presence of the G1896A mutation and the BCP variant by the A1762T/G1764A mutation.
Liver biopsy
Liver biopsy was performed on all patients and histopathological assessment was carried out by an experienced liver pathologist who was blinded to the clinical data. Ultrasound-guided biopsy was performed using a 16-gauge Menghini needle (Hepafix; B Braun Melsungen, Melsungen, Germany) under local anaesthesia. Tissue specimens were first prepared for histopathology examination and the other portions of the biopsy stored at −80℃ or in an RNA stabilization solution at −20℃ (RNALater; Ambion, Austin, Texas, USA) for analysis, including both cccDNA and pgRNA quantification. Fibrosis stage was assessed using the METAVIR scoring system. 19,20
Intrahepatic total and cccDNA quantification
Determination of HBV cccDNA levels was carried out by real-time PCR using a LightCycler 1.0 and LightCycler version 3.5 software (Roche Diagnostics). Reaction volumes (20 µl) consisted of 2 µl extracted DNA, 5 mM MgCl2, 0.5 µM primer mix and 0.2 µM hybridization probes. Selective HBV cccDNA primers consisted of two upstream primers (CCC1 5′-GCGGWCTCCCCGTCTGTGCC-3′ and CCC3 5′-GTCTGTGCCTTCTCATCTGC-3′) and a downstream primer (CCC2 5′-GTCCATGCCCCAAAGCCACC-3′). FRET hybridization probes were 5′-GTTCACGGTGGTCTCCATGCGACGT-FL-3′ and 5′-LCR640-AGGTGAAGCGAAGTGCACACGGWCC-3′. Total intrahepatic DNA was measured using primers RC1 5′-CTCGTGGTGGACTTCTCTC-3′ and RC2 5′-CAGCAGGATGAAGAGGAA-3′ and FRET probes 5′- CACTCACCAACCTSYTGTCCTCCAA-FL-3′ and 5′ = LCR640-TGTCCTGGYTATCGCTGGATGTGTCT-3′. Quantification standards were derived by dilution of a linearized plasmid (pHBV EcoR1) containing a greater than full-length HBV genome of genotype D. This standard has been previously titrated against the WHO international HBV reference standard to correlate quantification values. Variation in the amounts of liver tissue was normalized by quantifying β-globin in each sample with the Roche DNA control kit. The relaxed circular HBV DNA was calculated as total HBV DNA – cccDNA.
Intrahepatic pregenomic (Pg) RNA measurement
The RNA extraction was performed using RNeasy Mini Kit (Qiagen, Germany). Fresh frozen liver tissues that were previously stored in RNALater tubes were disrupted by homogenization and the lysate was purified according to the manufacturer’s instructions. A further purification of RNA was carried out using MicroPoly(A) Purist kit (Life Technologies, CA, USA) to remove any DNA carryover. Levels of hepatic RNA were normalized using the commercially available human G6PDH Housekeeping Gene Set (Roche Diagnostics). After reverse transcription of RNA to cDNA using High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, CA, USA), amplification and quantification of pgRNA was as previously described. 21
Replicative activity, virion productivity, and subviral particle productivity
As well as quantification of HBV seromarkers and intrahepatic markers, there were several ratios derived from these markers. Replicative activity was a ratio of pgRNA production per cccDNA molecule (pgRNA/cccDNA). Virion productivity was calculated as rcDNA levels per cccDNA molecule (rcDNA/cccDNA). Subviral particle productivity was the quantitative serum HBsAg levels divided by cccDNA levels (qHBsAg/cccDNA).
Statistical analysis
Characteristics of the study subjects were presented descriptively as proportion or median. Comparison between HBeAg-positive and -negative were tested using the chi-squared test for categorical data and the Mann–Whitney U-test for skewed numerical data. Mean differences of log-transformed values were calculated using t-test. Pearson’s correlation test was applied on the log-transformed values of intrahepatic and serum markers. A p-value <0.05 was considered significant. Analyses were done using the statistical software SPSS version 15 for Windows PC (SPSS, Chicago, IL, USA).
Results
Characteristics of the study subjects
Characteristics of the study subjects
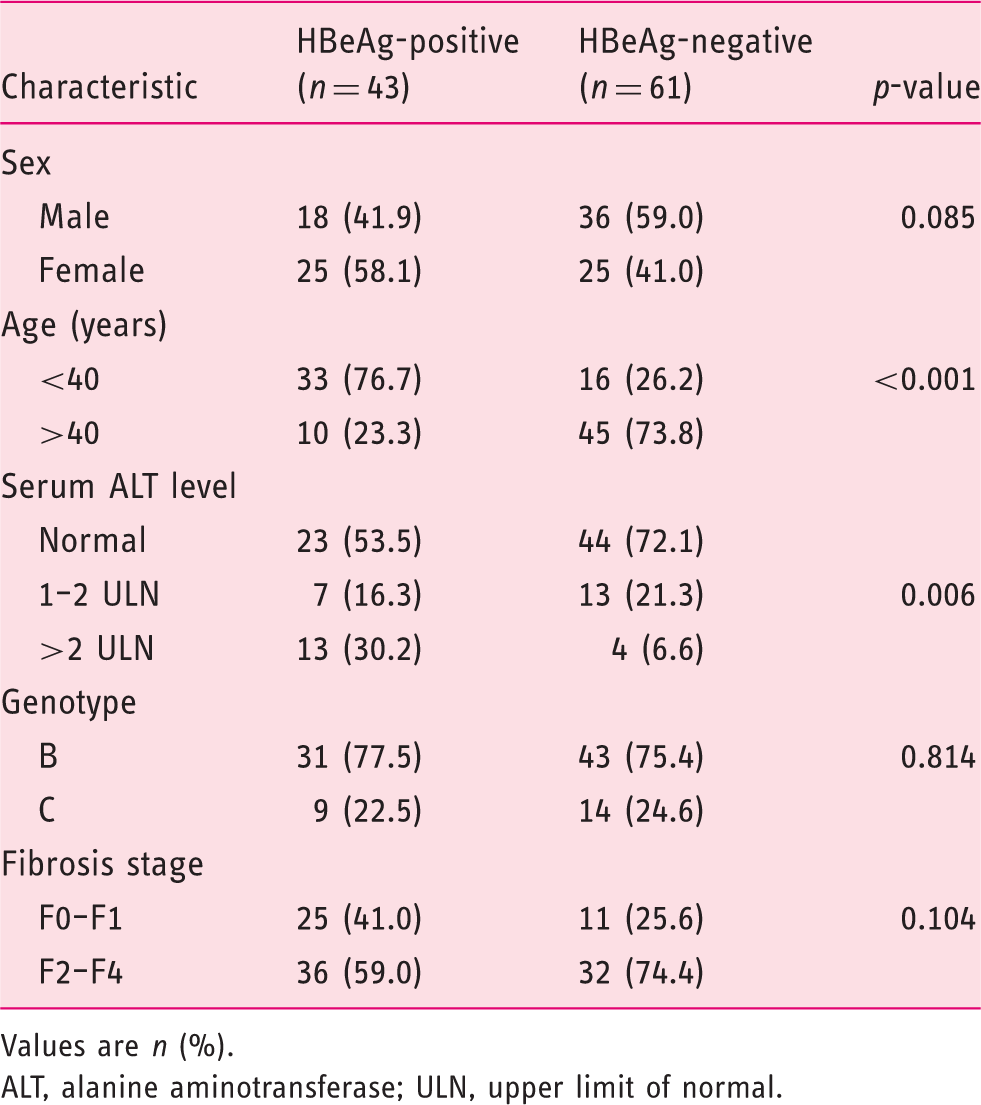
Values are n (%).
ALT, alanine aminotransferase; ULN, upper limit of normal.
Comparison between HBeAg-positive and -negative patients
Baseline characteristics of HBV markers
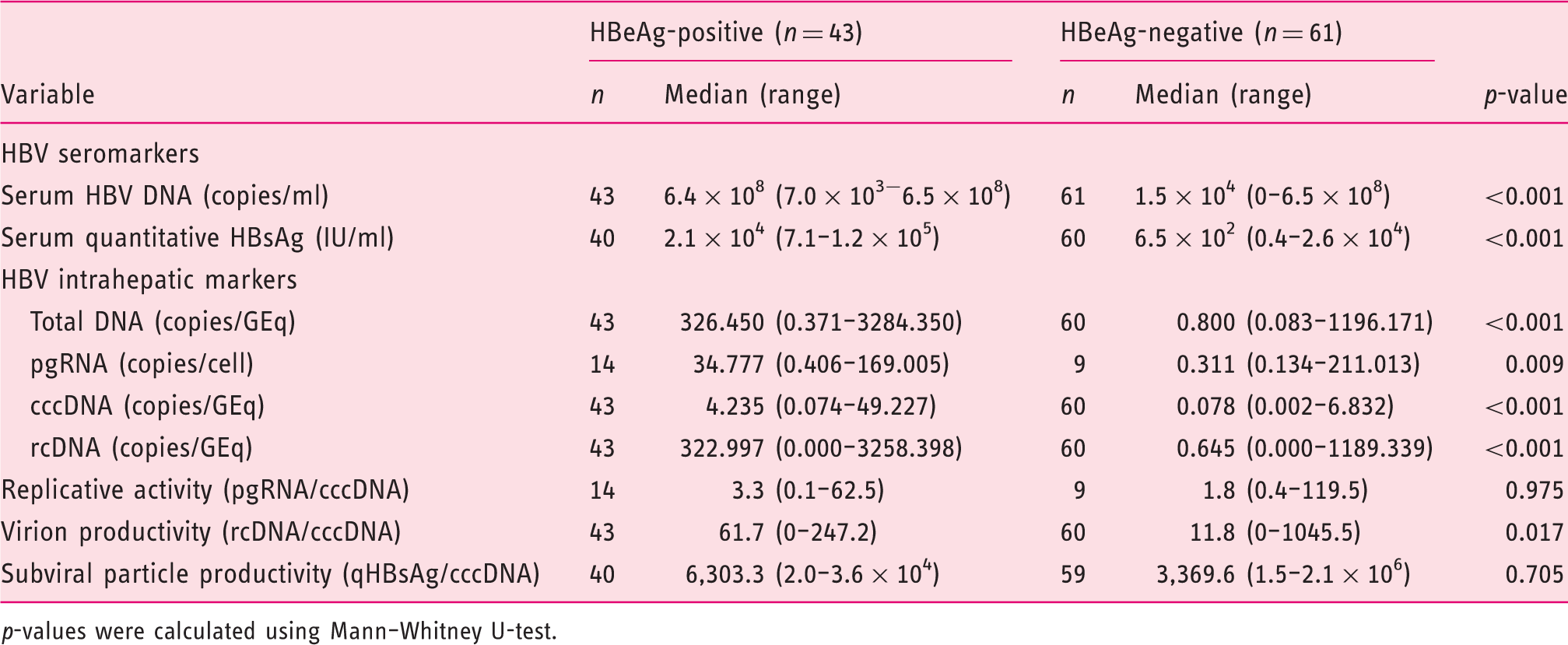
p-values were calculated using Mann–Whitney U-test.
Correlation between serum HBV DNA levels and intrahepatic HBV markers
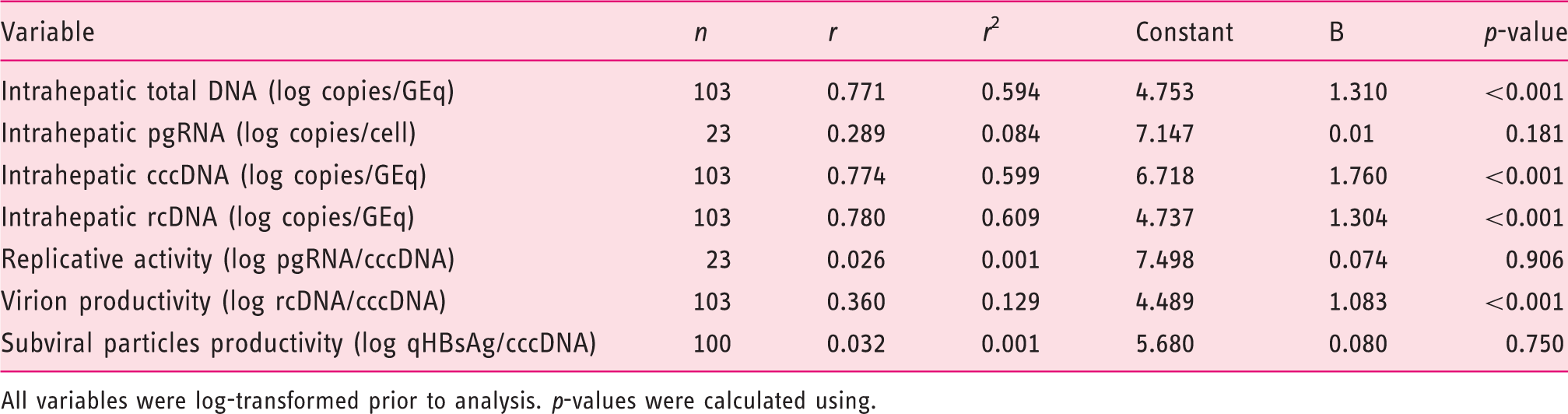
Serum HBV DNA levels can be predicted by the regression equation: serum HBV DNA (log copies/ml) = Bx + constant, where x represents each intrahepatic marker in their log values (Table 3). Serum HBV DNA was strongly correlated with intrahepatic total DNA (r = 0.771, r
2 = 0.594), intrahepatic cccDNA (r = 0.774, r
2 = 0.599), and intrahepatic rcDNA (r = 0.780, r
2 = 0.609). However, subgroup analysis showed only moderate correlation between intrahepatic total HBV DNA and serum HBV DNA levels in HBeAg-positive (r = 0.504, r
2 = 0.253) and HBeAg-negative (r = 0.651, r
2 = 0.423) patients (Figure 1a). Similar findings were observed between intrahepatic HBV cccDNA and serum HBV DNA levels in HBeAg-positive (r = 0.606, r
2 = 0.368) and HBeAg-negative (r = 0.552, r
2 = 0.305) patients (Figure 1b) and also between intrahepatic rcDNA and serum HBV DNA levels in HBeAg-positive (r = 0.454, r
2 = 0.206) and HBeAg-negative (r = 0.658, r
2 = 0.432) patients (Figure 1c). Serum HBV DNA levels showed weak or no correlations with intrahepatic pgRNA, RA, VP, and SVP productivity.
Correlations between intrahepatic total HBV DNA and serum total HBV DNA levels (a), intrahepatic cccDNA and serum HBV DNA levels (b), and intrahepatic rcDNA and serum HBV DNA levels (c). Correlation between intrahepatic markers and serum HBV DNA levels All variables were log-transformed prior to analysis. p-values were calculated using.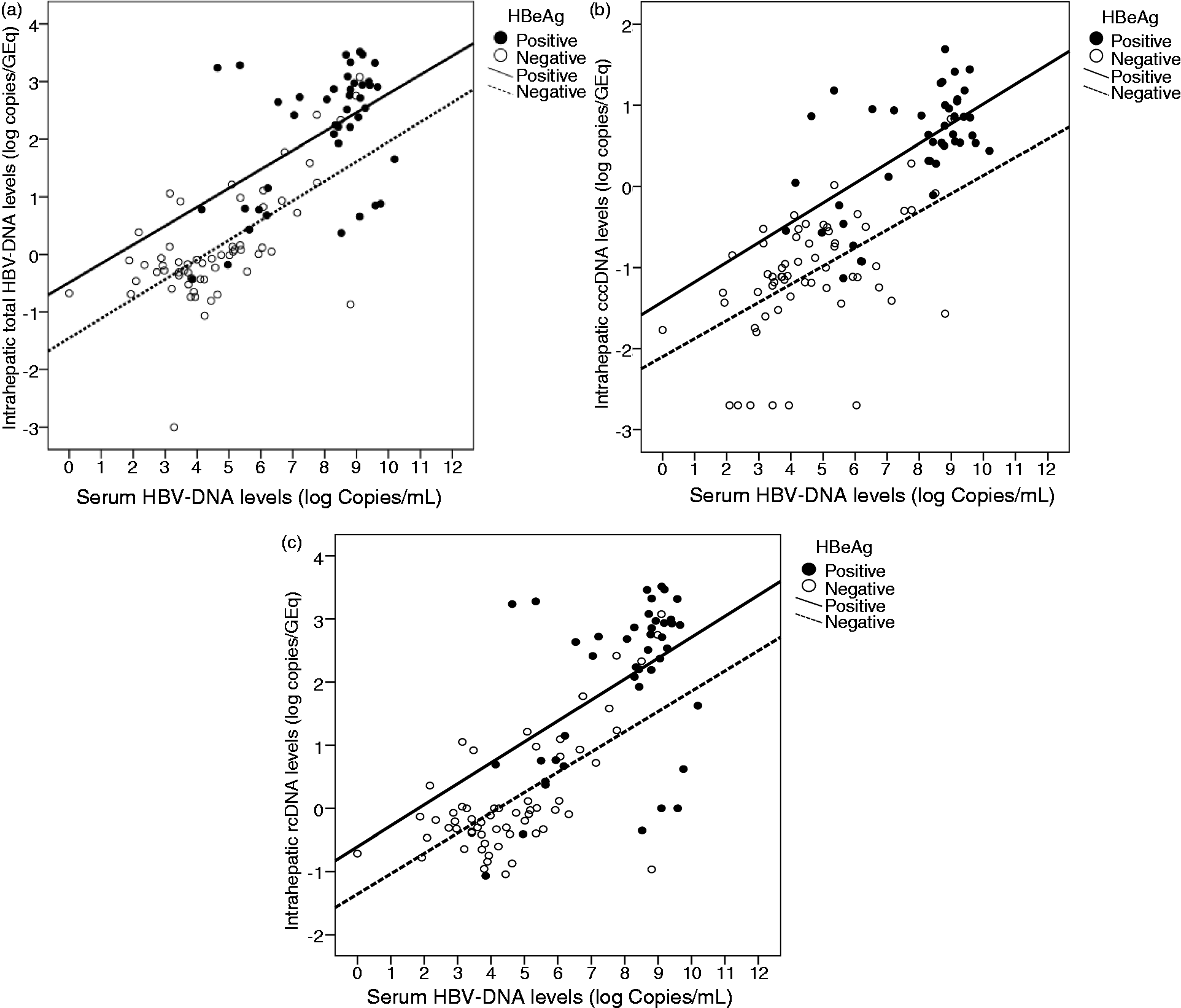
Correlation among serum quantitative HBsAg and other HBV markers
Correlation between intrahepatic markers and serum quantitative HBsAg levels
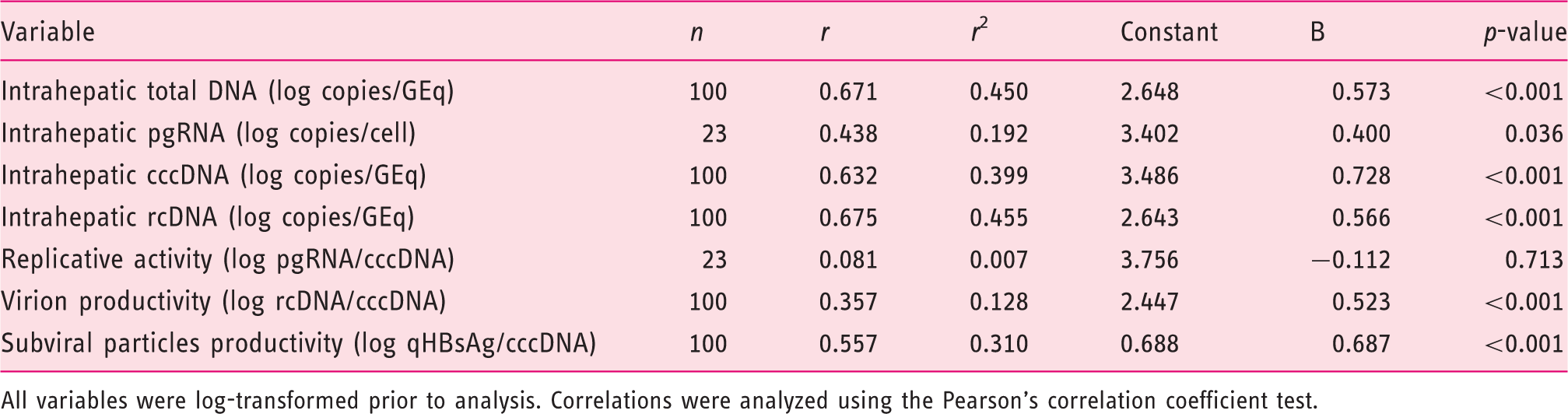
All variables were log-transformed prior to analysis. Correlations were analyzed using the Pearson's correlation coefficient test.
Difference levels of HBV markers between fibrosis F0–F1 and F2–F4
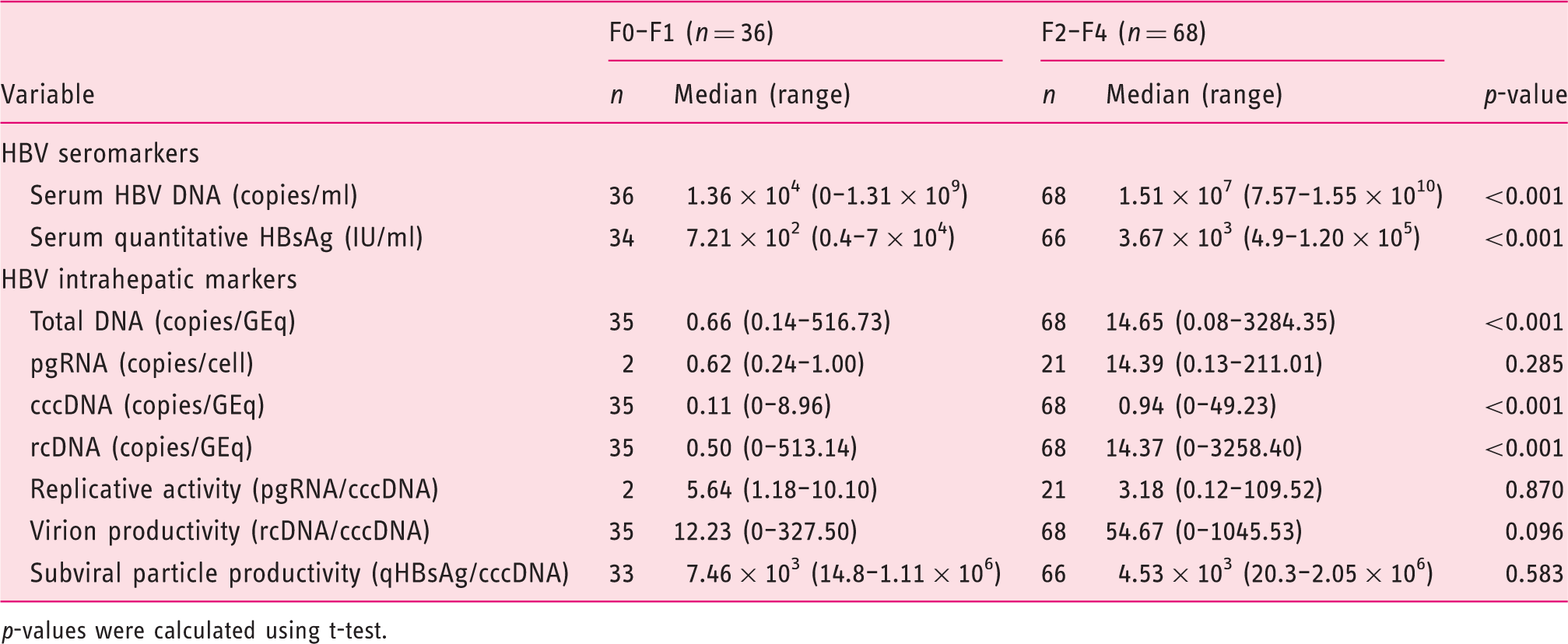
p-values were calculated using t-test.
PC/BCP mutation
Difference levels of intrahepatic HBV markers based on PC/BCP mutation both in HBeAg-positive and -negative patients

Discussion
To our knowledge, this is the first study to comprehensively document the relationship between intrahepatic HBV DNA replicative intermediates and quantitative HBV seromarkers (the ‘old’ HBV DNA level and the ‘new’ serum quantitative HBsAg levels) in CHB patients predominantly infected with genotype B. Patients aged less than 40 years were significantly more likely to be HBeAg positive, which is consistent with the predominant mother-to-infant mode of transmission. 22
Viral load, quantitative serum HBsAg levels, and all HBV intrahepatic markers were significantly higher in HBeAg-positive patients. These findings are consistent with previously reported studies 21,23 and antiviral therapy prospective studies. 9,10 Regarding the true replicative capacity, only VP (as defined by intrahepatic HBV rcDNA/cccDNA) provided the best indicator which was significantly higher in HBeAg-positive subjects. This suggest that VP may be a hallmark of immunotolerant or immunoclearance phase, when the patient is highly infectious. In contrast, RA and SVP productivity was similar, suggesting that, in HBeAg-negative subjects, pgRNA synthesis and viral capsid production may occur in the same proportion as with HBeAg-positive patients, but it is less likely to result in production of intact virions.
In this study, the term of ‘inactive carrier’ in some patients needs to be reevaluated, since even though these patients have low HBV DNA, they have been infected for such a long time and the data was collected with a cross-sectional design. Looking at the HBeAg-positive group, there were also patients in the immunotolerant phase; however, this might not influence the phenomenon since the transmission route might give a different perspective as most of our patients do not have a significant increase in alanine aminotransferase level.
This new insight might also imply that viral mutation in HBeAg-negative patients does not reduce viral capability to replicate its genetic material, but it may lack infective capacity. There are several mechanisms proposed to be involved in cccDNA regulation. First, mutations in the precore region and basal core promoter (PC/BCP) might influence cccDNA production through the transcription of pgRNA, which is under the control of the BCP. 24 However, unlike the previous report, 8 the PC/BCP mutation in our subjects was associated with higher VP despite the similar intrahepatic cccDNA levels. In addition, the proportion of PC/BCP mutation was not statistically different between HBeAg-positive and -negative patients (data not shown). This might be influenced by the predominant maternal transmission in developing countries. However, it is difficult to find out whether young patients were infected by mutant virus or already had a seroconversion at the time of diagnosis.
Secondly, the negative-feedback mechanism that regulates the recycling of nucleocapsid back to the nucleus 6 may contribute to cccDNA availability. The reduced VP in the HBeAg-negative patients could be meant that rcDNA in nucleocapsid are recycled back to the nucleus to replenish cccDNA pools. Immune-mediated mechanisms and host epigenetic factors, such as histone acteylation of the HBV viral minichromosome and HBV DNA methylation, have also been proposed to have a role in regulating cccDNA activity. 25
The availability of assays to quantify HBsAg has led to renewed interest in the potential to use the serum HBsAg level as an indirect measure of transcriptional activity. 26 Although the levels of HBsAg were considerably higher in HBeAg-positive patients in our study, the ratio of serum HBsAg per cccDNA (SVP productivity) was not significantly different between HBeAg-positive and -negative groups. This result was similar to a previous report 8 and implies that HBsAg production is influenced by intrahepatic HBV cccDNA levels. Our pgRNA quantification results may support this theory since the intrahepatic RA did not significantly differ between patients with high HBV DNA (as reflected by positive HBeAg) and low HBV DNA levels (as reflected by negative HBeAg). However, pgRNA purification and quantification is technically challenging and not in all cases could we achieve a successful result.
Correlation test between seromarkers and intrahepatic markers may explain whether seromarker concentrations can be predicted from intrahepatic marker values. The modest correlation observed between intrahepatic HBV DNA and serum HBV DNA levels in each HBeAg subgroup suggested that not all newly synthesized viral genome will be released into the bloodstream. This is supported by the weak correlation between serum HBV DNA levels and VP, implying that the rate of virion release cannot be predicted. Although HBeAg-positive patient produce more VP than HBeAg-negative, there is no evidence that high serum HBV DNA levels in a patient was caused by higher VP inside the hepatocyte. Furthermore, serum HBV DNA levels could not give any clue whether the virus is transcriptionally active (pgRNA level) or how much pgRNA is produced by a single cccDNA molecule (RA).
The moderate correlation among serum quantitative HBsAg and intrahepatic HBV DNA levels indicated that envelope production is only moderately dependent on the number of viral genome. Quantitative serum HBsAg levels also do not reflect the viral transcriptional activity and virion production rate. A recent study showed that a genomic variant at the PreS/S sequence might be responsible; no correlation between serum HBsAg and HBV DNA levels in patients infected with the preS/S mutant. 27
Altogether, this study showed that no seromarker could accurately reflect the true replicative activity of cccDNA. Seromarker changes do not represent changes in viral intrahepatic behavior. Patients with low seromarker concentrations are still showing a somewhat degree of intrahepatic active replication, especially those infected with mutant virus showing a HBeAg-negative phenotype.
This study also has limitations. First, serial liver biopsy was not done to evaluate the steps of viral replicative activity in a timely fashion as it should be in the natural history of CHB infection. However, serial intrahepatic cccDNA measurement for routine monitoring would not be an option since it is an invasive procedure with the potential for complications. Secondly, the pgRNA measurement could not be succeeded in large number of samples due to technical difficulties.
No international consensus is currently available on how to use the intrahepatic cccDNA and pgRNA assays. Our study may provide a better perspective for clinicians in the approach of CHB patient management. Combined serial measurements of quantitative serum HBsAg and HBV DNA levels are likely to remain the best way to monitor disease progression and treatment response in CHB patients receiving antiviral therapy. However, they do not tell physicians the true state of viral intrahepatic replication. Long antiviral therapy might be the only way to maintain viral suppression to prevent liver cirrhosis and hepatocellular carcinoma and repeated liver biopsy is still needed to evaluate liver tissue damage.
In conclusion, patients with positive HBeAg may show higher serum and intrahepatic markers and virion production, but the true replicative activity and subviral particle production may similar between HBeAg-positive and -negative subjects. Serum HBV DNA concentration and quantitative HBsAg could not accurately predict intrahepatic viral activity. Virion and SVP productions do not occur in parallel with replicative activity. Serial measurement of serum HBV DNA and quantitative HBsAg levels may be done to monitor treatment response, but they may not reflect the true viral intrahepatic replication. Liver biopsy is still the most accurate tool to monitor liver disease progression.
Footnotes
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Conflict of interest
The authors declare that there is no conflict of interest.
