Abstract
Objectives:
Ruxolitinib is used to treat myelofibrosis, polycythemia vera, and steroid-refractory graft-versus-host disease following allogeneic stem cell transplantation. This study aimed to determine the association between ruxolitinib and adverse events by evaluating case reports published between January 2014 and March 2024 in the Japanese Adverse Drug Event Report database.
Methods:
The signals for the ruxolitinib–adverse event association were identified using propensity score-adjusted reporting odds ratio analysis. Data obtained from the drug–gene interaction, drug signature, search tool for chemical interactions, and interaction reference index databases were used to construct a drug–gene interaction network. Functional and pathway enrichment analyses were performed using the Disease Ontology Semantic and Enrichment and ReactomePA R packages.
Results:
The propensity score-adjusted reporting odds ratio for ruxolitinib-associated adverse events was as follows: anemia, 18.49 (95% confidence interval (CI): 16.15–21.16); myelosuppression, 4.70 (95% CI: 3.54–6.24); pancytopenia, 1.97 (95% CI: 1.23–3.16); cardiac failure, 2.29 (95% CI: 1.60–3.28); hepatic function abnormal, 1.60 (95% CI: 1.15–2.23); herpes zoster, 6.40 (95% CI: 4.35–9.41); pneumonia, 2.96 (95% CI: 2.35–3.73); renal impairment, 1.34 (95% CI: 0.94–1.90); sepsis, 5.14 (95% CI: 3.75–7.05); interstitial lung disease, 0.33 (95% CI: 0.21–0.52); deep vein thrombosis, 0.32 (95% CI: 0.07–1.44); hemorrhage, 1.99 (95% CI: 1.05–3.75). We also assessed 3015 human genes that directly or indirectly interact with ruxolitinib. The molecular complex detection plug-in of Cytoscape was used to detect 24 clusters. Several genes were enriched in the biological processes of “anemia” and “bacterial infections,” identified as significant ruxolitinib-related disease terms.
Conclusions:
This retrospective analysis using the Japanese Adverse Drug Event Report database indicated potential associations between ruxolitinib and adverse events, including anemia and bacterial infections. Future research should explore the underlying pharmacological mechanisms using functional enrichment analysis of ruxolitinib-associated genes related to blood toxicity and bacterial infections.
Keywords
Introduction
Ruxolitinib, which primarily inhibits Janus kinase (JAK)1 and JAK2, is used to treat various diseases, including myelofibrosis (MF), polycythemia vera (PV), and steroid-refractory graft-versus-host disease following allogeneic stem cell transplantation.1 –5 PV is a clonal myeloproliferative neoplasm attributed to mutations in the JAK2 gene and is primarily characterized by an increase in red blood cell mass. 1 MF is a progressive, chronic myeloproliferative neoplasm that adversely impacts the daily lives of patients. 2 In patients with MF, the levels of pro-inflammatory cytokines (signaling through JAK1 and JAK2) are elevated. 2
JAK inhibitors are therapeutic agents targeting the cytokine signaling pathways, playing important roles in inflammatory diseases and immune system dysfunctions. 5 The JAK family comprises four enzymes, namely, JAK1, JAK2, JAK3, and tyrosine kinase 2 (TYK2), involved in distinct cytokine signaling cascades. JAK1 primarily regulates inflammatory and innate immune responses; JAK2 is essential for erythropoiesis, myelopoiesis, and platelet production; JAK3 supports lymphocyte proliferation and homeostasis; TYK2 mediates antiviral responses.5 –9
JAK inhibitors, including tofacitinib, baricitinib, and upadacitinib, are associated with various adverse events (AEs), including an increased risk of infection, thrombosis, liver dysfunction, and hematologic abnormalities.10 –13 Similarly, ruxolitinib is linked to various AEs, such as an increased infection risk, hematologic abnormalities, and liver dysfunction; hematologic abnormalities occur more frequently than with other JAK inhibitors.1 –5 Previous studies using the Spontaneous Reporting System (SRS) and bioinformatics have explored reproductive toxicology, 14 digoxin-associated anemia, 15 neuropsychiatric AEs of montelukast, 16 and gentamicin-associated hearing loss. 17 In this study, we examined ruxolitinib-associated AEs using SRS and identified genes responsible for characteristic hematological AEs, infections, and drug interactions using bioinformatics.
The pharmacological mechanisms underlying most AEs remain unclear. Most drugs exert their pharmacological effects and result in AEs through interactions with multiple proteins encoded by different genes. Analysis of drug–gene interactions has improved the understanding of drug toxicity. Integrated analysis using SRS (Food and Drug Administration (FDA) Adverse Event Reporting System (FAERS), and Japanese Adverse Drug Event Report (JADER) database) data and drug–gene interactions can provide deep insights into AEs.14 –17
In this study, we explored the ruxolitinib-associated disproportionality of AEs using propensity score-adjusted reporting odds ratio (PSW-ROR) analysis, leveraging data from the JADER database. Furthermore, we constructed a drug–gene interaction network using genes associated with ruxolitinib to gain insights into the toxicologi-cal mechanisms underlying ruxolitinib-related AEs, and conducted functional enrichment analysis of these genes to identify the potential toxicological mechanisms of ruxolitinib-related AEs.
Methods
Data source
Ruxolitinib was introduced in Japan in 2014; therefore, in this study, JADER data from January 2014 to March 2024 were downloaded from the Pharmaceuticals and Medical Devices Agency (PMDA) and the Japanese Regulatory Authority website (www.pmda.go.jp). All data from the JADER database were cleaned and anonymized in PMDA. The JADER database comprises the following four tables: patient demographic information (DEMO table), drug information (DRUG table), AEs (REAC table), and primary disease (HIST table). A relationship database was created using FileMaker Pro 17 Advanced software (FileMaker, Inc., Santa Clara, CA, USA) based on the American Standard Code for Information Interchange entity relationship diagram. 18 Drug data included the following role codes assigned to each drug according to its association with AEs: “suspected,” “concomitant,” and “interacting drugs.” Data in the SRS database have been reported by healthcare professionals. Our analyses, restricted to the “suspected drugs,” provide the most robust results regarding the association of certain drugs with AEs.
Definition of AEs
AEs were coded using the Medical Dictionary for Regulatory Activities (MedDRA, www.meddra.org) version 23.1. To evaluate ruxolitinib-associated AEs, we employed system organ classes (SOCs) with the preferred terms (PTs) related to each SOC summarized in Table 1.
Crude ROR and inverse probability of treatment weighting-adjusted ROR of adverse events of ruxolitinib in the JADER from January 2014 to March 2024.
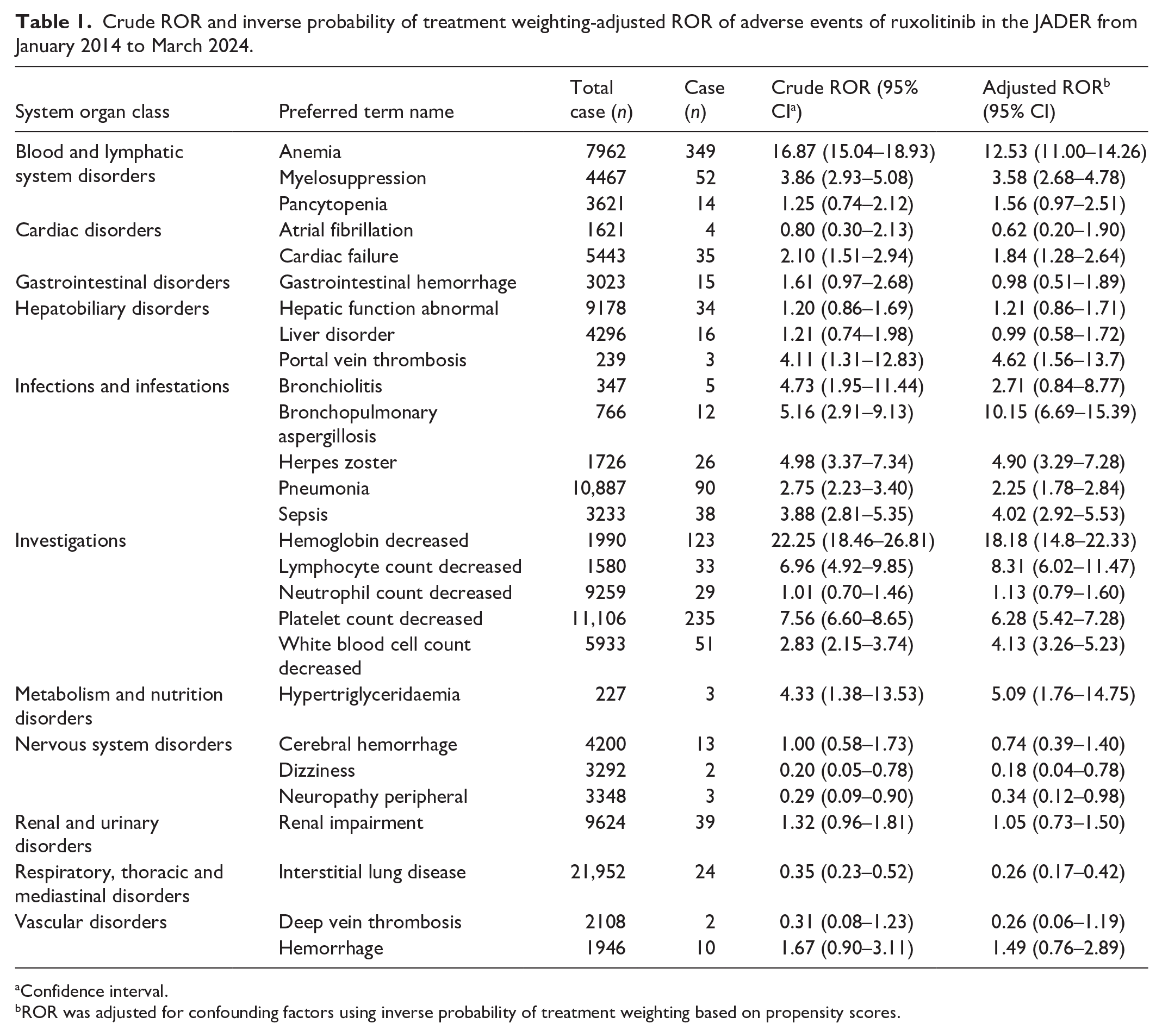
Confidence interval.
ROR was adjusted for confounding factors using inverse probability of treatment weighting based on propensity scores.
Signal detection
We used the crude ROR for administrative safety evaluation to examine the association between ruxolitinib and AEs. The crude ROR was used to calculate the odds of reporting an AE for a drug relative to all other AEs associated with that drug, compared to the odds for all other drugs in the JADER database. 19
This ratio is derived from a two-by-two contingency table, and results are presented as point estimates along with the corresponding 95% confidence interval (CI). A signal was considered positive if the lower limit of the 95% CI exceeded one and there were two or more reports. 19
Crude ROR calculations may be influenced by confounding factors related to differences in patient background. The crude ROR can be partially adjusted using multiple logistic regression analysis, offering the possibility of controlling for covariates.20,21 Propensity score (PS) methods have been recently used with stabilized inverse probability of treatment weighting (stabilized IPTW) to adjust for potential confounders.22 –26 We applied a PS method using stabilized IPTW to adjust for potential confounding factors in estimating adjusted RORs of AEs associated with ruxolitinib and refined the safety signal with a dedicated correction to detect possible confounding factors in the database. PS methods are statistical techniques employed to reduce confounding in observational studies by constructing matched pairs of treated and untreated individuals with similar PSs, thereby balancing covariates between groups. However, a primary limitation of PS matching is the potential reduction in sample size due to the exclusion of unmatched subjects.
In contrast, stabilized IPTW retains all subjects in the analysis by assigning weights based on their PSs. The stabilized weight for each individual is defined as:

where P(T) denotes the marginal probability of recei-ving the treatment, and P(T∣X) represents the PS conditional on covariates X. This approach mitigates the influence of extreme weights while preserving the full sample size, thereby enhancing statistical efficiency and reducing bias.
PS was estimated using a logistic regression model, in which the treatment status (treated or not treated with ruxolitinib) was regressed on the following covariates: sex, age category (<60 years vs ⩾60 years), and reporting year. The IPTW was calculated as follows:
For patients treated with ruxolitinib:

For patients not treated with ruxolitinib:

After applying stabilized IPTW, adjusted RORs were estimated using a weighted logistic regression model with IPTW as the frequency weight. ROR was derived from model coefficients, and 95% CIs were obtained using the standard error of the estimated coefficients.
All analyses were conducted using Python (version 3.12.3) (Python Software Foundation, Wilmington, DE, USA). The following Python libraries were utilized: Pandas (version 1.4.4) (Pandas Development Team via NumFOCUS, Austin, TX, USA) and NumPy (version 1.21.5) (NumPy Developers via NumFOCUS, Austin, TX, USA) for data processing, statsmodels (version 0.13.2) (statsmodels Development Team via NumFOCUS, Austin, TX, USA) for multiple logistic regression and generalized linear models, and scikit-learn (version 1.0.2) (scikit-learn Development Team via NumFOCUS, Austin, TX, USA) for PS estimation.
Drug–gene interaction network
The drug–gene interaction network was constructed using data from multiple sources: drug–gene interactions from the drug–gene interaction database (DGIdb, https://www.dgidb.org), drug signatures from the drug signatures database (DSigDB, http://dsigdb.tanlab.org), and broader interaction data from the “search tool for interactions of chemicals” (STITCH, https://stitch.embl.de). Genes indirectly associated with ruxolitinib were identified using iRefIndex 20.0 (“9606. mitab,” https://irefindex.vib.be). 27 The molecular complex detection (MCODE) algorithm (version 2.0.3) (Bader Lab, University of Toronto, Toronto, ON, Canada), 28 a plug-in in Cytoscape version 3.10.1 (Cytoscape Consortium, San Diego, CA, USA), was used to identify highly interconnected regions within protein–protein interaction networks, 29 with the following criteria: degree cutoff, 2; node score cutoff, 0.2; k-core, 2, and maximum depth from the seed, 100. Functional analysis and visualization of genes and gene clusters were performed using the R software (version 4.2.0) (R Foundation, Vienna, Austria). We used the Disease Ontology Semantic and Enrichment analysis (DOSE) package (YuLab-SMU, Southern Medical University, Guangzhou, China), 30 which implements methods for measuring semantic similarities between Disease Ontology terms and gene products. Accordingly, simple clusters, including triangles and squares, were excluded from the obtained cluster, and the remaining clusters were subjected to DOSE analysis conducted using the “enrichDO” function of the DOSE package (version 3.24.2) (YuLab-SMU, Southern Medical University, Guangzhou, China) in R with a p-value cutoff of 0.05, a p-value adjustment method of “BH,” and a q-value cutoff of 0.05. We performed a pathway enrichment analysis using ReactomePA, 31 designed for Reactome pathway-based analysis, to investigate the pathways associated with gene clusters. Pathway analysis was performed using the “enrichPathway” function from the ReactomePA package (version 1.42.0) (YuLab-SMU, Southern Medical University, Guangzhou, China) with a p-value cutoff of 0.05.
Results
ROR analysis
The JADER database included 583,504 reports from January 2014 to March 2024. After excluding cases with missing age or sex information, 502,961 reports (86.2%) were included in the analysis. Within this dataset, ruxolitinib was reported as the suspected drug in 1285 cases, with 703 cases (54.7%) of male patients and 582 (45.3%) of female patients. Additionally, 179 cases (13.9%) were patients aged < 60 years, whereas 1106 cases (86.1%) were patients aged ⩾ 60 years. The crude RORs of AEs associated with ruxolitinib were as follows: anemia, 21.45 (95% CI: 20.19–22.79); myelosuppression, 4.82 (95% CI: 4.21–5.53); pancytopenia, 1.41 (95% CI: 1.11–1.81); cardiac failure, 2.90 (95% CI: 2.47–3.40); hepatic function abnormal, 1.02 (95% CI: 0.86–1.22); herpes zoster, 6.01 (95% CI: 4.98–7.25); pneumonia, 3.57 (95% CI: 3.22–3.96); renal impairment, 1.85 (95% CI: 1.59–2.14); sepsis, 4.17 (95% CI: 3.56–4.89); interstitial lung disease, 0.39 (95% CI: 0.32–0.48); deep vein thrombosis (DVT), 0.37 (95% CI: 0.18–0.76); hemorrhage, 2.48 (95% CI: 1.88–3.28; Table 1).
Standardized mean differences (SMDs) were calculated for sex, age category, and reporting year to evaluate covariate balance following IPTW application. SMD below 0.1 indicates an acceptable balance. 23
Following IPTW adjustment, SMD values were 0.123 for the reporting year, 0.029 for sex, and 0.046 for the age category. While the SMD for the reporting year was just over 0.1, we deemed this acceptable as the distribution was not significantly imbalanced visually or clinically.
In the initial analysis, unstabilized IPTW values had a mean of 1.980 (min: 1.000; max: 2110.505) and a median (Q1–Q3) of 1.002 (1.001–1.003), whereas stabilized IPTW values had a mean of 1.000 (min: 0.395; max: 5.392 and a median (Q1–Q3) of 1.000 (0.999–1.001). These stabilized weights improved the balance and stability of the analysis.
The PSW-ROR of AEs associated with ruxolitinib was as follows: anemia, 18.49 (95% CI: 16.15–21.16); myelosuppression, 4.70 (95% CI: 3.54–6.24); pancytopenia, 1.97 (95% CI: 1.23–3.16); cardiac failure, 2.29 (95% CI: 1.60–3.28); hepatic function abnormal, 1.60 (95% CI: 1.15–2.23); herpes zoster, 6.40 (95% CI: 4.35–9.41); pneumonia, 2.96 (95% CI: 2.35–3.73); renal impairment, 1.34 (95% CI: 0.94–1.90); sepsis, 5.14 (95% CI: 3.75–7.05); interstitial lung disease, 0.33 (95% CI: 0.21–0.52); DVT, 0.32 (95% CI: 0.07–1.44); hemorrhage, 1.99 (95% CI: 1.05–3.75; Table 1).
Drug–gene interaction network
Our initial search across DGIdb, DSigDB, and STITCH databases identified 14 seed genes that interacted with ruxolitinib, namely CRLF2, CSF3R, JAK1, JAK2, JAK3, MPL, PARP1, PARP2, PARP3, PARP4, PLAUR, SRC, TYK2, and VHL, all of which were from strain 9606. The mitab dataset in the iRefIndex was integrated into a network comprising 18,270 nodes (genes) and 791,934 edges (interactions).
Genes that directly or indirectly interacted with the 14 seed genes were integrated into a network of 3015 nodes and 159,496 edges. Using the MCODE plug-in, 24 clusters were identified in the network. To elucidate their biological significance, functional enrichment analysis of these clusters was performed using a pathway analysis. The top two clusters related to ruxolitinib are shown in Table 2 and Figure 1; the remaining 22 are shown in Supplemental Data (Supplemental material).
Clusters of networks analyzed by MCODE a analysis on ruxolitinib-related genes.

Network scoring: Include loops = false, Degree cutoff = 2, Cluster finding: node score cutoff = 0.2, Haircut = true, Fluff = false, K-core = 2, Max depth from seed = 100.

Gene interaction networks obtained using the molecular complex detection plug-in in cytoscape. (a) Cluster 1 (99 genes), (b) Cluster 2 (241 genes).
For Cluster 1, the genes interacting with ruxolitinib were enriched in pathways such as “L13a-mediated translational silencing of ceruloplasmin expression,” “nonsense-mediated decay (NMD),” “NMD enhanced by the exon junction complex (EJC),” “eukaryotic translation initiation,” “cap-dependent translation initiation,” and “formation of a pool of free 40S subunits” (Table 3 and Figure 2). Cluster 1 was enriched in anemia-associated genes, including RPL5, RPL11, RPL27, RPL35, RPL35A, RPS10, RPS19, RPS27, RPS29, and RPS15A (Table 4 and Figure 3).
Pathway enrichment analysis of ruxolitinib-associated gene clusters using the ReactomePA a package.

p-value cutoff of 0.05.

Dot plot of the pathway analysis performed using the ReactomePA package. (a) Cluster 1 and (b) Cluster 2.
Disease ontology enrichment of ruxolitinib-related genes using the disease ontology semantic and enrichment a package.

p-value cutoff of 0.05, p-value adjustment method of “BH,” and q-value cutoff of 0.05.

Dot plot for disease ontology semantic and enrichment analysis using the disease ontology semantic and enrichment package. (a) Cluster 1 and (b) Cluster 2.
Cluster 2 was enriched in bacterial infection-associated genes, including interleukin-5 (IL-5), colony-stimulating factor 2 (CSF2, also known as GM-CSF), C-C motif chemokine receptor (CCR)2 and CCR5, Fc gamma receptor IIA (FCGR2A), and Fc gamma receptor IIIA (FCGR3A; Table 4 and Figure 3). In Cluster 2, the genes were enriched in various pathways including “PIP3 activates AKT signaling,” “intracellular signaling by second messengers, RAF/MAP kinase cascade, MAPK1/MAPK3 signaling, PI3K events in ERBB2 signaling,” “interleukin receptor SHC signaling, MAPK family signaling cascades,” and “interleukin-3, interleukin-5, and GM-CSF signaling” (Table 3 and Figure 2).
Data from the DOSE package were used to characterize diseases associated with ruxolitinib. Genes involved in anemia (adjusted p-value = 0.0003) in Cluster 1 and primary bacterial infectious disease (adjusted p-value = 0.0022) in Cluster 2 were significantly enriched (Table 4 and Figure 3).
Discussion
In this study, we propose a methodology to apply safety signals—specifically, disproportionality indices such as RORs, which are widely used in pharmacovigilance—to bioinformatics to identify genes potentially associated with AEs. AEs are associated with the pharmacological effects of drugs; therefore, these genes can potentially serve as therapeutic targets for managing AEs. This will help to facilitate the development of novel treatments or broaden the application of existing drugs. This study presents a practical approach within the exploratory domain of translational research and partially addresses the limitations of crude ROR by integrating PS methods and multiple logistic regression analysis. To the best of our knowledge, this is the first study to adjust RORs using PSW through IPTW, a novel contribution to the field.
Ruxolitinib is a JAK inhibitor used widely to treat MF and PV. However, its associated AEs may limit ruxolitinib’s therapeutic efficacy. Reports from the JADER database have linked ruxolitinib to major AEs. The lower limits of the 95% CIs of crude RORs related to the SOC of “blood and lymphatic system disorders,” “infections and infestations,” and “investigations” were >1, and the signal was significant.
In Phase 3 studies of PV and MF, the most frequent AEs leading to dose adjustment or interruption of ruxolitinib treatment were anemia and thrombocytopenia.1,2 Additionally, many patients with MF develop anemia during the disease and require red blood cell transfusion.
Conversely, bleeding episodes, attributed to either thrombocytopenia or qualitative platelet dysfunction, can complicate the clinical course of MF. 32 In the JADER database, significant signals were detected in the RORs associated with gastrointestinal hemorrhage and hemorrhage. These findings necessitate the careful management of hematopoietic disorder risks in patients receiving ruxolitinib therapy. Infections, such as pneumonia, sepsis, and herpes zoster, are also frequently reported common AEs associated with JAK inhibitors besides ruxolitinib, including tofacitinib and baricitinib, which are used for chronic inflammatory diseases.10 –13 Since the immunosuppressive effects of ruxolitinib can increase the risk of infection, infection control measures and early therapeutic intervention are essential.
Owing to the growing concern regarding thromboembolic events in patients undergoing treatment with JAK inhibitors, the FDA has mandated a “boxed warning” for most of these drugs currently available. 33 However, ruxolitinib lacks such a mandate. Setyawan et al. have detected a signal for portal vein thrombosis (PVT) in the FAERS database, 34 consistent with our findings from the JADER analysis. In contrast, no signal was detected for DVT in either study. Given the observed overreporting of PVT, the signal of which is based on three cases with possible concerns about random noise, associated with ruxolitinib in the present study, additional analyses are required. In the JADER dataset, including reports from January 2013 to March 2024, PVT was reported only in patients treated with ruxolitinib, not in those receiving other JAK inhibitors (tofacitinib, peficitinib, baricitinib, filgotinib, and upadacitinib; Supplemental Table 4). This is due to potential confounding by indication (e.g., PVT risk in myeloproliferative neoplasms). A direct comparison and analysis of PVT risk among JAK inhibitors was not feasible. Stratified analysis (e.g., PVT in ruxolitinib vs non-ruxolitinib-treated MF/PV patients) or comparison of PVT incidence with relevant epidemiological data will provide deeper insights.
A comprehensive analysis of thrombosis risk among JAK inhibitors using JADER suggested older age (>60 years) as a risk factor for more extensive embolic and thrombotic events across all JAK inhibitors examined. 35
JAK inhibitors are associated with increased risk of infection, thrombosis, liver dysfunction, and hematologic abnormalities, among others.10 –13 The results of this disproportionality analysis are consistent with those previously reported.10 –13 Moreover, gene enrichments in the biological processes of anemia and bacterial infection identified in this study correspond to the pharmacological effects of ruxolitinib.
Integrated analysis using FAERS data and drug–gene interaction analysis data provides novel insights into AEs.14 –16 Previously, we employed analogous methodologies to analyze montelukast and identified 1144 interacting human genes, generating 14 clusters, which included genes enriched in mood disorders. 16 Lin et al. 15 conducted a drug–gene interaction network analysis similar to our study, focusing on digoxin-associated anemia using FAERS. Wu et al. 14 integrated FAERS data with drug–gene network analysis to investigate variations in the reproductive toxicity of alopecia drugs. In this study, we employed the aforementioned comparable method,14 –16 wherein drug–gene interactions were queried in public databases to elucidate the toxicological mechanisms underlying ruxolitinib-related AEs. Specifically, 3015 human genes interacting with ruxolitinib were identified. We generated 24 clusters using the MCODE software (Bader Lab, University of Toronto, Toronto, ON, Canada). The results yielded two clusters that included diseases likely associated with ruxolitinib.
Cluster 1 was enriched with anemia-associated genes, including RPL5, RPL11, RPL27, RPL35, RPL35A, RPS10, RPS19, RPS27, RPS29, and RPS15A (Table 4 and Figure 3). Pathways related to intracellular protein synthesis, primarily in the endoplasmic reticulum, were identified and located downstream of JAK. Ruxolitinib-associated anemia likely occurs due to JAK2-mediated inhibition of erythropoietin (EPO) signaling in red blood cell progenitor cells. The EPO binds to the EPO receptor, activating JAK2.36,37 This, in turn, phosphorylates STAT5, promoting the proliferation and differentiation of erythroid progenitor cells.38,39 Furthermore, ruxolitinib reduces the allele burden of JAK2 V617F mutations and suppresses the autonomous proliferation of EPO-independent erythroid progenitor cells.40,41 A somatic mutation in JAK2 V617F is found in >90% of patients with PV and half of those with MF.42 –45 ROR analysis in JADER detected a signal for ruxolitinib-associated red blood cell depletion. Our functional enrichment analysis suggests that pathways related to intracellular protein synthesis in the endoplasmic reticulum indirectly regulate AEs related to ruxolitinib, even if the gene is not a direct drug target.
Cluster 2 was enriched in genes involved in bacterial infections and malignancy, including IL-5, CSF2, CCR2, CCR5, FCGR2A, and FCGR3A. Activation of eosinophils by IL-5 is crucial for immune responses, such as those elicited in allergic diseases and parasitic infections. 46 IL-5 binds to the IL-5 receptor to activate the JAK2-mediated signaling pathway and regulates eosinophil differentiation, proliferation, survival, and activation. 47 CCR2, a G protein-coupled receptor, is primarily expressed on monocytes and T cells and induces their migration to sites of inflammation. 48 The signaling pathways of CCR5, another G protein-coupled receptor, involve phosphatidylinositol 3-kinase/protein kinase B and JAK/signal transducer and activator of transcription, key in inflammatory responses and immune regulation. 49
However, CCR2 and CCR5 may show cross-talk with the JAK-STAT pathway.49,50 FCGR2A, expressed on macrophages and neutrophils, induces phagocytosis and antibody-dependent cellular cytotoxicity by binding to IgG antibodies. 51 Meanwhile, FCGR3A, expressed on natural killer cells (NK) and macrophages, activates cell-mediated immune responses via IgG binding. 51 JAK inhibition increases the risk of infection because of its mechanism of action.
Signals for pneumonia, sepsis, and shingles have been detected in the JADER database. Pneumonia is a frequently reported AE associated with JAK inhibitors.9 –13 JAK inhibition suppresses IL-6 and IL-2 signaling, decreasing the activity of T cells and NK and increasing the risk of infection.52,53 While JAK inhibition increases the risk of infection through immunosuppression, the source of the increased risk of infection includes the effects of underlying diseases and concomitant medications. JAK inhibitors are suspected to increase the risk of malignant tumor development, although the data remain inconclusive.32,54
Limitation
This study, which is based on spontaneous reports obtained from the JADER database, has some limitations. First, potential biases might have been introduced due to over-reporting, under-reporting, and missing data. Second, external factors, such as safety communications and market dynamics, might have influenced the reporting patterns. Finally, given that the JADER dataset contains limited patient and clinical data (e.g., comorbidities and concomitant medications), our findings should not be interpreted with caution. Spontaneous reports of AEs, such as “anemia” in this study, may be overrepresented due to clinicians’ heightened awareness of hematologic risks. Additional biases include notoriety bias, wherein the number of spontaneous reports increases following safety alerts issued by regulatory authorities.55 –57 Such warnings may increase RORs. Another relevant phenomenon is the Weber effect,58 –60 which describes the tendency for spontaneous reporting of AEs to sharply increase following a drug’s approval, subsequently plateau, and eventually decline over time; however, this effect is not consistently observed across all drugs. 60 To mitigate these biases, PS methods can be employed to balance groups based on potential confounding variables.61,62 Furthermore, several studies have attempted to adjust ROR estimates by incorporating factors such as reporting year, age, and sex into the model using multivariate logistic regression analyses.63 –65
In Japanese health insurance, individuals ⩾65 years or older are classified as elderly; this age classification is also seen in the package insert for ruxolitinib. 66 However, in the JADER database, age information has been modified and published in 10-year increments by the PMDA to protect personal information, making it difficult to accurately classify individuals at exactly 65 years of age. Defining the elderly population as those aged ⩾ 70 years may be more appropriate for ruxolitinib, given its characteristics. Nevertheless, in this study, individuals ⩾60 years old were classified and analyzed as elderly.
The ROR is merely an indicator of disproportionality and only identifies statistical associations, not causal relationships, and does not establish causation nor account for actual exposure rates. In this study, given the limited individuals’ background information (e.g., concomitant medications, underlying diseases, and patient characteristics), we calculated a partially adjusted PSW-ROR by controlling for age, sex, and reporting year.
Using only these three covariates may not fully control for all potential confounders. The models for estimating PS and calculating IPTW inherently involve estimation error and uncertainty; therefore, caution is essential to avoid overinterpretation of disproportional signals. Alternately, some approaches have been proposed to address these issues in a high-dimensional context.67 –69 It may be possible to quantify the potential impact of reporting bias using sensitivity analysis or external validation (e.g., comparing JADER signals to FAERS or published clinical trial data). Future investigations with prospective study designs or multivariable analyses with comprehensive covariate data should validate these findings and clarify the risk profiles of ruxolitinib and other JAK inhibitors.
To date, the drug–gene interactions previously examined have not been validated in experimental models or in vitro or in vivo experiments due to our limited knowledge of disease-related proteins and their interactions. Clusters 1 and 2 may explain the drug’s mechanism of action. These might not necessarily be off-target, unexpected AEs. In other words, the drug is effective, and dose-effect relationships should be assessed, rather than addressed by drug–gene interaction analysis. Therefore, the association between ruxolitinib and this gene must be experimentally confirmed.
Initially, we identified 14 proteins that interacted with ruxolitinib; however, the seed genes suggested by databases, such as DGIdb, used in this study are not direct pharmacological targets, and many may be indirectly affected by the drug. We utilized protein-protein interaction networks to integrate these 14 affected proteins into a larger network (iRefIndex). Our findings revealed that the network was enriched in genes associated with anemia and bacterial infection. Specifically, RPL5, RPL11, RPL27, RPL35, RPL35A, RPS10, RPS19, RPS27, RPS29, and RPS15A were associated with anemia, and IL5, CSF2, CCR2, CCR5, FCGR2A, and FCGR3A with bacterial infections. Furthermore, even within small clusters, we observed specific associations. For cerebrovascular disease, a cluster containing four genes included three associated genes: COL4A1, ELANE, and MMP12. Although these genes are not direct targets of ruxolitinib, future studies are necessary to determine whether specific “hotspots” within such protein-protein interaction networks can distinguish between drugs associated with anemia and those that lead to bacterial infections.
The drug–gene network analysis used in this study relies on public databases, such as DGIdb and STITCH, which may include non-causal relationships, limiting conclusions concerning mechanisms without experimental validation. Nevertheless, this approach, which uses gene enrichment, provides novel insights into ruxolitinib-associated AEs and will facilitate future research. Given the nature of exploratory network analysis, over-interpretation of clusters as direct toxicity mechanisms should be avoided, and functional validation, including in vitro and in vivo, is required to clarify gene and pathway associations.
Conclusion
Ruxolitinib is associated with significant AEs that require careful consideration. Our retrospective analysis of the JADER database suggested anemia, likely due to the suppression of erythropoiesis, as a major AE, underscoring the importance of vigilant management in patients. Pneumonia, sepsis, and other infections linked to the immunosuppressive effects of the drug have been frequently reported. Genetic analysis revealed that genes potentially influencing anemia and infection through interactions with ruxolitinib, including those related to the endoplasmic reticulum and immune response genes, such as IL5, CSF2, CCR2, CCR5, FCGR2A, and FCGR3A, were enriched. Due to the limitations of spontaneous JADER reports, such as reporting biases, incomplete patient information, and unverified drug–gene interactions, the findings should be interpreted cautiously. Further studies for validation and risk mitigation remain warranted.
Supplemental Material
sj-jpg-1-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-jpg-1-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-jpg-2-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-jpg-2-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-jpg-3-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-jpg-3-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-jpg-4-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-jpg-4-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-xlsx-10-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-xlsx-10-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-xlsx-11-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-xlsx-11-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-xlsx-12-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-xlsx-12-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-xlsx-13-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-xlsx-13-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-xlsx-5-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-xlsx-5-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-xlsx-6-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-xlsx-6-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-xlsx-7-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-xlsx-7-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-xlsx-8-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-xlsx-8-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Supplemental Material
sj-xlsx-9-smo-10.1177_20503121251348420 – Supplemental material for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks
Supplemental material, sj-xlsx-9-smo-10.1177_20503121251348420 for Analysis of adverse events with ruxolitinib using real-world datasets and drug-interaction networks by Hideyuki Tanaka, Mika Maezawa, Mizuki Tanaka, Ryogo Umetsu, Sakiko Hirofuji, Koumi Miyasaka, Satoshi Nakao, Yuka Nokura, Moe Yamashita, Nanaka Ichihara, Kana Sugishita, Tomofumi Yamazaki, Kohei Shiota, Hirofumi Tamaki, Kazuhiro Iguchi and Mitsuhiro Nakamura in SAGE Open Medicine
Footnotes
Acknowledgements
Not applicable.
Ethical considerations
Ethical approval was not sought for this study because the study was a database-related observational study and did not require the direct involvement of any research subjects. All results were obtained from data openly available online from the Pharmaceuticals and Medical Devices Agency website (![]() ). All the data from the Japanese Adverse Drug Event Report database were fully anonymized by the relevant regulatory authority before access was obtained.
). All the data from the Japanese Adverse Drug Event Report database were fully anonymized by the relevant regulatory authority before access was obtained.
Author contributions
H.T., M.M., M.T., R.U., and M.N. contributed to the overall study concept and design. H.T., M.M., S.H., K.M, S.N., Y.N., and M.N. performed data extraction and statistical analysis. H.T., M.M., and M.N. wrote the manuscript. M.T., R.U., S.H., K.M., S.N., Y.N., M.Y., N.I., K.S., T.Y., K.S., H.T., and K.I. revised the article to provide meaningful intellectual content. All authors have read and reviewed the manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was partially supported by JSPS KAKENHI (grant numbers 21K06646 and 21K11100). The funders had no role in the study design, data collection, analysis, decision to publish, or preparation of this article.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The datasets used and analyzed during the current study are available from the corresponding author upon reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
