Abstract
Objectives:
Concern for late detection of bacterial pathogens is a barrier to early de-escalation efforts. The purpose of this study was to assess blood, respiratory and urine culture results at 72 h to test the hypothesis that early negative culture results have a clinically meaningful negative predictive value.
Methods:
We retrospectively reviewed all patients admitted to the medical intensive care unit between March 2012 and July 2018 with blood cultures obtained. Blood, respiratory and urine culture results were assessed for time to positivity, defined as the time between culture collection and preliminary species identification. The primary outcome was the negative predictive value of negative blood culture results at 72 h. Secondary outcomes included sensitivity, specificity, positive predictive value and negative predictive value of blood, respiratory and urine culture results.
Results:
The analysis included 1567 blood, 514 respiratory and 1059 urine cultures. Of the blood, respiratory and urine cultures ultimately positive, 90.3%, 76.2% and 90.4% were positive at 72 h. The negative predictive value of negative 72-h blood, respiratory and urine cultures were 0.99, 0.82 and 0.97, respectively. Antibiotic de-escalation had good specificity, positive predictive value and negative predictive value for finalized negative cultures.
Conclusion:
Negative blood and urine culture results at 72 h had a high negative predictive value. These findings have important ramifications for antimicrobial stewardship efforts and support protocolized re-evaluation of empiric antibiotic therapy at 72 h. Caution should be used in patients with clinically suspected pneumonia, since negative respiratory culture results at 72 h were weakly predictive of finalized negative cultures.
Keywords
Introduction
Unnecessary broad-spectrum antibiotics increase adverse events in critically ill patients.1–4 Antibiotic de-escalation is well-documented as both safe and effective;5–9 a retrospective study of patients with sepsis and septic shock found that de-escalation of empiric antibiotics was associated with lower mortality. 10 Mechanisms of this mortality benefit may include that clinically appropriate antibiotic de-escalation decreases selective pressure for multi-drug resistant organisms and reduces antibiotic-related adverse events, including Clostridioides difficile infection.1,11,12
Despite the known benefits of streamlining antimicrobial therapy, several factors prevent timely antimicrobial de-escalation (e.g. clinical status, inconclusive microbiological results or results complicated by prior antibiotic administration and concern for late detection of bacterial pathogens). 13 While current laboratory standards recommend a 5-day incubation period for blood cultures before finalizing culture results, approximately 70%–90% of blood cultures are positive within 48 h in a general medical population, creating the potential for earlier de-escalation.14–16
Pardo et al. 15 first assessed time to positivity (TTP) for antimicrobial de-escalation in a mixed medical population and observed that culture results at 48 h had a 97.3% sensitivity (Se) and 99.8% negative predictive value (NPV) for finalized negative cultures. This finding was consistent with previous results demonstrating that 98% of all aerobic blood stream infections (BSIs) and 74% of gram-negative BSIs were detected at 48 h.14–17 A more recent study found that 85.3% of blood cultures were positive by 24 h. 18 These studies lend support to the safety of the clinical practice of antibiotic de-escalation in clinically stable patients with negative cultures at 48 h in a general medical population. However, while the findings from Pardo et al. 15 and Lambregts et al. 18 demonstrate significant potential for antibiotic management in the intensive care unit (ICU), data derived from non-ICU patients cannot be extrapolated to critically ill patients. Bacterial pathogens and resistance patterns in the ICU differ given the high rate of antimicrobial consumption, burden of intravascular devices and high frequency of nosocomial infections.19,20 Furthermore, severity of illness makes the consequences of inappropriate antimicrobial coverage more severe. 21 Melling et al. 22 assessed time to first culture positivity for methicillin-resistant Staphylococcus aureus (MRSA) in blood and respiratory cultures of critically ill patients receiving empiric coverage for MRSA. Investigators found that 85% of respiratory cultures and 97.6% of blood cultures ultimately positive for MRSA were positive at 48 h. 22 Despite these data, more information is needed regarding other sources of infection, including urinary, as well as resistant pathogens other than MRSA.
The purpose of this study was to assess blood, respiratory and urine culture results at 72 h as a predictor of finalized negative cultures to test the hypothesis that early negative culture results have a clinically meaningful NPV in a critically ill patient population. Given the acuity of the studied patient population, the 72-h time point was selected in order to increase the amount of clinical information available and decrease the likelihood of inappropriate de-escalation.
Methods
Study design
This study was conducted at Augusta University (AU) Medical Center, a 478-bed academic tertiary care center in Augusta, Georgia. AU Medical Center has 75 adult critical care beds divided among the trauma, surgical, medical, neuroscience and cardiac ICUs. All patients admitted to the medical intensive care unit (MICU) between March 2012 and July 2018 who had a blood culture obtained were retrospectively reviewed. Blood, respiratory and urine cultures were reviewed for TTP, defined as the time between culture collection and preliminary species identification. Growth at 24, 48 and 72 h was determined by the presence of any bacterial growth at the predefined time interval (Figure 1). The study protocol was reviewed and approved by the AU Medical Center Institutional Review Board (IRB), and the need for informed consent was waived.
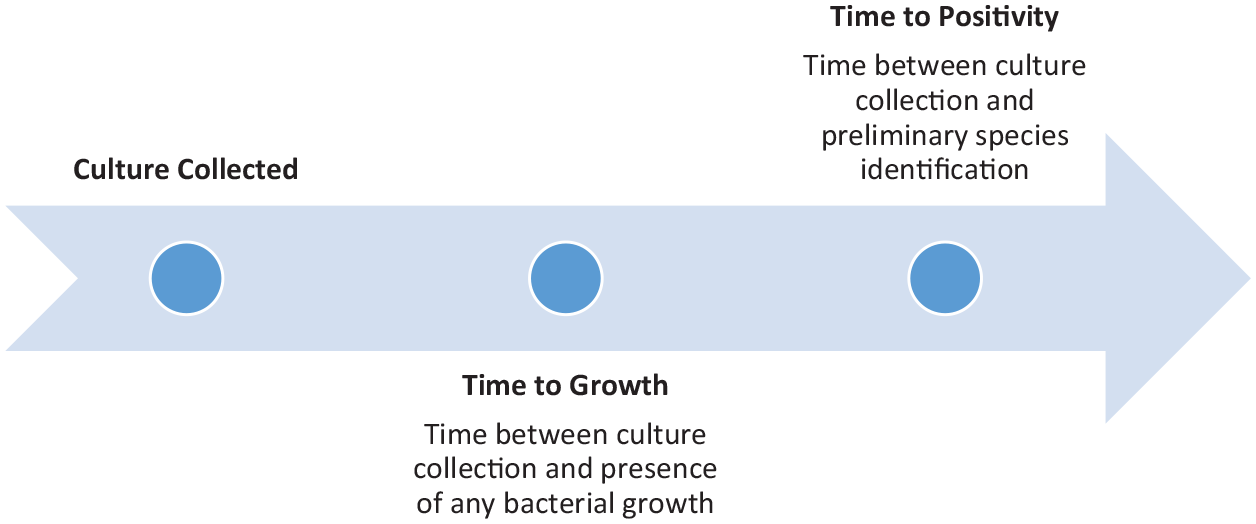
Definitions based on culture positivity.
Data collection
A report of all patients admitted to the MICU within the pre-specified time period was generated using admission orders to the MICU, and then patients were screened using current procedural terminology (CPT) charge codes to determine whether blood cultures were obtained. Patients were included if they were greater than 18 years of age and admitted to the MICU between March 2012 and July 2018 with blood cultures drawn. Patients were excluded if they had blood cultures positive for fungi or acid-fast bacilli or a culture isolate previously identified within a given admission. Multiple episodes of bacteremia per patient admission were permitted for distinct isolates. Patients were reviewed in reverse chronological order until the pre-specified number of blood culture results was obtained. Respiratory and urine culture data were derived from the cohort of patients with blood cultures. Respiratory cultures included bronchoscopic techniques, blind bronchial sampling and endotracheal aspiration.
The electronic medical record was reviewed to assess patient demographics, clinical outcomes, and blood, respiratory and urine culture data. Patient demographics included antibiotic administration prior to culture, clinical criteria for de-escalation, immune status, length of stay and mortality. Antibiotic administration was defined as any antibiotic administration in the 24 h prior to drawing blood cultures. Clinical criteria for de-escalation were defined a priori by the presence of all of the following at 72 h: negative blood culture, immunocompetent, normothermic (temperature 36–38°C, without cooling blanket, acetaminophen or non-steroidal anti-inflammatory drugs in the past 24 h), white blood cell count 4–12 × 103 cells/mcL and decreasing doses of vasopressors or vasopressor requirement of less than 0.2 mcg/kg/min of norepinephrine or equivalent. 23 Patients were deemed immunocompetent if they had no history of diagnosis of human immunodeficiency syndrome (HIV), acquired immunodeficiency syndrome (AIDS), anatomic or functional asplenia, hematologic or solid organ malignancy, history of organ transplant or rheumatologic/autoimmune condition receiving chronic immunosuppressive therapy. Culture data included suspected source of infection, pathogen or contaminant based on documentation in the medical record.
For patients with bacterial growth, bacterial species identified and TTP were recorded. Bacterial growth at 24, 48 and 72 h was determined by assessing the presence of any signs of bacterial growth on culture at this time. The primary objective of this study was the NPV of negative blood culture results at 72 h. Secondary objectives included the Se, specificity (Sp) and positive predictive value (PPV) of negative 72-h blood, respiratory and urine culture results. The NPV of culture results at 72 h in patients meeting clinical criteria for de-escalation, and TTP of MRSA, Pseudomonas aeruginosa, and all other species were also assessed. Time to growth was defined as the time between culture collection and the presence of any bacterial growth on culture. TTP was defined as the time between the culture being obtained and the time at which the species received preliminary identification (e.g. preliminary identification as MRSA) (Figure 1). Contaminants were defined by positive culture growth determined by the primary treatment team not to be an infectious organism as evidenced by documentation in the medical record or cessation of antibiotic treatment prior to or at the time of culture finalization. Cultures that were negative at 72 h and at finalization were considered true negatives. Cultures that were negative at 72 h but ultimately positive were considered false negatives.
Microbiology
The BacT/Alert® three-dimensional (3D) automatic blood culture system was used at the beginning of the study period, and was later replaced by the BacT/Alert® Virtuo® system (bioMerieux). Both systems work using colorimetric technology to detect microbial growth. The BacT/Alert® Virtuo® system maintains instrument temperature to reduce time to detection by approximately 20%. 24 At the beginning of the study period, AU Medical Center used the BacT/Alert® FA FAN (aerobic) and BacT/Alert® FN FAN (anaerobic) culture media, which were later replaced by the BacT/Alert FA FAN® (aerobic) and BacT/Alert FN FAN® (anaerobic) culture media (bioMerieux). The Verigene® system was used from 2015 onward to rapidly identify gram-positive and gram-negative organisms. Respiratory and urine cultures were both plated on blood agar and MacConkey agar; respiratory cultures were also plated on Columbia CNA and chocolate agar. Culture data are updated at regular intervals until culture finalization and completion of susceptibility testing.
Statistical analysis
A sample size of 1567 subjects was calculated to estimate the NPV of 72-h blood culture results to predict finalized negative cultures. Assumptions included a Se of 90%, Sp of 99%, prevalence of positive blood cultures of 10%, alpha level of 0.05, power of 80% and a lower bound on the NPV of 95%. 15 Descriptive statistics were calculated for all variables. Frequencies and percentages were used for categorical variables and mean values and standard deviations were used for continuous variables. Two-way tables were used to estimate Se, Sp, PPV and NPV of growth at 48 and 72 h for final growth for MRSA, P. aeruginosa, non-MRSA/non-P. aeruginosa species and all species on all culture types. Se, Sp, PPV and NPV were also calculated for the subgroup of patients meeting clinical criteria for de-escalation at 72 h. All statistical analyses were performed using R version 4.0.2 (R Core Team, 2020) with reproducible research using R Markdown (Allaire et al., 2020). 31 Confidence intervals for the median were calculated using DescTools package.
Results
The analysis included a total of 1567 blood, 514 respiratory and 1059 urine cultures. Of the 206 positive blood cultures, 120 (58.3%) were considered true infection, and 86 (41.7%) were considered contaminants. Similarly, of 307 positive respiratory cultures, 34 (11.1%) were considered contaminants, and of the 272 positive urine cultures, 43 (15.8%) were considered contaminants. Cultures that were positive and considered true infection represented 7.7%, 53.1% and 21.6% of all blood, respiratory and urine cultures analyzed, respectively.
Patient characteristics
Antibiotics were received prior to blood culture in 36.5% of patients. Of those receiving antibiotic coverage prior to blood culture, 51% of patients had coverage for MRSA and 66.3% had coverage for P. aeruginosa. Antibiotics were received prior to 56.2% of respiratory cultures and 38.9% of urine cultures. The median TTP for all positive blood cultures was 90 h (95% confidence interval (CI) = 83–99 h), for urine was 61 h (95% CI = 59–67 h), and for respiratory cultures was 73 h (95% CI = 70–78 h). Patient characteristics are represented in Table 1.
Patient characteristics.

SD: standard deviation; ICU: intensive care unit; IQR: interquartile range.
1567 blood cultures obtained from 1300 patients.
Pathogen-specific analysis
Of the blood, respiratory and urine culture results that were ultimately positive, 90.3%, 76.2% and 90.4% were positive for preliminary bacterial growth at 72 h. The mean TTP for P. aeruginosa, MRSA and non-MRSA/non-P. aeruginosa isolates on blood culture was 102.3 ± 27.4 h, 77.6 ± 24.4 h and 101.5 ± 51 h, respectively. The cumulative number of cultures with growth based on organism and culture type is reported in Table 2. The Se, Sp, PPV and NPV of 72-h blood and urine culture results for MRSA, non-MRSA/non-P. aeruginosa species and any species are reported in Table 3. Due to the low frequency of P. aeruginosa in blood (n = 3) and urine cultures (n = 12) and MRSA in urine cultures (n = 1), there is little confidence in the estimates of Se, Sp, PPV and NPV for these isolates. For respiratory cultures, P. aeruginosa and MRSA had good Sp and PPV but poor Se and NPV. For non-MRSA/ non-P. aeruginosa species and any species, 72-h culture results had good Se, Sp, PPV and NPV.
Growth at different time points by organism for all positive cultures.

MRSA: methicillin-resistant Staphylococcus aureus.
Sensitivity, specificity, positive predictive value and negative predictive value for final growth at 72 h based on culture type.

Se: sensitivity; CI: confidence interval; Sp: specificity; PPV: positive predictive value; NPV: negative predictive value; MRSA: methicillin-resistant Staphylococcus aureus; N.D.: not defined.
Clinical criteria analysis
Overall, 266 patients (17%) met all of the clinical criteria for de-escalation. The Se, Sp, PPV and NPV for patients meeting antibiotic de-escalation criteria at 72 h are represented in Table 4. Antibiotic de-escalation had good Sp, PPV and NPV, but poor Se for finalized negative cultures. This suggests that meeting criteria for de-escalation at 72 h was a strong predictor of finalized negative cultures; however, patients with finalized negative cultures did not always meet the clinical criteria for de-escalation at 72 h. Due to the low frequency of P. aeruginosa isolates in blood and urine cultures and MRSA in urine cultures, there is little confidence in the estimate of antibiotic de-escalation for these isolates.
Sensitivity, specificity, positive predictive value and negative predictive value for patients meeting antibiotic de-escalation criteria at 72 h based on all culture results (n = 266).

Se: sensitivity; CI: confidence interval; Sp: specificity; PPV: positive predictive value; NPV: negative predictive value; MRSA: methicillin-resistant Staphylococcus aureus.
Discussion
In the first study to calculate the NPV of negative blood, respiratory and urine cultures in critically ill patients, negative blood and urine culture results at 72 h had a high NPV for finalized negative cultures. Negative respiratory culture results at 72 h were weakly predictive of finalized negative cultures; therefore, negative respiratory cultures at 72 h should not guide antimicrobial de-escalation for critically ill patients with suspected pneumonia. Patients with negative cultures also meeting clinical criteria for de-escalation at 72 h had a high Sp, PPV and NPV for finalized negative cultures. The prevalence of a given disease impacts the positive and NPVs, whereas Se and Sp are independent of prevalence. This may explain the discrepancy between the low Se and high NPV observed in both the culture results at each of the pre-specified time periods and in the subgroup of patients meeting clinical criteria for de-escalation. Despite the high rate of false negatives, leading to low Se, the NPV of clinical criteria for de-escalation was high due to an overall low prevalence of positive blood (13.4%), respiratory (59.7%) and urine (25.7%) cultures. The results of this analysis suggest that little additional microbiologic information is gained by waiting until culture finalization and supports de-escalation practices.
For non-MRSA/non-P. aeruginosa and any species, negative 72-h blood, respiratory and urine culture results had high Se, Sp, PPV and NPV for finalized negative cultures. Negative 72-h respiratory cultures had high Se, Sp, positive and NPV for all species, with the exception of P. aeruginosa and MRSA, which had poor Se. In addition, the high NPV of 72-h culture results persisted in the subgroup of patients who met clinical criteria for de-escalation. Together, this information suggests that utilizing 72-h culture results to assess for de-escalation is safe in clinically stable ICU patients.
Critically ill patients require initial broad-spectrum therapy including coverage for MRSA and P. aeruginosa to ensure appropriate coverage is provided; however, continuation of broad-spectrum therapy is not always warranted and is associated with adverse events.25,26 Unnecessary antibiotic exposure has been associated with antimicrobial resistance and Clostridioides difficile infection.11,12 Epidemiologic studies indicate an increase in resistance, with up to 17.2% and 13.7% of Klebsiella and Escherichia coli isolates, respectively, exhibiting the extended spectrum beta-lactamase phenotype. 27 In a database study of patients with gram-negative hospital or ventilator-acquired pneumonia, 11.8% of patients were infected with carbapenem-resistant organisms. This is particularly concerning given the association between development of multi-drug resistant infections and an increase in mortality, likely due to an increase in initial inappropriate therapy.28,29
The high NPV of blood and urine culture results suggests that enough clinical information exists at 72 h to assess for de-escalation of antibiotics. This conclusion is further strengthened by the high Sp, PPV and NPV of patients meeting clinical criteria for de-escalation in addition to negative cultures. The ability to identify true negative cultures prior to culture finalization has broad implications for antimicrobial stewardship efforts and supports protocolized re-evaluation of empiric therapy at 72 h. This timepoint may be useful as a set point for antibiotic time outs, an antimicrobial stewardship tool which prompts providers to reassess the appropriateness of antibiotic therapy. Indeed, antibiotic time outs are recommended by both the Centers for Disease Control and Prevention and The Joint Commission to increase the appropriateness of antibiotic prescribing, and the implementation of formalized antibiotic time outs have been associated with a decrease in inappropriate antibiotic use in a generalized medical population.28–30
This study has several limitations. This study was powered based on the rate of all positive culture results, rather than the rate of positive cultures representing true bacterial infection. Contaminants were not excluded in order to limit the introduction of bias due to the retrospective nature of this review. Over one-third of patients received antibiotics prior to the culture being obtained, and antibiotic administration prior to culture was unable to be determined in an additional 15.7% of patients with blood cultures who were transferred from outside institutions, which can limit pathogen detection. The inclusion of patients with previous antibiotic exposure may have increased the TTP and decreased the NPV of negative culture results at 72 h. Given the known impact of previous antibiotic exposure on TTP, patients should be assessed for previous antibiotic exposure prior to assessing for de-escalation and longer periods of culture observation may be warranted.
Very few patients met the clinical criteria for de-escalation as pre-specified in the study design. This is largely attributable to missing information due to the retrospective nature of the study. In addition, due to the low number of P. aeruginosa isolates on blood and urine cultures, conclusions regarding this species were unable to be made. The single-center design may limit widespread generalizability as time to blood culture positivity may be impacted by culture media and instrument type. Institutional changes in laboratory instrumentation within the study period may have affected TTP during the study period. The transition from the BacT/Alert® 3D automatic blood culture system to the BacT/Alert® Virtuo® may have increased rate of bacterial recovery in the presence of antibiotics and biased the secondary outcome of time to positive preliminary identification. Additional demographic information with respect to severity of illness would assist in generalizing this information to other patient populations, including source of bacteremia. Finally, this study focused on microbiologic criteria and did not assess patient outcomes or how a protocol based on NPV might impact antimicrobial stewardship efforts.
Conclusion
Negative blood and urine culture results at 72 h are predictive of finalized negative cultures in critically ill patients. The results of this study support evaluation for antimicrobial de-escalation in critically ill patients who are immunocompetent, normothermic and on decreasing doses of vasopressors at this time point. Negative respiratory culture results at 72 h were weakly predictive of finalized negative cultures, and therefore, caution should be used in considering de-escalation in critically ill patients with suspected pneumonia. Further research evaluating the use of 72-h culture results to assess for de-escalation on patient outcomes is warranted, with particular focus on resistant organisms.
Supplemental Material
sj-docx-1-smo-10.1177_20503121211040702 – Supplemental material for Time to blood, respiratory and urine culture positivity in the intensive care unit: Implications for de-escalation
Supplemental material, sj-docx-1-smo-10.1177_20503121211040702 for Time to blood, respiratory and urine culture positivity in the intensive care unit: Implications for de-escalation by Lindsey A Sellers, Kathryn M Fitton, Margaret F Segovia, Christy C Forehand, Kevin K Dobbin and Andrea Sikora Newsome in SAGE Open Medicine
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
Ethical approval for this study was obtained from the Augusta University Institutional Review Board (IRB registration no. 00009620 and IRB approval no. 1312519).
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the National Center for Advancing Translational Sciences (NCATS) of the National Institutes of Health (NIH) under Award nos UL1TR002378 and KL2TR002381.
Informed consent
A waiver of HIPAA authorization and waiver of informed consent were approved as part of the IRB application.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
