Abstract
Background:
Diagnosing invasive mold infections (IMI) is challenging because they typically occur as opportunistic infections in immunocompromised patients who often present with non-specific symptoms. Furthermore, no single test can definitively diagnose IMI, and a proven diagnosis often requires invasive sampling. This sampling can be unsafe and difficult to perform, especially in the immunocompromised population most at risk for these infections.
Objectives:
The objective of this study is to assess the clinical utility of plasma microbial cell-free DNA next-generation sequencing (mcfDNA-NGS) for diagnosing invasive mold infections in the context of conventional diagnostic methods.
Design:
Retrospective observational study at a quaternary care center (2017–2025).
Methods:
The charts of 30 patients with mold-positive mcfDNA-NGS (Karius™ Spectrum; Redwood City, CA, USA) were reviewed, with IMI adjudicated per 2020 EORTC/MSGERC criteria. Provider documentation, medication orders, and patient outcomes were used to assess clinician interpretation of mcfDNA-NGS result accuracy (true positive vs false positive) and its impact on diagnostic and therapeutic decision-making. Turnaround time (TAT) and molecules per microliter (MPM) were summarized.
Results:
IMI final classifications were proven (6), probable (8), possible (5), and unclassified (11). Overall, 23/30 (77%) results were true positives. Among these 23 patients with true positive results for mold, 5 (22%) received a new clinical adjudication of IMI that had been entirely missed by conventional diagnostic testing. Furthermore, mcfDNA NGS provided species-level pathogen identification in 9 of the 23 (39%) true positive cases where conventional testing detected fungal elements or elevated biomarkers but could not identify the specific organism. Median TAT was 102 h; median MPM 657. MPM did not differ between true and false positives (p = 0.86). mcfDNA-NGS changed diagnostic classification in 7/30 (23%) and antimicrobial management in 16/30 (53%).
Conclusion:
mcfDNA-NGS provided noninvasive, actionable information, informing diagnosis and therapy. Future studies should define optimal stewardship and cost-effectiveness.
Plain language summary
Some serious infections are caused by molds that are hard to treat. Usual tests can take a long time or may require risky procedures, such as lung or tissue biopsies. A new blood test called microbial cell-free DNA sequencing (mcfDNA-NGS) looks for pieces of fungus directly in the bloodstream, helping doctors identify infection faster. We looked at 30 patients with mold infections at our hospital. The blood test helped doctors make or change a diagnosis in 1 out of 5 patients and adjust antifungal treatment in nearly half. It also gives results within about four days. This study showed that mcfDNA-NGS blood test can give helpful information quickly and safely, especially for people whose immune systems are weak. More studies are needed to learn how best to use this test and how it affects patient recovery and costs.
Background
Invasive molds, such as Aspergillus, Mucor, Fusarium, and Scedosporium, cause significant morbidity and mortality in immunocompromised patients.1,2 Early recognition is critical given the aggressive and disseminated nature of invasive mold infections (IMI), particularly in immunocompromised hosts. However, the diagnosis of IMI remains challenging, given their nonspecific clinical presentations. Moreover, no single diagnostic modality achieves both high sensitivity and specificity, necessitating the integration of clinical, radiologic, and mycologic data.3,4
The 2020 European Organization for Research and Treatment of Cancer/Mycoses Study Group Education and Research Consortium (EORTC/MSGERC) defines invasive mold infection as proven, probable, or possible based on host factors, clinical features, and mycologic evidence. 5 Proven infection requires either culture of mold from a sterile site or histopathologic visualization of hyphae or yeast-like forms from a sterile site. Probable disease necessitates compatible host factors, clinical features, and mycologic data, whereas possible disease is defined by host and clinical findings alone. 5
Mycologic testing remains central to diagnosis; however, it is hindered by several key limitations. Histopathology can confirm tissue invasion by the observation of microscopic fungal elements; however, obtaining tissue requires invasive biopsy that may be contraindicated in critically ill patients.6,7 Although culture is the cornerstone of diagnosis, it is slow (up to 4 weeks), technically demanding, and prone to contamination and overgrowth.8,9 Because molds are ubiquitous, a positive culture from non-sterile specimens may represent colonization rather than infection, particularly in non-immunocompromised hosts. 10 These limitations underscore the ongoing need for diagnostic tools that are faster, less invasive, and more accurate.
Non-culture-based diagnostic assays have become increasingly prominent. For example, serologic tests such as galactomannan (GM) and 1,3-β-D-glucan (BDG) offer noninvasive adjuncts to diagnosis.4,5 Serological GM detection, specific to Aspergillus and occasionally Fusarium, has demonstrated a sensitivity of 60%–80% and a specificity of 80%–95%. 4 BDG detects multiple fungal pathogens but lacks specificity and is best used as a supportive test. PCR-based assays further enhance specificity but require initial clinical suspicion and a standardized protocol. 4
Plasma microbial cell-free DNA (mcfDNA) using next-generation sequencing (NGS) technology has recently emerged as a promising diagnostic modality for invasive mold infection.11,12 This technology enables broad hypothesis-free pathogen detection from a plasma sample, without prior knowledge of the causative organism. After automated extraction and library preparation, mcfDNA fragments are sequenced and compared against curated microbial databases to identify potential pathogens. 11 Given its noninvasive nature and capacity for broad detection, mcfDNA-NGS represents a potentially transformative tool for diagnosing IMI in complex high-risk patients.
Objectives
This study aimed to evaluate the clinical utility of mcfDNA-NGS for detecting IMI in a quaternary care setting. Specifically, the objective was to determine how mcfDNA-NGS results influenced diagnosis and the management of patients with suspected IMI. Secondary objectives included describing the spectrum of mold species detected, turnaround times (TATs), and concordance between mcfDNA-NGS findings and traditional diagnostic modalities.
Design
This retrospective observational study was conducted at Baylor St. Luke’s Medical Center, a quaternary care center in Houston, Texas. Eligible patients in the cohort were those with positive mcfDNA-NGS results for mold between 2017 and 2025 (Figure 1). Any additional pathogens identified by mcfDNA-NGS were documented when present (Table 1). Given the absence of compiled data for initial suspected diagnoses at time mcfDNA NGS testing, we did not include negative results for patients who were suspected of IMI prior to testing. This study complied with the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines (Supplemental Table 1). 13

Flowchart showing inclusion and exclusion criteria for creating the cohort for this study.
Microbiological testing.

Blood culture positivity indicates mold growth from blood and is distinct from mcfDNA NGS, which detects circulating cell-free fungal DNA without organism growth.
BAL, bronchoalveolar lavage; BDG, (1, 3)-β-D-Glucan assay; BK virus, Human polyomavirus 1; CMV, cytomegalovirus; CSF, cerebrospinal fluid; CT, computed tomography; EBV, Epstein–Barr Virus; GM, Galactomannan assay; GMS, Gomori Methenamine Silver stain; HSV-1, Herpes Simplex Virus Type 1; mcfDNA NGS, microbial cell-free DNA next-generation sequencing; MPM, molecules per microliter; NA, not applicable; spp., Species (plural).; TAT, turnaround time.
Methods
For each patient, charts were reviewed from the admission during which mcfDNA-NGS testing occurred through the present date of review. Patient plasma specimens were sent to Karius (Redwood City, CA, USA) for analysis using the Karius™ Spectrum test, a plasma-based assay that identifies more than 1000 pathogens by detecting circulating microbial cell-free DNA. For each specimen, mcfDNA was isolated, sequenced, and compared with reference genomic databases to determine the presence of organisms.12,14
Results were reported as molecules per microliter (MPM), which refers to the quantity of DNA sequencing reads attributable to a specific organism detected in each microliter of plasma. 15 MPM values are determined by comparing the number of sequencing reads attributed to a given organism with those of an internal control. 11 Researchers only documented MPM values for mold pathogens identified via mcfDNA NGS (Table 1). Reference ranges varied by mold type, with many thresholds omitted due to the high risk of invasive disease at any detectable level (Table 1). Clinical TAT was calculated from the time of test order to result availability in the electronic medical record (Table 1).
Data collection
Medical records were systematically reviewed for patient demographics, host factors, clinical presentation, imaging findings, mycologic evidence, antimicrobial management, and clinical outcomes (Tables 1–3). Besides mcfDNA NGS, additional diagnostics (e.g., imaging, microbiology, serology, and histopathology) were ordered at clinician discretion, and documented by reviewers. One investigator (R.B.) performed the initial chart review, and a second investigator (M.A.M.) independently verified accuracy. Discrepancies were resolved by consensus.
Patient characteristics and clinical features.

AML, acute myeloid leukemia; BLT, bilateral lung transplant; CF, cystic fibrosis; CNS, central nervous system; COPD, chronic obstructive pulmonary disease; CT, computed tomography; EORTC/MSGERC, European Organization for Research and Treatment of Cancer/Mycosis Study Group Education and Research Consortium; HbA1c, hemoglobin A1c; ILD, interstitial lung disease; MRI, magnetic resonance imaging; SLE, systemic lupus erythematosus; s/p, status post.
Clinical impact of mcfDNA NGS testing.
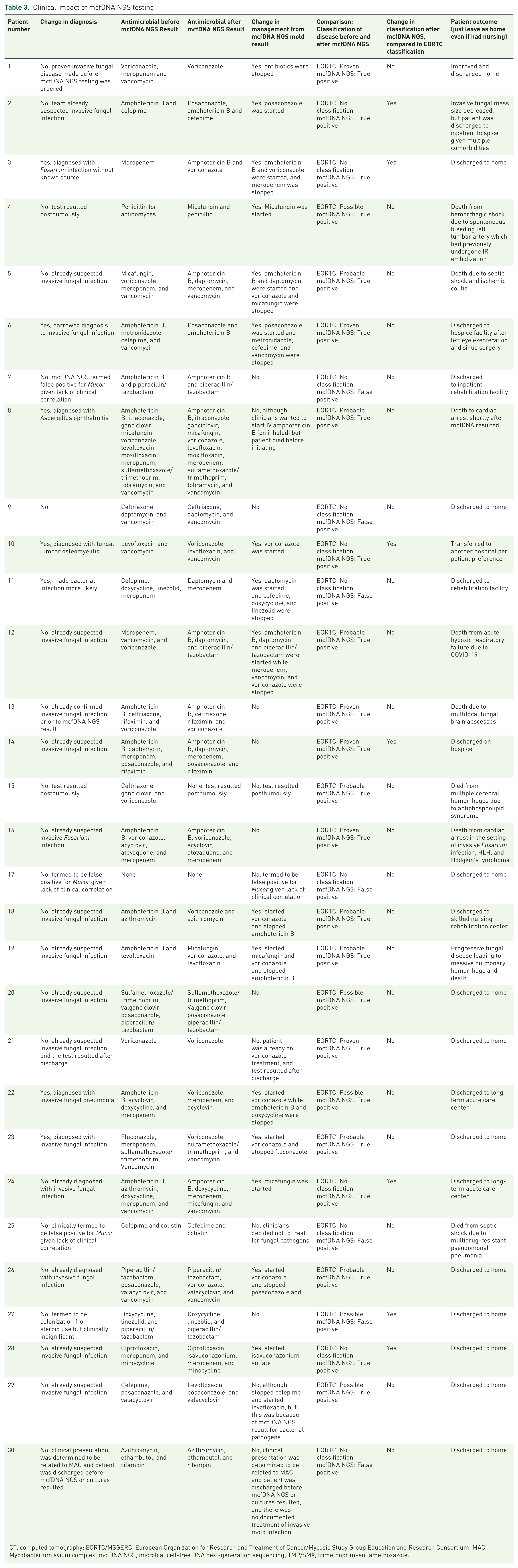
CT, computed tomography; EORTC/MSGERC, European Organization for Research and Treatment of Cancer/Mycosis Study Group Education and Research Consortium; MAC, Mycobacterium avium complex; mcfDNA NGS, microbial cell-free DNA next-generation sequencing; TMP/SMX, trimethoprim–sulfamethoxazole.
Variable definitions
Mycologic evidence was assessed according to EORTC/MSGERC; definitions included: (1) culture of mold from a sterile site (e.g., blood, cerebrospinal fluid, tissue biopsy), (2) histopathologic visualization of fungal hyphae from sterile sites, (3) culture from non-sterile sites (e.g., bronchoalveolar lavage, sputum, wound specimens), (4) microscopy from non-sterile sites, and (5) positive GM antigen testing (Table 1). 5 Although BDG results were documented, BDG positivity does not meet the EORTC/MSGERC definition as mycologic criteria (Table 1). 5 Furthermore, mold-specific PCR assays were not available at the quaternary institution, and as a result, no corresponding PCR data were recorded.
Host factors were defined according to the EORTC/MSGERC criteria and included: (1) prolonged corticosteroid therapy (⩾0.3 mg/kg for ⩾3 weeks within 60 days prior to testing), (2) treatment with T-cell or B-cell immunosuppressants within 90 days, (3) solid organ transplantation, (4) hematologic malignancy, and (5) neutropenia (Table 2). 5 Additional immunocompromising conditions not included in EORTC/MSGERC criteria—specifically uncontrolled diabetes and decompensated cirrhosis—were also documented given their clinical relevance to IMI risk (Table 2).16–18 The American Diabetes Association defines uncontrolled diabetes as a hemoglobin A1c (a measure of average blood glucose levels over the preceding 2–3 months) exceeding 7%. 18 According to the American Gastroenterological Association, decompensated cirrhosis is characterized by the emergence of clinically evident complications, including ascites, hepatic encephalopathy, and/or variceal hemorrhage from the esophagus or stomach. 19
Clinical features were defined per EORTC/MSGERC criteria and included pulmonary involvement (nodules, opacities, or consolidation on chest computed tomography (CT)), sinonasal or orbital involvement (extension across bony barriers on imaging), and central nervous system lesions (focal abnormalities on CT or magnetic resonance imaging (MRI); Table 2). 5 Extrapulmonary manifestations such as soft tissue involvement, osteomyelitis, and cutaneous findings were also documented, though cutaneous findings are not recognized as clinical features under EORTC/MSGERC criteria (Table 2).
Disease classification and outcome adjudication
Each case was classified according to the EORTC/MSGERC definitions of invasive mold infection (Table 2). 5 The primary reviewer (R.B.) classified cases by comprehensive chart review, including patient demographics, previous diagnoses, presenting symptoms, working diagnoses at the time of mcfDNA-NGS testing as documented in clinical notes, vital signs, laboratory results, procedures, and official radiographic interpretations by institutional radiologists. Proven infection required either culture of mold from a sterile site or histopathologic visualization of hyphae in sterile tissue (Table 1). 5 Probable infection required the presence of host factors as defined by EORTC/MSGERC, compatible clinical features, and mycologic evidence (Table 2). 5 Possible infection was defined by the presence of host factors and clinical findings without mycologic confirmation (Tables 1 and 2). 5 Cases that did not meet criteria for proven, probable, or possible infection were categorized as unclassified (Table 2).
Independent of EORTC/MSGERC classification, mcfDNA-NGS results were adjudicated as true positive or false positive based on treating clinician documentation in the electronic medical record (Tables 1–3). Evidence supporting true infection versus colonization was also corroborated by subsequent antimicrobial management decisions and documented clinical improvement or deterioration (Table 3).
Diagnostic impact was defined as any change in disease classification explicitly attributed to mcfDNA-NGS results in clinician documentation (Table 3). Therapeutic impact was defined as any antimicrobial modification (initiation, discontinuation, or change in antifungal agent) explicitly attributed to mcfDNA-NGS results in clinical notes (Table 3).
Patient outcomes were categorized as: discharged home, transferred to rehabilitation or long-term acute care facility, discharged to hospice, or death (Table 3). For deceased patients, documentation was reviewed to determine whether IMI was listed as the cause of death (Table 3).
Statistical analysis
Quantitative variables were assessed for normality using visual inspection of histograms and the Shapiro-Wilk test. Given the small sample size (n = 30) and non-normal distribution of key variables, all continuous data were reported as median with interquartile range (IQR) or range, as appropriate (Table 4). TAT was measured in hours as a continuous variable. Due to the wide range of MPM values (spanning several orders of magnitude) and variation in reference ranges across mold species, MPM was not dichotomized or categorized for primary analyses (Table 4).
Comparison of MPM values by clinical classification.

IQR, interquartile range; MPM, molecules per million; N, number of samples.
Categorical variables were reported as frequencies and percentages. MPM values were compared between true positive and false positive cases to assess whether quantitative fungal DNA burden could distinguish infection from colonization. The Mann–Whitney U test was used, given the non-parametric distribution of MPM data. A two-tailed p-value < 0.05 was considered statistically significant. No adjustment for multiple comparisons was performed, considering the exploratory nature of this analysis.
Given the small sample size and descriptive nature of this study, formal multivariable adjustment for confounding was not performed. However, potential confounders were addressed through comprehensive clinical adjudication. The determination of true versus false positive mcfDNA-NGS results incorporated multiple factors, including host status, clinical presentation, imaging findings, conventional mycologic testing, clinical course, response to therapy, and treating clinician assessment (Tables 1–3). This integrative approach minimized the potential for confounding by indication (i.e., patients with more severe illness receiving more intensive diagnostic workup) and ensured that classifications reflected the totality of clinical evidence rather than any single factor.
Key potential confounders documented included: (1) non-EORTC/MSGERC immunocompromising conditions (uncontrolled diabetes, decompensated cirrhosis and (2) availability of tissue diagnosis (Tables 1–3). These factors were considered qualitatively when interpreting the results.
Diagnostic and therapeutic impact were descriptively examined by: (1) presence or absence of tissue confirmation, (2) EORTC/MSGERC classification category, and (3) specific mold species detected (Table 3). Formal statistical testing for interactions was not performed due to limited sample size and power.
Not every patient underwent all diagnostic modalities. These missing data reflect real-world clinical practice patterns where diagnostic testing was ordered at clinician discretion based on clinical suspicion, patient stability, and procedural risk. Missing diagnostic tests were not assumed to be negative; rather, the absence of testing was documented (Tables 1 and 2). This approach is appropriate for a descriptive cohort study and reflects the pragmatic clinical utility of mcfDNA-NGS in situations where conventional diagnostics are incomplete or unavailable.
All 30 patients were followed from their index hospitalization to final disposition (discharge home, transfer to rehabilitation/long-term acute care, discharge to hospice, or death; Table 3). No patients were lost to follow-up during the index hospitalization (Table 3).
Results
There were 981 patients who underwent mcfDNA NGS testing at Baylor St. Luke’s Medical Center between 2017 and 2025 (Figure 1). Of these, 340 had negative mcfDNA NGS results, 555 were positive for non-mold pathogens on mcfDNA NGS testing, and 56 had no result due to cancellation, courier delay, or inadequate sample (Figure 1). There were no duplicate tests. Thirty patients tested positive for at least one mold pathogen on mcfDNA NGS testing and were included in this study (Figure 1).
The median age was 54.8 years (range: 22–82), and 11 (36.7%) were female (Table 2). Eighteen patients (60%) had at least one predisposing factor for IMI according to the 2020 EORTC/MSGERC criteria, with 11 (36.7%) exhibiting more than one risk factor (Table 2). Twelve (40.0%) received T-cell or B-cell immunosuppressants within 90 days before testing (Tables 2 and 5). 5 Eight (26.6%) had received prolonged corticosteroid therapy, defined as a therapeutic dose of ⩾0.3 mg/kg for ⩾3 weeks within the 60 days prior to testing (Tables 2 and 5). Six (20%) were solid organ transplant recipients; four (13.3%) had hematologic malignancies, and five (16.7%) were neutropenic (Tables 2 and 5). Of the remaining 12 patients that did not have host factors according to the 2020 EORTC/MSGERC criteria, four (36%) had uncontrolled diabetes (patients 1, 2, 21, and 28) and three (25%) had decompensated cirrhosis (patients 3, 14, 24; Tables 1 and 2).
Aggregate data summary.

Some patients had >1 host factor.
Uncontrolled diabetes and decompensated cirrhosis are not included among EORTC/MSGERC host criteria but were present in several true IMI cases.
BDG, 1,3-β-D-glucan; CNS, central nervous system; EORTC/MSGERC, European Organization for Research and Treatment of Cancer/Mycosis Study Group Education and Research Consortium; IMI, invasive mold infection; mcfDNA, microbial cell-free DNA; NGS, next-generation sequencing.
Sixteen patients (53.3%) demonstrated pulmonary involvement based on chest CT findings (Table 5). Three patients (10%) had radiologic evidence of sinus and/or orbital involvement, and three (10%) had focal central nervous system (CNS) lesions (Table 5). Cutaneous involvement was observed in five patients (16.6%), including two with eschars (patients 16 and 18); however, cutaneous findings are not defined as clinical features per the 2020 EORTC/MSGERC criteria (Table 5). 5 Three additional patients (10%) presented with soft tissue or bone involvement, including osteomyelitis and localized extremity lesions (Table 5).
Eighteen patients (60.0%) had mycologic evidence of infection, as defined by the 2020 EORTC/MSGERC criteria (Table 1). 5 Mold was cultured from sterile sites in three cases, including epidural tissue specimens for patients 1 and 21 and cerebrospinal fluid for patient 13 (Tables 1 and 5). Patient 13 also had fungal elements visualized on microscopy of the CSF (Table 1). In three additional patients, molds were identified on histopathology of sterile specimens, including angioinvasive hyphae and fungal elements visualized in orbital tissue (Tables 1 and 5).
Mycological evidence from non-sterile specimens was identified in fifteen (50%) patients. Of these, twelve fulfilled EORTC/MSGERC mycologic criteria; the remaining three patients (patients 20, 25, and 28) had positive BDG as their sole mycologic finding, which does not meet the EORTC/MSGERC criteria (Table 1). 5
Ten patients had mold growth from cultures of non-sterile sites: four from extremity wound cultures, three from sinus cultures, two from bronchoalveolar lavage, and one from lung tissue biopsy (Tables 1 and 5). Of these, patient 3 also had mold cultured from a different, sterile site (Tables 1 and 5). Fungal hyphae were visualized on microscopy of non-sterile sites in seven patients, two of which also had fungal hyphae identified in sterile site specimens (Tables 1 and 5). Twelve patients had positive BDG results, and six patients had positive galactomannan assays (Tables 1 and 5).
The median TAT for NGS testing was 102 h (interquartile range (IQR), 74–123; Table 1). mcfDNA-NGS identified five major mold classifications, the most common being the Aspergillus genera, detected in 17 patients, followed by molds within the Mucorales order (in seven patients; Table 1). The median MPM value for mold detections was 657 (IQR, 62–2293). The highest mean MPM level was observed among patients with proven infections according to EORTC/MSGERC criteria (Table 4). 5 However, there was no statistically significant difference in the MPM values between true- and false-positive cases (p = 0.86; Table 4).
Based on the 2020 EORTC/MSGERC definitions, six patients met the criteria for proven IMI, eight for probable infection, and five for possible infection; the remaining 11 patients did not meet criteria for any classification of IMI (Tables 2 and 5). 5 True positive mold infections were confirmed in 23 of 30 patients (77%) based on integration of host factors, clinical presentation, imaging findings, mycologic evidence, clinical course, and treating clinician assessment (Tables 1 and 3). The mcfDNA-NGS results in all proven and probable cases were clinically confirmed as true positive (Table 3). Of the five patients with possible invasive mold classifications, only one (patient 27) was determined to represent colonization rather than true infection (false positive; Table 3).
Among the 11 unclassified cases, five results were ultimately determined to be true positives (Table 3). Four of these lacked classic host factors as defined by the EORTC/MSGERC guidelines but were immunocompromised due to uncontrolled diabetes (patients 2 and 28) or decompensated cirrhosis (patients 3 and 24; Table 2). The final true positive patient, unclassified by EORTC/MSGERC criteria due to absent clinical findings, had MRI evidence of lumbar osteomyelitis (Tables 2 and 3). 5 The remaining six patients that were not classified were considered false positives due to colonization (Tables 1 and 4).
Tissue confirmation was obtained in 13 patients (43.3%); five patients had positive cultures alone, one had fungal elements identified on microscopy alone, and the remaining seven patients had both culture and microscopy confirmation (Table 1). Of these 13 patients, 12 were classified as true positives and one as false positive (Tables 1 and 3). The remaining 17 patients had no tissue confirmation; of these, 10 were considered true positives based on non-invasive findings, including positive galactomannan antigen testing, concordant mcfDNA NGS results, and clinical course as interpreted by treating physicians (Tables 1 and 3).
Among all 30 patients, mcfDNA-NGS results led to a change in diagnostic classifications in seven cases (23.3%) and modification of antimicrobial management in 16 cases (53.3%; Table 3). Following hospitalization, 18 patients were discharged, including 14 who returned home and four who were transferred to rehabilitation or long-term acute care facilities (Table 3). Twelve patients died or were discharged to hospice, with IMI definitively listed as the cause of death in three cases (patients 13, 16, and 19; Table 3).
Discussion
This is the first study to systematically evaluate the clinical impact of mcfDNA-NGS on the diagnosis and management of IMI among patients with varied causes of immunosuppression. Prior work has established the diagnostic potential of plasma mcfDNA-NGS for fungal disease, but few studies have examined how the test influences clinical decision-making. Our findings demonstrate that mcfDNA-NGS can meaningfully influence patient care, leading to diagnostic changes in 23.3% and therapeutic modification in 53.3% of cases (Table 4).
Other studies have shown that plasma mcfDNA-NGS can detect fungal DNA in patients with proven and probable IMI, with reported sensitivities ranging from 44% to 68% and specificities from 95% to 100%.20–22 Moreover, because tissue biopsy is often unsafe or infeasible in patients with hematological malignancy, transplantation, or critical illness, plasma mcfDNA-NGS offers a noninvasive alternative when conventional sampling cannot be performed.23,24 This advantage was corroborated by our findings, in which seventeen patients (57%) lacked tissue confirmation, of which 10 (58.8%) were clinically adjudicated as true positives based on concordant mcfDNA-NGS results and supporting radiologic or serologic evidence. A case-control study evaluating mcfDNA sequencing in high-risk immunocompromised patients reported higher sensitivity for proven versus probable invasive fungal disease (60.0% vs 37.1%) 25 Although formal sensitivity calculations were not possible due to the absence of a control group, our results were concordant with prior findings: false positive results were more common in categories with lower diagnostic certainty according to EORTC/MSGERC criteria.
Quantitative dynamics of fungal mcfDNA have been evaluated in several longitudinal studies. Heldman et al. 23 and Karam et al. 26 demonstrated that MPM levels rose before clinical diagnosis and declined with antifungal therapy. In our series, the highest MPM levels occurred in patients with proven infections. However, there was no significant correlation between MPM values and clinical classification (true vs false positive), indicating that quantitative mcfDNA burden alone does not distinguish infection from colonization. These findings support prior observations that MPM is most informative when interpreted longitudinally rather than as an absolute cutoff. 23
Our data also highlight persistent gaps between molecular diagnostics and classification frameworks. Of the 11 patients who were unclassified under the EORTC/MSGERC criteria, five were determined to have true positive IMI (Table 4). To receive a classification, patients must have specific host factors under the EORTC/MSGERC criteria. 5 Four patients remained unclassified despite having significant immunocompromising conditions: two had uncontrolled diabetes, and two had decompensated cirrhosis. Both diagnoses are known to cause immunodeficiency leading to IMI, but were excluded from the criteria.5,16–19,27–31
One exhaustive case–control review of 925 patients with zygomycosis since 1885 found that 36% of patients with IMI had diabetes as their sole host risk factor, representing the most common underlying condition for IMI. 27 This discovery corresponds with our findings, as one unclassified patient had uncontrolled diabetes and invasive mucormycosis. The other patient with pulmonary aspergillosis also lacked recognized EORTC/MSGERC host factors, yet this presentation has been documented in case series of patients with uncontrolled diabetes and no other risk factors.28,29 A systematic review and meta-analysis further demonstrated that ICU patients with diabetes mellitus were more likely to develop invasive aspergillosis than those without diabetes. 30
Regarding cirrhosis and IMI, a study of 185 patients identified 19 cases of IMI after excluding all other types of immunodeficiency. 16 The two patients in our cohort with cirrhosis who had true positive IMI but lacked EORTC/MSGERC-defined host factors had decompensated cirrhosis. This aligns with a meta-analysis showing that IMI was more common in patients with decompensated cirrhosis compared to those with compensated cirrhosis, whose IMI rates did not differ from the general population. 17 Another report examining hematologic malignancy associated with pulmonary IMI emphasized that the rigid definitions may underestimate disease in the molecular era. 32 Updating the consensus framework to incorporate mcfDNA-NGS, as well as decompensated cirrhosis and uncontrolled diabetes as host factors, could be essential to accurately capture the full spectrum of modern fungal disease.
Careful attention should be paid to the potential for false positivity when interpreting the mcfDNA-NGS results. Around a quarter (7/30) of the patients in this study had false positive results, which is consistent with other reports. Benamu et al. 33 reported irrelevant mcfDNA in 29% of the patients with febrile neutropenia. The false positive rate of approximately 25% in our cohort underscores the need for judicious interpretation. Multiple mechanisms may contribute to false positivity, including transient mold DNAemia from mucosal barrier disruption, airway colonization releasing cell-free DNA without tissue invasion, residual DNA from previously treated infections, and potential low-level contamination despite rigorous laboratory protocols. 33 To optimize clinical utility, mcfDNA-NGS results should be interpreted within a comprehensive diagnostic framework. Positive results, particularly low-level detections, require correlation with host factors (including non- EORTC/MSGERC immunocompromising conditions such as uncontrolled diabetes and decompensated cirrhosis), compatible clinical and radiologic findings, and conventional mycologic evidence when available. Serial MPM monitoring may help distinguish active infection from colonization or clearance, though our data suggest single MPM values alone do not reliably differentiate these states. Results obtained during or shortly after antifungal therapy should be interpreted cautiously, as they may represent treated disease rather than active infection. Antimicrobial stewardship programs should establish institutional guidance for mcfDNA-NGS utilization, including pretest probability assessment, multidisciplinary result interpretation involving infectious diseases expertise, integration with rather than replacement of conventional diagnostics, and quality improvement tracking to refine testing algorithms over time. Such stewardship efforts are essential to maximize diagnostic yield while minimizing overinterpretation and unnecessary antimicrobial exposure.
This study has several limitations. Only patients with positive mcfDNA NGS results were included because data on clinical suspicion for IMI at the time of test ordering were not systematically documented, precluding inclusion of all tests ordered for suspected IMI. Furthermore, since only patients with positive mcfDNA-NGS results were included, sensitivity and specificity could not be determined. Additionally, this study did not include a formal a priori sample size calculation. The sample size was determined by the availability of all mold-positive mcfDNA-NGS cases at our institution during the study period (n = 30), rather than by statistical power considerations. Given the exploratory nature of this investigation and the lack of established effect size estimates for mcfDNA-NGS diagnostic and therapeutic impact in IMI at the time of study design, a convenience sampling approach was utilized. The resulting sample size limits statistical power for subgroup analyses, precludes robust evaluation of test performance across different patient populations or mold species, and increases the risk of type II error when comparing quantitative parameters such as MPM values between groups.
PCR testing was not available at the study institution and therefore was not performed in any patient. This limitation is significant because PCR is included in the EORTC/MSGERC IMI criteria and, according to a systematic review, has demonstrated high sensitivity for detecting invasive mold infections, especially Aspergillus species. 34
Additionally, cost-effectiveness was not evaluated, so the financial and operational impact of incorporating mcfDNA-NGS testing into routine practice remains uncertain. Despite these limitations, our clinician-verified documentation of how mcfDNA-NGS results influenced diagnostic and therapeutic decisions provides real-world evidence bridging analytic validation with bedside implementation.
The generalizability of this study’s findings is moderate and context-dependent. As a single-center, retrospective observational cohort conducted at a quaternary academic medical center, the results are most applicable to tertiary and quaternary care settings that manage complex, immunocompromised patients and have access to infectious diseases expertise, advanced imaging, and antifungal stewardship support. External validity is supported by the real-world nature of test ordering and interpretation, the heterogeneous patient population with varied immunosuppressive conditions and clinical presentations, and the inclusion of a broad range of invasive mold pathogens beyond Aspergillus alone. However, generalizability is limited by the small sample size, inclusion of only patients with mold-positive mcfDNA-NGS results, reliance on clinician adjudication rather than a uniform microbiologic gold standard, and institution-specific diagnostic workflows and TATs. Additionally, findings are specific to a single commercial mcfDNA-NGS platform and may not directly translate to other metagenomic assays. Consequently, while the results provide clinically relevant insights into the utility of mcfDNA-NGS in high-risk hospital settings, caution is warranted when extrapolating these findings to community hospitals, lower-risk populations, or screening contexts, underscoring the need for larger, multicenter prospective studies to confirm broader applicability.
In conclusion, plasma mcfDNA-NGS is an important advance in non-invasive mold diagnostics. Our findings show that mcfDNA-NGS results can guide diagnosis and management, highlighting the need for prospective studies to define its optimal clinical and economic role.
Supplemental Material
sj-docx-1-tai-10.1177_20499361261437009 – Supplemental material for Clinical impact of microbial cell-free DNA next-generation sequencing for invasive mold infection—a single-center retrospective observational study
Supplemental material, sj-docx-1-tai-10.1177_20499361261437009 for Clinical impact of microbial cell-free DNA next-generation sequencing for invasive mold infection—a single-center retrospective observational study by Rebecca Berger, Nina Howard, Sarah Grant, Fernando Centeno, Todd Lasco and Mayar Al Mohajer in Therapeutic Advances in Infectious Disease
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
