Abstract
Background:
Metagenomic next-generation sequencing (mNGS) testing identifies thousands of potential pathogens in a single blood test, though data on its real-world diagnostic utility are lacking.
Objectives:
Determine the diagnostic utility of mNGS testing in practice and factors associated with high clinical utility.
Design:
Retrospective cohort study of mNGS tests ordered from June 2018 through May 2020 at a community teaching hospital.
Methods:
Tests were included if ordered for diagnostic purposes in patients with probable or high clinical suspicion of infection. Exclusions included patient expiration, hospice care, or transfer outside of the institution. Utility criteria were established a priori by the research team. Two investigators independently reviewed each test and categorized it to either high or low diagnostic utility. Reviewer discordance was referred to a third investigator. The stepwise multiple regression method was used to identify clinical factors associated with high diagnostic utility.
Results:
Among 96 individual tests from 82 unique patients, 80 tests met the inclusion criteria for analysis. At least one potential pathogen was identified in 58% of tests. Among 112 pathogens identified, there were 74 bacteria, 25 viruses, 12 fungi, and 1 protozoon. In all, 46 tests (57.5%) were determined to be of high diagnostic utility. Positive mNGS tests were identified in 36 (78.3%) and 11 (32.4%) of high and low diagnostic utility tests, respectively (p < 0.001). Antimicrobials were changed after receiving test results in 31 (67.4%) of high utility tests and 4 (11.8%) of low utility tests (p < 0.0001). In the multiple regression model, a positive test [odds ratio (OR) = 10.9; 95% confidence interval (CI), 3.2–44.4] and consultation with the company medical director (OR = 3.6; 95% CI, 1.1–13.7) remained significantly associated with high diagnostic utility.
Conclusion:
mNGS testing resulted in high clinical utility in most cases. Positive mNGS tests were associated with high diagnostic utility. Consultation with the Karius® medical director is recommended to maximize utility.
Plain language summary
Our institution has utilized a meta-genomic test that identifies bacteria, DNA-based viruses, fungi and protozoa from blood sample in hospitalized patients to support diagnostics in select clinical cases. We evaluated the utility of these tests in an adult and pediatric population. We found that 58% of the 96 tests from 82 unique patients produced a pathogen. Overall, a majority (58%) of tests were deemed to be of high utility which directly resulted in changes in antimicrobial therapy, selection of duration of therapy, direction for new diagnostics, or avoidance of further need for diagnostics. Positive tests and consultation with the medical director of the laboratory were both associated with high utility of the tests.
Introduction
Diagnostic modalities available to practicing clinicians span a wide variety of techniques, including microbial culture, mass spectrometry, pathogen-specific serology, and nucleic acid identification through polymerase chain reaction.1–3 However, such modalities have limitations depending on assay variability, laboratory personnel experience, and provider gestalt when suspecting infection. With these traditional diagnostic techniques, pathogen detection may not always be feasible when faced with fastidious organisms, high-risk biopsy interventions with associated sampling errors, and sample sterilization from antimicrobials. Metagenomic next-generation sequencing (mNGS) is a shotgun approach to diagnosis, utilizing cell-free DNA-based genome assembly and subsequent identification of gene targets within a sample that does not depend on growth in culture or targeting using specific primer molecules. 4 Clinically, mNGS has shown to have predictive and prognostic implications for various malignancies as well as a modality for pathogen identification in infected hosts – particularly for scenarios where the site of infection is difficult to sample for traditional pathogen identification.5–10 Moreover, multiple types of pathogens including viral, fungal, bacterial, and parasitic isolates can be recovered using a single assay allowing for an unbiased approach to clinical diagnosis. 11 These qualities make mNGS attractive among providers; however, current data reveal limitations in guiding usage, clinician misinterpretation, and maximizing clinical utility in presumably infected hosts. 12 Specific limitations to the mNGS testing methodology include high susceptibility to contamination at the time of collection and sample preparation and the need to weed out host cell-free DNA, which adds complexity to the sequencing process and may complicate result interpretation or lead to over-treatment of non-pathogenic organisms. 12
Positive tests have demonstrated high sensitivity in the literature ranging from 74% to 95%, and have been associated with longer hospital stays and increased 28-day mortality.7,13–15 However, the utility of test results leading to clinically actionable interventions has yet to be fully delineated, as few studies to date have evaluated the patient care impact that mNGS testing in suspected bloodstream infections has on antimicrobial utilization.4,16–18 The purpose of this study is to determine the clinical utility of positive and negative mNGS tests and better characterize the clinical applications by identifying factors associated with high and low diagnostic utility. Although not a diagnostic accuracy study intended to assess specificity and sensitivity, we aimed to determine the factors associated with the high or low utility to provide a framework for clinicians to effectively utilize mNGS. We hypothesize that there will be discrete factors associated with high or low utility which will aid in the applicability of mNGS in pathogen identification and clinical decision-making.
Methods
Karius® test and reference standards
Five milliliters of whole blood was collected in a BD Vacutainer (PPT) or K2-ethylenediamineteraacetic acid tube and separated via centrifugation within 6 h of the initial draw. Fresh (within 4 days of initial collection) or frozen samples were sent to the Karius Incorporated® laboratory (Redwood City, CA, USA). Testing was performed by Karius® per routine testing protocol. 19 A positive test was defined by the identification of at least one organism on the report. Comparison diagnostic tests including bacterial and fungal cultures, histopathology, microscopy, mass spectrometry, pathogen serology, and broad range and multiplex polymerase chain reactions were performed under quality-controlled laboratory metrics per institutional protocols. These represent the reference standard and were chosen based on the suspected source of infection per clinician discretion, with all data being collected retrospectively.
Study design
We conducted a retrospective cohort study of all mNGS samples sent for Karius® testing from Prisma Health – Midlands Richland Campus (Columbia, SC, USA), a 641-bed academic medical institution and 174-bed free-standing Children’s Hospital, between 1 June 2018 and 31 May 2020. Individual tests were included for data collection and review if they were ordered for clinical diagnostic purposes for inpatients who had a pre-test infectious diagnosis or clinical suspicion for infection documented in provider notes. Tests were excluded if the patient expired, was admitted to hospice service, or was transferred to an outside hospital system prior to receiving the result of the Karius® test. Data collection was performed by the principal investigator. Two researchers (ARW and WZW) conducted an independent review of all tests meeting inclusion. Researchers categorized each test as either ‘high’ or ‘low’ diagnostic utility using criteria adapted from a prior study (Table 1). 17 These criteria reflect current antimicrobial and diagnostic stewardship initiatives to optimize antimicrobial use and minimize costs associated with diagnostic testing. The designation was based on the treating team’s interpretation and subsequent treatment decisions after receiving the test results. A variety of traditional diagnostic tests were performed as described above in addition to mNGS, based on the treating clinician’s discretion. These results were used as part of the criteria to determine clinical utility as described in Table 1. Variability in results between traditional diagnostic tests and mNGS was reflected in whether mNGS was deemed high versus low utility, including negative or indeterminate traditional diagnostics when compared with a positive mNGS result. Positive traditional diagnostic tests were defined as positive bacterial or fungal cultures, broad range or multiplex polymerase chain reaction, histopathology, serology, or microscopy when an organism was identified and deemed to be the etiology of the patient’s infection by the treating clinician. These measures are in accordance with the STARD 2015 guidelines for reporting diagnostic accuracy studies. 20 After review of all tests meeting inclusion, any discrepant designations of diagnostic utility between the two researchers were escalated to a third researcher (PBB) for final independent review and utility designation.
Diagnostic utility criteria.

Outcomes
The primary outcome of interest was the proportion of tests determined to be high and low diagnostic utility. Tests were grouped into high- and low-utility cohorts and additional analyses were conducted to identify clinical, microbiological, and demographical characteristics associated with high or low diagnostic utility.
Statistical analysis
Comparisons of categorical and continuous variables were made using chi-squared/Fisher’s exact test and Student’s t-test, respectively. To identify characteristics associated with high or low clinical utility, stepwise multiple logistic regression was used to identify clinical variables associated with high and low diagnostic utility. Variables with a p value of less than 0.25 in the univariate regression model were included in the multiple regression model. Odds ratios (ORs) and 95% confidence intervals (CIs) were reported to demonstrate the strength of association between each variable and diagnostic utility. A formal power calculation was not performed due to the descriptive nature of the primary outcome.
Results
In all, 96 individual mNGS tests were ordered during the study period and assessed for eligibility, representing 82 discrete patients. In total, 16 tests failed to meet inclusion (Figure 1) with the primary reason for exclusion being outpatient testing, resulting in 80 total tests meeting inclusion. In all, 46 tests (57.5%) were deemed high utility, and 34 tests (42.5%) were deemed low utility, which represented 34 and 31 unique patients, respectively. The study population included 42 male and 40 female patients, with an average age of 18 and 17 years in the high- and low-utility cohorts, respectively. Nine patients (25%) had invasive hardware at baseline in the high-utility cohort compared to 14 (45.2%) in the low-utility cohort, showing a trend toward statistical significance favoring low diagnostic utility (p = 0.083). In addition, there was a trend toward statistical significance for drug-induced immunocompromise that favored high diagnostic utility (p = 0.085). Apart from these two characteristics, there were no differences in baseline patient characteristics between high- and low-utility groups. Additional patient demographics and characteristics are provided in Table 2. Table 3 provides the distribution of pathogens identified in the high- and low-utility cohorts. The total number of organisms identified in the high- and low-utility cohorts were 94 and 18, respectively (p < 0.0002), which included bacteria (59 versus 15, p = 0.091), viruses (24 versus 1, p = 0.070), fungi (10 versus 2, p = 0.952), and protozoa (1 versus 0, p = 1) showing no statistically significant difference in breakdown of pathogens but significantly more in the high-utility group.

Inclusion of eligible tests for diagnostic utility evaluation.
Baseline characteristics.

Microbiology distribution.

Of the 57.5% of tests that met the criteria for high diagnostic utility, 25% resulted in the initiation of appropriate antimicrobial therapy and 24% resulted in the de-escalation of initial therapy. For the 42% of tests that were characterized as low diagnostic utility, 70% of those tests had no impact on antimicrobial therapy. The results from high- and low-utility tests are shown in Figures 2 and 3, respectively. Test characteristics for all tests ordered are shown in Table 4, which included 78.3% of positive tests being high utility compared to only 32.4% of positive tests determined to be low utility (p < 0.001). Consult with the Karius® director was found to be associated with high diagnostic utility (p < 0.039). A high-utility test was associated with a more frequent change in antimicrobial therapy (p < 0.0001), largely driven by the de-escalation of therapy (67.7% versus 25%, p = 0.0002).

High diagnostic utility criteria.

Low diagnostic utility criteria.
Test characteristics.

Determined not to be in response to Karius® test result.
Variables included in the univariate regression analysis are shown in Table 5. A positive test (p < 0.0001), consult with the Karius® director (p = 0.081), increase in qPitt bacteremia score from baseline at the time of test (p = 0.083), being on antimicrobials at the time of test (p = 0.218), and repeating a test (p = 0.113) all met criteria for inclusion in the multiple regression for association with high utility. 21 The presence of diabetes mellitus met inclusion for association with low utility (p = 0.126). Subsequent multiple regression analysis depicted in Figure 4 revealed that a positive test (p = 0.0003) and a consult with the Karius® medical director (p = 0.043) retained statistical significance for an association with high diagnostic utility, while the presence of diabetes mellitus retained statistical significance for an association with low diagnostic utility (p = 0.039).
Univariate regression analysis.

CI, confidence interval; OR, odds ratio.
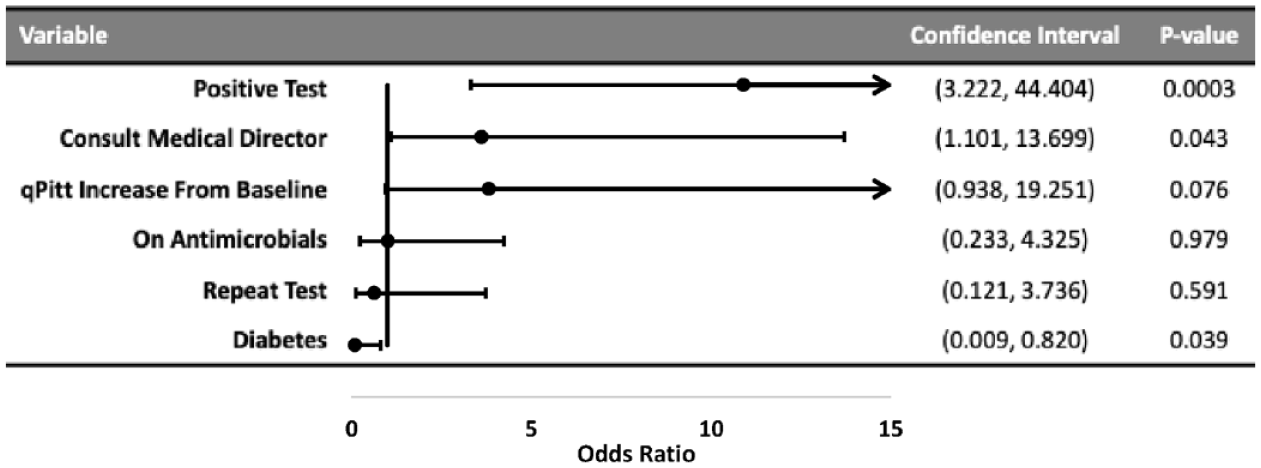
Multiple regression analysis.
Discussion
With the ease of testing and a broad range of organisms that may be identified, including those that are difficult to isolate in traditional culture, mNGS has emerged as an enticing tool to aid in the diagnosis of infectious syndromes. The majority of current literature evaluating its clinical use has focused on specific populations, including pediatrics, post-transplant, and hematologic/oncologic patients, as well as for specific infectious diseases, such as culture-negative endocarditis, pneumonia, and fever with neutropenia.18,22–24 Currently, a paucity of data exists evaluating the clinical utility of mNGS testing in real-world practice, making the overall value of this tool poorly studied. This retrospective cohort study of a mixed patient population found that Karius® tests resulted in high diagnostic utility in a majority of cases (58%), which differed significantly from a previous study in which only 7.3% of tests yielded a positive impact. 24 This is primarily due to differences in study design of the aforementioned study as a large majority of tests resulted in either no impact or indeterminant impact (89%). These two categories were not included in our study design to allow for a direct comparison of groups to identify patient subsets or clinical characteristics that may increase the diagnostic utility. Notably, Hogan et al. 17 reported that when only looking at tests that were determined to have a ‘positive’ or ‘negative’ impact (nine tests in total), 66% had a positive impact which is similar to our finding of 58% high diagnostic utility.
A documented criticism of shotgun sequencing for the diagnosis of infectious syndromes is the risk of contamination or detection of either non-viable or non-clinically relevant organisms which could potentially lead to unnecessary treatment. 12 A key finding of our study is the overall impact the Karius® test had on antimicrobial therapy. Most tests determined to be high diagnostic utility were designated as such based on the action items of antimicrobial de-escalation or discontinuation in 54% of tests. Notably, the proportion of high-utility tests that identified an infectious organism (as opposed to a negative test) was significantly higher compared to the low-utility cohort. With positive tests retaining independent association with high diagnostic utility in the multivariate analysis, this suggests that the primary use of positive tests at our institution is to de-escalate antimicrobials. This is supported by previous studies evaluating the utility of mNGS testing for specific infectious indications (i.e. lower respiratory tract infections, pediatric endocarditis); but to our knowledge, this is the first report of reduction in antimicrobial use as a primary finding within a diverse population and suspected infection sources.13,23 While antimicrobial stewardship outcomes, such as antimicrobial days of therapy or time to appropriate antimicrobial therapy, were not collected in our study by design, this finding is hypothesis generating in that positive mNGS tests may reduce antimicrobial consumption, though a study designed to test this outcome is needed.
At the time of this study, Karius® Inc. has offered clinical consultation with all test reports – including those with a negative result. To our knowledge, this is the first study to include whether consultation with a Karius® Inc. representative is associated with high or low diagnostic utility. While the formal report only includes organisms of which the detected cell-free DNA meets a concentration and quality threshold using a proprietary algorithm, hundreds of organisms may be detected by metagenomic sequencing within a given sample. In our experience, the correlation of test findings with patients’ clinical picture has rarely, albeit impactfully, led reports to be amended with clinically relevant organisms. Specifically for tests that failed to meet certain test standards (e.g. human genome background interference) but the cell-free DNA was detected at high concentrations.
In several cases, tests that do not meet quality standards have resulted in potential life-saving interventions. As a brief example, a critically ill 59-year-old gentleman with acute on chronic respiratory failure secondary to community-onset pneumonia continued to clinically worsen despite antibiotics targeted at both community-acquired pathogens as well as Pseudomonas aeruginosa and methicillin-resistant Staphylococcus aureus. Atypical pneumonia coverage was initially withheld secondary to a negative Legionella urine antigen test. Traditional infectious workup, including culture and rapid diagnostics for bacterial, viral, and fungal pathogens, remained negative. On hospital day 6, he continued to clinically decline with the development of gross hemoptysis. A Karius® test was ordered at that time and returned positive for Legionella pneumophilia, for which his antimicrobial regimen was adjusted to target. He ultimately improved with definitive therapy for Legionella pneumonia and was discharged on hospital day 14.
Our study does carry certain limitations given its retrospective nature. We were unable to reliably determine the pre-test intent of the ordering physician; therefore, tests ordered with an infection rule-out intent may have resulted in more antibiotic changes than those with an infection rule-in intent, thus being more likely to result in high diagnostic utility by our definition. Second, given that commercially available mNGS testing is a relatively new diagnostic tool, there was likely a period in our study when treating clinicians were not as comfortable with making treatment decisions in response to test results due to a lack of clinical experience. This may have led to more tests resulting in low diagnostic utility early in our study period as our utility criteria were largely based on action items or changes made to therapy. Another limitation was the timing of sampling relative to the onset of infectious symptoms and initiation of antimicrobial treatment. mNGS was a second or even last-line diagnostic option in nearly all cases in our study and its place in aiding diagnosis tended to fall after conventional methods failed to provide pathogen identification. While, in theory, cfDNA can be detected in serum samples despite pre-treatment with antimicrobials, it is not known whether the duration of pre-treatment will influence mNGS test results. Therefore, we may have had an increased rate of false-negative results as compared to protocols that used mNGS testing earlier in the clinical timeline. It was for this limitation as well that we were unable to calculate sensitivity and specificity due to an overall lack of gold-standard positive results. Lastly, with our relatively small sample size and a mixed patient population, we were likely unable to enroll enough patients with specific comorbidities to detect a statistical association with high or low diagnostic utility.
Conclusion
Our study demonstrates that mNGS testing used to aid in the diagnosis of infectious diseases results in high diagnostic utility in most cases when ordered and interpreted by infectious disease specialists. There is potential that Karius® tests accompanied by a consultation with a Karius® Inc. representative may reduce antimicrobial consumption in medically complex cases, though this hypothesis should be assessed in subsequent studies. More data are needed to assess the most appropriate place in therapy where mNGS testing should be employed to maximize sensitivity while facilitating the timely initiation of appropriate antimicrobial therapy.
