Abstract
Optimal care of patients requiring long-term outpatient parenteral or oral antimicrobial therapy by infectious diseases (ID) specialists is facilitated by an accurate microbiologic diagnosis. Close collaboration between ID specialists and the clinical microbiology laboratory for routine or specialized molecular testing can result in more accurate diagnoses, streamlined antimicrobial regimens, and improved patient outcomes.
Introduction
Like antimicrobial stewardship, precision medicine aims to target the right treatment to the right patient at the right time. 1 While precision medicine typically refers to treatments tailored to a patient’s own genetic profile, in infectious diseases (ID), molecular characteristics of microbes may have a significant impact on a patient’s clinical outcome, and often dictate antimicrobial therapy. Patients with deep-seated or chronic infections often have multiple comorbidities and require complex surgical and medical management. These patients are at risk for prolonged hospitalizations and readmissions due to treatment failures. 2 Long-term outpatient parenteral antimicrobial therapy (OPAT) or complex outpatient antimicrobial therapy including either oral or parenteral therapy (COpAT) overseen by ID specialists (pharmacists and physicians) is a common treatment modality for such patients; however, optimal selection of the most targeted regimens, whether oral or parenteral, requires timely and accurate microbiologic diagnosis. For instance, infective endocarditis (IE), a common OPAT diagnosis, is classified as ‘blood culture negative’ in approximately 10% of cases from industrialized regions, either due to bacteria whose growth is inhibited by prior antibiotics, or microorganisms which cannot be isolated by routine culture techniques. 3 As such, identification of Coxiella burnetii, Bartonella species, or Tropheryma whipplei by polymerase chain reaction (PCR) assay or other molecular techniques, was added as a new Major Criterion to the 2023 Duke-International Society for Cardiovascular Infectious Diseases Criteria for IE. 3 Therefore, close collaboration with the clinical microbiology laboratory to facilitate routine or specialized testing can maximize the likelihood of optimal patient outcomes. However, this collaboration should include a discussion about the availability and cost of proposed testing. Matrix-assisted laser desorption ionization–time of flight mass spectrometry (MALDI-TOF MS) is most economical, followed by target-specific PCR assays. Broad-range PCR and genomic sequencing remains considerably more expensive, therefore not yet widely available, but may become more cost-effective with time, like prior technologies (e.g. MALDI). 4 The benefit of obtaining an accurate diagnosis to optimize patient care may justify the cost of specialized testing, and ID and microbiology colleagues typically work together to enable the best possible patient outcomes.
Herein, we use fictionalized clinical scenarios based on experiences from our institution to provide examples of optimal collaboration between ID specialists and clinical microbiology laboratories to assist in the management of diagnostically challenging patients requiring extended antimicrobial regimens. Molecular testing options are summarized (Table 1), as are suggestions for collaborative workflows (Figure 1).
Molecular tests for microbial identification and pros and cons of each.
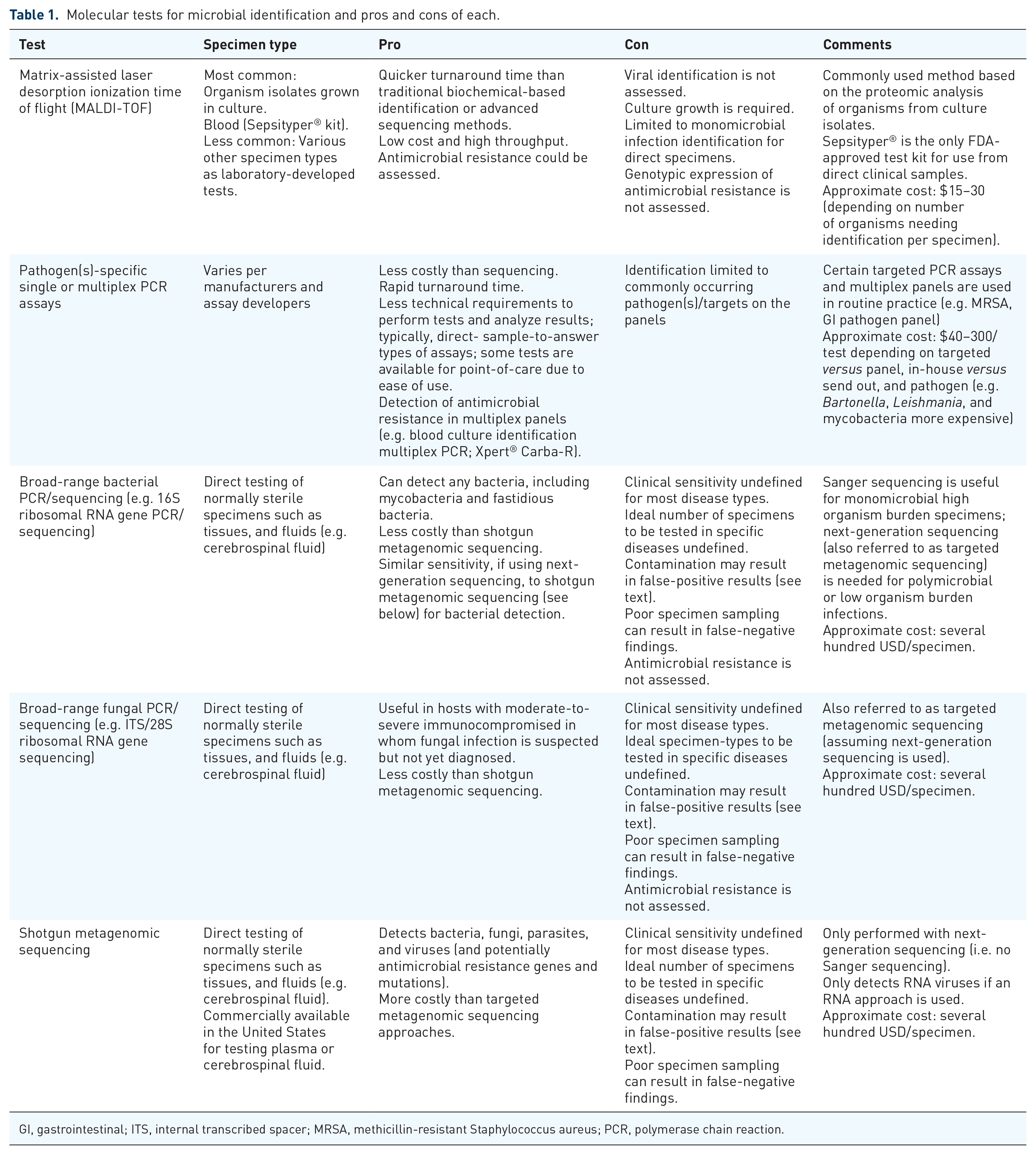
GI, gastrointestinal; ITS, internal transcribed spacer; MRSA, methicillin-resistant Staphylococcus aureus; PCR, polymerase chain reaction.
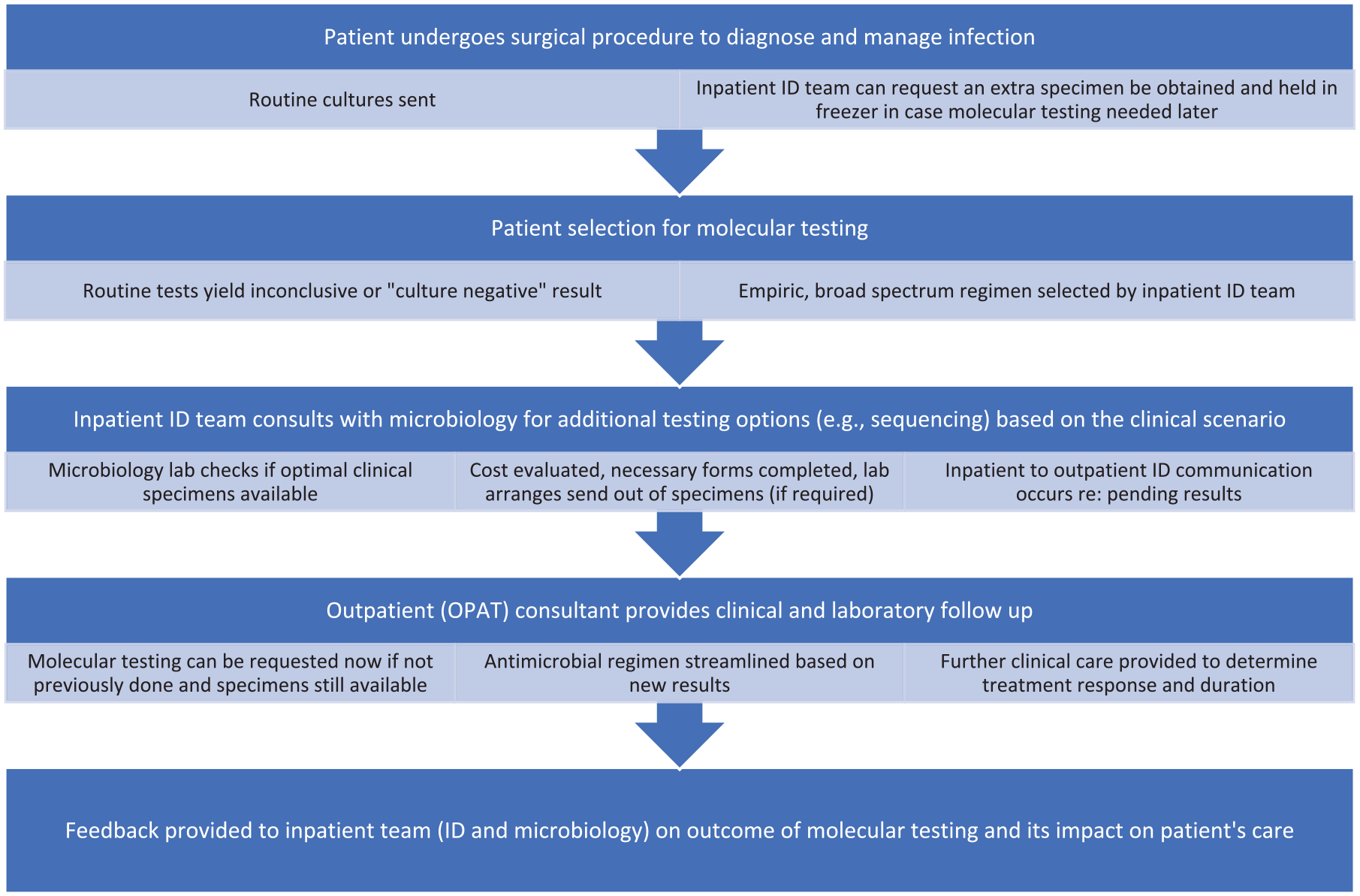
Proposed workflow for streamlined infectious diseases-microbiology collaboration.
Case 1: Methicillin-susceptible Staphylococcus aureus bacteremia and vertebral osteomyelitis, a classic OPAT scenario
A 68-year-old female with controlled type 2 diabetes mellitus, a body mass index of 35 kg/m2, and a history of L4-S1 spinal fusion for lumbar spinal stenosis 5 years prior presented to the emergency department with severe low back pain, fever, and tachycardia. She denied recent trauma or invasive procedures to the spine. Examination of her lower back revealed a healed surgical site without open wounds but tenderness to palpation of paraspinal muscles surrounding the surgical site. Two sets of blood cultures were collected, and she was initiated on empiric intravenous vancomycin and piperacillin/tazobactam. Within 36 h, blood cultures yielded Gram-positive cocci in clusters, most consistent with Staphylococci, and the treating team was notified. MRI revealed discitis/osteomyelitis and a spinal epidural abscess; broad-spectrum antibiotics were maintained. Microbial identification by MALDI-TOF (matrix-assisted laser desorption/ionization time-of-flight mass spectrometry, Bruker Sepsityper®) of the positive blood culture broth identified Staphylococcus aureus 4 h after the culture flagged positive. Results of antimicrobial susceptibility testing ultimately revealed methicillin-susceptible S. aureus (MSSA). The treating team was informed, the ID service consulted, and the patient de-escalated to IV nafcillin and oral rifampin on hospital day 3. The patient received immediate irrigation and debridement of her spinal hardware infection; an epidural abscess was evacuated, and the spinal hardware retained for stability. Blood cultures cleared after source control. A PICC line was placed, and the patient was discharged on hospital day 10 on long-term IV cefazolin and oral rifampin. She tolerated the regimen well, and her clinical symptoms and inflammatory markers had improved at completion of IV therapy.
MSSA bacteremia and vertebral osteomyelitis is a commonly encountered scenario by ID specialists regardless of the practice setting. It is well-established that ID consult involvement in severe infections like S. aureus bacteremia is lifesaving and ensures that patients receive appropriate diagnostic and therapeutic management. 5
Current best practice recommendations indicate that laboratories should adopt rapid diagnostic methods for microorganism identification from positive blood culture broths. 6 Rapid identification can be achieved using MALDI-TOF MS, either using a direct blood culture bottle testing strategy such as the MALDI-TOF Sepsityper® kit (as done here) or after rapid short incubation of high inoculum subcultures; these approaches expedite appropriate management of bacteremia and aid in detection of rare and unusual organisms. 6 Current best practice recommendations also advise that laboratories use rapid diagnostic methods for antibacterial resistance detection from positive blood culture broths. 6 Several methods for rapid assessment as to whether S. aureus is methicillin-resistant or not, directly from positive blood culture bottles, are available to facilitate timely escalation or de-escalation of antimicrobial therapy, as appropriate. 7 Rapid identification of microorganisms detected in blood and their associated antimicrobial susceptibilities facilitates appropriate antibiotic therapy especially when coupled with an electronic medical record alert or comment ‘nudge’ within the report pertaining to appropriate therapy and/or ID consultation to optimize management, which may include a biofilm-active agent such as rifampin for hardware-associated infections, such as this one. 8 Although the role of rifampin for staphylococcal hardware infections remains unsettled, a recent expert review continued to support its use.9,10
Case 2: Mycobacterial infection in an immunocompromised and pregnant host
A 30-year-old pregnant female at 13-weeks gestation with a history of indeterminate colitis, on infliximab therapy for 2 years, and latent tuberculosis treated with 4 months of rifampin prior to initiation of infliximab, presented with 4 months of a progressively painful and ulcerated mass of her left first proximal interphalangeal joint. She was born in Bangladesh and emigrated to New York 13 years ago, where she resides with her spouse and three children. She is a homemaker, and sometimes prepares meals requiring cleaning and removing scales of fish acquired at a local market. Physical examination of the left index finger revealed an exophytic lesion over the dorsum of the proximal interphalangeal joint measuring 3 cm × 2 cm with surrounding vesicles. Additional tender subcutaneous nodules were noted on the left forearm. Dermatopathology of punch biopsies reported pseudoepitheliomatous epidermal hyperplasia and markedly inflamed granulation tissue suggestive of infection. Periodic acid-Schiff-diastase and Gomori methenamine silver stains were negative as were bacterial and fungal cultures. Fite stain was negative as were initial mycobacterial cultures. Due to the patient’s clinical findings, compromised immune status and exposure history, the consulting ID team requested that the biopsy specimen undergo specialized send-out testing. Based on this request, the clinical microbiology laboratory coordinated advanced molecular testing for Bartonella species, Leishmania species, and non-tuberculous mycobacteria at a reference laboratory. A week later, Mycobacterium marinum was identified by direct PCR followed by sequencing of the hsp65 and rpoB genes. Dual therapy with azithromycin and ethambutol was started and the patient received periodic laboratory and clinical monitoring to ensure an optimal outcome (COpAT). At her 6-week ID follow-up, the subcutaneous lymphadenopathy had nearly resolved, and the finger lesion significantly improved. She was treated for two additional months beyond resolution of the finger lesion without recurrence of infection despite chronic infliximab therapy.
Routine microbiologic testing was negative for bacteria (including mycobacteria) and fungi, suggesting that the diagnosis could have been missed in the absence of a detailed exposure history and direct communication with the clinical microbiology laboratory to coordinate specialized testing for unusual pathogens. Upon diagnosis, selection of an optimal antimicrobial regimen (with consideration for the patient’s pregnancy status) and close clinical monitoring by an ID specialist led to cure of the patient’s infection.
Case 3: Organ transplant candidate with multiple pulmonary nodules
A 60-year-old woman with type 2 diabetes mellitus, end-stage renal disease, and a previous 30-pack-year smoking history, underwent computed tomography (CT) of the thorax for lung cancer screening prior to deceased donor renal transplantation, with the finding of innumerable bilateral pulmonary nodules suspicious for metastatic disease. A positron emission tomography scan revealed numerous F-fluorodeoxyglucose (FDG) avid pulmonary nodules (several measuring up to 1.1 cm) with additional non-avid (<1 cm) nodules as well as increased FDG activity in the right hilum. She underwent right middle and lower lobe wedge resection. Pathology was negative for malignancy but noted multiple necrotizing granulomas with yeast forms with weakly positive mucicarmine and Fontana-Masson stains morphologically suggestive of Cryptococcus species. Aerobic and anaerobic tissue cultures were negative as was serum cryptococcal antigen. Given the presumptive diagnosis of cryptococcal pneumonia, she was started on renally adjusted fluconazole induction therapy for 2 weeks followed by consolidation therapy with plans to complete 6–12 months per IDSA guidelines. 11
CT of the thorax at 4 and 7 months after initiation of fluconazole showed only slight improvement. Given suboptimal radiographic response, lack of growth from tissue culture, and negative cryptococcal serum antigen, additional testing was pursued. Serum Aspergillus galactomannan, serum (1–3)-β-D-glucan, Coccidioides serology, urine Histoplasma antigen, and repeat serum cryptococcal antigen were negative. Broad-range fungal PCR that targets the 18S ribosomal RNA gene, internal transcribed spacer (ITS)1 and ITS2, followed by Cryptococcus-specific gene amplification and sequencing, confirmed the presence of Cryptococcus neoformans var. grubii. All other fungal PCRs were negative.
The patient remained on fluconazole with plans to complete 12 months of therapy, and received periodic laboratory and clinical monitoring to ensure an optimal outcome (COpAT). Her persistent pulmonary nodules were thought to represent scarred granulomas and sequelae of prior infection rather than an ongoing active fungal disease. Despite a weakly positive mucicarmine stain, the lack of growth on routine fungal cultures and negative cryptococcal antigen prompted the ID specialists to pursue advanced molecular testing to establish a definitive diagnosis. Confirmatory PCR testing helped avoid unnecessary broadening of antifungals to more costly or toxic agents, such as posaconazole or amphotericin. Diagnostic confirmation also limited the duration of treatment to 12 months, rather than a more prolonged or potentially indefinite course of antifungal therapy.
Case 4: ‘Culture negative’ periprosthetic joint infection
A 70-year-old male presented to the emergency department with history of recurrent periprosthetic joint infection (PJI). He had a history of controlled diabetes, and hypertension, and had undergone left total knee arthroplasty complicated by PIJ with methicillin-resistant S. lugdunensis managed with explant of the prosthesis, placement of a vancomycin/tobramycin spacer, and 6-week of IV vancomycin at an outside hospital. He had delayed wound healing concerning for ongoing infection and was therefore referred to our medical center for repeat irrigation and debridement and replacement of the vancomycin/tobramycin spacer. Tissue cultures showed Escherichia coli (resistant only to ampicillin and ampicillin/sulbactam). The consulting ID team recommended a 6-week course of high-dose IV ceftriaxone. The patient clinically improved and there was an appropriate decline in inflammatory markers at the end of treatment. Subsequent synovial fluid cultures aspirated off antibiotics were negative. He then received repeat total knee arthroplasty; all tissue cultures were negative. Unfortunately, 4 months later, the patient developed a recurrence of pain and swelling of the left knee with rising inflammatory markers. X-ray of knee suggested loosening of the prosthesis. He was taken to the operating room for debridement of a purulent effusion, removal of a loose femoral component, polyethylene liner exchange, and placement of antibiotic-impregnated beads. Tissue cultures were all negative although pathology showed numerous neutrophils suggestive of infection.
The patient was initially treated with IV vancomycin and piperacillin/tazobactam but developed acute kidney injury which extended his hospitalization by a week. Due to lack of microbiologic diagnosis by conventional techniques, synovial tissue was sent for broad-range PCR and next-generation sequencing (NGS) for bacteria, fungi, or acid-fast bacilli (AFB) from the University of Washington Molecular Microbiology Reference Laboratory. Results revealed low levels of Streptococcus anginosus, Haemophilus influenzae, Stenotrophomonas maltophilia, and Candida albicans. High-dose oral daily fluconazole was added to the patient’s regimen of high-dose daily ceftriaxone. The patient completed 8 weeks of IV ceftriaxone/oral fluconazole therapy and then transitioned to chronic oral antibiotic suppression; was doing well 6-month later.
This case stresses the importance of close monitoring to assess treatment response and to determine the clinical significance of detected microorganisms. Moreover, feedback to the inpatient ID consultants and microbiology laboratory on the patient’s clinical progress is key to improving our collective understanding of the optimal role of specialized testing (Figure 1).
PJI, like IE, is a common OPAT diagnosis, with 5–35% of cases characterized as culture negative due to antecedent antimicrobial therapy, inadequate use of diagnostics, or presence of fastidious organisms. 12 Improved culture methods, such as sonication to increase yield from biofilms on implant surfaces, and tissue cultures in blood culture bottles, have improved microbiologic yield, but some cases remain microbiologically negative. 13 Multiplex PCR assays utilize predefined targets, so not all pathogens are detected. 14 NGS is increasingly used for PJI diagnosis, but its clinical utility as an up-front diagnostic is not well-defined. Shotgun metagenomic next-generation sequencing (mNGS) can identify pathogens not detected by culture but is complex and costly. 16S ribosomal RNA gene-based targeted metagenomic next-generation sequencing (tNGS) has similar performance as mNGS of sonicate fluid for microbial detection in culture negative PJIs with advantages of turnaround time, cost, and ease of data analysis. 12
In this case, advanced molecular testing revealed multiple microorganisms, including C. albicans, which led to addition of an antifungal agent; multiple bacterial species of unclear significance were also detected (e.g. bacterium of the phylum Bacteroidota). Sequencing techniques are highly sensitive,14,15 and microbial DNA present at low concentrations is easily detected. Microbial DNA can arise from pathogens in the specimen or introduced during specimen collection (from the patient’s normal microbiota, normal microbiota of healthcare providers collecting the specimen, devices and containers used for specimen collection, the normal microbiome of the laboratory or personnel performing testing in the laboratory, reagents, etc.). Distinguishing contaminants from true pathogens can pose a challenge in some circumstances. In this case, coverage of all listed microorganisms detected would have been difficult and may have predisposed the patient to recurrent acute kidney injury due to multiple concurrent antimicrobials. Careful interpretation of results of microbiologic testing, whether culture-based or molecular, is needed as microorganisms detected can be contaminants or true pathogens. The clinical significance of detected microorganisms can be ascertained by assessing the amount of microorganism detected, and whether it is detected in more than one specimen. Direct communication between the clinical team and the testing laboratory can be helpful.
Conclusion
Close partnership between ID specialists and the clinical microbiology laboratory can meaningfully improve the care of OPAT patients by expediting time to appropriate antimicrobial therapy, facilitating early de-escalation to the most effective and targeted long-term oral or IV regimen, and in some cases, escalation of therapy to cover pathogens previously unknown. This partnership facilitates pursuit of advanced testing, including sequencing-based testing, when routine testing does not yield a microbiologic diagnosis or the patient is labeled as having a ‘culture negative’ infection, often resulting in treatment with a more broader spectrum regimen. However, no test is perfect; test interpretation should ideally occur in conjunction with the doctoral-level microbiologist performing the advanced testing if clinical significance of the associated results is unclear or if testing yields unexpected results. Confirmatory testing may include performance of an organism-specific PCR assay or serologic testing. ID specialists are encouraged to reach out to the microbiologist performing the test and vice versa.
Herein, we have provided several examples of collaborative efforts between ID specialists and the clinical microbiology laboratory to enhance both diagnostic and antimicrobial stewardship of complex infections requiring long-term antimicrobial therapy with the goal of optimizing patient outcomes. Effective collaboration between ID and clinical microbiology can result in ‘precision medicine’ for the OPAT patient population.
