Abstract
Purpose:
To create a model predictive of an individual’s risk of developing a de novo multidrug-resistant (MDR) infection while in the intensive care unit (ICU).
Methods:
This is a case-control study in which 189 ICU patients diagnosed with their first infection with an MDR organism were compared on the basis of demographic, past medical and clinical variables to randomly selected ICU patients without such an infection, era-matched in a 2:1 ratio. A prediction tool was derived using multivariate logistic regression.
Results:
Five features remained predictive of developing an infection with a drug-resistant pathogen: hospitalization within a year [adjusted odds ratio (OR) 2.14], chronic hemodialysis (3.86), underlying oxygen-dependent pulmonary disease (1.86), endotracheal intubation within 24 h (2.46) and reason for ICU admission (respiratory failure 2.89, non-respiratory failure, non-shock presentation 1.85). Using a scoring system (0–7 points) based on the adjusted OR, risk categories were derived (low: 0–2 points, intermediate: 3–4 points and high risk: 5–7 points). The negative predictive value at a score cutoff of 2 is excellent (88.9%).
Conclusions:
A clinical prediction rule comprised of five easily measured ICU variables reasonably discriminates between patients who will develop their first MDR infection versus those who will not.
Introduction
In the medical intensive care unit (MICU), infections with multidrug-resistant (MDR) bacteria are increasingly commonplace1–3 and associated with substantial morbidity4–6 and cost. 7 The first methicillin-resistant Staphylococcus aureus (MRSA) and extended-spectrum beta-lactamases (ESBL) Klebsiella isolates were identified in 1960 8 and 1983, 9 respectively; they now represent over 50% 10 and up to 20% 11 of infections with those organisms. Infection with the drug-resistant variants of these organisms is associated with increased mortality, not only due to a patient’s burden of chronic illness, but difficulty with eradication 12 and toxicity of required antimicrobials 13 as well. Additionally, some drug-resistant pathogens may be more virulent 14 and can potentially present with different patterns of illness. 15
Unfortunately, ICU patients with the clinical syndrome of sepsis often do not have a pathogen identified for several days, if at all. 16 Traditional culture techniques have no impact on the choice of empiric antimicrobials, which are based on the suspected site of infection, presumed risk of harboring an MDR organism, local antibiograms and often, severity of illness. Culture data presently only serve to alter antimicrobials after a substantial part of the clinical course has passed. Even if the identified organism was empirically treated, if MDR and the patient’s first to be documented, healthcare workers may have been entering and exiting the room with hand hygiene only instead of the recommended gown and glove contact isolation precautions. This potentially places neighboring patients at risk for nosocomial transmission – known to be commonplace in the ICU.17,18
This highlights the unmet need for methods of earlier organism detection. Although polymerase chain reaction (PCR)-based techniques to identify particular organisms or resistance genes are conceptually attractive, most are experimental19,20 with unclear performance characteristics21,22 and impact patient outcomes will require further study. Accordingly, none of these novel techniques are included in consensus guideline approaches of organism detection.23,24 An adjunct prediction tool describing the likelihood of harboring an MDR organism based fully on patient characteristics rather than microbiologic techniques may supplement decision making for cases in which culture data are pending, negative, or equivocal. Traditional risk prediction models have classified individuals into dichotomous ‘community’ or ‘hospital’ risk profiles, for the most part focusing on demographics and medical history 25 largely ignoring clinical presenting features. Newer models have begun to highlight the importance of the patient’s clinical course.23,26 The importance of such considerations is magnified as drug resistance is increasingly reported in community-derived infections. 27
Therefore, we attempt to identify surrogates for the development of infections with MDR bacteria, identifiable early upon admission to the ICU and inclusive not only of demographics and past medical history, but also features of the present clinical course. In addition, rather than dichotomizing risk we sought to create risk categories, to better serve the clinician in practice.
Methods
Design
This is a single center, retrospective case-control study of patients admitted to the Medstar Georgetown University Hospital MICU over a 6-year period from 2008 to 2014. We leveraged information kept by hospital epidemiology in order to identify patients diagnosed with an infection requiring implementation of contact isolation precautions during their stay. The potential causative organisms include MDR Gram-negative rods (as defined by the CDC) along with clinical MRSA and vancomycin-resistant enterococcus (VRE) infections. It is not our practice to routinely swab for MRSA and VRE on ICU admission; however, patients were included for a positive result of a MRSA nares or VRE rectal swab if it was performed in order to assist with the choice of antimicrobial therapy for infection at a distant site. Excluded were those with prior or pre-existing MDR infections on ICU arrival.
The control group, a concurrent population that did not develop an MDR infection, was selected randomly among the remaining ICU patients in a 2:1 ratio. We chose to frequency match this group by year in order to account for natural changes in the prevalence of MDR organisms over time. 28 Conversely, we felt it appropriate to not match based on additional criteria as the recorded variables (listed below) were all thought of as unique, potentially clinically relevant exposures. Instead, we attempted to address confounding with a relatively large sample and subsequent multivariate analysis. The study was approved by the Institutional Review Board of Medstar Georgetown University Hospital.
Data
Recorded were variables potentially associated with the development of an MDR bacterial infection based on clinical relevance and prior studies.29,30 The three categories this encompassed were demographic information (age, gender, residence in a healthcare facility prior to hospitalization), past medical history (hospitalization within a year, surgery within a year, diabetes mellitus, cirrhosis, long-term oxygen use irrespective of the specific pulmonary diagnosis, New York Heart Association (NYHA) class III or higher congestive heart failure (CHF), hemodialysis, cancer diagnosis, immunosuppression, indwelling central line, indwelling foley) and clinical variables measured during the first 24 h of the patient’s ICU course [reason for ICU admission, endotracheal intubation, central venous or arterial catheterization, severity of illness, hospital length of stay (LOS) prior to ICU admission, duration of antibiotics prior to culture positivity].
Indwelling central lines were defined as peripherally inserted central catheters, chemotherapy ports, and hemodialysis lines placed prior to the hospitalization. A patient was considered immunosuppressed if they carried a diagnosis of AIDS, were concurrently receiving chemotherapy, or had an organ transplant in the past. The reason for ICU admission was the predominant presenting feature (chosen among respiratory failure, circulatory collapse, post-cardiac arrest, encephalopathy, or other) as recorded by the admitting intensivist. The severity of illness was measured with the Acute Physiology And Chronic Health Evaluation (APACHE II) score. We gathered data from medical record review: physician notes, summaries of recent hospital stays, nursing flowsheets along with laboratory, culture and radiographic data.
Analysis
Summary statistics describe the frequency and mean of each categorical and continuous variable, respectively. Unadjusted odds ratios (OR), 95% confidence intervals (CI) and p-values using the Wald test were derived from the coefficients of a univariate logistic regression model, with the development of a drug-resistant bacterial infection as the outcome. Variables with an unadjusted association with the outcome were subsequently included as candidates in stepwise multivariate logistic regression model construction. In the process of model creation, we attempted to balance accuracy with complexity, penalizing the inclusion of additional terms if not balanced by substantially improved predictive power. A receiver operating characteristic (ROC) curve was constructed to describe overall model discrimination via the area under the curve or c-statistic.
In order to simplify risk prediction for patient level application and facilitate description of test characteristics, we developed a clinical prediction rule. 31 To do so, we weighted variables based on their adjusted OR to generate a ‘score’ attributable to each trait. Subsequently, we scored the entire cohort, to describe test accuracy and generate clinically relevant score ‘cutoffs’. The analysis was performed with SAS 9.4 (SAS Institute, Cary, NC, USA).
Results
Patients
We identified 249 patients with a new contact isolation requirement during the study period. Of those, 60 were excluded because of a prior MDR infection on careful chart review or incomplete information in the medical record. The characteristics of those 189 patients with MDR infections and the 378 randomly selected era-matched control patients are shown in Table 1. The ‘reason for ICU admission’ is a categorical variable for which unadjusted OR are derived via comparison to circulatory collapse. Overall, the MDR group was more ill (APACHE II score 23.3 ± 8.4 versus 19.3 ± 8.1), more likely to be admitted to the ICU for respiratory failure [OR 3.1 (95% CI: 1.98–4.81)], and be intubated within 24 h of arrival to the ICU [OR 2.47 (95% CI: 1.73–3.53)].
Characteristics of ICU patients who developed an MDR bacterial infection versus control patients.
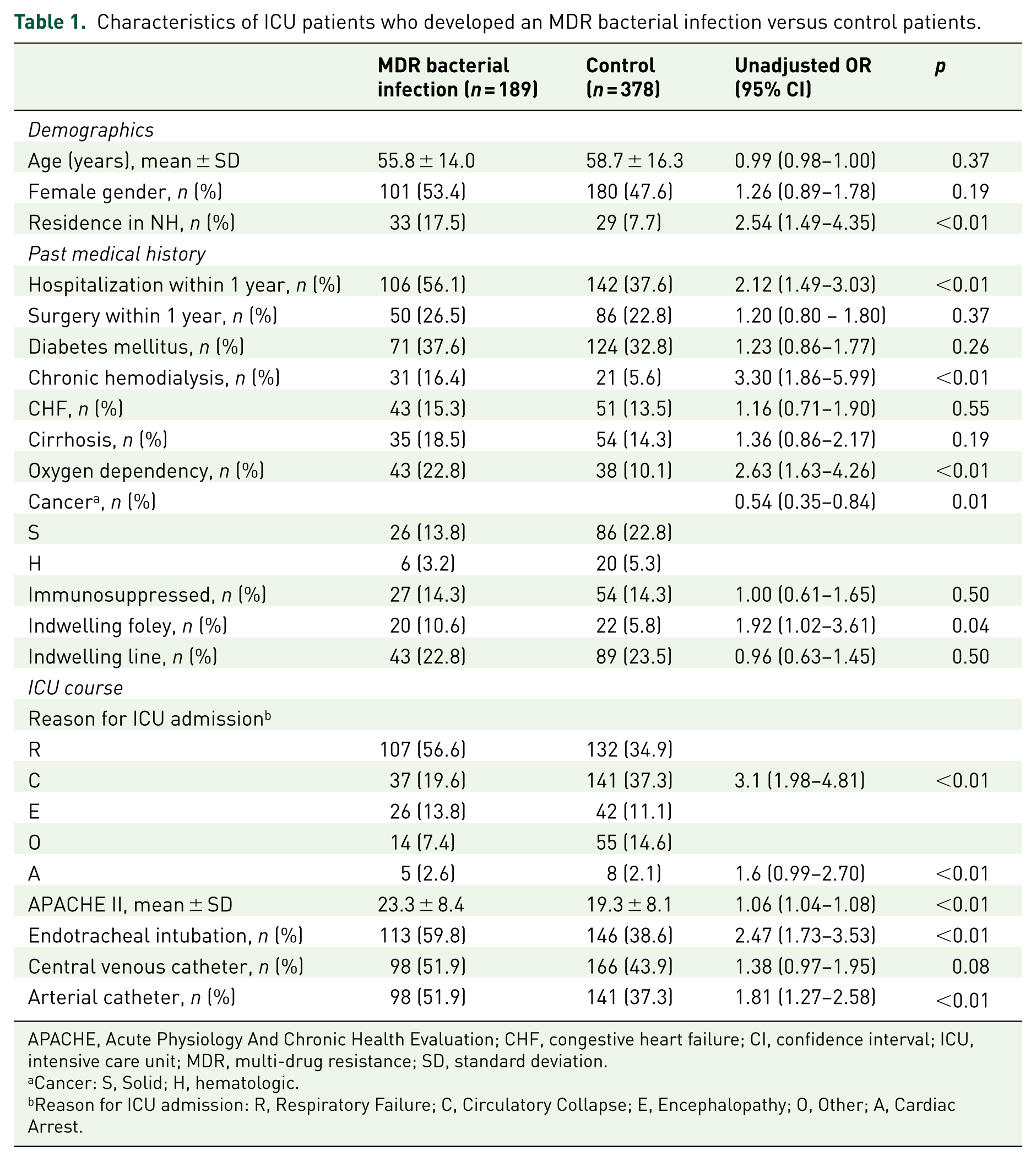
APACHE, Acute Physiology And Chronic Health Evaluation; CHF, congestive heart failure; CI, confidence interval; ICU, intensive care unit; MDR, multi-drug resistance; SD, standard deviation.
Cancer: S, Solid; H, hematologic.
Reason for ICU admission: R, Respiratory Failure; C, Circulatory Collapse; E, Encephalopathy; O, Other; A, Cardiac Arrest.
Pathogens
Table 2 shows the identified organisms and frequency. Most cultures were respiratory (51%, from sputum, endotracheal tube aspirates, or bronchoalveolar lavage), followed by urinary (19%) then blood (11%). A smaller number were skin and soft tissue infections (8%), and the remainder were swab cultures used to help guide antimicrobial therapy for infection at a distant site.
Causative organisms.

Methicillin-resistant Staphylococcus aureus
Vancomycin-resistant enterococcus.
Modeling
The 10 patient characteristics that differed between the MDR and control groups were introduced into stepwise multivariate logistic regression modeling. Five remained predictive, shown in Table 3 with adjusted OR. To describe overall model discriminatory ability, we constructed an ROC curve and calculated the c-statistic: 0.73. The APACHE II score also retained predictive ability after adjusted analysis; however, it minimally improved model performance (c-statistic improved to .76). Given its cumbersome nature to calculate, we formulated our model without its inclusion.
Adjusted OR.

CI, confidence interval; ICU, intensive care unit; OR, odds ratio.
A scoring system with point values derived from adjusted OR resulted in all variables being valued at one point except for chronic hemodialysis [OR 3.86 (95% CI: 2.04–7.29)] and admission to the ICU for respiratory failure [OR 2.89 (95% CI: 1.78–4.70)], each scored at two points.
Clinical prediction rule
The entire cohort of 567 patients was then ‘scored’ with possible values ranging from 0 to 7. Subsequently, we examined test characteristics at each score cutoff. The sensitivity, specificity and positive and negative predictive values are shown in Table 4 at each score cutoff. The low-, intermediate- and high-risk groups were developed based on the maximal (or near-maximal) positive and negative predictive values. Of the studied patients, 187 were in the low-risk category, 262 were intermediate, and 117 were high risk.
Test characteristics at each score.

NPV, negative predictive value; PPV, positive predictive value.
At a score cutoff of 2, the negative predictive value is 88.9%, whereas at a score cutoff of 5 the positive predictive value is 71.4%. The model, therefore, more reliably rules out those who have a very low potential for developing an MDR infection than it does predicting who is at very high risk of developing one.
Discussion
MDR organisms are acquired via antibiotic selection pressure 32 or preexisting colonization due to residency in a high-risk environment. 33 They are disturbingly commonplace in the ICU 2 and patients without a history of MDR bacteria acquire them with some regularity. Among a group that is MDR naive, we have demonstrated that five easily measured characteristics readily apparent upon ICU admission reasonably discriminates between patients who will and will not develop their first MDR infection.
Our approach of combining MDR Gram-negative rods together with MRSA and VRE is unusual, but one we find to be potentially relevant. Several prior studies have studied MRSA18,34 or MDR Gram-negative organisms30,35,36 separately. The shared quality of infections with those organisms, however, is drug resistance and potential for nosocomial transmission. Additionally, the clinical syndrome of sepsis often occurs without a clear source of infection with substantial overlap in presentation between those categories of pathogens, and empiric antimicrobial strategies generally target both Gram-positive and Gram-negative organisms. Therefore, we elected to study one’s risk of having either.
While some of the variables in our model are known risk factors for harboring MDR organisms, others are less well described – namely home oxygen use. We chose to record this because of an anecdotally observed association at our institution. One proposed mechanism for ventilator-associated pneumonia is bacterial colonization within a biofilm of the endotracheal tube and ventilator tubing; 37 similarly, we speculate that home oxygen tubing and humidifiers may harbor bacteria. There is scant data regarding the sterility of home oxygen equipment, some of which suggests that it may not be sterile.38,39 Requiring oxygen therapy certainly indicates more severe underlying cardiopulmonary illness, and, such patients are more likely exposed to antimicrobials as well – potentially leading to colonization of equipment with drug-resistant pathogens. Current oxygen prescribing guidelines are derived from older literature in patients with COPD40,41 and widely applied toward patients with other cardiopulmonary diagnoses. Given recent data suggesting the non-utility of oxygen therapy in patients with COPD 42 and our demonstrated association of oxygen use with MDR infections, perhaps the medical community should carefully re-evaluate the indications for home oxygen and investigate equipment sterility.
The major strength of this study is the inclusion of all patients over a several year period in a busy, tertiary care MICU with new drug-resistant infections, with only clerical exclusionary criteria. Additionally, we measured many characteristics per patient including a thorough description of a patient’s demographics, past medical history, and present clinical course (including vital signs, laboratory data, and acute organ dysfunction as reflected by the APACHE II score). The variables in the final model are plausibly biologically related to the development of MDR infections. Additionally, the potential value of a clinical decision rule over a list of risk factors is the ability to assign a predictive ‘weight’ to each variable. 31 We hope this may be helpful in reclassifying ICU patients toward or away from an MDR risk profile. The implications are severalfold, including the decision to institute gown and glove contact isolation precautions along with candidacy for targeted de-escalation of antimicrobial therapy.
Presently available data regarding the utility of contact isolation precautions to prevent nosocomial infection transmission are conflicting.43–45 As a result, guidelines are incongruous46,47 and practice patterns vary widely. 48 A potential utility of our findings is to identify a group of patients who are high risk for developing MDR organisms and selectively isolate, rather than including all-comers. Aiming this preventative intervention at those most likely to be implicated in nosocomial transmission of organisms may increase its chance of success.
Antibiotic de-escalation based on culture data is a cornerstone of antimicrobial stewardship programs, evidence based 49 and guideline recommended, 50 although poorly adhered to. 51 There are many reasons for this, mistrust of culture data and fear of changing a working regimen among them. Identifying patients at low risk for developing MDR infections via our model may assist clinicians in decisions regarding de-escalation.
Our study has several limitations. Hospital LOS is a known predictor for the acquisition of drug-resistant hospital flora. Unfortunately, there are a large number of patients in our tertiary-care ICU transferred from other institutions for whom the LOS from the original hospital could not easily be determined. Instead of including a censored variable in the model, we chose to eliminate it. Duration and choice of recent antimicrobial therapy is a substantial risk factor for acquiring drug-resistant organisms as well; 52 however, they are more easily measured in the ‘MDR’ group. Quantifying duration in the control sample would have been imprecise and comparing values between groups would have been difficult. We explored measuring empiric carbapenem and antifungal therapy prior to ICU arrival, but both occurred infrequently enough to not be a practical addition. Additionally, we have found in our clinical practice that obtaining such information is not always feasible, especially when the patient is transferred from another institution. Much of what we are measuring may simply represent surrogates for antecedent antibiotic use; if so, we find them to be more easily captured.
Malignancy was more often found in the control group, contrary to our expectation. There are several possible reasons for this observation. First, it may simply be an artifact of multiple comparisons. It may also be related to other characteristics of our group with cancer, namely that it was younger overall and many patients were admitted for sepsis due to drug-related neutropenia soon after chemotherapy initiation, often having less substantial comorbid illness or pre-existing healthcare exposure. This is speculative, however.
We do not have a validation sample. Though similar studies have arbitrarily split patients into derivation and validation groups, we felt that there wasn’t a meaningful enough distinction between subsets of our cohort to do so. Additionally, a model derived from a larger cohort of patients may benefit from improved validity; however, in order to be used clinically, it will have to be validated in a separate ICU population. This is especially important in models derived from non-random case-control populations like ours, in which true disease prevalence is not reflected, potentially leading to overestimation of model performance. 53
Finally, application of these results, once validated, will require an assumption about the similarity of patients with positive cultures and without 16 since the model is derived from those with positive cultures. Any epidemiologic study of bacterial infections is subject to the same consideration regarding external validity.
Conclusion
In patients who are MDR infection-naive, five easily observed characteristics (ICU admitting diagnosis, oxygen dependency, hemodialysis, intubation within 24 h and hospitalization within one year) distinguishes those who will and will not develop their first MDR infection in the ICU, with implications for antimicrobial therapy and empiric contact isolation precautions. The model is best suited to identify a low-risk group given its excellent negative predictive value (89%), and will require validation prior to clinical use.
Footnotes
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Georgetown University Department of Medicine Pilot Grant: Risk Factors for Acquiring Multi-Drug Resistant Infections within the Intensive Care Unit (PI: Daniel Jamieson).
Conflict of interest statement
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
