Abstract
Integrating real-world healthcare data is challenging due to diverse formats and terminologies, making standardization resource-intensive. While Common Data Models (CDMs) facilitate interoperability, they often cause information loss, exhibit semantic inconsistencies, and are labor-intensive to implement and update. We explore how generative artificial intelligence (GenAI), especially large language models (LLMs), could make CDMs obsolete in quantitative healthcare data analysis by interpreting natural language queries and generating code, enabling direct interaction with raw data. Knowledge graphs (KGs) standardize relationships and semantics across heterogeneous data, preserving integrity. This perspective review proposes a fourth generation of distributed data network analysis, building on previous generations categorized by their approach to data standardization and utilization. It emphasizes the potential of GenAI to overcome the limitations CDMs with GenAI-enabled access, KGs, and automatic code generation. A data commons may further enhance this capability, and KGs may well be needed to enable effective GenAI. Addressing privacy, security, and governance is critical; any new method must ensure protections comparable to CDM-based models. Our approach would aim to enable efficient, real-time analyses across diverse datasets and enhance patient safety. We recommend prioritizing research to assess how GenAI can transform quantitative healthcare data analysis by overcoming current limitations.
Plain language summary
This perspective review explores whether Artificial Intelligence (AI) can revolutionize healthcare data analysis by reducing the current reliance on Common Data Models (CDMs), which encompass the following elements:
• CDMs are approaches that standardize diverse healthcare to a single shared format to enable efficiencies in data management and analyses using the same analysis syntax and analytic tools.
• Although CDMs have strengths, they also have limitations, such as high costs, potential loss of important details, significant effort to produce and maintain, and delays in data availability due to lengthy data processing steps.
• With the rapid growth of healthcare data, effectively analyzing it is crucial for patient safety and public health.
• AI may offer an alternative solution by analyzing data directly in its original form, reducing costs, preserving data details, and enabling real-time insights that support better patient outcomes and safer medication use.
This review investigates challenges currently associated with CDMs and explores how AI, particularly generative AI, can directly analyze raw data without the need for standardization. We discuss the following:
• How AI can interpret complex questions and generate accurate answers from raw data, enabling more timely analyses of real-world data.
• While CDMs may still be necessary in the short term, AI has the potential to eventually replace them, improving patient care and safety outcomes by providing faster and more precise insights.
• This perspective could lead to new methods of using healthcare data to inform decision-making and enhance treatment outcomes.
• By adopting advanced AI technologies, healthcare providers and researchers can better understand treatment risks and benefits, make more informed decisions, and ultimately improve patient safety and public health.
Introduction
This manuscript represents a perspective review, synthesizing insights from literature and expert consensus to evaluate the evolving role of Common Data Models (CDMs) in distributed analytics.
Over the past several decades, electronic healthcare databases have evolved from simple medical record repositories into sophisticated tools for complex epidemiological research, informing regulatory, clinical, and policy decisions. 1 Alongside randomized controlled trials, registries, and spontaneous reports, they are pivotal for understanding and ensuring medication safety during drug development and clinical use.
These transformations can largely be categorized into generational developments that we have defined as follows (Figure 1). This generational framework is based on the authors’ synthesis of advancements in healthcare data analysis and their implications for real-world evidence generation. The first generation (initiated circa 1970) featured proprietary systems relying on single-database analyses without standardized formats.2 –4 The second generation (initiated circa 1991) introduced widely used coding standards and enterprise databases, 5 but data sharing involved individual contracts and bespoke cross-database comparisons or substantial computational resources for meta-analyses across multiple proprietary sources.6,7
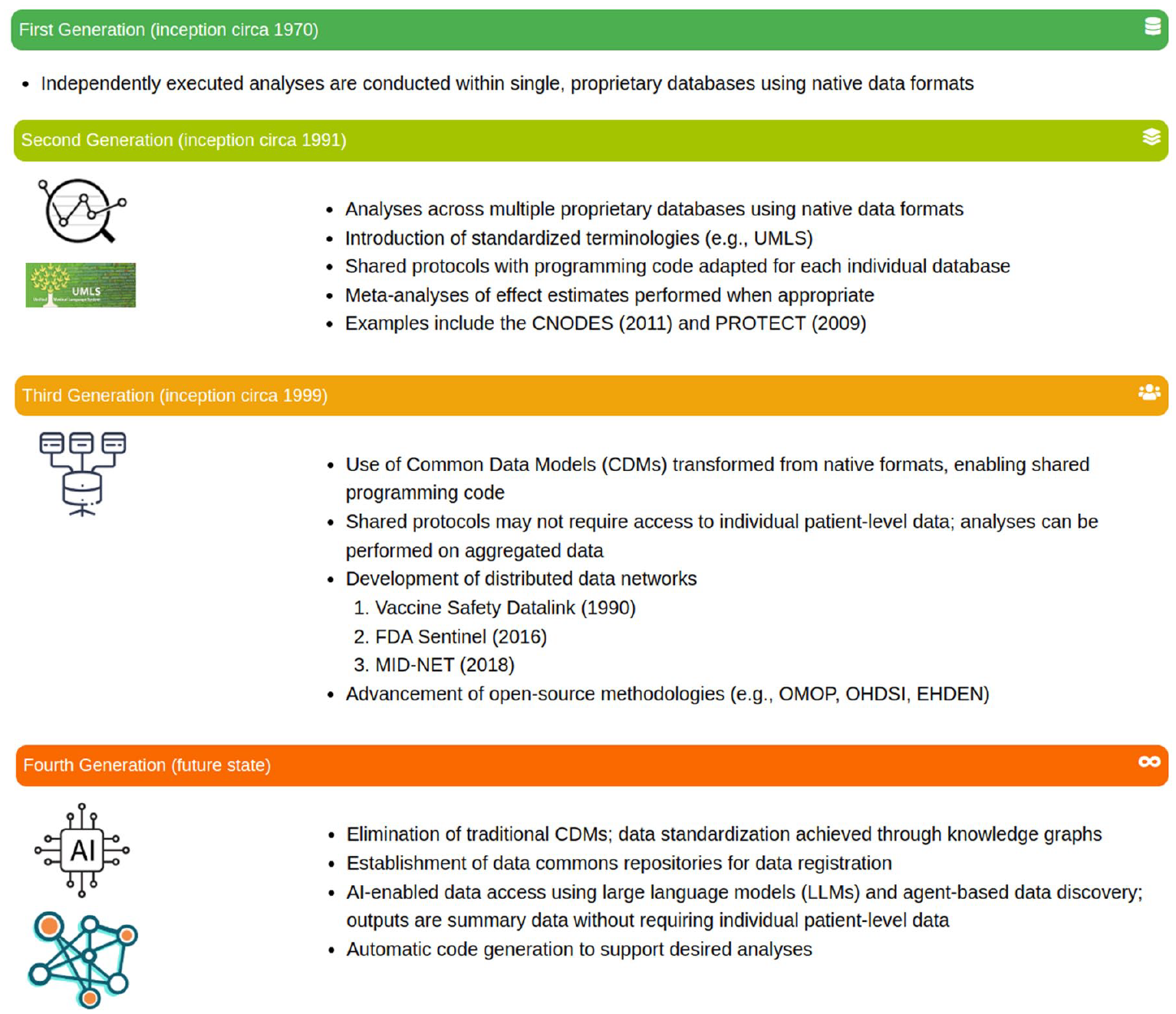
Four generations of quantitative analysis in healthcare data.
The third generation, starting in the late 1990s and gaining momentum after the FDA Amendments Act of 2007 (https://www.fda.gov/regulatory-information/selected-amendments-fdc-act/food-and-drug-administration-amendments-act-fdaaa-2007) and the launch of Mini-Sentinel in 2009, 8 is characterized by the adoption of CDMs. 9 Early CDMs like the Vaccine Safety Datalink’s (VSD’s)10,11 demonstrated the feasibility of CDMs. While the Observational Medical Outcomes Partnership’s (OMOP’s) CDM (https://www.ohdsi.org/data-standardization/) popularized distributed data networks (DDNs) and open-source methodologies, greatly improving interoperability and streamlining analyses across varied databases.5,8,12,13 Once a database was converted to a given CDM external standardized analyses could be conducted with the agreement of the database holder across the database with the need to only share summarized results. Similarly, standardized tools and programs developed for the CDM could be readily used by the database holder on their own data.
However, despite their significant contributions, CDMs are not without limitations, particularly when scaling analyses across increasingly complex and heterogeneous data sources. The advent of generative artificial intelligence (GenAI) and knowledge graphs (KGs) heralds a potential fourth generation in electronic healthcare data utilization. These technologies promise to integrate, analyze, and interpret multiple datasets—whether for distributed analyses, multi-site studies, or other complex use cases—without requiring a single standardized format. KGs are an evolving research tool and will enable increasingly effective use of GenAI in enabling analyses across different data formats.
This paper explores the relevance of CDMs considering these emerging innovations, examining whether GenAI can address current challenges and render traditional CDMs less central to future healthcare data analyses.
Background
The role of CDMs
History and development of DDNs and CDM utilization
The development of DDNs has evolved through two primary approaches: the common protocol approach and the CDM approach. However, sometimes a hybrid of the two approaches was used where a study-specific CDM was implemented (e.g., earlier studies in the Asian Pharmacoepidemiology Network (AsPEN)). Early networks, such as the Canadian Network for Observational Drug Effect Studies (CNODES; https://www.cnodes.ca/), and the Pharmacoepidemiological Research on Outcomes of Therapeutics by a European Consortium (PROTECT; https://www.imi.europa.eu/projects-results/project-factsheets/protect), adopted the common protocol approach. This method allowed each participating site to retain data in its original format while following standardized study protocols and centrally developed code.14,15 While this approach enabled data custodians to maintain control and minimized the transformation workload, it also introduced significant challenges in data standardization. Harmonizing data across sites proved time-intensive, increasing the complexity of cross-database analyses and limiting scalability, particularly in scenarios requiring rapid responses. 16
To address these limitations, the CDM approach was introduced to standardize data structures across multiple sites. By transforming data into a uniform format before analysis, CDMs aimed to enhance comparability and efficiency in multi-database research. However, the adoption of CDMs also presented trade-offs, including resource-intensive data transformation processes, potential information loss, and challenges in achieving consistency across different CDMs. 17 These trade-offs highlight the ongoing need for innovative solutions to bridge gaps in interoperability and efficiency within DDNs.
Global perspectives on CDM usage
Internationally, various federated DDNs use CDMs to enhance health-related data analyses. An early example is the VSD project,10,11 which established a CDM to enable secure, federated querying across multiple health maintenance organizations. This model facilitated vaccine safety surveillance while addressing concerns about data privacy and confidentiality. Additional examples include the Medical Information Database Network (MID-NET), 18 the AsPEN (https://www.aspensig.asia/), TriNetX, 19 and CNODES. 14
Several other global networks have adopted CDMs to standardize health data integration. Networks like the OMOP and the Observational Health Data Sciences and Informatics (OHDSI; https://www.ohdsi.org/) program promote a highly structured CDM to support cross-network interoperability. 20 Others, such as the Sentinel Initiative (https://www.sentinelinitiative.org/), NorPEN (https://www.norpen.org/), IMI ConcePTION (https://www.imi-conception.eu/), PCORnet (https://pcornet.org/), and the DARWIN-EU (https://www.darwin-eu.org), European Health Data and Evidence Network (EHDEN; https://www.ehden.eu/), employ different CDMs tailored to specific regulatory and research needs. 21 Meanwhile, initiatives like DARWIN-EU facilitate extensive data integration across diverse European sources using the OMOP CDM.
Despite the advantages of standardization, concerns remain regarding information loss when mapping diverse datasets to a standardized model. These concerns drive ongoing refinement of CDMs to balance standardization with the need for preserving the granularity and semantic integrity of source data.22,23 Alternative approaches, such as the Generalized Data Model (GDM), have been proposed to maintain data in their native formats while leveraging clinical codes and hierarchical structures. 22 However, while these alternatives mitigate information loss, they also introduce complexities in data analysis and may extend the timeline required to generate insights.
Challenges in CDM implementation
The adoption and implementation of CDMs present several challenges, as identified through literature reviews and expert discussions in DDN analysis. The growing size and diversity of data networks intensify the challenges of converting data to a standard CDM, compounded by the constantly changing healthcare landscape. 24 Integrating heterogeneous data sources complicates the creation of universally applicable CDMs essential for drug safety surveillance. 25 Ensuring semantic consistency is crucial to avoid misinterpretations in drug safety analysis.
Kent et al. highlight the difficulties in standardizing data across healthcare systems, where variability in data collection methods can compromise analysis reliability. 26 Overcoming technological, methodological, regulatory, and ethical challenges adds complexity to CDM implementation. Analysts also struggle with accessing data through various vendor tools and employing different strategies, which complicates analyses and prolongs insight generation. 27 In addition, evolving CDMs and their ecosystems can lead to discrepancies in outputs, further complicating comparisons across different models.3,28,29 Other CDM implementation challenges, whether using the pragmatic or generic approach, are defined as follows:
Information loss and data semantics: Transitioning data into a CDM can lead to substantial information loss, especially if the CDM does not perfectly represent the source data.30 –34 Garza et al. emphasized that unless a CDM perfectly represents the source data, information loss will occur, potentially altering the original data semantics. 35 This issue is pronounced with registries, which are heterogeneous in data collection and structure. 36
Resource intensity of data transformation: Transforming and maintaining data in a CDM is resource-intensive, often deterring organizations with limited resources. 22 Frequent updates and ensuring compatibility across different CDM versions add to the complexity and workload. 34
Variability in data sources: Standardization is challenging due to variability in healthcare delivery and data capture across regions and time, risking the loss of critical data nuances. 22 Unclear assumptions during data conversion can lead to overconfident decisions based on flawed interpretations, underscoring the need for transparent documentation and critical evaluation of the transformation process.
Diverse CDMs and analytical tools: Variations in CDMs and their associated analytical ecosystems can yield differing results, complicating decision-making. 28 Careful selection of a particular model and its tools is essential, as different models may impact the generated evidence.
These challenges have prompted the exploration of alternative approaches to data integration and standardization. One such alternative is the GDM proposed by Danese et al.. 22 The GDM focuses on retaining the original semantic representation using clinical codes in their native vocabularies and preserving hierarchical information and provenance. By avoiding transformation into a standardized CDM, the GDM aims to reduce information loss and maintain data integrity.
However, while the GDM addresses some challenges of traditional CDMs, it introduces its own limitations. Analyses using the GDM can be time-consuming, taking longer from ideation to execution. 22 This extended timeline may impact the timeliness of research findings, especially in fast-moving fields where rapid insights are critical. The complexity of working with heterogeneous data in native formats may require more sophisticated analytical tools and expertise, potentially limiting accessibility for some organizations.
Challenges with terminologies and ontologies in real-world data networks
In real-world data (RWD) networks, integrating and analyzing data from diverse sources is often complicated by variations in medical terminologies and ontologies. Common terminologies used historically in healthcare include coding systems such as the International Classification of Diseases (e.g., ICD-9 and ICD-10), Systematized Nomenclature of Medicine Clinical Terms (SNOMED CT), and Logical Observation Identifiers Names and Codes (LOINC). Many other terminologies are employed based on specific subject areas, such as lab values (e.g., LOINC for laboratory tests) and billing codes (e.g., Current Procedural Terminology). These terminologies are essential for coding diagnoses, procedures, and other clinical information within electronic health and medical records. 37 However, the diversity of these coding systems necessitates complex mapping and cross-referencing to achieve consistent data integration, posing significant challenges for data harmonization and interoperability in RWD networks.
Variability of terminologies across systems: Different healthcare systems and regions may use different coding systems or versions of the same system. For instance, while some systems use ICD-9-CM, others have transitioned to ICD-10-CM, and still others may use SNOMED CT or Read codes. 38 This variability poses significant challenges for data integration and interoperability within RWD networks.
Challenges in mapping between terminologies: Cross-mapping between different coding systems is necessary for integrating data from multiple sources. However, these mappings can be complex and may not capture the full semantic relationships between codes, leading to potential information loss or misinterpretation. 39 For example, mapping from ICD codes to SNOMED CT or Medical Dictionary for Regulatory Activities (MedDRA) may not always be straightforward due to differences in granularity and coding structures. 40
MedDRA and its integration with RWD terminologies: In drug safety surveillance, the MedDRA is the standard terminology used for coding adverse events. 41 Integrating MedDRA-coded data with RWD, which often uses ICD codes or other terminologies, requires accurate and reliable mapping. Inconsistencies or inaccuracies in these mappings can affect the identification and evaluation of safety signals. 42
Versioning and updates: Frequent updates and version changes in coding systems add another layer of complexity. Ensuring that mappings remain accurate over time requires continuous maintenance and updates, which can be resource-intensive. 43
Leveraging AI and KGs: AI and KGs offer promising solutions to these challenges. AI techniques can automate the mapping between different terminologies, enhancing accuracy and reducing manual effort. 44 KGs can represent complex relationships between medical concepts across different coding systems, facilitating interoperability and integration. 45
As we anticipate an increasing need for more multimodal quantitative surveillance, the ability to work rapidly and effectively across diverse data streams becomes ever more important. Therefore, standards applied to the collection and reporting of Individual Case Safety Reports and MedDRA, along with their links to RWD terminologies like ICD codes, are essential for ensuring consistency and reliability in data integration and analysis. 46
By addressing these challenges through the adoption of AI-driven solutions and adherence to standardized terminologies, RWD networks can improve data interoperability and enhance the quality of healthcare analytics, ultimately contributing to better patient outcomes and more informed public health decisions.
Additional challenges faced in multi-database analyses
Multi-database analyses leverage data from various sources, offering significant benefits for rapid-cycle analyses and safety signal detection in healthcare. However, their widespread adoption faces challenges for data analysts, custodians, and stakeholders who require rapid, actionable, and trustworthy outputs to impact public health and patient care.
The continual search for fit-for-purpose RWD
Identifying ideal RWD sources that accurately represent target populations is inherently challenging. Each database captures only a subset of the population under specific healthcare settings, introducing biases related to demographics, geography, and healthcare access. Data must be fit for specific study purposes and sufficiently recent to avoid misleading insights about current practices and outcomes. 47 Researchers must select data sources closely aligned with their objectives, carefully weighing strengths and limitations to minimize bias.
For data custodians, maintaining databases in a CDM or multiple CDMs poses a significant workload, especially if the CDM does not align with their core database model. Adopting a CDM that does not perfectly map to the primary data use may require compromises on data utility or adopting a simpler CDM closer to specific databases. This challenge explains why new CDMs continue to be proposed despite long-standing models like Sentinel and OMOP .22,48
Ensuring trustworthiness requires considering how extensively databases and their conversions need to be documented and preserved for future reanalysis. For data providers, extensive documentation of transformations can be burdensome. While replication—achieving consistent results using the same methods on similar data—can suffice, 49 stakeholders may demand reproducibility to ensure validity, 50 requiring detailed documentation and preservation, adding to custodians’ workload.
Trustworthiness also depends on producing rapid, actionable outputs impacting public health and patient care. Data custodians must balance timely data updates with maintaining CDMs, which may not align with their primary operations. This tension underscores the importance of developing more adaptable data models or alternative approaches that reduce providers’ workload while ensuring data remains fit-for-purpose.
Privacy concerns and data sharing complexities
Privacy regulations, such as the General Data Protection Regulation 51 in the European Union and the Health Insurance Portability and Accountability Act 52 in the United States, impose stringent controls on how personal health information (PHI) is collected, stored, and shared. While protecting patient confidentiality, these regulations complicate data sharing, often causing delays and increased costs. 53 Data custodians must invest substantial resources to ensure compliance, limiting their ability to share data promptly.
Logistical complexities in data sharing, especially internationally, are considerable. Differing regulations and standards across countries complicate the harmonization of data collection and analysis procedures,15,54 necessitating sophisticated data governance structures. For stakeholders requiring rapid and trustworthy outputs, these delays can hinder timely decision-making for public health and patient care.
Commercial concerns and data format limitations further restrict data availability, as highlighted by Walker et al. 55 Privacy concerns particularly affect access to PHI, such as unstructured clinical notes, limiting the depth of possible analysis.
Decentralization versus centralization of data
To balance comprehensive analysis with privacy concerns, initiatives like the FDA’s Sentinel network maintain decentralized patient-level data while centralizing summary data for analysis. 53 This model permits detailed analyses locally, sharing only aggregated results centrally. For data custodians, this allows control over sensitive data, ensuring compliance and preserving primary database purposes.
However, decentralization can restrict the depth and speed of analysis stakeholders desire. Sensitive data remains at the source, potentially slowing the generation of rapid, actionable insights needed for public health decisions and patient care. Stakeholders must rely on custodians’ capacity and willingness to perform analyses promptly and accurately. Furthermore, clear governance frameworks and secure access protocols are essential to ensure sensitive data are only used and shared in compliance with regulatory and ethical standards.
Technical challenges in data aggregation and synthesis
Combining data from multiple databases poses significant technical challenges. Variations in data capture methodologies, variable definitions, coding systems, and missing data impact the quality and comparability of the aggregated dataset. Researchers must utilize sophisticated harmonization techniques—often requiring complex algorithms and substantial computational resources—to ensure the combined data accurately reflects underlying realities.
For data custodians, the workload to keep a database updated in a CDM or different CDMs is considerable, especially if these models are not integral to their primary operations. Adopting a CDM not perfectly aligned with primary data use can necessitate compromises or additional resources. This is why new CDMs continue to be proposed, aiming for closer alignment with specific datasets. 48
For stakeholders seeking rapid and trustworthy outputs, technical challenges in data aggregation can cause delays and affect reliability. Ensuring data transformations and analyses are well-documented and reproducible enhances trust but adds complexity and time to deliver actionable insights.
Despite these challenges, data custodians recognize two primary benefits of converting to CDMs: (1) the ability to use tools developed within an ecosystem designed for a given CDM and (2) the capability to readily participate in multiple database studies based on that common structure. However, we believe that in an AI-enabled future, these benefits may be more easily attainable without relying on traditional CDMs. Advanced AI techniques could facilitate data integration and analysis across heterogeneous databases, reducing custodians’ burden and accelerating the delivery of actionable insights to stakeholders.
Discussion
Having examined the evolution of the CDM and its current role in PV leads us to wonder what comes next. As we alluded to in the introduction, we believe there is a strong role to play for GenAI and KGs to enable the fourth generation of safety surveillance. Below, we will explore the building blocks necessary to move us into the fourth generation, as well as explore potential challenges and considerations the PV community must face as we move into the next phase of technology to assist in PV-related activities.
Building blocks of the fourth generation enabled safety surveillance
We propose a fourth generation of AI-enabled data analysis to simplify multi-database analysis and ease data availability for custodians. Key elements may include the following: (1) establishing data commons for data registration, (2) utilizing AI-enabled data access via agent-based discovery, (3) standardizing data through KGs, (4) supporting automatic code generation for analyses, and (5) managing operational aspects like master service agreements to protect personally identifiable information, facilitate reanalyses, and ease access while respecting data localization laws and collaboration requirements. This approach could reduce the need to transform native data into CDMs, enabling efficient, real-time analysis across diverse datasets. To realize this potential, we recommend prioritizing research that assesses how GenAI can accelerate the effective use of electronic health data to enhance patient safety.
The evolution of drug safety surveillance into its fourth generation requires a paradigm shift toward more dynamic and interconnected data ecosystems. Central to this shift is the ability to seamlessly integrate and analyze data from diverse sources—whether in-house or external. This integration first relies on the concept of data “discoverability,” ensuring that data systems are easily findable and comprehensively described by metadata that details their contents and relevance to specific studies. Notably, although ontologies for RWD metadata are important to enable this paradigm shift, the extent to which these metadata exist and are available is insufficient. Processes, potentially supported with GenAI tools, may support the development and make available ontologies for RWD metadata, as are financial support and a standardized approach to ensure metadata maintenance and updating. Recent frameworks like the DIVERSE framework and the MINERVA metadata list offer promising strategies for standardizing metadata across diverse data sources.56,57 Such tools are factors that may support interoperability and reduce bias, particularly in distributed analytics.
DDNs enhance statistical power by aggregating data from multiple sources, enabling robust analyses that can detect rare adverse events and improve generalizability across diverse populations and settings. 58
However, integrating varied data sources presents challenges, especially for pharmacovigilance (PV). Safety analyses often use data not optimized for population-level studies, leading to issues in data availability and technical barriers due to different formats and coding systems, which can introduce selection bias. 59 The Structured Process to Identify Fit-For-Purpose Data (SPIFD) framework addresses these challenges by systematically assessing data reliability and relevance to mitigate bias. 60
Data commons for data registration
Data commons provide shared platforms co-locating data, storage, and computing resources with common APIs and tools. 61 They offer centralized environments (e.g., cloud-based tools like Google Colab; https://colab.research.google.com/) where diverse datasets can be managed and analyzed using standardized methods. However, in healthcare, privacy concerns often prevent the open sharing of sensitive data. We envision a data commons model allowing controlled metadata sharing and supporting decentralized analysis, preserving data privacy and autonomy. 62 Standardizing metadata and providing secure tools can enhance data integration efficiency and facilitate robust research findings.
However, CDMs, as seen in initiatives like Sentinel, effectively protect privacy and maintain local data control, important for political and cultural acceptance. The data commons approach must ensure comparable data privacy and governance to be acceptable.
While integrating data commons could benefit the fourth-generation approach by facilitating data registration and discovery, they are not essential. Core objectives—leveraging AI for data access, standardizing data via KGs, and automating analyses—could be achieved without centralized data commons, provided robust interoperability and data governance mechanisms exist.
From centralized to decentralized study data management
Initially centralized, the data commons framework has demonstrated significant potential in improving data accessibility and utility across various fields. As highlighted by Guha et al., this framework enables the pooling of vast amounts of public data, making it accessible via standardized APIs. 63 While centralization has its benefits, drug safety is now transitioning toward a decentralized model where data can remain in its native environment but still be fully accessible and integrated into broader safety surveillance systems. Advancements in Cloud APIs and visualization tools have facilitated this shift, enabling real-time analysis and reporting of safety signals, which could reduce drug-related risks.
Looking ahead, integrating AI—including machine learning for predictive analytics and natural language processing for mining unstructured data (such as social media)—will further enhance the effectiveness of drug safety surveillance. This proactive approach not only addresses current surveillance needs but also helps predict and mitigate risks associated with new pharmaceuticals, ultimately improving patient safety on a global scale.
Architectural framework for a decentralized data commons
The proposed architectural framework for a decentralized data commons must support key capabilities, such as data authentication (verifying the publisher and trustworthiness of the data through mechanisms like digital signatures, like those used in open-source software), detailed metadata descriptions (including information about coding scheme versions and the language of the source), and alignment with KGs. This framework should enable data to be described in its native language, and when paired with a large language model (LLM), it allows for the interpretation and integration of diverse data sets without the need for conventional CDM conversions.
Potential challenges and considerations
Transitioning to a decentralized data management model introduces several challenges. These include the risk of data loss if a provider withdraws their data from the commons, the potential introduction of low-quality or malicious data by bad actors, and the complexities associated with ensuring continuous data integrity and security. Addressing these challenges requires robust mechanisms for data verification, quality control, and stakeholder cooperation to preserve the integrity and usefulness of the data commons.
Standardizing data with KGs in drug safety
The integration of GenAI with KGs represents a significant advancement in addressing key challenges in drug safety, including interoperability, data integration, and predictive analysis. KGs are structured representations that connect entities (e.g., drugs, diseases, clinical trials) through meaningful relationships, transforming complex, multi-dimensional data into actionable insights. By organizing data into interconnected networks, KGs potentially offer a framework for enhancing safety surveillance and enabling predictive, preventive, and personalized medicine. While the current literature on the application of KGs in pharmacovigilance remains sparse, with limited examples demonstrating their utility, this emerging area holds promise for advancing the field and addressing critical gaps in drug safety.
Integration across diverse datasets
An important challenge in drug safety is integrating heterogeneous RWD—including clinical, genomic, and chemical datasets—while adhering to FAIR (Findable, Accessible, Interoperable, and Reusable) principles. Currently, many analyses are conducted in silos, requiring significant levels of domain expertise to develop queries that can address specific drug safety questions.
The PheKnowLator project demonstrates the potential of KGs to integrate diverse data modalities, such as genomic, proteomic, clinical, and chemical information, into a unified and FAIR-compliant framework. 64 By building on examples like PheKnowLator, the combination of KGs and GenAI could automate the harmonization of RWD with other high-dimensional datasets. This synergy holds promise for enhancing routine pharmacovigilance activities—such as adverse event detection and monitoring—by enabling faster and more timely access to comprehensive and supporting evidence.
Predictive models using KGs
The PlaNet system by Brbić et al. exemplifies the combination of AI and KGs to address safety prediction challenges in clinical trials. 65 PlaNet uses two distinct KGs—one for clinical data and the other for biological and chemical relationships—to predict safety outcomes and adverse events more effectively. By leveraging predictive modeling, PlaNet demonstrates a framework with the potential to improve the accuracy of safety assessments, potentially offering a scalable solution for identifying potential risks earlier in the drug development process.
Validation and quality assurance
Despite these advances, ensuring the reliability of AI-driven KG systems remains a critical challenge. Quality assurance processes, such as benchmarking KG models against established safety datasets and validating predictive outputs, are essential to building trust in their applications. For example, frameworks that compare GenAI predictions to validated adverse event databases can help measure their accuracy, consistency, and reproducibility. Incorporating such validation mechanisms will be crucial as KGs and AI are further integrated into routine safety surveillance systems.
Toward real-time drug safety surveillance
The potential for KGs to enable real-time data querying, as demonstrated by LinkedIn’s use of KG technologies with sub-millisecond response times, 66 signals a potential, major shift in pharmacovigilance. Applying similar capabilities to drug safety surveillance could allow regulators and researchers to query complex datasets dynamically and efficiently. This is particularly relevant for responding to emerging safety signals in real time, improving decision-making for clinical and regulatory stakeholders.
Automatic code generation for PV
Innovative applications of GenAI
GenAI, particularly LLMs, shows promise in transforming healthcare data analysis. LLMs interpret natural language queries and generate coherent responses, bridging the gap between human inquiry and machine-readable data.67 –69 Techniques like retrieval-augmented generation (RAG) further enhance LLMs’ accuracy for data-validated responses.70,71 LLMs can automate complex data interactions, including generating SQL queries for epidemiological research.72,73
GenAI can transform federated RWD networks by enabling dynamic evolution of data structures and optimizing queries, reducing the operational burden on data providers. Re-envisioning the extract, transform, and load (ETL) process as a business context document allows for real-time queries executed directly against raw data, bypassing predefined CDM transformations. 74 This approach could enhance the flexibility of data pipelines.
Such advancements allow researchers to ask complex questions across RWD databases, whether in native formats or CDMs. GenAI facilitates comparisons of analyses performed on native versus CDM-transformed data, improving the scope and accuracy of real-world evidence.
For example, assessing a pharmaceutical product’s adverse events globally currently requires access to diverse data formats. A GenAI-driven interface could query these sources worldwide, including databases in the United States, EU, Japan, China, and more. Assuming data privacy and governance issues are resolved, researchers could estimate and compare incidence rates across formats and regions, enhancing global result comparability.
These developments suggest a future where GenAI simplifies data handling and broadens analytical capabilities. By offering rapid insights into PV and patient safety, GenAI could extend tools developed for CDM ecosystems across various RWD sources, enhancing overall evidence generation.
Enhancing data interoperability with AI and KGs
GenAI and KGs offer promising avenues for data interoperability without relying solely on traditional CDMs. AI-enabled, agent-based data discovery can facilitate real-time analysis across diverse datasets while preserving privacy. 61 KGs standardize data by capturing relationships inherent in heterogeneous sources, reducing information loss and maintaining integrity. 75
Data commons repositories for data registration can streamline sharing by providing standardized metadata, 62 and automatic code generation tools can enhance code portability and replicability. 76 Brat et al. argue that LLMs’ ability to interpret diverse data formats may reduce the reliance on traditional data models, prompting a reevaluation of data interoperability strategies in healthcare. 77
Collaboration among data custodians, analysts, regulators, and developers is essential to create governance frameworks that balance accessibility with privacy and ethical considerations, moving toward more efficient use of RWD and improving patient outcomes.
Emerging techniques like federated learning and secure multiparty computation provide frameworks for privacy-preserving data analysis, helping ensure sensitive data remains secure while enabling collaborative research.
Challenges and future directions
Current challenges in CDM implementation
The growing size and diversity of data networks intensify the challenges of converting data to a standard CDM, compounded by the constantly changing healthcare landscape. 24 Integrating heterogeneous data sources complicates the creation of universally applicable CDMs essential for drug safety surveillance. 25 Ensuring semantic consistency is crucial to avoid misinterpretations in drug safety analysis.
Kent et al. highlight the difficulties in standardizing data across healthcare systems, where variability in data collection methods can compromise analysis reliability. 26 Overcoming technological, methodological, regulatory, and ethical challenges adds complexity to CDM implementation. Analysts also struggle with accessing data through various vendor tools and employing different strategies, which complicates analyses and prolongs insight generation. 27 In addition, evolving CDMs and their ecosystems can lead to discrepancies in outputs, further complicating comparisons across different models.28,29
Future directions
Addressing these challenges requires innovative approaches that leverage emerging technologies.
Enhancing ontological consistency with AI and KGs in biomedical data
Addressing the complexities of DDNs requires careful consideration of the terminologies and ontologies that underpin the description and categorization of data within these systems. Traditional management of medical terminologies often involves cross-walk maps created by CDMs, but these solutions have inherent limitations, particularly in areas like drug safety, where data must be meticulously accurate, current, and contextually meaningful.
Li et al. introduced FHIR-generative pre-trained transformer (GPT), an innovative approach that combines LLMs with the Fast Healthcare Interoperability Resources (FHIR) standard to enhance health data interoperability. 78 By leveraging LLMs’ natural language understanding capabilities, FHIR-GPT can effectively bridge different data formats and terminologies, enabling more seamless data exchange and integration across healthcare systems. 79
The integration of GenAI and KGs presents new opportunities to overcome these limitations. These technologies offer more dynamic and contextually aware methods for managing the evolution of medical terminologies, ultimately improving the consistency and utility of biomedical data across different systems.
The role of generative AI in enhancing ontological consistency
GenAI has the potential to fundamentally improve the management of biomedical terminologies by dynamically harmonizing mismatched terminologies across DDNs. This technology can leverage its ability to analyze vast amounts of structured and unstructured data to identify and resolve inconsistencies in real time.
The reason GenAI holds such promise in this domain is its capacity to generate contextually appropriate mappings that go beyond the rigid structures of traditional crosswalks. Using AI to understand the underlying semantics and relationships between different medical codes, it can provide more accurate and flexible mappings between systems like MedDRA and ICD. This could streamline the integration of data from disparate sources, allowing for faster, more accurate data analysis without the risk of misclassification or oversimplification.
In the observational research landscape, where the quality and reliability of RWD are paramount, GenAI can help standardize the ontological frameworks that underpin these data. This standardization can improve the interoperability of healthcare data systems, ensuring that patient outcomes, safety signals, and treatment efficacy data are consistently represented across multiple platforms and coding standards. Ultimately, this reduces the potential for errors in drug safety surveillance and accelerates the ability to respond to new medical challenges.
GenAI helps to create a dynamic ecosystem where medical terminologies can evolve in sync with the rapid pace of biomedical research, ensuring that healthcare systems are better equipped to handle future pandemics, emerging diseases, and evolving medical knowledge.
Generative AI and the future of observational research
The advent of GenAI holds significant potential to revolutionize observational research by enhancing the ETL process and enabling more dynamic data analyses. This technology facilitates direct querying of diverse RWD databases, significantly streamlining the research process by reducing the need for extensive data preparation, which is crucial for applications like effective signal detection. 80
GenAI, particularly the use of LLMs, is rapidly advancing in automating and supporting complex computer coding tasks. Recent studies have demonstrated that LLMs when trained with user guides can effectively provide textual responses for coding rules necessary for data entry into databases.74,81 This suggests a future where LLMs could be trained on specific database structures or data model architectures, enabling them to convert and adapt code in real time for any targeted database system.
These capabilities would dramatically increase the efficiency of database analyses, allowing for optimal coding that minimizes information loss during data conversion processes. This advancement could eliminate the labor-intensive and repetitive tasks associated with data conversion and updates in CDMs or native database structures. Furthermore, this approach would facilitate the effective reuse of code across different DDNs, making it easier to apply proven analytical tools across various data systems without significant reconfiguration.
GenAI tools have the potential to support other aspects necessary for the implementation of observational studies. For example, GenAI can assist with defining, evaluating, and monitoring the consistency of phenotypes over time, as well as summarize information from unstructured observational data for phenotype validation. In addition, the interpretation of study diagnostics to inform study design choices and ultimately choose a design based on a framework that enables causal inference 82 may be supported with GenAI.
Integrating GenAI into observational research will necessitate rigorous quality assurance processes to ensure data integrity and reliability. Leveraging existing frameworks, such as those developed by the OHDSI for quality-assuring CDMs, could provide a foundation for developing AI-specific quality assurance standards. In addition, applying comprehensive quality management systems that are crucial in PV will be essential to build trust and verify the accuracy of AI-generated insights in real-world applications.
By addressing these challenges and ensuring robust validation protocols, GenAI can significantly lower the barriers to efficient and effective observational research, paving the way for more timely insights into patient outcomes and drug safety.
Conclusion
While CDMs have a current role in record linkage and certain analytical use cases, the advent of AI-driven methods promises a transformative shift in distributed analytics, potentially rendering traditional models less central.
The expansion of electronic healthcare databases and DDNs offers unprecedented opportunities for health data analysis. These tools can significantly improve the identification of drug-related risks and benefits, empowering healthcare professionals to make informed decisions that enhance patient safety. As these systems evolve, they will shape the future of healthcare by leveraging insights from routine clinical data.
Integrating advanced technologies like GenAI and KGs could transform the management and analysis of RWD. Although CDMs are unlikely to be replaced immediately, these technologies indicate a future where CDMs may become less central. KGs can facilitate the analysis of diverse RWD datasets in their native formats, addressing challenges associated with data standardization, privacy, and multi-database analyses.
Tackling ontological challenges from various coding systems is crucial. KGs, supported by GenAI and human-in-the-loop validation, could enable multimodal analyses of RWD alongside other data sources like clinical trials and adverse event reports. This harmonization can lead to more comprehensive and accurate analyses.
GenAI has the potential to revolutionize interactions between data providers and researchers by dynamically managing data structures in federated and decentralized networks. This reduces operational burdens, while KGs provide a robust framework for integrating diverse biomedical data, allowing for deeper insights into drug safety surveillance and personalized medicine.83,84
As we adopt these emerging technologies, it is crucial to do so responsibly. 85 Fostering collaboration among data scientists, clinicians, and regulatory bodies, along with ethical AI use, will help maximize the benefits of big data for patient outcomes and public health. Advanced sandboxing techniques and non-PHI summaries enable analysis without direct patient-level data access, enhancing privacy protections within this new framework.
Despite the long-standing use of CDMs, ongoing discussions in the literature emphasize that no single model offers a perfect solution. A key question is not just how many data sources have been converted into a CDM, but how many remain unavailable and why. Evaluating the value of CDMs and exploring alternative approaches remain central to the discourse, as demonstrated by recent work like that of Tsai et al. 86
Complexities in accessing multiple data sources—such as the need to use different vendor interfaces for datasets with unique structures—highlight the limitations of current approaches. 27 These challenges could be mitigated if AI-driven solutions allow analysts to focus directly on the data itself, without being constrained by specific data models or interfaces. By enabling a data model-agnostic focus, AI has the potential to simplify and enhance the analytical process across diverse data environments.
In conclusion, the convergence of GenAI and KGs represents a promising frontier in the evolution of drug safety surveillance. By embracing these technologies responsibly and collaboratively, we can address current limitations, unlock deeper insights from complex datasets, and ultimately improve patient outcomes and public health.
