Abstract
Background:
Young adults with diabetes in Asia represent a heterogeneous group. Using traditional clinical criteria to preselect individuals for testing for maturity-onset diabetes of the young (MODY) may exclude a large proportion from testing. High-sensitivity C-reactive protein (hs-CRP) has shown promise as a biomarker to differentiate hepatic nuclear factor 1 alpha (HNF1A)-MODY from type 2 diabetes. We aimed to compare the use of hs-CRP as a biomarker versus traditional criteria, to guide testing for HNF1A-MODY among a cohort of young adults with diabetes in Singapore.
Methods:
A total of 252 adults (age of onset ⩽45 years) and 20 children with diabetes were recruited. Using traditional criteria (family history of diabetes and onset of diabetes ⩽25 years) and an hs-CRP cut off of ⩽0.5 mg/l, 125 and 37 adults, respectively, were identified for HNF1A gene testing. All children underwent HNF1A gene testing.
Results:
Five adults (5/143, 3.5%) with HNF1A-MODY were identified. There were no HNF1A gene mutations among the children. Traditional criteria correctly identified all five HNF1A-MODY individuals (5/125, 4%), while applying an hs-CRP level of ⩽0.5 mg/l selected just 1 of these 5 for HNF1A gene testing (1/37, 2.7%). None of those with a positive GAD antibody or undetectable C-peptide level had HNF1A-MODY.
Conclusion:
The use of hs-CRP to guide screening for HNF1A-MODY among Asian young adults with diabetes did not improve the diagnostic yield. Applying a combination of age of onset of diabetes under 25 years and a family history of diabetes alone could guide targeted HNF1A-MODY screening in Asians, with an expected yield of 4% diagnosed with HNF1A-MODY among those screened.
Introduction
Maturity-onset diabetes of the young (MODY) is an important subtype of young-adult-onset diabetes. An accurate diagnosis of MODY has implications for treatment, allowing the selection of the best treatment modality targeted towards the gene defect. In hepatic nuclear factor 1 alpha (HNF1A)-MODY, the correct diagnosis allows a switch from insulin (first-line treatment in type 1 diabetes) or metformin (first-line treatment in type 2 diabetes) to sulphonylureas or meglitinides with sustained improved control. 1 Diagnosing MODY also has follow-on implications for the person’s family in terms of risk stratification and disease prognostication. 2
The prevalence of MODY is estimated at 1–2% of all diabetes among White populations, 3 with GCK-, HNF1A-, HNF4A- and HNF1B-MODY accounting for majority of the cases of MODY.4,5 The low prevalence, and cost and time involved in genetic testing renders whole-population screening nonfeasible and not cost effective. 6 The proportion of MODY among those with young-adult diabetes reported from China, Japan, Hong Kong and Korea are varied, at 5–8% for HNF1A-MODY, 2.5–4% for GCK-MODY, and 0–4% for HNF4A-MODY.7 –10 However, these studies performed targeted gene testing among highly preselected groups which will have a much higher likelihood of MODY. What appears consistent is that lowering the age of onset of diabetes as a selection criterion appears to increase the pick-up rates of MODY among those screened. 11
This seems to support a combination of family history and age of onset of diabetes ⩽25 years as the main selection criteria for genetic testing for MODY. 12 However, in European cohorts, reliance on these criteria alone would have excluded 50% of those with MODY from genetic testing.4,13 Among Asians, overlapping clinical phenotypes presents an additional tier of difficulty to distinguish between diabetes subtypes. The prevalence of glutamic acid decarboxylase antibodies (GADA) in Asian type 1 diabetes is much lower (41.5% versus 60–90%) compared with White populations; 14 insulinopenia in an Asian with negative GADA would not necessarily rule out type 1 diabetes. Asians develop type 2 diabetes at a lower body mass index (BMI) and at a younger age; 15 an Asian who is lean and young at the onset of diabetes is still more likely to have type 2 diabetes rather than MODY. Recruiting Asians for genetic testing of MODY appears particularly challenging.
In 2008, common variants in HNF1A were shown to be associated with alterations in high-sensitivity C-reactive protein (hs-CRP) levels in genome-wide association studies,16,17 suggesting a potential use for hs-CRP as a biomarker for HNF1A-MODY. hs-CRP levels were subsequently found to be significantly lower in HNF1A-MODY compared with other forms of diabetes, including autoimmune diabetes, type 2 diabetes, and glucokinase gene (GCK)-MODY.18,19 In a multicenter European study, 20 hs-CRP was found to discriminate between HNF1A-MODY and type 2 diabetes with high sensitivity and specificity. However, the use of hs-CRP as a biomarker to guide screening for HNF1A-MODY has not be assessed in unselected diabetes populations or other ethnicities.
In this study, we aimed to compare the use of hs-CRP as a biomarker with traditional criteria (combination of family history and age of onset of diabetes ⩽25 years) to guide testing for HNF1A-MODY among a cohort with young-adult-onset diabetes (diabetes onset ⩽45 years) in Singapore.
Methods
This study was conducted at Singapore General Hospital (SGH), and KK Women’s and Children’s Hospital (KKH), tertiary referral centers in Singapore, from August 2012 to October 2014. Ethics approval was granted by the institutional review board overseeing both hospitals. Informed consent was taken from all study participants or their parents. Consecutive persons with diabetes meeting the inclusion criteria who consented to take part in the study (n = 252) were recruited. Eligible consenting participants referred by their treating clinicians were also recruited. To enrich our target population for MODY, we recruited adults with younger-onset diabetes (⩽45 years), and children with diabetes onset under 18 years, with negative or unknown islet antibody status. These criteria were chosen in an effort to select individuals with young-onset non-type 1 diabetes, since beta cell autoantibody positivity is known to be an uncommon occurrence (<1%) in MODY. 21 While this group will inevitably include those with GADA-negative type 1 diabetes, we felt it was important to include these individuals in recruitment, given the heterogeneity of diabetes aetiologies and the overlapping phenotypes in Asians. 22 People with known type 1 diabetes and pregnant women were excluded.
A total of 252 adults and 20 children were recruited. Detailed medical history and anthropometric measurements were recorded. hs-CRP, glycosylated haemoglobin (HbA1c), random C-peptide, glucose, and lipid panel were measured. GADA was analysed in the core laboratory (IASP 2015 Lab number: 1501, Lee Kong Chian School of Medicine) by radioligand assay in the whole cohort. GADA positivity was determined as the 99.5th percentile of 1192 healthy German control participants (age range 18–70 years; mean age 39.7 years) and 145 healthy Singaporean control subjects (age range 20–69 years; mean age 49.1 years), and expressed as arbitrary NIDDK units (DK units/ml). More than or equal to21 DK units/ml for GADA was defined as positive.
Those with initial hs-CRP more than 10 mg/l had a repeat hs-CRP measured after 6 weeks to exclude the influence of recent infections. Those with persistently raised hs-CRP remained within the cohort, since they may qualify for gene analysis based on traditional criteria. hs-CRP was measured using a turbidimetric assay on a Beckman Coulter DxC800 analyser (lower limit of detection of 0.2 mg/l). HbA1c was measured using cation exchange high-performance liquid chromatography (Bio-Rad Laboratories, Hercules, California, USA) on a Variant II analyser (Diabetes Control and Complications Trial aligned), while C-peptide was measured using chemiluminescence assays (Siemens, Tarrytown, NY, USA) on an ADVIA Centaur XP analyser.
HNF1A genetic analysis was done at the SGH Department of Clinical Research, Genetics laboratory. Genomic DNA was extracted from peripheral leukocytes using standard procedures. All 10 coding exons of the HNF1A gene were amplified by polymerase chain reaction. Single-strand sequencing was then carried out in both forward and reverse directions using standard methods on an ABI3100 (Applied Biosystems, Waltham, Massachusetts). Sequences were compared using a published sequence (using NCBI Reference Sequence NG_0117131.2) to detect mutations. ClustalW2-Multiple Sequence Alignment software EMBL-EBI, Wellcome Genome Campus, Hinxton, Cambridgeshire, UK was used to carry out the DNA analysis.
HNF1A genetic analysis was conducted in the following groups:
(1) Adults with diabetes onset ⩽ 25 years and a family (parental) history of diabetes (traditional criteria group);
(2) Adults with hs-CRP level ⩽ 0.5 mg/l and diabetes onset ⩽ 45 years (hs-CRP criterion group);
(3) All children.
Family history was defined as the presence of diabetes among either parent. We chose an hs-CRP cut off of ⩽0.5 mg/l based on a multicentre European study that used different hs-CRP assays across several cohorts of HNF1A-MODY subjects and suggested that an upper limit threshold of 0.5 mg/l was sufficient to distinguish HNF1A-MODY from other subtypes of diabetes. 20
Variants that were previously reported as implicated in monogenic diabetes were assigned as pathogenic. The pathogenicity of novel missense mutations was determined by cosegregation of the mutation with diabetes, in comparison with the ExAC v3.0 and gnomAD databases, the correlation with phenotype and evidence from in silico analyses.
All statistical analyses were carried out using IBM SPSS Software version 23 (IBM, IBM Corp, Armonk, NY, USA). Data are presented as mean with standard deviations (mean ± SD), median with interquartile range (IQR) [Median (IQR)], or proportions (%). Independent t tests, Mann–Whitney U test and Z tests were used as appropriate. A two-sided p value of ⩽0.05 was considered significant. For the paediatric cohort, BMI Z scores were derived according to the age–sex-specific BMI centile charts of World Health Organization (WHO). 23
Results
A total of 272 (252 adults, 20 children) participants were recruited with an ethnic mix representative of the Singapore population. 24 A total of 1033 adult participants out of 5387 who attended our diabetes outpatient service during the recruitment period met the inclusion criteria. The recruited adult cohort were younger (34.7 years ± 10.1 versus 50 years ± 12.6, p < 0.01), had an earlier onset of diabetes (24.6 years ± 9.2 versus 35 years ± 8, p < 0.01) and were leaner (BMI 26.8 kg/m2 ± 4.8 versus 28.6 kg/m2 ± 5.3, p < 0.01) compared with the eligible cohort. A total of 20 children with diabetes were recruited from 75 eligible children with antibody-negative diabetes out of 400 children diagnosed less than 18 years of age. The recruited paediatric group were younger (15.4 years ± 1.8 versus 16 years ± 1.4, p = 0.02), had the same age of onset (11.9 years ± 2.5 versus 12.3 years ± 2.7, p > 0.05) and similar BMI Z scores [median (IQR):1.2 (0.2, 2.2) versus 1.5 (0.7, 2.4), p = 0.55] compared with the eligible cohort.
The majority of adults (87.7%) had a family (parental) history of diabetes, while a third had a history of diabetes in three generations (Table 1). The prevalence of GADA positivity was expectedly low (5.9%) since type 1 diabetes was an exclusion criterion. Consistent with this was a detectable C-peptide (mean C-peptide 1221 ± 740.2 pmol/l). A high proportion was overweight (69.9% had BMI >23 kg/m2), 25 suggesting that the majority had young-onset type 2 diabetes. Also, 49% (125/252) of the adult cohort met traditional criteria, while hs-CRP identified 14.6% (37/252) for HNF1A screening; 19 adults satisfied both criteria. The traditional criteria group had a younger age of diabetes onset, higher BMI and higher triglyceride levels compared with the hs-CRP criterion group (all p ⩽ 0.05, Table 1 and 2). There were no differences in the hs-CRP levels across the three ethnic groups.
Baseline characteristics of adult and paediatric cohorts and subgroups.
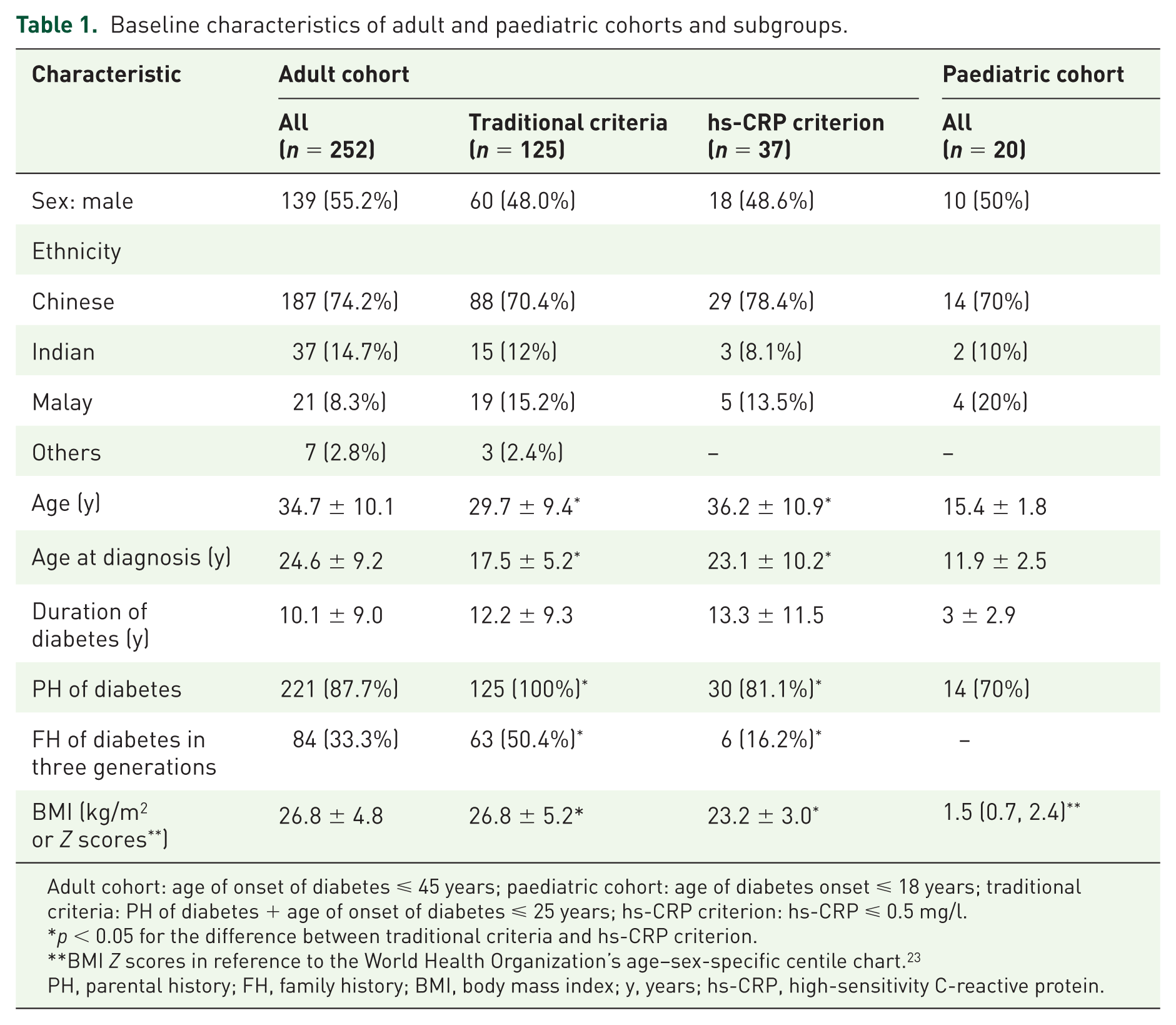
Adult cohort: age of onset of diabetes ⩽ 45 years; paediatric cohort: age of diabetes onset ⩽ 18 years; traditional criteria: PH of diabetes + age of onset of diabetes ⩽ 25 years; hs-CRP criterion: hs-CRP ⩽ 0.5 mg/l.
p < 0.05 for the difference between traditional criteria and hs-CRP criterion.
BMI Z scores in reference to the World Health Organization’s age–sex-specific centile chart. 23
PH, parental history; FH, family history; BMI, body mass index; y, years; hs-CRP, high-sensitivity C-reactive protein.
Baseline biochemical parameters.
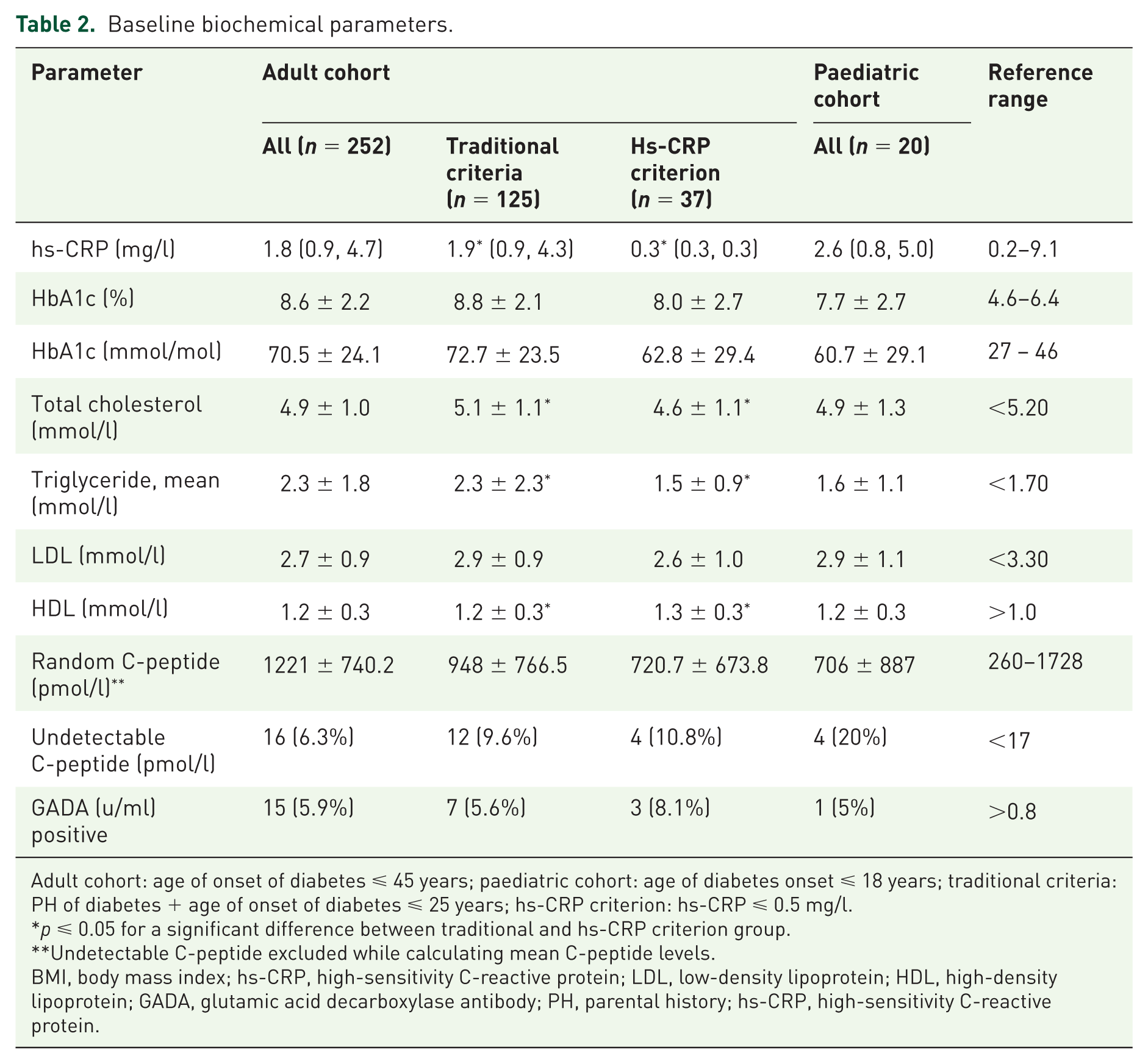
Adult cohort: age of onset of diabetes ⩽ 45 years; paediatric cohort: age of diabetes onset ⩽ 18 years; traditional criteria: PH of diabetes + age of onset of diabetes ⩽ 25 years; hs-CRP criterion: hs-CRP ⩽ 0.5 mg/l.
p ⩽ 0.05 for a significant difference between traditional and hs-CRP criterion group.
Undetectable C-peptide excluded while calculating mean C-peptide levels.
BMI, body mass index; hs-CRP, high-sensitivity C-reactive protein; LDL, low-density lipoprotein; HDL, high-density lipoprotein; GADA, glutamic acid decarboxylase antibody; PH, parental history; hs-CRP, high-sensitivity C-reactive protein.
Only two children had hs-CRP levels less than 0.5 mg/l. BMI of child participants was categorized according to the Singapore Health Promotion Board’s age-specific BMI centile chart. 26 A BMI <3rd centile was classified as severely underweight, 3rd to <5th centile as underweight, 5th to <90th centile as acceptable weight, 90th to <97th centile as overweight and > 97th centile as obese. Among the 20 children recruited for the study, 13 (65%) had acceptable weight, 3 (15%) were overweight and 4 (20%) were obese (>97th centile). Using the WHO age–sex-specific BMI standard deviation scores (SDS), in which SDS ⩾ 1 defines overweight and SDS ⩾ 2 defines obese status, 5 (25%) were overweight while 6 (30%) were obese. 23 Just one child had a positive GADA.
In total, 163 (143 adults and 20 children) underwent HNF1A Sanger sequencing.
Traditional versus high-sensitivity C-reactive protein criteria
Traditional criteria identified 125 out of 252 for screening for HNF1A-MODY. Out of these 125 individuals, 5 (4%) were found to have HNF1A-MODY. Using an hs-CRP criterion of <0.5 mg/l, 37 out of 252 individuals were screened of which 1 (2.7%) was found to have HNF1A-MODY. None in the paediatric cohort were found to have HNF1A-MODY.
Among the 143 adult participants who underwent HNF1A sequencing, 14 (9.8%) had undetectable C-peptide. All of these had onset of diabetes ⩽ 25 years of age and 50% had diabetes onset ⩽ 10 years; all except one had a BMI < 25 kg/m2, 9/14 had negative GADA, suggesting a significant proportion with GADA-negative type 1 diabetes among this group.
Characteristics of participants with hepatic nuclear factor 1 alpha maturity-onset diabetes of the young
Sanger sequencing of the HNF1A gene identified five adults with HNF1A-MODY (Table 3). The proportion of HNF1A-MODY among the adults tested was 3.5% (5/143). All these individuals fulfilled the traditional criteria of a positive family history of diabetes mellitus (DM) and age of diabetes onset under 25 years (range of age of onset of DM: 9–24 years). None of the participants with HNF1A-MODY had positive GADA. Four out of five had hs-CRP levels > 0.5 mg/l and BMI levels which would place them in the overweight category using Asian BMI cut offs (BMI > 23 kg/m2). 25 Diabetes duration ranged from 1 to 17 years. Four out of five of these individuals were on insulin treatment and could potentially have a trial of switch of therapy to sulphonylureas or meglitinides.
Characteristics of adults with hepatocyte nuclear factor 1 alpha maturity-onset diabetes of the young.
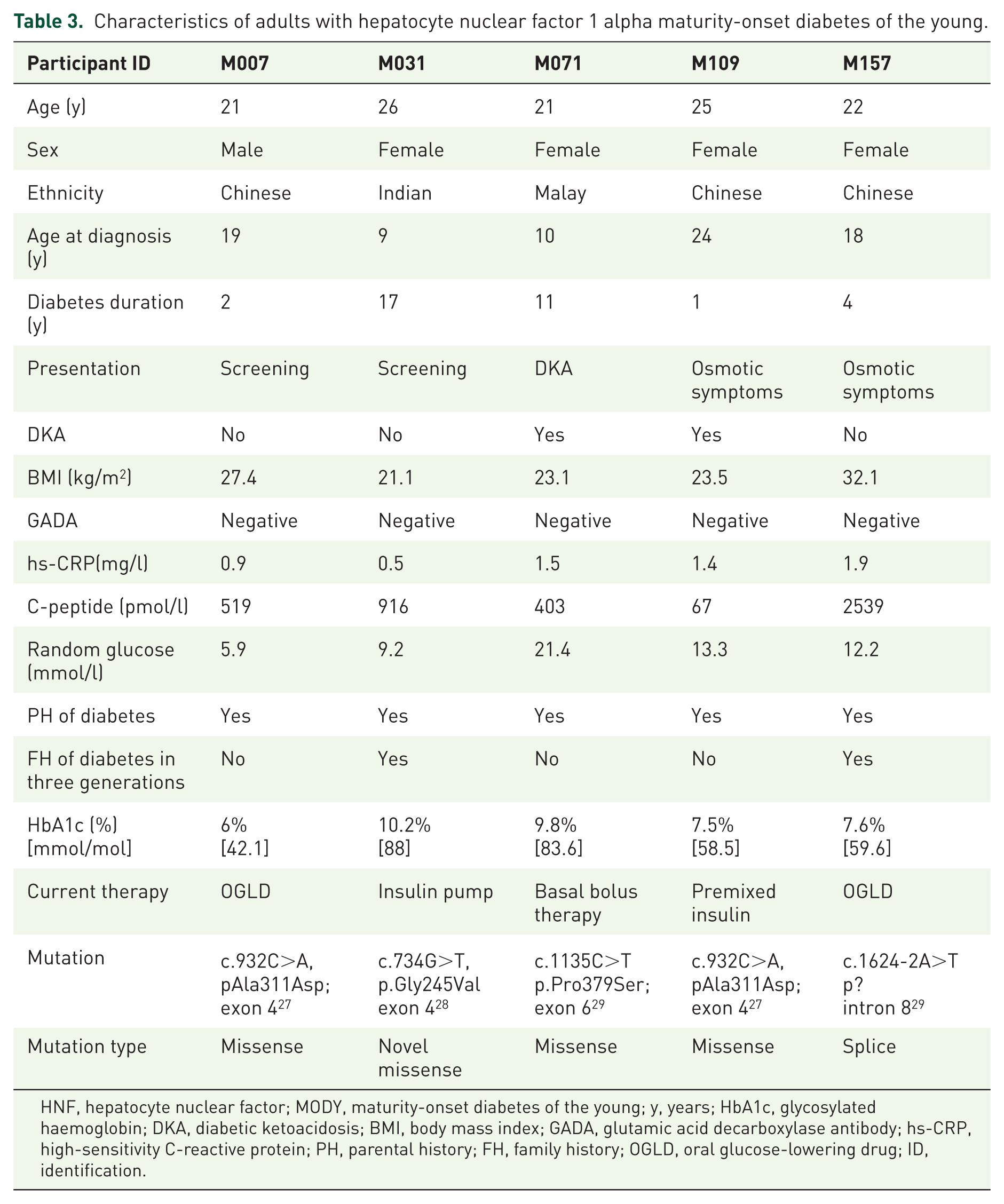
HNF, hepatocyte nuclear factor; MODY, maturity-onset diabetes of the young; y, years; HbA1c, glycosylated haemoglobin; DKA, diabetic ketoacidosis; BMI, body mass index; GADA, glutamic acid decarboxylase antibody; hs-CRP, high-sensitivity C-reactive protein; PH, parental history; FH, family history; OGLD, oral glucose-lowering drug; ID, identification.
Of the four heterozygous variants reported herein, one was novel and three were rare pathogenic/likely pathogenic variants. To validate their rareness and novelty, we screened two recently available large population sequencing reference databases, The Exome Aggregation Consortium 30 and The Genome Aggregation Database. 31 ExAC v0.3.1 and gnomAD v2.0 contain 60,706 and 123,136 exomes aggregated respectively from unrelated individuals having different ethnic backgrounds. 32 We checked allelic frequency according to the ethnicity of the participants, especially in the East Asian (EAS) and South Asian populations. The novel missense variant HNF1A, c.734G>T had no presence in either database. Of the other HNF1A variants, the missense variant c.1135C>T had 7 allele counts (ACs) out of 15,381 individuals in the South Asian population, splice site variant c.1624-2A>T and missense variant c.932C>A had 2 and 1 ACs, respectively, out of 8615 individuals in the EAS population in gnomAD database. Allele frequencies of these variants are summarized in Table 4.
Hepatocyte nuclear factor 1 alpha variants and characterization.
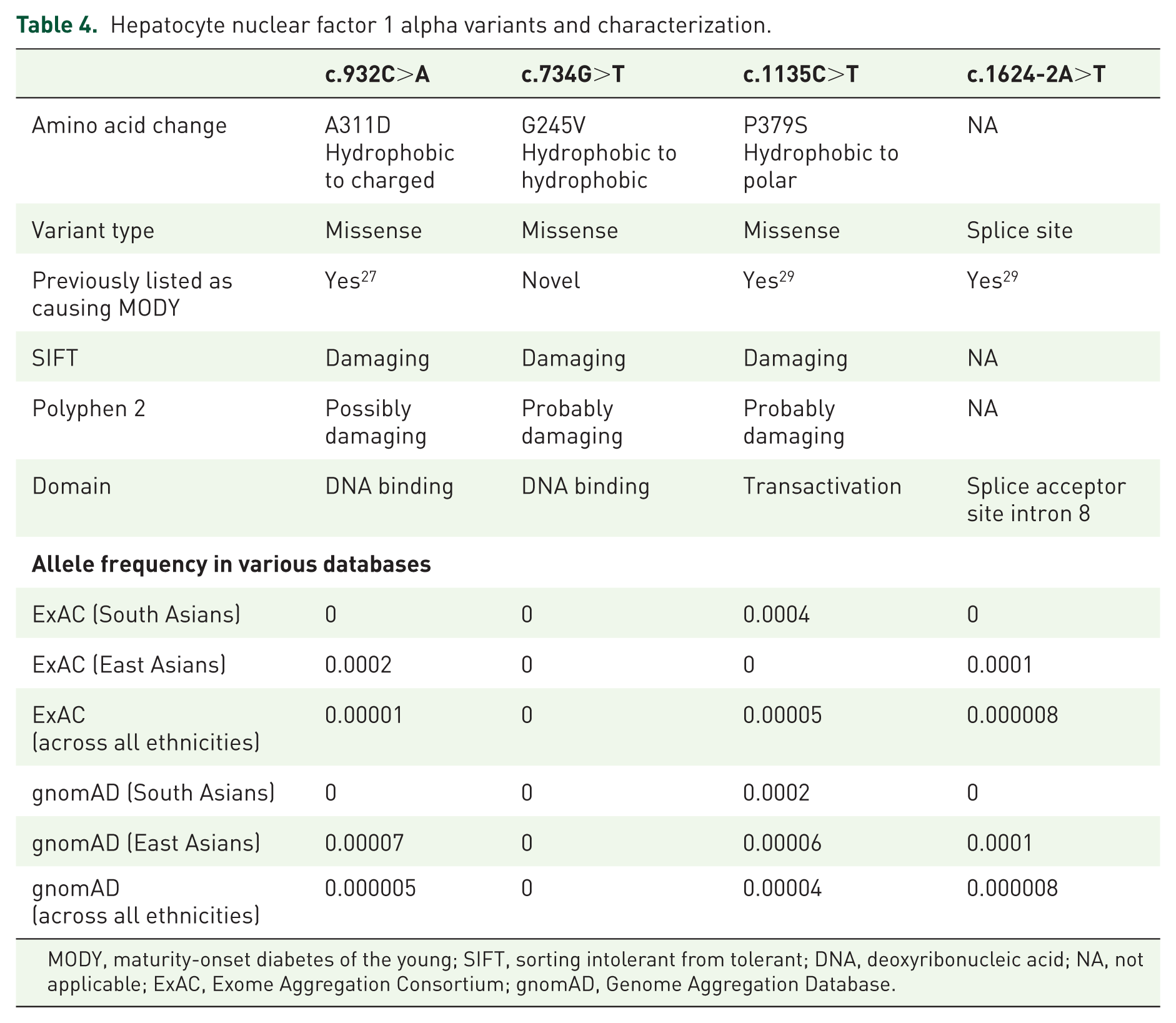
MODY, maturity-onset diabetes of the young; SIFT, sorting intolerant from tolerant; DNA, deoxyribonucleic acid; NA, not applicable; ExAC, Exome Aggregation Consortium; gnomAD, Genome Aggregation Database.
With reference to Table 3, participant M031 had a novel missense variant in exon 4 (c.734G>T) resulting in an amino acid change p.Gly245Val. This variant was predicted to be pathogenic by Sorting Intolerant Form Tolerant (SIFT), Polyhen2, Align Grantham Variation Grantham Distance (GVGD) scores, and was not present in the ExAC v3.0 database. Another amino acid change at the same residue (c.733G>A) p.Gly245Arg has been previously reported in a South Asian with young-onset (<25 years), non-insulin-treated diabetes with a three-generation family history. 28 This codon is part of the evolutionarily conserved domain and mutations in this region can disrupt the ability for dimerization of HNF1A.
Participant M031 (current age 26 years) was labelled with ‘type 1 diabetes’ at age 9 years, although never had diabetic ketoacidosis (DKA) and GADA was negative at diagnosis. She was lean (BMI 21.1 kg/m2), and despite receiving insulin for 17 years, she had endogenous insulin production (C-peptide 916 pmol/l). Her father was diagnosed with insulin-requiring diabetes at the age of 16 (current age 63 years) and required a coronary artery bypass graft in his thirties. He had undergone laser therapy for diabetic retinopathy but did not have microalbuminuria. Her mother had type 2 diabetes diagnosed in the late forties. HNF1A sequencing revealed the presence of the same mutation in her father but not her mother. The cosegregation of this novel missense variant with young-onset diabetes and its position within the HNF1A gene suggests that this variant is likely pathogenic. In view of preceding suboptimal glycaemic control and a desire to start a family soon, she was switched to insulin pump therapy with much improved glycaemic control (HbA1c 6.6%, 48.6 mmol/mol).
Participant M157 had a high-confidence loss-of-function rare splice acceptor variant HNF1A c.1624-2A>T, p?,29,33 which was found in 1/163 cases in this sequenced cohort. This variant has only been observed once in the ExAC (60,706 exomes) and twice in the gnomAD (123,136 exomes) databases. Despite twice the number of sequenced exomes in the gnomAD database compared with the ExAC database, the minor allele frequency (MAF) remained unchanged in the EAS (MAF = 0.0001) and global population (MAF = 8.139 × 10−6). This indirectly reflects the rarity of this variant within the EAS and across the global population.
M157 was diagnosed with diabetes at the age of 18 years. She described her body build at diagnosis as ‘normal weight, perhaps slightly overweight’ compared with peers. By age 20 years, she had bilateral laser therapy for diabetic retinopathy. She was first seen at our hospital following an admission (21 years) for a hepatic abscess. She is an only child, her father had insulin-treated diabetes since the age of 13 years, but had passed away from cardiac causes (age 52 years). Her mother also had diabetes diagnosed in her forties, was overweight at diagnosis and is on oral medications with no known diabetes-related complications. Her mother declined predictive gene testing but is likely to have type 2 diabetes. Although familial cosegregation was not possible, M157 did respond well to oral sulphonylureas (HbA1c fall from 9.9% to 7.5%). This splice site variant is likely to be pathogenic in M157, resulting in HNF1A-MODY, although her obesity status (BMI 32.1 kg/m2) is likely to contribute to an additional diagnosis of type 2 diabetes.
The rest of the variants in M007, M109, M071 have previously been reported to be implicated in MODY;27,29 participants M007 and M109 had a previously reported pathogenic missense variant in exon 4 (c.932C>A, pAla311Asp). 27 Both M007’s parents had later-onset diabetes (mother at age 51 years, father at age 58 years) and were on oral medications with no known complications. He had two siblings who were both reportedly without diabetes, although they had not undergone testing. Despite being approached several times and offered genetic testing, he declined on behalf of his family members. M109 was diagnosed with diabetes at age 24 years; GADA and Islet Cell Antibody (ICA) were both negative. Her father, and both paternal grand-father and uncle were known to have been diagnosed with diabetes in their late thirties. She had been initiated on insulin, but moved away and was lost to follow up prior to these results being available.
Participant M071 had a previously reported pathogenic missense variant in exon 6 (c.1135C>T p.Pro379Ser). 29 Participant M071 (current age 21 years) was labelled with ‘type 1 diabetes’ at the age of 11 years when she presented with DKA. Her GADA and ICA at presentation were negative. Random C-peptide at initial presentation was 432.9 pmol/l (glucose 13.2 mmol/l). She was started on basal-bolus insulin therapy with regular and isophane insulin and had a subsequent admission for DKA due to nonadherence to insulin therapy. Yet, her random C-peptide 10 years after diagnosis was 399.6 pmol/l, which would suggest she did not have type 1 diabetes. Her BMI at 14 years was 22.6 kg/m2 and current BMI was 23.1 kg/m2. There was a strong family history of diabetes, with both parents presenting with diabetes in their mid-forties. They were intermittently adherent to oral glucose-lowering drugs (OGLDs) and follow-up visits; genetic testing was offered but not taken up. Given her slim body habitus through this period, it was unlikely that her young-onset diabetes was related to type 2 diabetes. The positive family history of diabetes was not discriminatory, given that a family history of diabetes was so prevalent in this cohort (87.7%, Table 1). Participant M071 subsequently became pregnant and was maintained on a basal-bolus regimen with insulin analogues throughout pregnancy, achieving an HbA1c of 6.9% (52 mmol/mol) during this period. She had premature rupture of membranes and had an emergency C-section at 28 weeks into her pregnancy. Her son is currently 8-months old, and well.
M071’s first cousins (III-4 and III-5) also had young-onset diabetes. III-5 was also a participant (M044) of the study; both of his parents had diabetes, with his mother (unrelated to M071) being diagnosed with young-onset diabetes after developing gestational diabetes in her third pregnancy. III-5 tested negative for the same HNF1A variant. He was obese with a BMI of 31 kg/m2 and likely had young-onset type 2 diabetes.
Discussion
The aim of the study was to assess the utility of hs-CRP-based criteria in guiding targeted screening for HNF1A-MODY among Asians. Traditional criteria identified all HNF1A-MODY individuals but at the expense of sequencing many more individuals. While hs-CRP selected fewer individuals for HNF1A testing, it did not improve upon the diagnostic yield and would have excluded a significant proportion of those with HNF1A-MODY from diagnostic genetic testing. Although one individual with HNF1A-MODY was identified using the hs-CRP criterion, this individual also met the traditional criteria and would have been identified nonetheless. The overall proportion of HNF1A-MODY in the group tested based on traditional criteria was 4% (5/125).
These findings would be consistent with a recent French study 34 which compared hs-CRP levels among a large group of HNF1A-MODY individuals against those with familial young-onset type 2 diabetes and those with type 2 diabetes. While hs-CRP levels were lower in HNF1A-MODY compared with the latter two groups, using just clinical criteria alone were sufficient to discriminate between HNF1A-MODY versus type 2 diabetes, and the addition of hs-CRP did not appear to add value to this discrimination.
There could be various reasons why hs-CRP was not found to be useful in this Asian cohort. hs-CRP levels are influenced by ancestry, independent of BMI, age and assay type used. 35 Although we did not find any interethnic differences in hs-CRP among the three Asian ethnicities, it is possible that the hs-CRP levels of Asians with HNF1A-MODY are not low enough to allow differentiation from type 2 diabetes. A higher hs-CRP cut off may have identified more subjects with HNF1A-MODY, however, that would necessitate genetic testing of a larger number of subjects.
Young Asians with diabetes clearly present a very heterogeneous group with overlapping clinical features; identifying individuals that are neither consistent with type 1 diabetes or type 2 diabetes becomes a challenge. Family history on its own as a screening criterion is clearly not discriminatory, given that 88% in this Asian cohort had a family (parental) history of diabetes. The majority of this cohort had very-young-onset type 2 diabetes (mean 24.6 ± 9.2 years), and an overweight status. hs-CRP levels rise with higher BMI and accompanying insulin resistance. 36 Accordingly, the higher BMI and insulin resistance in this cohort could have reduced the utility of hs-CRP. Using BMI at diagnosis would have been more informative, but few would be able to recall their BMI at diagnosis. Nevertheless, four out of five adults with HNF1A-MODY were overweight (BMI > 23 kg/m2). Since individuals with HNF1A-MODY are not resistant to developing obesity and insulin resistance, hs-CRP may be less helpful in selecting these individuals. Conversely, current BMI level should not be used as the sole criterion to exclude an individual from genetic screening.
Another challenge in correctly assigning etiology is the low prevalence of islet autoimmunity in Asian type 1 diabetes. 14 The overall prevalence of antibody in this cohort was 5.9%, and GADA positivity was not associated with any significant differences in HbA1c, BMI or random C-peptide levels. By implication, this cohort would include those with antibody-negative type 1 diabetes, lowering the overall proportion of MODY within the target screening population. Indeed, close to 10% (14/143) had undetectable C-peptide, with negative GADA in 9/14 of these individuals. It is expected that antibody-negative type 1 diabetes individuals will be leaner and more likely to have lower hs-CRP levels, hence may have been over-represented in the low-hs-CRP group. Two out of five individuals with HNF1A-MODY had had a history of DKA in the past. DKA has been previously reported in individuals with HNF1A-MODY. 37 A previous history of DKA should therefore not exclude an individual from HNF1A-MODY screening if there are other suggestive features. Clearly, trying to distinguish between young-onset type 2 diabetes, antibody-negative type 1 diabetes, and MODY among Asians is not straightforward, and predictive calculators based on White populations would require validation before its use in the Asian setting.
Among the 20 children, no HNF1A-MODY was identified. A total of 35% of the cohort was overweight or obese and only one child had a positive GADA, implying the majority of these children had young-onset type 2 diabetes. Given the shorter duration of diabetes in this group, BMI at recruitment will be more reflective of BMI at diagnosis. Unlike in adults, children with a raised BMI are unlikely to have MODY. The sampling of a larger group of children will be needed to confirm this.
To the best of our knowledge, this is the first study performing targeted screening for HNF1A-MODY using traditional criteria and hs-CRP in a large multiethnic Asian cohort. The proportion of HNF1A-MODY in the traditional criteria group was 4% [minimum prevalence 5/252 (1.9%)]. This may seem lower than the reported proportion of 5–8% from other Asian countries.7 –10 However, these earlier studies assessed highly selected groups and reported novel missense variants as mutations where there could still be uncertainty over their pathogenicity. We are mindful that our study is not population based. The minimum prevalence of 1.9% of HNF1A-MODY would be similar to the prevalence of monogenic diabetes found in a recent population-based study in the UK 38 (prevalence of monogenic diabetes of 2.5%, of which <1% had HNF1A- MODY, in those with diabetes under 20 years).
Limitations include the assessment of only GADA as the islet autoantibody implicated in Asian type 1 diabetes and the possible inclusion of antibody-negative type 1 diabetes in our cohort. Although the prevalence of GADA positivity is consistently low in Asian type 1 diabetes, it remains the most prevalent antibody. 39 While conceding that MODY has been found in individuals previously misdiagnosed as having type 1 diabetes, and beta cell autoantibody positivity has been found in those with MODY, this was an uncommon occurrence (<1%). 40 Nevertheless, selection for genetic testing was not based on the antibody status. Participants were selected for HNF1A sequencing based on single-generation positive family history of diabetes rather than two generations. However, this selection criterion allowed wider inclusion for testing. This was a nonrandom sample, in that only consecutive or referred eligible subjects who consented for the study were recruited. Only HNF1A was analysed in both cohorts; other possible monogenic causes in these groups cannot be excluded. Lastly, as HNF1A analysis was performed using Sanger sequencing, large genomic rearrangements or deletions will not be detected using this approach. In addition, not all the 252 samples were sequenced. These two reasons could potentially underestimate the proportion of HNF1A-MODY in this population.
Conclusions
Young Asians with diabetes represent a heterogeneous group with overlapping clinical phenotypes that poses a challenge to physicians in terms of classification, risk stratification and optimal management. In Asians, individuals with type 2 diabetes present younger and with lower BMI, while those with HNF1A-MODY may present with higher BMI and more insulin resistance. hs-CRP does not appear to improve the diagnostic yield beyond the use of traditional clinical criteria for preselecting Asians for HNF1A genetic analysis. Using traditional criteria identified more HNF1A-MODY subjects but at the expense of sequencing three times more individuals.
