Abstract
Rotator cuff tendon injuries often lead to shoulder pain and dysfunction. Traditional treatments such as surgery and physical therapy can provide temporary relief, but it is difficult to achieve complete healing of the tendon, mainly because of the limited repair capacity of the tendon cells. Therefore, it is particularly urgent to explore new treatment methods. In vitro experiments were performed to explore the mechanism of differentiation of umbilical cord mesenchymal stem cells (UCMSCs) to tendon cells and to evaluate their potential in promoting rotator cuff injury repair. Growth factors such as CTGF, GDF-6, and GDF-7 were used to induce the differentiation of UCMSCs, and gene expression changes during the differentiation process were analyzed by single-cell sequencing. Hes1 overexpression and animal models were constructed to study its role in UCMSCs differentiation and rotator cuff injury repair. CTGF was the optimal factor for inducing the differentiation of UCMSCs into tendon cells. With increasing induction time, UCMSCs exhibited obvious tendon cell characteristics, such as changes in cell morphology and increased expression of tendon-specific proteins (MKX, SCX, and TNC). Single-cell sequencing analysis revealed key cellular subpopulations and signaling pathways during differentiation. Furthermore, overexpression of the Hes1 gene significantly promoted the differentiation of UCMSCs to tendon cells and showed its therapeutic effect in rotator cuff injury repair in an animal model. This study confirmed the potential of UCMSCs in tendon injury repair, especially the critical role of Hes1 in promoting UCMSCs differentiation and rotator cuff tendon-bone healing, which provides a theoretical basis and experimental rationale for the development of new cellular therapeutic strategies.
Background
Rotator cuff tendon injuries, as a common disorder of the musculoskeletal system, not only limit the quality of patients’ daily life and work, but also place a significant financial burden on the healthcare system.1,2 Rotator cuff tears are a common cause of shoulder pain and dysfunction. The function of tendons is to connect bones and muscles, transmit forces generated by muscle contraction to the bones, cause joint movement and body motion, and maintain joint stability. Tendon injuries and muscle injuries caused by overuse or large loads of sharp stimuli are common types of athletic injuries in sports training and everyday life.3,4 With the aging of the world’s population and the increasing number of people participating in sports, over 30 million skeletal muscle system injuries occur worldwide every year, which is a common disease in the field of sports medicine, with 30%–50% of all sports injuries being tendon related. 5
Despite advances in current treatments, complete healing of tendon injuries remains a challenge for most patients. Traditional conservative treatment methods, such as corticosteroid injections, low-intensity pulsed ultrasound and shock wave therapy, have achieved certain success in relieving pain and promoting initial healing.6,7 However, these therapies often fail to effectively promote the complete recovery of tendon tissues and may have side effects, such as tissue degeneration and immunosuppression caused by the long-term use of corticosteroids. 8 In addition, although surgical treatment can repair the ruptured parts through direct suture or transplantation, its effect also has certain limitations. Surgery can only restore the basic structure of the tendon and cannot reconstruct its functional tissues. Moreover, the recurrence rate after surgery is relatively high, especially after high-intensity or overloaded exercise. 9 The high recurrence rate after surgery has always been a scientific problem faced by sports medicine doctors, mainly because tendon cells have a low proliferation ability and the blood supply in the damaged area is insufficient, resulting in the limited self-repair ability of tendon tissues, a slow repair process and often accompanied by the formation of scar tissues.10–12 Therefore, the current treatment methods still cannot effectively meet clinical needs.
Amidst the rapid development in the field of medicine and biotechnology, cell therapy, as an emerging therapeutic approach, demonstrates the potential to cure tendon injuries. 13 Mesenchymal stem cells have strong self-replication and multidirectional differentiation ability, and can differentiate into osteoblasts, chondrocytes, tendon cells, and many other cells under specific induced conditions.14,15 Stem cell technology is one of the most promising biological approaches to promote rotator cuff healing, and has shown excellent results in in vitro and animal experiments, as well as in a number of clinical studies.16,17 With the deeper understanding of stem cells and other multi-energy cell types, there is a growing optimism about their potential application in tissue engineering and regenerative medicine. Umbilical Cord Mesenchymal Stem Cells (UCMSCs), as an emerging stem cell type, are considered as an ideal cell source especially suitable for translation to clinical applications due to their wide source, easy accessibility and low immunogenicity.18,19 UCMSCs were relatively stable in vitro proliferation and long-term storage, no adverse effects were found, and there was a low incidence of graft rejection and graft-versus-host disease, and a very low incidence of infection after transplantation into animal models. 20 Wang et al. 21 found that exosomes from human umbilical cord mesenchymal stem cells promote early damaged endometrial repair through miR-202-3p mediated extracellular matrix (ECM) formation. Thus, UCMSCs are stem cells with pluripotency that can be applied to organ regeneration and have a wide range of potential for tissue engineering and regenerative medicine. All in all, stem cell therapy has effectively overcome the limitations of tendon self-repair by enhancing cell proliferation, promoting the differentiation of tendon cells, improving the local microenvironment, reducing immune rejection reactions and enhancing tissue function recovery.
The study aims to investigate the molecular mechanisms underlying the induction and differentiation of umbilical cord mesenchymal stem cells into tendons in vitro, identify key signaling pathways and genes related to tendon differentiation, and explore whether targeted induction of tendons in vivo could be achieved by regulating key genes involved in tendon differentiation. This study reveals the complexity of UCMSCs differentiation and the role of key regulatory genes, providing clinically relevant solutions for the treatment of tendon lesions.
Materials and methods
Main materials and reagents
Rat umbilical cord mesenchymal stem cells (UCMSCs, CP-R302, Pricella, China), 293T cells (Pricella, China), DMEM medium (L110KJ, Yuanpei, China), Fetal bovine serum(AC03L055, life-ilab, China), GDF-6 (HY-P79333, MedChemexpress, USA), GDF-7 (92004ES10, Yisheng, China), CTGF (HY-P72154, MedChemexpress Biotechnology, USA), Goldenstar™ RT6 cDNA Synthesis Kit Ver.2 (TSK302M, Tsingke, China), qPCR Mix (SYBR Green I) kit and PCR primer (TSE002, Tsingke, China), RIPA lysis buffer (Beyotime, China), TNC (A1927, abclonal, China), MKX (ab236400, abcam, UK), SCX (ab58655, abcam, UK), GAPDH (A19056, abcam, UK), Hes1 (ab71559, abcam, UK), Hras (ab32417, abcam, UK), GAPDH Rabbit mAb (A19056, abclonal, China), Lentiviral Packaging Kit (41102ES10, Yisheng, China), Fetal bovine serum (26050070, Thermo, USA), trypsin (LP0042, Thermo, USA), PBS (10010001, Thermo, USA). Anti-CD34 antibody (PE), Anti-CD44 antibody (APC), Anti-CD45 antibody (APC-Cy7) Anti-integrin β1 (CD29)-FITC conjugate antibody were purchased from BD Pharmingen, USA.
UCMSC culture and group induction
Rat UCMSCs were cultured in DMEM medium containing 10% fetal bovine serum with 1% penicillin and 1% streptomycin incubated at 37℃ in a humidified incubator containing 95% air and 5% CO2. Depending on the inducer, the UCMSCs were divided into control group (DMEM medium), GDF-6 group (25 ng/ml GDF-6), GDF-7 group (25 ng/ml GDF-7), and CTGF group (25 ng/ml CTGF). Observation and detection were performed 14 days after induction. After determining the best inducer, the stem cells were treated at different time points (0, 3, 7, and 14 days) and the corresponding samples were collected for testing.
Lentivirus packaging and stable cell line establishment
The constructed plasmid was transfected into 293T to collect lentivirus. Lentivirus infection was carried out when the UCMSCs reached around 50%–80%. It was then replaced with fresh complete culture medium containing 5 μg/ml puromycin. When all cells had been killed by puromycin, the stable transformed strains were obtained.
Take the well-grown constructs of empty vector and Hes1-overexpressing UCMSCs stably transfected cells, and prepare the cell suspension. The above cell suspensions were mixed well and plated according to the grouping. After the cells were wall-plated, the cells were grouped according to the following groups: (1) Overexpression empty vector group (oe-NC): the constructed Hes1 overexpression empty UCMSCs stable transplants cells were cultured for 14 days in DMEM complete medium; (2) Hes1 overexpression vector group (oe-Hes1): the constructed Hes1 overexpression UCMSCs stable transplants cells were cultured for 14 days in DMEM complete medium; (3) CTGF-induced + oe-NC vector group (CTGF + oe-NC): constructed empty UCMSCs stable transplants cells, cultured with DMEM complete culture medium containing a final concentration of 25 ng/ml CTGF for 14 days, changing the liquid every 3 days, cultured up to 14 days to collect samples according to the subsequent detection.
Establishment and treatment of a rat rotator cuff tear model
The SPF-grade male SD rats (8 weeks old) were purchased from Byrness weil biotech Ltd (Chongqing, China) and were housed individually in a clean and open room maintained at a stable temperature (22℃) on a 12/12 h light/dark cycle. The study was conducted in accordance with the experimental norms and standards approved by the Animal Welfare and Research Ethics Committee. After 1 week of adaption, rats were randomly divided into four groups.
Supraspinatus tendon injury and repair was performed in rats bilaterally.Briefly, the supraspinatus tendon was sharply detached from the humeral head and repaired to its original attachment site by placing a modified Mason-Allen stitch on the tendon and passing a suture through the bone tunnel in the humeral head below the growth plate.Sham operated group was sutured immediately after incision of supraspinatus tendon without rotator cuff tear. Animals either received no treatement (sham operated group, model group) or UCMSCs stabilized cell line treatment. After treatment, the rats were anesthetized by intraperitoneal injection of sodium pentobarbital, and the tissues of the supraspinatus tendon were obtained and stored for future use.
Single-cell RNA sequencing (scRNA-seq)
The tested cells were washed, resuspended, and prepared into single-cell suspension of appropriate concentration. According to the number of target cells for the corresponding up-sampling, after the up-sampling, observe whether the Gel Beads in Emulsions (GEMs) can be formed normally, if so, the GEMs will be sucked out and transferred to the PCR tube for reverse transcription and library construction. After the library check was qualified, sequencing was performed on the machine.
Quality control, dimensionality reduction, and cluster analysis
For the processing of single-cell data, we first performed data quality control analysis to remove doublet as well as low-quality cells, Seurat for downstream analysis, louvain algorithm for cluster analysis of normalized data, t-distributed stochastic neighbor embedding (tSNE) algorithm for visualization of cell clustering in two-dimensional space, and SingleR for automatic annotation of cell types.
Mimetic sequencing analysis
The proposed temporal analysis was performed using Monocle3 software, which first generates cell clusters by dimensionality-decreasing clustering, then annotates the cell types by finding Marker genes and inferring the cell developmental trajectories based on the changes in the expression of the genes in each cell.
Gene differential expression and functional enrichment analysis
Marker genes for all clusters were analyzed using the wilcoxon algorithm, and genes that were highly expressed specifically in each cluster, had logFC > 0.25 and were expressed in at least 20% of the cells were selected as differentially expressed genes. Differences between groups were analyzed by FindMarks function for each cell clusters and differential volcano plots were drawn. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were performed using the ClusterProfiler package.Pathways with p_adj < 0.05 were considered to be significantly enriched.
Real-time quantitative polymerase chain reaction (qPCR)
Total RNA was extracted using Trizol reagent. According to Goldenstar ™ RT6 cDNA Synthesis Kit Ver.2 instruction manual for reverse transcription synthesis of complementary DNA (cDNA). Real-time quantitative PCR (qPCR) was prepared using a 2× T5 Fast qPCR Mix (SYBR Green I) kit to prepare a reaction system. The reaction was then performed in a real-time fluorescence quantitative PCR instrument to detect the Ct values of Hes1, MKX, SCX, and TNC. The housekeeping gene GAPDH was used for normalization, and the relative expression level of the gene was calculated using the 2−ΔΔCt method. The primers used were displayed in Table 1.
Primer information.

Western blot (WB)
Total cellular protein was extracted using RIPA lysate. The protein extract was diluted with one-fourth of the 5× SDS loading buffer and heated to 100℃ for 6 min for denaturation. Samples were separated by sodium dodecyl sulfonate polyacrylamide gelelectrophoresis and then transferred onto a polyvinylidene fluoride (PVDF) membrane. The primary antibodies TNC, MKX, SCX, and GAPDH were diluted 1:1000 and incubated at 4°C overnight. The next day, the secondary antibody GAPDH Rabbit mAb was diluted 1:2000 and incubated at room temperature for 1 h. The membranes were covered evenly using ECL exposure solution and detected in a nucleic acid protein gel imager. Gray scale values of the bands were determined using ImageJ.
Flow cytometry
Cells from each experimental group were collected separately by trypsin, centrifuged, supernatant discarded and resuspended. Set up isotype control: take a tube of uninduced UCMSCs cells and add 5 μL IgG2a-FITC Isotype Control from murine myeloma, 5 μL Mouse IgG1 κ, PE, Isotype Control, 5 μL Mouse IgG2b, APC, Isotype Control and 5 μL Mouse IgG1 κ, APC-Cy7, Isotype Control, shake and mix well, incubate at room temperature away from light, centrifuge, discard the supernatant and resuspend. In the experimental group, one tube of cells was taken separately, 5 μL of Anti-integrin β1 (CD29)-FITC conjugate antibody was added to each tube, shaking and mixing, incubated at room temperature away from light, centrifuged, supernatant discarded and resuspended. Repeat the above steps, add 5 μL Anti-CD34 antibody (PE), 5 μL Anti-CD44 antibody (APC), 5 μL Anti-CD45 antibody (APC-Cy7) in a single addition, shake and mix well, incubate at room temperature away from light, centrifugation, discarding the supernatant, and resuspension, and then use a flow cytometer (CytoFLEX, Beckman, USA).
Immunofluorescence
UCMSCs were inoculated on culture plates or slides and cultured to appropriate density. Cells were fixed with 4% paraformaldehyde for 20 min. Cells were treated with 0.1% Triton X-100 for 5 min to increase membrane permeability. Closed with 5% bovine serum albumin (BSA) for 1 h to reduce non-specific binding. Add specific primary antibodies (e.g. anti-COL I, COL III antibodies) and incubate overnight at 4°C. Subsequently, fluorescently labeled secondary antibodies were added and incubated at room temperature and washed again to remove unbound secondary antibodies. Finally, the nuclei were stained using DAPI, blocked with an anti-fluorescence quencher and the fluorescent signals were observed and recorded by fluorescence microscopy to analyze the expression and localization of the antigen.
Hematoxylin-eosin (HE) staining
The supraspinatus tendons were fixed with 4% paraformaldehyde, embedded them in paraffin and sectioned into 5 μm thick slices. The sections were sequentially deparaffinized in xylene and rehydrated with 100%, 95%, 88%, and 75% ethanol. After hematoxylin and eosin staining were done, the sections were analyzed under the microscope to the histological structure of supraspinatus tendon.
Masson staining
The supraspinatus tendons were fixed, embedded, sectioned to appropriate thickness, and adhered to slides. The sections were sequentially placed in xylene and gradient ethanol for deparaffinization and rehydration. The slices were first stained with turquoise blue and Mayer hematoxylin staining solution for 3 min. After staining with mayer hematoxylin, 1% hydrochloric acid alcohol was used for differentiation. Then the slices were sequentially immersed in Ponceau staining solution, phosphomolybdic acid solution and aniline blue solution for 10, 10, and 5 min. Finally, after dehydration and transparency with alcohol and xylene, the slices were sealed with neutral gum. The slices were examined microscopically and the images were captured and analyzed.
Biomechanical analysis
After obtaining the tissues of the humerus-supraspinatus site, all the peritendinous tissues except the supraspinatus tendon were excised and preserved in 4% paraformaldehyde solution. For biomechanical testing, the shoulder tissues were securely fastened to an instrumented biomechanical tester (Instron 5548; Instron, Norwood, MA, USA), and the supraspinatus tissues were tested for the maximal failure load at a displacement rate of 5 mm/min. The loaded load at the time of supraspinatus tendon rupture was observed and recorded in detail as the maximum failure load (N). The linear slope of the load-displacement curve was used as the stiffness.
Statistics analysis
Single-cell sequencing analyses were performed using the R (version 4.3.2) language, and descriptive statistical values of the data were expressed as mean ± standard deviation, and the data were analyzed by one-way ANOVA and Tukey’s post-hoc tests with p < 0.05 as the level of significance of the difference using GraphPad prism 8.0 statistical software, and plotted using Origin 2017 software.
Results
The optimal inducer of CTGF for the differentiation of UCMSCs into tendon cells
To select the inducer with the best differentiation effect, we treated UCMSCs with GDF-6, GDF-7, and CTGF for 14 days. The induced cells are shown in Figure 1(a). Before induction, it presents a stem cell like appearance, with cells arranged in a spiral and radial pattern. The cell volume is large, irregular, and has many branches. After induction, compared with the control group, the cells treated with the inducer showed a spindle shaped morphology, with long protrusions extending from the cytoplasm of spindle shaped cells toward both poles, and the protrusions were obvious.

Analysis of the induction effect of BMP-12, GDF-7, and CTGF on the differentiation of UCMSCs: (a) morphological changes of UCMSCs after induction treatment, (b) mRNA expression of MKX, SCX, and TNC detected by qPCR, (c) bands and protein expression of MKX, SCX, and TNC detected by WB, and (d) COL-I and COL-III expression detected by immunofluorescence (400×). **p < 0.01 and *** p < 0.001 compared with the control group.
The results of qPCR and WB assays showed that the mRNA and protein expression levels of MKX, SCX, and TNC were significantly higher in the GDF-6, GDF-7, and CTGF treatment groups compared to the control group (p < 0.05), with the CTGF group showing the most significant induction effect (Figure 1(b) and (c)). The immunofluorescence results showed that the expression of COL I and COL III was increased after induction treatment with GDF-6, GDF-7, and CTGF, and the CTGF group had the best induction effect (Figure 1(d)). Therefore, we chose CTGF for subsequent induction experiments.
Differentiation of UCMSCs to tendon cells after CTGF induction
In order to further explore the induction effect of CTGF on UCMSCs, cells at different time points after induction (day 0, day 3, day 7, and day 14) were observed and detected accordingly. We can see that in terms of cell morphology changes, with the increase of induction time, the nucleus became smaller, the cytoplasm became larger, and the cell differentiation was obvious (Figure 2(a)). The flow cytometry results showed that the UCMSCs cultured in this study were positive for the expression of markers CD29 (positive rate > 93.7%) and CD44 (positive rate > 97.5%), and negative for CD34 (positive rate < 2.48%) and CD45 (positive rate < 1.42%) expression (Figure 2(b) and (c)). It verified the presence of tendon-lineage proteins in UCMSCs cultures undergoing tenogenic induction. The results of qPCR and WB showed that the mRNA and protein expression levels of MKX, SCX, and TNC on the 3, 7, and 14 days after CTGF induction were all significantly increased than those in the control group (p < 0.05), and the improvement effect was most obvious on the 14 day (Figure 3(a) and (b)). Immunofluorescence results demonstrated that COL I and COL III expression increased with induction time (p < 0.05, Figure 3(c)).

Comprehensive analysis of the changes in cell morphology and immunophenotype during the induction process of UCMSCs: (a) cell morphology changes at different time points after CTGF induction treatment, (b) the immunophenotypic expression and positivity of UCMSCs at different induction time were analyzed by flow cytometry, and (c) analysis of positive immunophenotype. ***p < 0.001, **p<0.01, and *p < 0.05 compared with the 0 day.

Analysis of the induction effect of CTGF on the differentiation of UCMSCs at different induction time periods: (a) mRNA expression of MKX, SCX, and TNC detected by qPCR, (b) bands and protein expression of MKX, SCX, and TNC detected by WB, and (c) COL-I and COL-III expression detected by Immunofluorescence (400×). ***p < 0.001 and *p < 0.05 compared with the 0 day.
ScRNA-seq analysis of potential mechanism and microenvironment of CTGF induced transformation of UCMSCs into tendon cells
ScRNA-seq is a powerful tool for studying cellular biology, characterizing cell heterogeneity, identifying cell types, and exploring intercellular communication and interactions. To further explore the potential effects of CTGF on induced tendon cells, scRNA-seq was performed on UCMSCs induced by CTGF on 0, 3, 7, and 14 days. The results demonstrated that the cells were clustered into four cell subpopulations (Figure 4(a)) and annotated as three types of cells: Attachment cell, stem cell, and tendon cell (Figure 4(b)). In the early stage of induction, attachment cells are mainly involved in cell attachment and migration, providing support for the subsequent differentiation process. On day 0, the cells are mainly in an undifferentiated state, possessing the potential for self-renewal and multi-directional differentiation. As the induction time extends, the proportion of stem cells significantly decreases starting from day 3, while the proportion of tendon cells increases significantly, indicating that the induction of CTGF promotes the differentiation of stem cells into tendon cells (Figure 4(c) and (d)). The results of the proposed chronological analysis indicate that over time, stem cells gradually differentiate into tendon cells (Figure 4(e)).

Single cell sequencing analysis of UCMSCs after CTGF induction at different induction time periods: (a) cell fractionation tSNE visualization, (b) cell type tSNE visualization, (c) cell type tSNE visualization at different time periods, (d) proportion chart of different cell types at different time periods, and (e) proposed time series analysis. Attachment cells: attachment cells play a crucial role in the early stage of cell culture. By coming into contact with the basement membrane or the surface of the culture medium, they facilitate cell attachment, migration, and expansion; stem cells: stem cells have the potential for self-renewal and multi-directional differentiation, enabling them to differentiate into various cell types; tendon cells: tendon cells are specialized connective tissue cells, mainly responsible for the formation and maintenance of tendons.
The results of differential gene analysis showed that, compared with day 0, 539 genes were significantly upregulated and 3508 genes were significantly downregulated on day 3. Among the upregulated genes, genes such as Acta1 and Acta2 play roles in the process of CTGF-induced differentiation of UCMSCs into tendon cells. They are involved in functions such as muscle contraction, cytoskeleton regulation, transcriptional regulation, cholesterol synthesis, and extracellular matrix construction, enabling cells to acquire contraction and movement abilities, determine differentiation fates, reshape their morphology, and construct the extracellular environment. On day 7, 723 genes were significantly upregulated and 3413 genes were significantly downregulated. Among the upregulated genes, genes like Hes1 and Sox9 are involved in functions such as transcriptional regulation, cholesterol and fatty acid synthesis, substance transport regulation, and cytokine signal transduction during the CTGF-induced differentiation of UCMSCs into tendon cells. They play roles in determining the differentiation direction, providing an energy and material basis, constructing the extracellular matrix, and maintaining the orderliness of differentiation. On day 14, 1353 genes were significantly upregulated and 1099 genes were significantly downregulated (Figure 5(a)). Among the upregulated genes, genes such as Lcn2 and Nr4a2 are involved in functions such as iron metabolism and antibacterial defense regulation, transcriptional regulation, cholesterol uptake, antioxidant stress response, immune regulation, mTOR signaling pathway regulation, and energy metabolism during the CTGF-induced differentiation of UCMSCs into tendon cells. They create a stable micro-environment for cells, guide the differentiation direction, and meet the energy and metabolic needs of cells. We took the intersection of the number of up-regulated genes in the 3 day versus 0 day, 7 day versus 0 day, and 14 day versus 0 day groups, screened 124 differential genes, and analyzed the functions of the genes (Figure 5(b)).

Differential gene and functional enrichment analysis of UCMSCs after CTGF induction at different induction time periods: (a) volcanic plot of differentially expressed genes at different time periods compared to 0 days, (b) intersection analysis of up-regulated genes in three comparison groups, (c) GO enrichment bubble plots compared to 0 days at different time periods, (d) KEGG upregulated enrichment bubble plot compared to 0 days at different time periods, and (e) mRNA expression of Hes1 detected by qPCR. ***p < 0.001 compared with the 0 day.
The GO enrichment analysis comparing 3 day and 0 day demonstrated the 10 most significantly enriched pathways, where Biological Process (BP) were mainly involved in muscle tissue development, ameboidal-type cell migration and regulation of cellular macromolecule biosynthetic process, Cellular Component (CC) were mainly associated with collagen-containing extracellular matrix, cell leading edge and nuclear envelope pathways, Molecular Function (MF) were mainly involved in transcription coregulator activity, ubiquitin-like protein transferase activity and RNA polymerase ll-specifc DNA-binding transcription factor binding pathway. binding and other pathways. GO enrichment analysis comparing 7 day and 0 day showed that BP were mainly involved in ameboidal-type cell migration, response to radiation and negative regulation of cell cycle pathways, CC were mainly involved in chromosomal region, membrane raft and membrane microdomain pathways, and MF were mainly involved in transcription coregulator activity, GTPase regulator activity and nucleoside- triphosphatase regulator activity and nucleoside-triphosphatase regulator activity pathways. GO enrichment analysis comparing 14 day and 0 day displayed that BP were mainly involved in pathways such as ameboidal-type cell migration, regulation of apoptotic signaling pathway and small GTPase mediated signal transduction, CC were mainly involved in nuclear envelope, intrinsic component of organelle membrane and collagen-containing extracellular matrix pathways, and MF were mainly involved in pathways such as phospholipid binding, GTPase regulator activity and nucleoside-triphosphatase regulator activity (Figure 5(c)). KEGG pathway analysis showed that the pathways significantly enriched on 3 day compared to 0 day included chemical carcinogenesis reactive oxygen species, oxytocin signaling pathway, and diabetic cardiomyopathy pathway. The pathways significantly enriched on 7 day compared to 0 day include protein processing intrinsic plasticity, PPAR signaling pathway, and parathyroid hormone synthesis, secretion and inhibition pathways. The pathways significantly enriched on 14 day compared to 0 day include the HlF-1 signaling pathway, ribosome, and mineral absorption pathways (Figure 5(d)).
We took the intersection of the number of upregulated genes in the 3 versus 0 and 7 versus 0 groups, screened out 80 differential genes, and analyzed the functions of these genes. The genes were sorted in descending order according to the log2FC value. The list of the top 15 differential intersection genes is shown in Table 2. Based on previous research findings, Hes1 can induce the differentiation of stem cells.22,23 In addition, among these upregulated KEGG pathways, we found that the Oxytocin signaling pathway ranked high and was involved in the process of stem cell differentiation. We analyzed the genes enriched in the Oxytocin signaling pathway and predicted the binding of the transcription factor Hes1 to the genes enriched in the Oxytocin signaling pathway through the JASPAR database. The prediction results showed that Hras in this pathway could bind to Hes1, and there were 13 binding sites. We hypothesized that after induction, Hes1 is upregulated in umbilical cord stem cells, activating the Oxytocin signaling pathway, thus affecting the tenogenic differentiation of stem cells. Moreover, the qPCR results showed that the mRNA expression level of Hes1 on the 3rd, 7th, and 14th days after CTGF induction was significantly higher than that of the control group (p < 0.05, Figure 5(e)). Therefore, we selected Hes1 for subsequent experiments.
Gene list (Top 15).

Hes1, as a key gene in CTGF promoting the differentiation of UCMSCs into tendon cells
By comparing the scRNA-seq results of the control group and the induced group, we identified Hes1 as a key gene for CTGF induced differentiation of UCMSCs into tendon cells. To further investigate the effect of Hes1 on the differentiation of UCMSCs into tendon cells, we constructed a Hes1 overexpression model. It can be seen from the cell images that the cells in the oe-Hes1 group showed an obvious differentiation state. While in the oe-NC + CTGF group, although the cells had a certain degree of differentiation, the degree of differentiation was not significant (Figure 6(a)). The results of qPCR and WB showed that the mRNA and protein expressions of Hes1, Hras, MKX, SCX, and TNC in the oe-Hes1 group were significantly higher than those in the oe-NC group (p < 0.05). The mRNA and protein expressions of Hes1, Hras, MKX, SCX, and TNC in the oe-NC + CTGF group were lower than those in the oe-Hes1 group but higher than those in the oe-NC group (Figure 6(b)–(d)). In addition, the expressions of COL I and COL III detected by immunofluorescence also showed similar results (Figure 6(e)). These result suggested that Hes1 is one of the key genes that can promote the differentiation of UCMSCs into tendon cells.

Analysis of the induction of the Hes1 gene on the differentiation of UCMSCs to tendon cells: (a) changes in cell morphology (scale bar: 100 μm), (b) WB protein bands, (c) protein expression of Hes1, Hras, MKX, SCX, and TNC detected by WB, (d) mRNA expression of Hes1, Hras, MKX, SCX, and TNC detected by qPCR, and (e) COL-I and COL-III expression detected by immunofluorescence (400×). ***p < 0.001 and **p < 0.01 compared with the oe-NC group.
Effect of Hes1 on rotator cuff tears at the animal level
To validate the role of Hes1 at the animal level, we constructed a rotator cuff tear model with SD rats and treated with UCMSCs-stabilized strains. Finally, we collected supraspinatus tendon to detect changes in the associated proteins and genes. As shown in Figure 7(a) and (b), compared with the sham group, the levels of Hes1, Hras, MKX, SCX, and TNC proteins were significantly decreased in the model group at 2 and 4 postoperative weeks (p < 0.05). Similarly, RT-PCR analysis also revealed that rotator cuff tear significantly decreased the mRNA expression of Hes1, Hras, MKX, SCX, and TNC (p < 0.05, Figure 7(c)), while treatment with oe-Hes1 stable UCMSCs significantly increased their expression, and oe-NC group also showed significant increase after 4 weeks of treatment. These results suggest that UCMSCs therapy can improve rotator cuff tears and that Hes1 is a key gene in regulating rotator cuff tears.

The effect of Hes1 gene on rotator cuff tear at the animal level: (a) WB protein bands in 2 and 4 weeks, (b) protein expression of Hes1, Hras, MKX, SCX, and TNC in 2 and 4 weeks detected by WB, and (c) mRNA expression of Hes1, Hras, MKX, SCX, and TNC in 2 and 4 weeks detected by qPCR. ***p < 0.001, **p < 0.01 compared with the sham group, and ###p < 0.001 compared with the model group.
Histological observations at 2 and 4 weeks post-surgery revealed that the tendon-bone interface in the sham-operated group revealed thick, regular, columnar collagen fibers, fewer capillaries, and less inflammatory cell infiltration. In contrast, the tendon-bone interface in the model group showed mesenchymal cells proliferation, a large amount of loose connective tissue, infiltration of inflammatory cells, capillary proliferation and fibroblasts proliferation. The tendon bone interface of the oe-NC group exhibited mesenchymal cell proliferation, along with loose connective tissue and inflammatory cell infiltration, as well as capillary proliferation. However the tendon bone interface of the oe-Hesl group showed thick, regular, and tight collagen fibers arranged in a columnar pattern, with a small amount of infiltration of adipocytes and inflammatory cells, and a large number of newly formed capillaries (Figures 8 and 9(a)). Masson staining result showed that the collagen volume fraction in the model group was significantly higher than the sham group at 2 and 4 weeks post surgery, while the collagen volume fraction in both the oe-NC and oe-Hes1 groups was significantly lower than model group (Figure 9(b)). Immunofluorescence detection results indicated that at the 2 and 4 weeks post-surgery, the expression of COL I and COL III in both the oe-NC and oe-Hes1 groups was significantly higher than in the model control group, with the oe-Hes1 group showing the most prominent increase (p < 0.05, Figures 10(a), (b), 11(a), and (b)). Biomechanical testing results showed that at the 2 and 4 weeks post-surgery, the ultimate failure load and stiffness in the oe-NC and oe-Hes1 groups were significantly higher than in the model control group, with the oe-Hes1 group showing a more significant increase (p < 0.05, Figure 12(a) and (b)).

HE staining observation of changes in rotator cuff tendon tissue at 2 and 4 weeks (The cytoplasm is varying shades of pink to peach, the intracytoplasmic eosinophilic granules are bright red, the collagen fibers are pale pink, the elastic fibers are bright pink, the red blood cells are orange, and the proteinaceous fluid is pink.).

Masson staining observation of changes in rotator cuff tendon tissue at 2 and 4 weeks: (a) Masson staining of different groups after 2 and 4 weeks and (b) collagen volume fractions at 2 and 4 weeks (Collagen fibers or proteins, mucus, cartilage appear blue, cytoplasm, muscle, cellulose, glial cells, and red blood cells appear red, and nuclei appear blue purple.). ***p < 0.001 compared with the sham group and ###p < 0.001 compared with model group.

Immunofluorescence detection analysis of different groups in 2 weeks: (a) Mean fluorescence intensity of COL-I and COL-III and (b) COL-I and COL-III expression detected by immunofluorescence (400×). ***p < 0.001 compared with the sham group and ###p < 0.001 compared with model group.
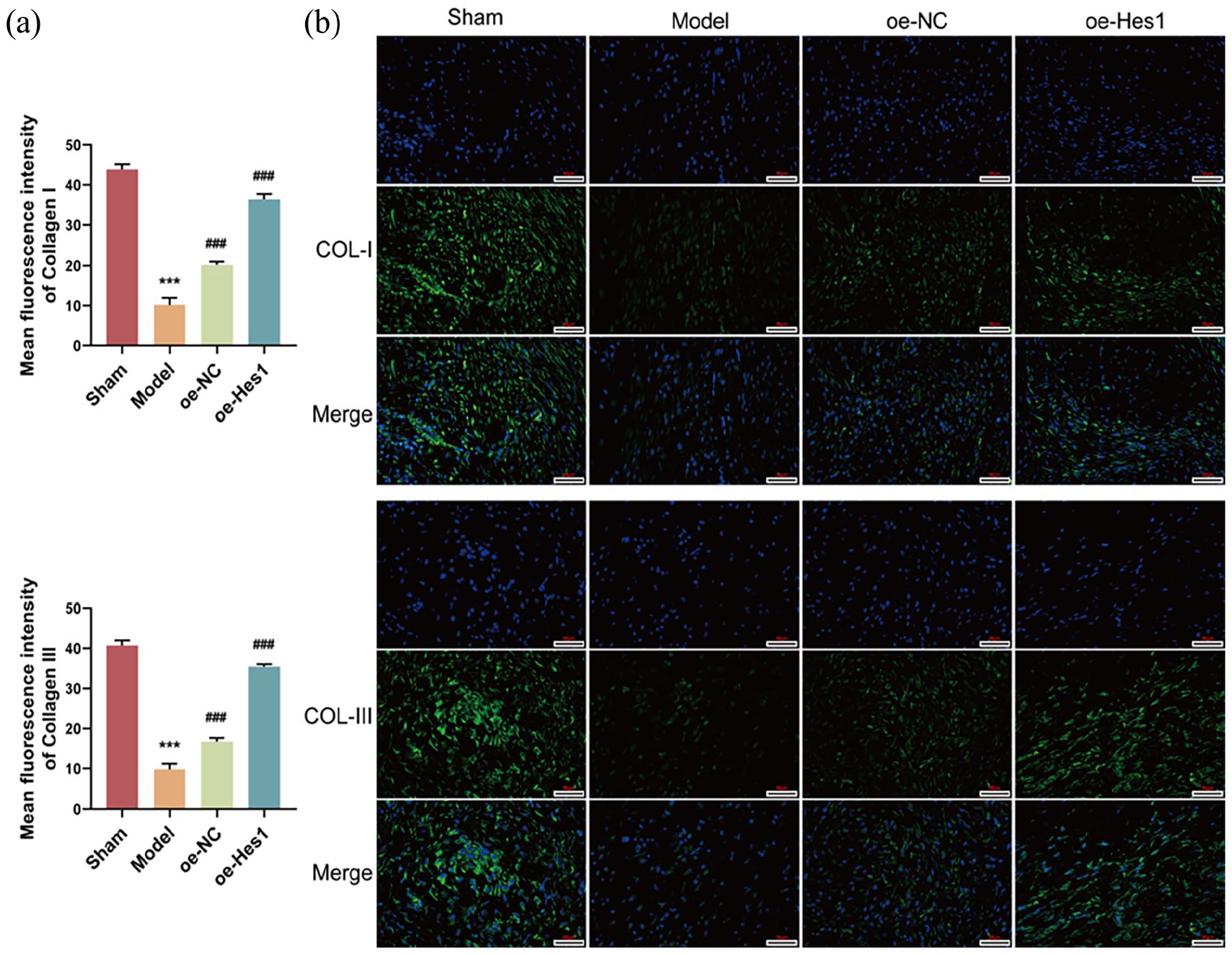
Immunofluorescence detection analysis of different groups in 4 weeks: (a) mean fluorescence intensity of COL-I and COL-III and (b) COL-I and COL-III expression detected by immunofluorescence (400×). ***p < 0.001 compared with the sham group and ###p < 0.001 compared with model group.

Biomechanical testing of tendon tissue changes at 2 and 4 weeks in different groups: (a) ultimate failure load (N) and (b) stiffness (N/mm). ***p < 0.001 compared with the sham group and ###p < 0.001 compared with model group.
Discussion
This study aims to explore the molecular mechanism of UCMSCs differentiating into tendon cells in response to the clinical needs of rotator cuff tendon injury treatment. Through optimized in vitro induction conditions, we successfully promoted the differentiation of tendon cells in UCMSCs and validated their potential to promote rotator cuff tendon bone healing in animal models. The findings of this study not only provide scientific basis for the feasibility of UCMSCs in tendon repair, but also point out the direction for future clinical translational research.
The ability of CTGF to induce UCMSCs toward a tendon cell phenotype is a significant advancement. Previous studies have explored the role of growth differentiation factors (GDFs) and bone morphogenetic proteins (BMPs) in tenogenic differentiation with varied success.18,24,25 Our work demonstrates that CTGF is more effective than GDF-6 and GDF-7, suggesting a more potent role in tendon healing. This is exemplified in studies where CTGF has been shown to enhance collagen production and improve the biomechanical properties of tendons,26,27 thereby highlighting its critical role in extracellular matrix remodeling. This may be related to the function of CTGF in regulating extracellular matrix formation and tendon cell maturation.
MKX is a transcription factor that plays a critical role in tendon and ligament genesis and development. It controls tendon cell proliferation, differentiation and ECM synthesis by regulating the expression of downstream target genes.28,29 MKX interacts with other transcription factors (e.g. SCX and TNC) to synergistically regulate tendon cell fate and function. SCX promotes tendon mechanical strength by regulating the expression of collagen and proteoglycans in the ECM.30,31 TNC, an extracellular matrix protein that plays an important role in tendon formation and repair, affects the mechanical properties of tendons by regulating the expression of other ECM proteins. 32 Hras, as an important signaling molecule, plays a key role in the proliferation and differentiation of tendon cells as well as in the synthesis of extracellular matrix, and the regulation of its activity has a potential impact on tendon tissue repair and regeneration. 33 Under CTGF induction, UCMSCs exhibited distinct morphological features of differentiation. Flow cytometry was used to verify changes in the surface markers of UCMSCs during the induction process, further confirming that these cells were indeed differentiating into tenocyte lineage. As the induction time increased (3, 7, and 14 days), the expression of MKX, SCX, TNC, and other markers continued to rise, with the most significant expression observed at day 14. This suggests that the differentiation of UCMSCs into tenocytes is a gradual process, and the longer the induction time, the more pronounced the differentiation effect.
The results of single-cell sequencing provided us with dynamic changes of the cells at different time points. Analysis revealed that, with the increase in induction time, the proportion of the stem cell population in UCMSCs gradually decreased, while the proportion of the tenocyte population significantly increased. This change indicates that UCMSCs, under the induction of CTGF, gradually differentiate from a stem cell state into tenocytes. In the differential gene analysis, we found that many genes related to tendon formation, such as Hras, were significantly upregulated, further confirming the key role of CTGF in cell differentiation. Moreover, GO enrichment analysis revealed significant enrichment of pathways related to biological processes such as muscle tissue development, cell migration, and transcriptional regulation, suggesting that these processes may play important roles in the differentiation of UCMSCs into tenocytes. According to the prediction of the JASPAR database, the transcription factor Hes1 can directly bind to the Hras gene in the oxytocin signaling pathway, and there are 13 binding sites. This binding may regulate the expression of the Hras gene, thereby affecting the expression of tendon cell markers (such as MKX, SCX, and TNC) and the differentiation process of tendon cells. Some studies have found that oxytocin and its receptor affect the functions of cells in bone tissue, such as osteoblasts, osteoclasts, and bone marrow mesenchymal stem cells, thus regulating the processes of bone growth, remodeling, and metabolism. 34 This further corroborates the possibility that Hes1 may exert its effect through this pathway.
In this study, we established a Hes1 overexpression model and found that it significantly promoted the differentiation of UCMSCs into tendon cells. We also observed that the mRNA and protein levels of Hes1, Hras, MKX, SCX, and TNC in the oe-Hes1 group were significantly higher than those in the control group. SCX and MKX, as key transcription factors for tendon cells, have been widely validated for their roles in tendon differentiation. Therefore, their upregulation in this study further supports the function of Hes1 in promoting UCMSCs tendon differentiation. In the animal experiments, we established a rotator cuff tear model in SD rats and explored the role of Hes1 overexpression in tendon repair. HE staining and Masson staining results showed that in the oe-Hes1 group, tendon injury repair was significantly accelerated, with tissue structure reconstruction being more complete and orderly compared to other groups. Immunofluorescence results revealed that at the tendon-bone interface in the oe-Hes1 group, collagen fibers were well-organized, and the expression levels of COL I and COL III were significantly higher than those in the model group. Numerous studies have shown that COL I and COL III are the major collagen types in tendon and bone tissues and play a crucial role in the formation of the tendon-bone interface and tissue repair.35,36 This suggests that Hes1 not only promotes UCMSC differentiation at the cellular level but also accelerates overall tissue repair. Further biomechanical testing showed that the ultimate failure load and stiffness in the oe-Hes1 group were significantly higher than those in the model group. The improvement in biomechanical performance indicates that Hes1 has significant potential in enhancing the mechanical stability and functional recovery of tendon tissue.
Moreover, our findings on the role of Hes1 in tendon regeneration are groundbreaking. Prior research has primarily focused on the role of Hes1 in neurogenesis and stem cell maintenance.37–39 Our study shifts this paradigm by demonstrating that Hes1 overexpression significantly promotes UCMSCs differentiation into tendon cells, thereby enhancing tendon repair. This is corroborated by findings where modulation of Hes1 has been linked to changes in cell fate decisions and differentiation capabilities, suggesting a broader role for Hes1 in regenerative medicine. The test results also suggest that UCMSCs therapy combined with growth factors such as connective tissue growth factor (CTGF) can further enhance the efficacy of UCMSCs in tendon repair.
However, this study also has certain limitations. Although in vitro experiments and animal models support the application of UCMSCs, their efficacy and safety in human treatment still need to be verified through large-scale clinical trials. Specifically, the application of UCMSCs first requires evaluation of its safety in a Phase I trial to ensure there is no toxicity and immune rejection. Subsequently, its efficacy is evaluated in a Phase II trial, and a Phase III trial is used to verify its effectiveness in a large-scale population. At the same time, the production of UCMSCs must strictly follow GMP (Good Manufacturing Practice) standards to ensure the quality and stability of the product. Although UCMSCs have low immunogenicity, the risk of immune reactions still needs to be evaluated for allogenic transplantation. The regulatory approval of cell therapy involves multiple aspects such as clinical trial design, ethical review, and product quality. In addition, the molecular mechanism by which Hes1 promotes the differentiation of UCMSCs still needs further study.
Conclusion
In this study, we successfully revealed the molecular mechanism of the differentiation of UCMSCs into tendon cells and evaluated their potential application in promoting rotator cuff tendon-bone healing. Through in vitro experiments, we found that CTGF was a potent inducer of the differentiation of UCMSCs into tendon cells. Single-cell sequencing analysis further revealed key cellular subpopulations and signaling pathways during differentiation, providing molecular-level insights into the targeted induction of UCMSCs. In addition, overexpression of Hes1 significantly promoted the differentiation of UCMSCs and showed the potential to improve the healing of rotator cuff injuries in animal models. These findings provide a theoretical basis and experimental rationale for the development of stem cell-based therapeutic strategies, which are expected to be a new approach for the treatment of rotator cuff tendon injuries. Future studies will focus on optimizing the induction conditions of UCMSCs and validating their therapeutic effects in a wider range of preclinical models to facilitate the translation of this therapy to clinical applications.
Footnotes
Acknowledgements
Not applicable.
Authors contributions
YS and YW conceived the study and designed and performed all single-cell sequencing and animal experiments. YX and JW conducted cell isolation, culture, and passage experiments. CY and ZH designed and performed cell experiments. FS performed the biological analysis and interpretation. TW wrote the manuscript and revised the manuscript with input from all authors. All authors have read and agreed to the published version of the manuscript.
Availability of data and material
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the Qingdao Natural Science Foundation (No: 24 - 4 - 4 - zrjj - 154 - jch) and the National Key Research and Development Program of China (No: 2023YFC2812004).
Ethics approval and consent to participate
All experimental procedures have been approved by the Animal Research Ethics Committee of Qingdao University Affiliated Hospital (Approval Ethics Review No. QYFY WZLL 28992). All methods were carried out in accordance with relevant guidelines and regulations. All methods are reported in accordance with ARRIVE guidelines.
Consent to participate
Not applicable.
