Abstract
The odontogenic differentiation of dental pulp stem cells (DPSCs), which is vital for tooth regeneration, was regulated by various functional molecules. In recent years, a growing body of research has shown that miRNAs play a crucial role in the odontogenic differentiation of human dental pulp stem cells (hDPSCs). However, the mechanisms by which miRNAs regulated odontogenic differentiation of hDPSCs remained unclear, and the application of miRNAs in reparative dentin formation in vivo was also rare. In this study, we first discovered that miR-3074-3p had an inhibitory effect on odontogenic differentiation of hDPSCs and antagomiR-3074-3p-conjugated PEI-AuNPs effectively promoted odontogenic differentiation of hDPSCs in vitro. AntagomiR-3074-3p-conjugated PEI-AuNPs was further applied to the rat pulp-capping model and showed the increased formation of restorative dentin. In addition, the results of lentivirus transfection in vitro suggested that FKBP9 acted as the key target of miR-3074-3p in regulating the odontogenic differentiation of hDPSCs. These findings might provide a new strategy and candidate target for dentin restoration and tooth regeneration.
Introduction
In recent years, human dental pulp stem cells (hDPSCs) have emerged as promising seed cells for tissue engineering due to their accessibility, minimal invasion, and multi-directional differentiation.1,2 Specifically, their ability to differentiate into odontoblasts has shown great potential in the field of tooth regeneration. 3 Odontogenic differentiation of hDPSCs was regulated by various functional molecules,4–6 among which microRNAs (miRNAs) have been reported to play a crucial role in coordinating odontogenic differentiation of hDPSCs in vitro.7–9 Despite this, the precise mechanism and role of miRNAs in the odontogenic differentiation of hDPSCs were not yet well understood. Furthermore, functional vectors were required to facilitate the uptake and function of miRNAs. While pure miRNAs cannot be directly taken up by cells, the application of functional vectors holds promise for promoting reparative dentin formation in vivo.
Gold nanoparticles (AuNPs) are small nanomaterials with a diameter ranging from 1 to 100 nm that are easy to synthesize and stable. It was reported that AuNPs could be conjugated with many functional parts, making them an ideal carrier for small molecule drugs or biological macromolecules, such as DNA, siRNA, and miRNA. 10 When modified with polyethylenimine (PEI), PEI-AuNPs provided miRNA more protection from enzymatic degradation, enhanced cellular uptake, and endosome escape.11,12 Previous studies have confirmed that miRNA-conjugated PEI-AuNPs had the ability to facilitate liver protection and corneal repair.13,14 Furthermore, it has been reported that miRNAs could promote osteogenesis or osseointegration when loaded by AuNPs.15–17 Since the process of odontogenesis is similar to that of osteogenesis and both physiological processes are involved in collagen formation and matrix calcification with the expression of the same mineralization-related genes and proteins, such as ALP, COL I, DMP-1, and Bmps,18,19 miRNA-conjugated PEI-AuNPs may be an effective strategy for tooth regeneration. However, whether it has the potential to restore dentin has not been reported.
FK506 binding protein 9 (FKBP9), a member of the immunophilin family, can bind to the immunosuppressive drug FK506. 20 FKBPs are known to participate in various biological processes, such as immunosuppression, tumor development, and metastasis.21–23 FKBP9, in particular, has been proven to play a key regulatory role in protein folding and transport, signal transduction, nerve growth, and other aspects. 24 However, its role in osteogenic differentiation or odontogenic differentiation has never been reported.
The mammalian target of rapamycin (mTOR) is a crucial signaling molecule that integrates various intracellular and extracellular signals to regulate cell growth and metabolism.25,26 Our previous study has demonstrated the involvement of the mTOR signaling pathway in regulating the mineralization process of odontoblast cells. 27 This finding suggested that there might be molecular pathways involved in the regulation of odontogenesis under the activation of the mTOR pathway.
This study aimed to screen for the potential regulator that might participate in the odontogenic differentiation of hDPSCs through mTOR pathway activation. We identified miR-3074-3p and demonstrated its ability to regulate the odontogenic differentiation of hDPSCs in vitro. In addition, antagmiR-3074-3p-conjugated PEI-AuNPs showed a promotive effect on odontogenic differentiation of hDPSCs in vitro and dentin restoration in vivo. Subsequent studies revealed that FKBP9 was an effective target of miR-3074-3p. The FKBP9-enhanced lentivirus transfection experiments further validated that miR-3074-3p regulated the odontogenic differentiation of hDPSCs by targeting FKBP9. Our findings provided a research basis for dentin restoration and tooth regeneration.
Materials and methods
Isolation and culture of hDPSCs
HDPSCs were obtained from third molars or premolars extracted for orthodontic reasons from patients under 25 years of age at Nanfang Hospital. The pulp tissue was decomposed using 1 mg/mL type I collagenase (Sorabio, Beijing, China) for 0.5 h at 37°C and adhered to the culture bottle. Subsequently, the cells that climbed from the tissues were passaged and cultured in a 5% CO2 environment at 37°C. HDPSCs were cultured in Dulbecco-modified Eagle medium (DMEM, Gibco, USA) supplemented with penicillin, streptomycin, and 10% fetal bovine serum. The cells from the second to sixth passages were used in this study. The culture and identification of hDPSCs were shown in Supplemental Figure 1.
Flow cytometric analysis
Flow cytometric analysis was carried out by entrusting a biotechnology company (Jiamai Biotechnology Co., Ltd, Guangzhou, China). HDPSCs were washed and collected into the flow tube. Then, PBS containing primary antibodies were added into the sample for incubation for 2 h at room temperature in the dark: fluorescein isothiocyanate (FITC)-conjugated anti-human Stro-1 (Proteintech, China, FITC-65184), allophycocyanin (APC)-conjugated anti-human CD45 (Biolegend, USA, 304011), or anti-human CD90 (Biolegend, USA, 328113) and phycoerythrin (PE)-conjugated anti-human CD146 (Proteintech, China, PE-65181), anti-human CD29 (Biolegend, USA, 303003), anti-human CD11b (Biolegend, USA, 393111), or anti-human CD34 (Biolegend, USA, 343505). No primary antibody incubated sample was used as a negative control. The cell suspensions were washed twice, resuspended in PBS, and analyzed with a Novocyte D2060R Flow Cytometer (Agilent, USA). Novoexpress 1.5.6 (Novoexpress Software, USA) was employed for data analysis and graph plotting.
Odontogenic differentiation and adipogenic differentiation of hDPSCs
To verify its odontogenic differentiation capacity, hDPSCs were induced using a mineralization-inducing medium (DMEM containing 10% FBS, 50 μg/mL ascorbic acid, 10 mM β-glycerophosphate, and 0.01 μM dexamethasone; Solarbio, Beijing, China) for 14 days. Cells were washed with 1× PBS and fixed with 4% paraformaldehyde for 15 min, followed by staining with Alizarin Red S. The same experimental conditions were used for subsequent mineralization experiments. To verify its adipogenic differentiation capacity, hDPSCs were induced using an adipogenic medium (DMEM supplemented with 500 mM isobutyl-methylxanthine, 0.5 M hydrocortisone, 60 mM indomethacin, 10 mM insulin, and 10% FBS; Solarbio, Beijing, China) for 21 days. Cells were washed with 1× PBS and fixed with 4% paraformaldehyde for 15 min, followed by staining with Oil Red O (Solarbio, Beijing, China).
MiRNA transfection
HDPSCs were cultured to about 75% confluence and transfected with miRNA mimic negative control (sense, 5′-UUUGUACUACACAAAAGUACUG-3′), miRNA inhibitor negative control (sense, 5′-CAGUACUUUUGUGUAGUACAAA-3′), miR-3074-3p mimic (sense, 5′-GAUAUCAGCUCAGUAGGCACCG-3′), and miR-3074-3p inhibitor (sense, 5′-CGGUGCCUACUGAGCUGAUAUC-3′; Xindai Bio, Guangzhou, China) through Lipofectamine 2000 (Invitrogen, USA) according to the manufacturer’s instructions. AntagomiR-NC (antagomiR NC #22) or antagomiR-3074-3p (sense, 5′-GAUAUCAGCUCAGUAGGCACCG-3′) conjugated with PEI-AuNPs was transfected according to the transfection amount of miRNA. MiRNA mimic negative control and miR-3074-3p mimic were transfected at the concentration of 50 nM. MiRNA inhibitor negative control, miR-3074-3p inhibitor, antagomiR-NC/PEI-AuNPs, and antagomiR-3074-3p/PEI-AuNPs were transfected at the concentration of 100 nM. The cells were transfected once every 2-3 days during mineralization induction.
Alkaline phosphatase (ALP) staining
After 7 days of mineralization induction, hDPSCs were washed with 1× PBS and fixed with 4% paraformaldehyde for 15 min. Then, cells were washed several times with deionized water and treated with prepared ALP staining buffer (Beyotime, Shanghai, China) at 37°C for 30 min. After washing several times with 1× PBS, the intensity of dark blue was assessed under an inverted microscope (Leica, Germany). The experiment was repeated three times separately.
Alizarin Red S staining
After 14 days of mineralization induction, hDPSCs were washed with 1× PBS and fixed with 4% paraformaldehyde for 15 min. Then, cells were washed several times with deionized water and treated with 0.2% Alizarin Red S staining buffer (Solarbio, Beijing, China) at 37°C for 30 min. After washing several times with 1× PBS, mineralization nodules were assessed under an inverted light microscope (Leica, Germany). The experiment was repeated three times separately.
Total RNA isolation and qRT-PCR analysis
Total RNA was extracted from the cells using Trizol reagent. The extracted RNA was then quantified, and 1000 ng RNA was reversed and transcribed into cDNA using the cDNA synthesis kit (Ruizhen Bio, Guangzhou, China) according to the manufacturer’s instructions. The reaction volume of 10 μL containing 5 μL SYBR, 0.4 μL Primer, 3.6 μL H2O, and 1 μL cDNA was used to perform qPCR on Light Cyler 96 (Roche, Switzerland). The internal reference for mRNA and miRNA was glyceraldehyde 3-phosphate dehydrogenase (GAPDH) and U6 RNA, respectively. The qPCR primers are shown in Table 1. 2−∆∆CT method was used to analyze the relative gene expression in each group. The experiment was repeated three times separately.
Sequences of primers for qRT-PCR.

Western blot analysis
Endogenous proteins were isolated from cells using RIPA supplemented with PMSF. The protein concentration was measured with a BCA protein determination kit (Fdbio Science, Hangzhou, China). Extracted proteins were denatured with loading buffer at the ratio of 4:1. Then 20 mg protein was quantified according to concentration and loaded onto 8% SDS-PAGE gel and transferred to 0.45 μm PVDF membrane. The membranes were blocked with 5% BSA and then incubated with primary antibody at 4°C overnight. After washing with TBST for 30 min, membranes were incubated with a secondary antibody for 1 h. Following another wash with TBST for 30 min, membranes were measured using G: BOX Chemi XX9 (Syngne, USA). The primary antibodies were as follows: anti-alkaline phosphatase (Abmart, China, T55421, 1:7000), anti-collagen I (Abmart, China, T61022, 1:1000), anti-DMP-1 (Abmart, China, PY8825, 1:2000), anti-DSPP (Santa Cruz, USA, sc-73632, 1:500), anti-GAPDH (Abmart, China, P60037, 1:5000). Finally, the protein band on membranes were analyzed using ImageJ. The experiment was repeated three times separately.
PEI-AuNPs and GELMA synthesis
A mixture of 6 mL of 14 mM HAuCl4 solution and 4.32 mL of 1% PEI solution was stirred by magnetic rotor at room temperature for 24 h. Clear wine-red gold nanoparticles sol was obtained. Then, 5 mL of the solution was then dialyzed with deionized water for 7 days to remove excess PEI. The dialyzed wine red gold nanoparticle sol was concentrated with polyethylene glycol solid to 1.25 mL. The concentrated solution was collected and stored at 4°C. When used, PEI-AuNPs were mixed evenly with miRNA according to the proportion indicated by agarose gel electrophoresis. Solid GELMA is dissolved in 1% 2959 photoinitiated solution at a ratio of 1:9 by ultrasonic vibration to obtain liquid 10% GELMA. The resulting solution was stored at room temperature and away from light.
Agarose gel electrophoresis
About 20 μM antagomiR-NC or antagomiR-3074-3p was mixed with PEI-AuNPs solution at different volume ratios. The ability of PEI-AuNPs to conjugate with miR-3074-3p was evaluated by conducting 1% agarose gel electrophoresis at room temperature for 15 min using an electrophoresis device (Bio-rad, America).
MiRNA labeling and cell uptaking
Cy3-labeled antagomiR-NC (antagomiR NC #22(Cy3)) or Cy3-labeled antagomiR-3074-3p (sense, Cy3-5′-GAUAUCAGCUCAGUAGGCACCG-3′) conjugated with PEI-AuNPs and free Cy3-labeled antagomiR-3074-3p were added to cells at the concentration of 100 nM according to the specific miRNA. After 12 h of incubation, cells were washed with 1× PBS and fixed with 4% formaldehyde for 15 min. Then, cells were stained with DAPI dye (Thermo, America). Samples were observed under confocal laser scanning microscopy (Leica, Germany).
Cell proliferation assay
HDPSCs were seeded on 96-well culture plates at the density of 2000 cells per well for incubation. Then, 10 μL CCK8 working solution was added into each well and co-incubated with cells for 1 h. The proliferation rate of cells was determined using a plate reader (Molecular Devices, USA) through 450 nm absorbance values. The procedure was conducted every day for the next 7 days to collect data on cell proliferation. The experiment was repeated three times separately.
Cell scratch assay
HDPSCs were seeded on 6-well culture plates (ABC Biotech, Hong Kong, China). After the hDPSCs were grown to confluency, scratches in each group were made using a 200 μL pipette tip. The samples were then observed under an inverted fluorescence microscope (Leica, Germany) at 0, 12, 24, and 72 h after the scratches. The experiment was repeated three times separately.
Evaluation of antagomiR/PEI-AuNPs Release from GELMA
A specific amount of cy3-antagomiR/ PEI-AuNPs conjugated with a specific amount of GELMA was dispersed in 400 μL DNase/RNase-Free water at room temperature. Then, a fixed amount of supernatant was obtained from the suspension daily for the next 7 days. The release of Cy3-antagomiR/ PEI-AuNPs from GELMA was evaluated by measuring the fluorescence intensity in the supernatants using a multimode plate reader (CYTATION 5, BioTek; excitation = 550 nm; emission = 615 nm).
Rat pulp-capping model
Eight-week-old Male Sprague Dawley (SD) rats were purchased from the Animal Center of the Southern Medical University (Guangzhou, China). All animal studies were conducted in accordance with the protocols approved by the Animal Care Committee of Southern Medical University, China (No. IACUC-LAC-20221017-008). The rat pulp-capping experiments were carried out according to Swanson’s research. 28 A class I defect was created at the central cusp of maxillary first molars using a portable electric dental engine. The antagomiR/PEI-AuNPs/GELMA complex was injected into the defect by an Insulin syringe and illuminated by ultraviolet light at 365 nm wavelength at a distance of 1 cm for the 30s. Then, the defect was sealed with glass ionomer cement and resin. The opposing tooth cusps were ground off to relieve occlusion. Thirty SD rats were equally divided into five groups. The maxillary first molar was only opened in blank control group, while in the other four groups, the maxillary first molar was capped with GELMA, PEI-AuNPs/GELMA, antagomiR-NC/PEI-AuNPs/GELMA or antagomiR-3074-3p/PEI-AuNPs/GELMA, respectively. The maxillary first molars were dissected at 1, 4, and 8 weeks after the surgery and fixed with 4% FPA for 1 week, and then demineralized in 4% EDTA at 37°C for 4 weeks for further dehydration, paraffin embedding, and sectioning.
Histological analysis
The maxillary first molar samples were fixed with 4% PFA and demineralized for 4 weeks. After further dehydration and paraffin embedding, the samples were sectioned at a thickness of 4 µm and subjected to hematoxylin and eosin (H&E) staining according to standard protocols. Stained sections assessment was performed using an ortho-microscope (Olympus BX 51).
Prediction of target genes of miRNA
The target genes of miRNA were predicted using the MIRDB (http://mirdb.org/), MirWalk (http://mirwalk.umm.uni-heidelberg.de/), and TargetScan (http://wargetsww.tcan.org/) databases.
Luciferase reporter assay
Luciferase reporter assay was carried out by entrusting a biotechnology company (Youming Biotechnology Co., Ltd, Guangzhou, China). Luciferase reporter vectors carrying Pmir-report-pten wild-type and Pmir-report-pten mutant were transfected into 293T cells coupled with miR-3074-3p mimics or corresponding controls. After 24 h of transfection, the cells were lysed with passive lysis buffer. Then, Firefly and Renilla luciferase activity was assessed.
Lentivirus transfection
FKBP9-enhanced virus was designed and provided by a biotechnology company (Sbo-bio Biotechnology Co., Ltd, Shanghai, China). HDPSCs were seeded on 6-well culture plates at a density of 1 × 105 cells per well. The cells were divided into three groups: untransfected group, empty virus group, and FKBP9-enhanced virus transfected group. Once cells were cultured to adhere to the wall, transfection was conducted with the optimal MOI of 100, as determined in the pre-experiment (sbo-bio, China). After 3 days of transfection, the cells were fixed with 4% PFA and stained with rhodamine dye (Thermo, America) and DAPI dye (Thermo, America), respectively. Then, samples were observed under a laser confocal microscope (Leica, Germany). When the cell confluence rate reached about 80%, the cells were passaged at a ratio of 1:3. The next day, 2 μg/mL puromycin was added to the culture medium, and the screening medium was refreshed every 2 days. The selected cells were used for the mineralization induction experiment.
Statistical analysis
All data were presented as means ± standard deviation (SD). Unpaired t-tests were performed for comparison between two groups, while analysis of variance was performed for comparison among multiple groups. The statistically significant differences were defined as *p < 0.05, very significant for **p < 0.01, and the most significant for ***p < 0.001. GraphPad Prism 7 (GraphPad Software, USA) was employed for data analysis and graph plotting.
Results
MiR-3074-3p might participate in the odontogenic differentiation of hDPSCs
The RNA sequencing results of odontoblasts in the mTOR pathway activation and NC groups revealed 46 differently expressed miRNAs between the two groups (Figure 1(a)). We screened eight most differentially expressed miRNAs from them (Figure 1(b)), among which miR-3074 exhibited the highest expression level (Figure 1(c)). We verified the expression of four different miRNAs in odontoblasts using qRT-PCR (Figure 1(d)) and observed a significant difference in the expression of miR-3074-3p between the two groups. These results indicated that miR-3074-3p might participate in the process of odontogenic differentiation.

MiR-3074-3p might participate in the odontogenic differentiation of hDPSCs: (a) Volcano map of 46 differently expressed miRNAs between NC group and TSC1 group, (b) heat map presentation of eight relative miRNAs with ⩾10-fold difference between NC group and TSC1 group, (c) eight mainly different miRNAs with ⩾10-fold Difference between NC group and TSC1 group, and (d) comparison of relative miR-3074-3p, miR-23b, let-7b-3p, let-7b-5p content between NC group and TSC1 group by qRT-PCR. Data were presented as means ± SD, n = 3/group, *p < 0.05. **p < 0.01. ***p < 0.001.
MiR-3074-3p regulated odontogenic differentiation of hDPSCs negatively
To determine the role of miR-3074-3p in odontogenic differentiation, we investigated its potential effects on the odontogenic differentiation of hDPSCs. Through transfection with miR-3074-3p mimics or miR-3074-3p inhibitors to hDPSCs, miR-3074-3p was overexpressed or silenced to explore whether miR-3074-3p could regulate the expressions of the odontogenic marker. The results of mineralization induction showed that the mRNA and protein levels of odontogenic markers, such as ALP, COL-1, DMP-1, and DSPP, were significantly repressed by miR-3074-3p overexpression and upregulated by inhibiting miR-3074-3p (Figure 2(a)–(c)). Similar trends were observed in the evaluation of ALP staining and Alizarin Red S staining (Figure 2(d)). These results suggested that miR-3074-3p effectively inhibited odontogenic differentiation of hDPSCs.
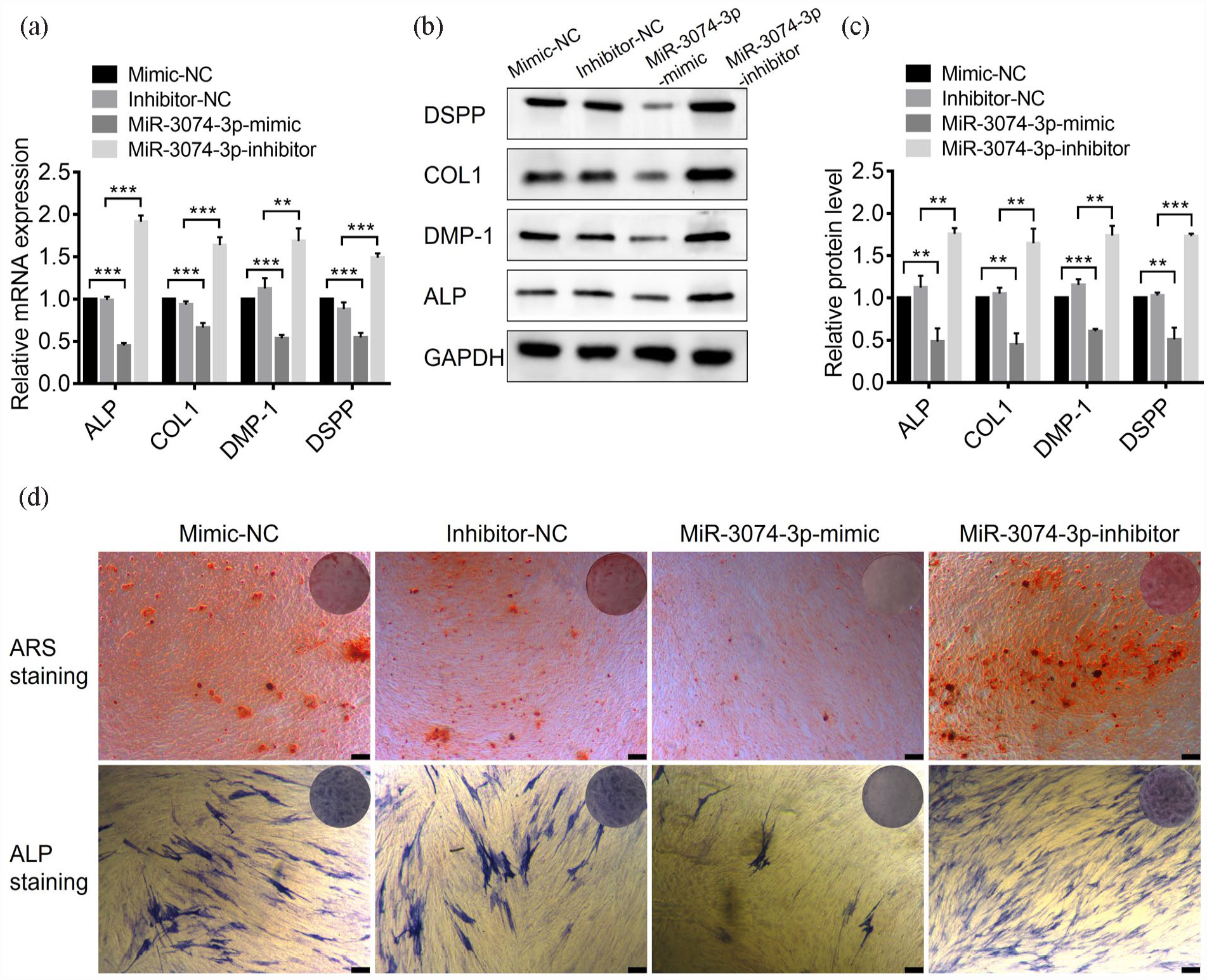
MiR-3074-3p inhibited the odontogenic differentiation of hDPSCs in vitro, and miR-3074-3p inhibitor could promote it: (a) The expression levels of the odontogenic marker genes (ALP, COL1, DMP-1, and DSPP) in hDPSCs treated with miR-3074-3p mimics or inhibitors were detected by qRT-PCR, (b and c) The expression levels of the odontogenic marker proteins (ALP, COL1, DMP-1, and DSPP) were detected and quantified by western blotting, and (d) ALP staining and Alizarin Red S staining were performed with the treatment of miR-3074-3p mimics or inhibitors (Scale bar = 300 μm). Data were presented as means ± SD, n = 3/group, *p < 0.05. **p < 0.01. ***p < 0.001.
Cellular delivery and biocompatibility of PEI-AuNPs to antagomiR-3074-3p
To achieve the continuous function of antagomiR-3074-3p in vivo, we selected PEI-AuNPs as the carrier for miRNA to enhance the miRNA’s ability to penetrate the cell membrane and exert its effects in this study. The synthesized PEI-AuNPs were approximately 30 nm in size under TEM (Figure 3(a)). The agarose gel electrophoresis result showed that both antagomiR-NC and antagomiR-3074-3p combined with PEI-AuNPs completely at the ratio of 3:2 or higher (Figure 3(b)). The result suggested that PEI-AuNPs could combine with antagomiR-NC or antagomiR-3074-3p at relatively low volume.

Fabrication of antagomiR/PEI-AuNPs and their biocompatibility: (a) the TEM images of PEI-AuNPs (scale bar = 50 nm), (b) agarose gel electrophoresis analysis of atagomiR-NC and antagomiR-3074-3p combined with PEI-AuNPs, (c) confocal microscopy images of hDPSCs after treatment with pure Cy3-antagomiR-3074-3p, Cy3-atagomiR-NC/PEI-AuNPs, and Cy3-antagomiR-3074-3p/PEI-AuNPs (scale bar = 50 μm), (d) the proliferation assay of hDPSCs after treatment with lipo2000, atagomiR-NC/PEI-AuNPs, and antagomiR-3074-3p/PEI-AuNPs, and (e) cell scratch assay of hDPSCs after treatment with lipo2000, atagomiR-NC/PEI-AuNPs, and antagomiR-3074-3p/PEI- AuNPs (scale bar = 250 μm). Data were presented as means ± SD, n = 3/group, *p < 0.05. **p < 0.01. ***p < 0.001, ns means no significance.
To confirm the successful cellular delivery of antagomiR-NC/PEI-AuNPs and antagomiR-3074-3p/PEI-AuNPs, they were marked with red fluorescence cy3 before transfection. The images of the laser confocal microscope revealed that cells treated with free Cy3-antagomiR-3074-3p showed no red fluorescence, while red fluorescence could be observed in cells treated with Cy3-antagomiR-NC-conjugated PEI-AuNPs or Cy3-antagomiR-3074-3p-conjugated PEI-AuNPs (Figure 3(c)). The result indicated that antagomiR-NC and antagomiR-3074-3p could be taken up by hDPSCs with relatively high efficiency with the aid of PEI-AuNPs.
To assess the cytocompatibility of antagomiR/PEI-AuNPs in vitro, we conducted cell proliferation assay and cell scratch assay. The cell proliferation assay showed that compared to traditional transfection reagents lipo2000, antagomiR-NC/PEI-AuNPs and antagomiR-3074-3p/PEI-AuNPs did not exhibit cytotoxicity at the working concentration (Figure 3(d)). The cell scratch assay showed that compared to the blank group, cells in antagomiR-NC/PEI-AuNPs and antagomiR-3074-3p/PEI-AuNPs groups exhibited a similar migration and proliferation rate while the lipo2000 group was slower at the gaps (Figure 3(e)). These results indicated that antagomiR-NC-conjugated PEI-AuNPs or antagomiR-3074-3p-conjugated PEI-AuNPs exhibited satisfactory cytocompatibility, making them promising for biomedical applications.
The promotive function of antagomiR-3074-3p/PEI-AuNPs in odontogenic differentiation of hDPSCs in vitro
To confirm the effects of antagomiR-3074-3p-conjugated PEI-AuNPs in vitro, we treated hDPSCs with antagomiR-NC/PEI-AuNPs and antagomiR-3074-3p/PEI-AuNPs continuously during mineralization induction. RT-qPCR and Western blot assays revealed that odontogenic markers were significantly upregulated by transfecting antagomiR-3074-3p/PEI-AuNPs at both mRNA and protein levels (Figure 4(a)–(c)). Moreover, antagomiR-3074-3p/PEI-AuNPs increased the ALP activity in ALP staining and deposition of mineralized nodules in Alizarin Red S staining (Figure 4(d)). These results suggested that antagomiR-3074-3p-conjugated PEI-AuNPs could promote the odontogenic differentiation of hDPSCs in vitro.
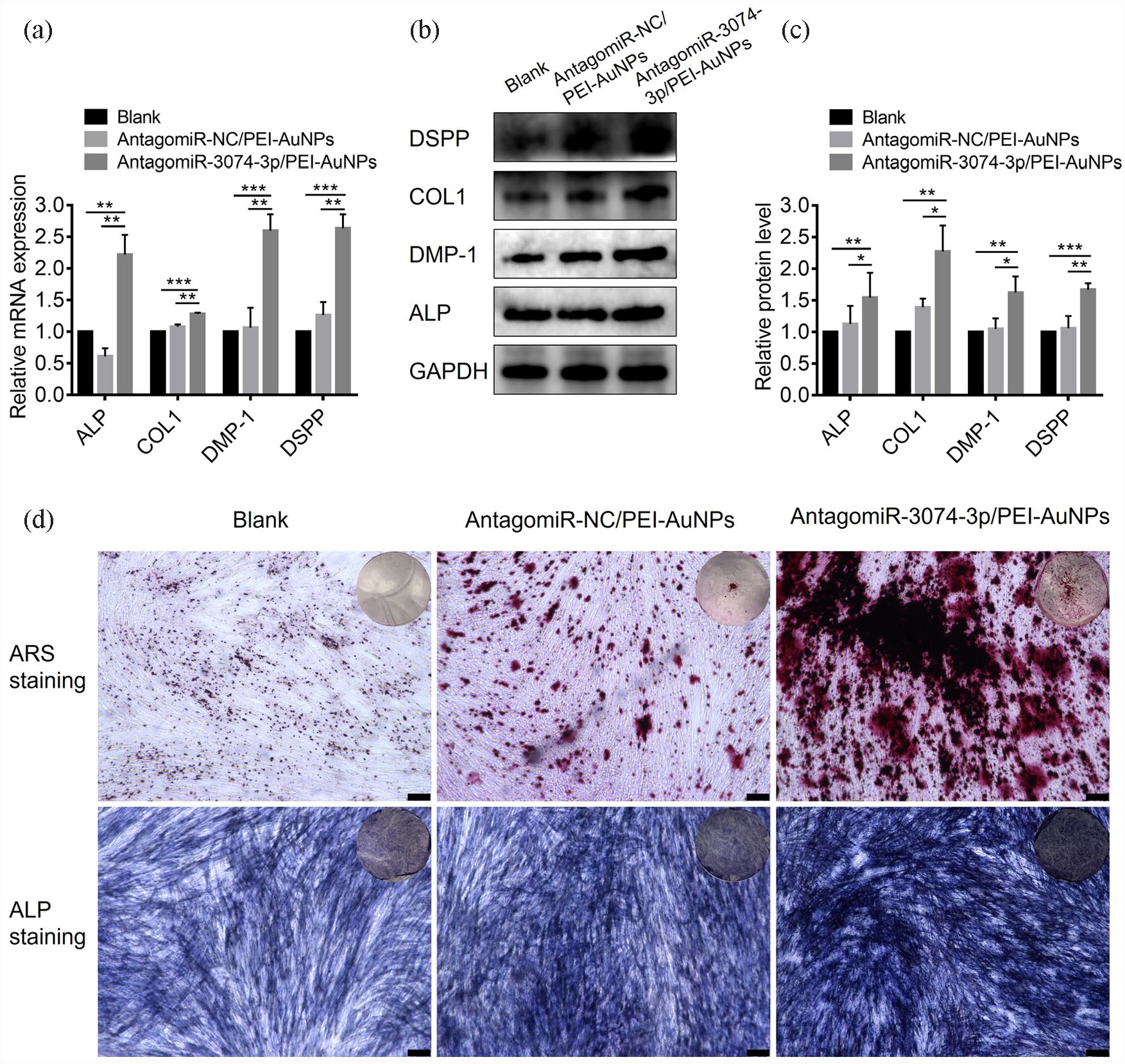
AntagomiR-3074-3p/PEI-AuNPs had a promotive effect on odontogenic differentiation of hDPSCs in vitro: (a) QRT-PCR analysis of mRNA levels of the odontogenic marker (ALP, COL1, DMP-1, and DSPP) in hDPSCs treated with antagomiR-NC/PEI-AuNPs or antagomiR-3074-3p/PEI-AuNPs, (b and c) western blotting and quantification analysis of protein levels of the odontogenic marker (ALP, COL1, DMP-1, and DSPP) in hDPSCs treated with antagomiR-NC/PEI-AuNPs and antagomiR-3074-3p/PEI-AuNPs, (d) ALP staining and Alizarin Red S staining were performed with the treatment of antagomiR-NC/PEI-AuNPs and antagomiR-3074-3p/PEI-AuNPs (scale bar = 250 μm). Data were presented as means ± SD, n = 3/group, *p < 0.05. **p < 0.01. ***p < 0.001.
AntagomiR-3074-3p/PEI-AuNPs/GELMA complex had promotive function in dentin restoration in vivo
To achieve sustained release of antagomiR-3074-3p/PEI-AuNPs in animal experiments, we chose to distribute them in GELMA. Biocompatible and three-dimensionally structural GELMA enabled continuous release of antagomiR-3074-3p/PEI-AuNPs in corresponding parts, thereby realizing their function. 29 We conducted the release experiment to determine the sustained-release effect of GELMA. The result showed that antagomiR-3074-3p/PEI-AuNPs or antagomiR-NC/PEI-AuNPs could achieve a slow release of approximately 80% within 7 days (Figure 5(a)). Previous studies have reported that GELMA can release more than 80% rapidly within 2 days in vitro. 30 The addition of gold nanoparticles extended the release period, thereby promoting the continuous action of antagomiR-3074-3p.

AntagomiR-3074-3p/ PEI-AuNPs/ GELMA could promote dentin formation in a rat pulp-capping model: (a) the release curve of cy3-antagomiR-NC/PEI-AuNPs or cy3-antagomiR-3074-3p/PEI-AuNPs from GELMA in 7 days, and (b–d) hematoxylin–eosin-stained sections of rat tooth perforation capped with GELMA, PEI-AuNPs/GELMA, antagomiR-NC/PEI-AuNPs/GELMA and antagomiR-3074-3p/PEI-AuNPs/GELMA complexes throughout 1, 4, and 8 weeks (4×, scale bar = 250 μm; 10×, scale bar = 100 μm). Arrow, the tooth perforation; PD, primary dentin; RD, reparative dentin. Data were presented as means ± SD, n = 3/group, *p < 0.05. **p < 0.01. ***p < 0.001, ns means no significance.
Then, we conducted the rat pulp-capping experiments. The results showed that when the rat tooth was opened only, there was no dentin repair at the tooth perforation, leading to gradual pulp necrosis in a large area with less secondary dentin formation in the pulp cavity within 8 weeks (Figure 5(d)). Although GELMA group showed less pulp necrosis, there was still no effective dentin repair at the tooth perforation in 8 weeks (Figure 5(d)). Loading of PEI-AuNPs led to reduced inflammation in 1 week (Figure 5(b)) and a small amount of mineralized deposit in 4 weeks (Figure 5(c)) at tooth perforation. Moreover, a significant increase in dentin formation was observed at the tooth perforation at 8 weeks in PEI-AuNPs/GELMA group and antagomiR-NC/PEI-AuNPs/GELMA group (Figure 5(d)). Further conjugation of antagomiR-3074-3p resulted in relatively complete dentin repair at the tooth perforation and closure of the tooth perforation at 4 and 8 weeks (Figure 5(c) and (d)). The newly formed dentin contained a stripe that distinguished it from the original dentin structure, indicating the formation of third dentin. Probably due to the long duration of the experiment, all experimental groups exhibited more secondary dentin formation and calcification in the pulp cavity, which reduced the pulp cavity volume. However, the pulp perforating hole exhibited the most complete dentin formation in antagomiR-3074-3p/ PEI-AuNPs/GELMA group. These results demonstrated the potential of antagomiR-3074-3p-conjugated PEI-AuNPs in promoting dentin restoration in vivo.
FKBP9 acted as the target gene of miR-3074-3p
To identify potential targets of miR-3074-3p, we used MIRDB, MirWalk, and TargetScan databases combined with the results of chipdegs. Three targets (FKBP9, Cav1, and Crebrf) were predicted to be regulated by miR-3074-3p (Figure 6(a)). Combined with high-throughput chip results, FKBP9 exhibited the most significant difference among odontoblasts in TSC1 group and NC group (Figure 6(b)). Additionally, the network database has predicted partial complementary sequences between the FKBP9 and miR-3074-3p (Figure 6(c)). Therefore, FKBP9 might be a potential target of miR-3074-3p.

FKBP9 acted as the target gene of miR-3074-3p: (a) the prediction of the target gene of miR-3074-3p from TargetScan, MIRDB, and MirWalk databases combined with the results of chipdegs, (b) comparison of expression of FKBP9 between NC group and TSC1 group in odontoblasts, (c) schematic diagram of the complementary sequence between the FKBP9 3′-UTR and miR-3074-3p, (d) luciferase reporter assay of co-transfection of miR-3074-3p and two types of FKBP9 3′-UTR, (e) the expression levels of the FKBP9 gene in hDPSCs after transfected with miR-3074-3p mimics or inhibitors were determined by qRT-PCR, (f and g) Western blotting determined and quantified the expression levels of FKBP9 protein in hDPSCs after being transfected with miR-3074-3p mimics or inhibitors. Data were presented as means ± SD, n = 3/group, *p < 0.05. **p < 0.01. ***p < 0.001.
To determine whether miR-3074-3p targets FKBP9 directly, we conducted the luciferase assay. The result revealed that miR-3074-3p repressed the FKBP9 significantly while mutations of the putative miR-3074-3p site abrogated responsiveness to miR-3074-3p (Figure 6(d)). Subsequently, the potential association between expression levels of FKBP9 and miR-3074-3p was verified through transfection of miR-3074-3p mimic or inhibitor to hDPSCs. The result showed that endogenous FKBP9 mRNA and protein levels in hDPSCs were reduced or enhanced (Figure 6(e)–(g)). Taken together, these results demonstrated that miR-3074-3p was capable of inhibiting the expression of FKBP9 in hDPSCs.
FKBP9 has a promotive effect on odontogenic differentiation of hDPSCs
To clarify the effect of FKBP9 on the odontogenic differentiation of hDPSCs, we designed FKBP9-enhanced lentivirus to transfect hDPSCs. Lentivirus empty vector and FKBP9 enhanced lentivirus were all transfected successfully into hDPSCs, as shown in fluorescence images (Figure 7(a)). The transfection results showed that mRNA and protein levels of FKBP9 in hDPSCs could be successfully enhanced by transfection of FKBP9-enhanced lentivirus (Figure 7(b)–(d)). At the same time, the results of mineralization induction after transfection showed that the mRNA and protein levels of odontogenic markers in hDPSCs were significantly increased by transfection of FKBP9-enhanced lentivirus (Figure 7(e)–(g)). Similar results were observed in the evaluation of ALP staining and Alizarin Red S staining (Figure 7(h)). These results suggested that overexpression of FKBP9 promoted odontogenic differentiation of hDPSCs.
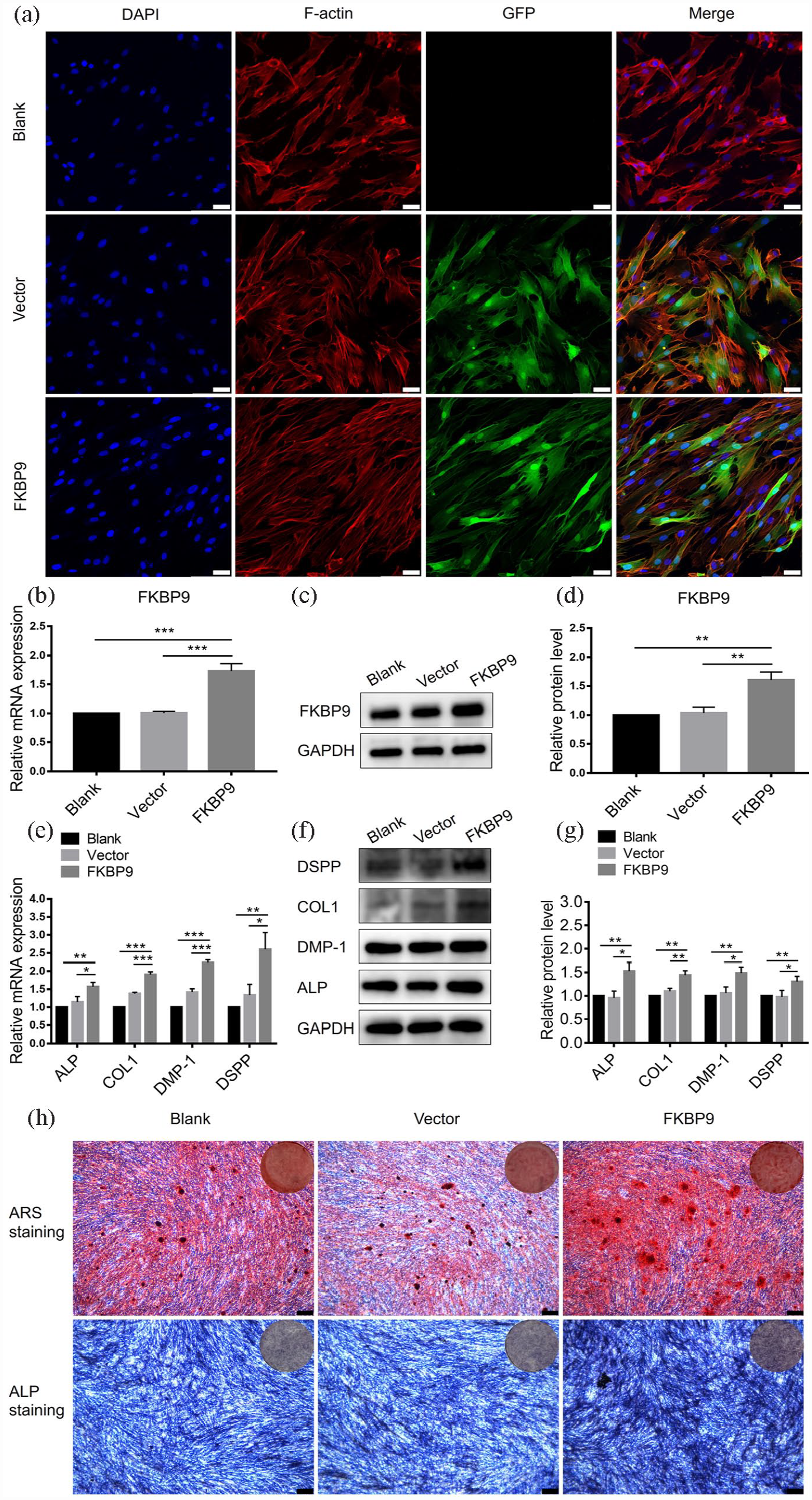
FKBP9 had the promotive effect of odontogenic differentiation of hDPSCs: (a) confocal microscopy images of hDPSCs after treatment with lentivirus empty vector or FKBP9 expression plasmid (scale bar = 25 μm), (b) after treatment with lentivirus empty vector or FKBP9 expression plasmid, the expression levels of the FKBP9 gene in hDPSCs were detected by qRT-PCR, (c and d) the expression levels of FKBP9 protein in hDPSCs were detected and quantified by western blotting, (e) the expression levels of the odontogenic marker genes (ALP, COL1, DMP-1, and DSPP) in hDPSCs were detected by qRT-PCR after treatment with lentivirus empty vector or FKBP9 expression plasmid, (f and g) the expression levels of the odontogenic marker protein (ALP, COL1, DMP-1, and DSPP) were determined by quantified Western blotting, (h) ALP staining and Alizarin Red S staining were performed with the treatment of lentivirus empty vector or FKBP9 expression plasmid (scale bar = 250 μm). Data were presented as means ± SD, n = 3/group, *p < 0.05. **p < 0.01. ***p < 0.001.
MiR-3074-3p inhibited the odontogenic differentiation of hDPSCs by repressing FKBP9
To further elucidate whether miR-3074-3p inhibited odontogenic differentiation of hDPSCs by targeting FKBP9, we conducted the rescue experiment. FKBP9-enhanced lentivirus was transfected to hDPSCs treated with miR-3074-3p mimics. As the mRNA and protein expression level of the odontogenic marker indicated, the decrease in odontogenic differentiation induced by miR-3074-3p mimics treatment was effectively rescued by overexpression of FKBP9 (Figure 8(a)–(c)). Similar results were observed in the evaluation of ALP staining and Alizarin Red S staining (Figure 8(d)). These results illustrated that miR-3074-3p inhibited the odontogenic differentiation of hDPSCs by repressing FKBP9.
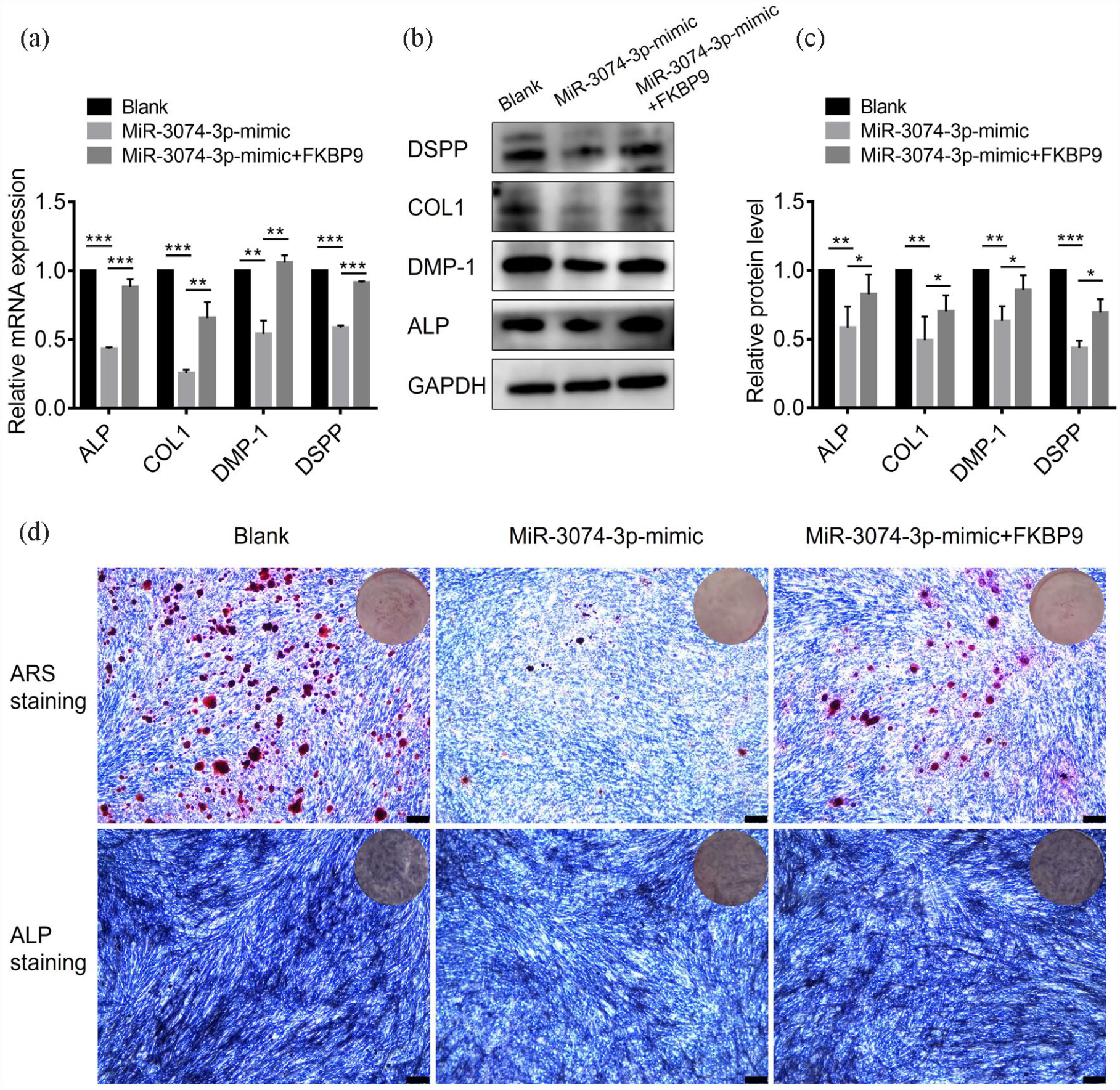
MiR-3074-3p inhibited the odontogenic differentiation of hDPSCs by repressing FKBP9: (a) QRT-PCR analysis of mRNA levels of the odontogenic marker (ALP, COL1, DMP-1, and DSPP) in hDPSCs treated with miR-3074-3p mimics or miR-3074-3p mimics combined with lentivirus FKBP9 expression plasmid, (b and c) Western blotting and quantification analysis of protein levels of the odontogenic marker (ALP, COL1, DMP-1, and DSPP) in hDPSCs treated with miR-3074-3p mimics or miR-3074-3p mimics combined with lentivirus FKBP9 expression plasmid, (d) ALP staining and Alizarin Red S staining were performed with the treatment of miR-3074-3p mimics or miR-3074-3p mimics combined with lentivirus FKBP9 expression plasmid (scale bar = 250 μm). Data were presented as means ± SD, n = 3/group, *p < 0.05. **p < 0.01. ***p < 0.001.
Discussion
DPSCs have been considered suitable seed cells for tissue engineering for years. 31 So far, DPSCs have been proven to have other multipotent differentiation potentials, such as osteoblasts, myoblasts, epithelial cells, neural cells, and adipocytes.32–34 Furthermore, DPSCs play an important role in tooth development and restoration due to their ability to differentiate into odontoblasts. The application of functional molecules to enhance the odontogenic differentiation potential of DPSCs is the key to achieving effective tooth restoration and regeneration.
A previous study by our team indicated that mTOR pathway activation significantly enhanced dentin formation. 27 In this study, we conducted further RNA sequencing to investigate the potential involvement of miR-3074-3p in the odontogenic differentiation of hDPSCs. MiR-3074, first reported in 2015, 35 was mainly involved in tumor development, cell metabolism, and tissue repair. Previous studies have reported that miR-3074-5p was mainly involved in the occurrence and development of the reproductive process.36,37 In addition, miR-3074-3p was reported to participate in the inflammatory response and repair of nerve injury.38,39 However, its role in odontogenesis remained less explored. A number of evidences indicated that miRNAs were closely related to the metabolism of stem cells. 40 In this regard, miR-3074-3p was reported to regulate the process of myoblast differentiation and had the potential to improve muscle regeneration. 41 Therefore, the potential of miR-3074-3p in tooth regeneration could not be ignored. Our study showed that treatment with miR-3074-3p mimic or inhibitor could considerably change odontogenic markers in hDPSCs, such as ALP, COL-1, DMP-1, and DSPP, at both mRNA and protein levels. ALP activity in ALP staining and deposition of mineralized nodules in Alizarin Red S staining also changed remarkably. These results suggested that miR-3074-3p was involved in the odontogenic differentiation of hDPSCs, which conformed to the biological characteristics of miRNA, representing a new function of miR-3074-3p.
Although many studies have confirmed the effect of miRNAs on odontogenic differentiation of hDPSCs,3,7–9 these studies mainly focused on in vitro studies, with few experiments conducted in vivo. The formation of reparative dentin by odontoblasts differentiated from mesenchymal stem cells in dental pulp is the basis of the repair function of the pulp-dentin complex. Swanson et al. confirmed that reparative dentin formation at the tooth opening could be used to explore the odontogenesis function of small molecules in vivo. 28 Gold nanoparticles had good biocompatibility and were suitable for carrying various small molecule functions. It has been reported that miRNAs could promote bone formation when loaded with gold nanoparticles.15–17 Carrying miRNA through gold nanoparticles was a reasonable strategy for tooth regeneration. Nevertheless, whether miRNA could promote dentin formation while loading gold nanoparticles remained unclear. In this study, we synthesized PEI-AuNPs that could carry miRNAs. AntagomiR-3074-3p-conjugated PEI-AuNPs not only effectively enhanced the mRNA and protein expression level of odontogenic markers in hDPSCs in vitro but also promoted the formation of restorative dentin in rat tooth perforation under the condition of slow release from GELMA. Severe pulp necrosis occurred in the absence of pulp capping material, resulting in no dentin formation at the pulp opening. With the addition of gold nanoparticles, there was a tendency to restore dentin at the pulp opening, although it was still incomplete. AuNPs have been reported to increase the osteogenic differentiation of hMSCs and positively affect the osteogenic differentiation of periodontal ligament stem cell (PDLSC) sheets.42,43 Consistent with this evidence, our results implied that gold nanoparticles had certain osteogenic functions in stem cells. After the addition of antagomiR-3074-3p, complete restorative dentin was formed at the pulp opening in 4 and 8 weeks, indicating the promotive function of antagomiR-3074-3p-conjugated PEI-AuNPs in dentin formation in vivo. Taken together, these results not only strongly illustrated the odontogenesis effect of antagomiR-3074-3p/PEI-AuNPs in vivo but also proposed antagomiR-3074-3p-conjugated PEI-AuNPs as an effective strategy for dentin restoration.
As a small molecule with regulatory function, miRNA regulates metabolism mainly by regulating the expression of target genes. Lee et al. 41 demonstrated that miR-3074-3p promoted myoblast differentiation by targeting Cav1. In our study, we first found that FKBP9 participated in the odontogenic differentiation of hDPSCs, and it was proved to be the target for miR-3074-3p to regulate the odontogenic differentiation of hDPSCs. As a member of FKBPs, FKBP9 has been shown to participate in various metabolic processes. Li et al. 44 reported that FKBP9 might be involved in the growth and metabolism of chickens when promoted by roxarsone. Brown et al. 45 reported that FKBP9 played a significant role in protein folding, and deficiency of FKBP9 led to susceptibility to neuropathy and neurodegenerative diseases. This evidence suggested that FKBP9 was closely related to protein metabolism. Besides, numerous studies have shown that the high expression of FKBP9 was closely related to the occurrence and development of glioblastoma multiforme with high levels of protein metabolism.46–48 However, the role of FKBP9 in the odontogenic differentiation of hDPSCs and its relationship with miR-3074-3p remains unexplored. In this study, through FKBP9-enhanced lentiviral transfection in vitro, the expression level of odontogenic markers was improved, and ALP activity in ALP staining and deposition of mineralized nodules in Alizarin Red S staining were raised. Furthermore, the overexpression of FKBP9 rescued the effect of miR-3074-3p. These results indicated that FKBP9, as the target of miR-3074-3p, regulated the odontogenic differentiation of hDPSCs. It could be speculated that FKBP9 might somehow promote the expression of odontogenic proteins. Interestingly, Xu et al. pointed out that FKBP9 deletion would accumulate misfolded or unfolded proteins and cause UPR (unfolded protein response), further leading to endoplasmic reticulum stress and cell death. 48 Despite this, few related studies have investigated the role of FKBP9 in endoplasmic reticulum stress, and whether FKBP9 promoted odontogenic differentiation of hDPSCs through this mechanism needed further exploration.
In this study, we reported for the first time the involvement of miR-3074-3p in the odontogenic differentiation of hDPSCs. In vivo experiments revealed that antagomiR-3074-3p-conjugated PEI-AuNPs in GELMA effectively restored dentin, thereby presenting potential therapeutic applications in dentin restoration and tooth regeneration. In addition, our findings highlighted the crucial role of FKBP9 in the odontogenic differentiation of hDPSCs for the first time, providing a foundation for further studies on stem cell differentiation related to FKBP9. However, research on miR-3074-3p was still limited, and whether FKBP9 regulates the metabolism of hDPSCs through endoplasmic reticulum stress required further investigations.
Conclusion
In summary, the present study demonstrated that miR-3074-3p played a vital role in regulating the odontogenic differentiation of hDPSCs by targeting FKBP9. Furthermore, we found that antagomiR-3074-3p/AuNPs exhibited remarkable odontogenic potential in vitro and in vivo. Our findings will provide a new strategy and a potential target for DPSCs regarding dentin restoration or tooth regeneration.
Supplemental Material
sj-docx-1-tej-10.1177_20417314231184512 – Supplemental material for Enhanced effects of antagomiR-3074-3p-conjugated PEI-AuNPs on the odontogenic differentiation by targeting FKBP9
Supplemental material, sj-docx-1-tej-10.1177_20417314231184512 for Enhanced effects of antagomiR-3074-3p-conjugated PEI-AuNPs on the odontogenic differentiation by targeting FKBP9 by Tao Jiang, Shenghong Miao, Jingjie Shen, Wenjing Song, Shenglong Tan and Dandan Ma in Journal of Tissue Engineering
Footnotes
Acknowledgements
We appreciate each participant providing assistance in this study.
Author contributions
Tao Jiang and Shenghong Miao participated in the design of this study, and they both carried out the study. Tao Jiang wrote and revised the manuscript. Jingjie Shen performed material preparation. Wenjing Song guided the preparation of the material. Shenglong Tan guided the revision of the manuscript. Dandan Ma guided the design and conduction of this study and manuscript editing. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (81970930).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
