Abstract
Background:
Interleukin-7 receptor (IL7R) mutation has been demonstrated to be an adverse prognostic factor in acute lymphoblastic leukemia (ALL) patients. However, the effects of the IL7R mutation on acute myeloid leukemia (AML) have rarely been reported. Here, we investigated IL7R mutations and their effects on AML patients.
Methods:
A total of 346 newly diagnosed AML patients from January 2017 to July 2020 at Nanfang Hospital were analyzed in this study. A genomic panel of 167 gene targets was detected by next-generation sequencing.
Results:
Among 346 patients, 33 (9.5%) AML patients carried IL7R mutations. With a median follow-up of 50.7 months (95% confidence interval (CI) 17.3–62.2), the 5-year overall survival (OS) rates were 51.5% (95% CI 37.0%–71.0%) and 72.2% (95% CI 67.4%–77.3%; p = 0.008), the 5-year event-free survival (EFS) rates were 36.1% (95% CI 23.2%–57.1%) and 58.1% (95% CI 52.9%–63.8%; p = 0.005), the 5-year non-relapse mortality (NRM) were 21.4% (95% CI 8.5%–38.2%) and 6.2% (95% CI 3.7%–9.5%; p = 0.004) in the IL7R mutant (IL7R MUT ) group and non-IL7R mutant (IL7R WT ) group, respectively. There is no significant difference in the disease-free survival (75.1% vs 73.5%, p = 0.885) and cumulative incidence of relapse (25.7% vs 25.2%, p = 0.933) between IL7R MUT and IL7R WT group. Furthermore, patients who underwent hematopoietic stem cell transplantation (HSCT) still had more adverse outcomes in the IL7R MUT group than in the IL7R WT group (5-year OS: 61.9% vs 85.3%, p = 0.003). In the TET2 (p = 0.013) and DNA methyltransferase 3A (DNMT3A; p = 0.046) mutation subgroups, the presence of IL7R mutations was associated with worse OS than in AML patients without IL7R mutations.
Conclusion:
Our study demonstrated that the IL7R mutation is associated with an inferior prognosis for AML patients. Patients with IL7R mutations have higher NRM, shorter OS, and EFS than patients without IL7R mutations, even patients who have undergone HSCT. Future larger and multicentric prospective studies will be explored.
Plain language summary
With the development of NGS, more and more cytogenetic and molecular markers have been found to be associated with prognosis of ALL. IL7R mutation is associated with an inferior prognosis for AML patients. Patients with IL7R mutation have higher NRM, shorter OS and EFS than patients without IL7R mutation, even patients who have undergone HSCT.
Keywords
Background
Acute myeloid leukemia (AML) is a clonal malignant proliferative disease of myeloid blasts in the hematopoietic system.1–3 With the development of next-generation sequencing (NGS), an increasing number of cytogenetic and molecular markers have been found to be associated with the pathogenesis and prognosis of AML.4–9 European Leukemia Net (ELN), a genetic risk stratification, has been widely used in the clinical and plays a pivotal role in guiding appropriate treatment. 6 However, several studies have suggested that the clinical outcomes are heterogenous within the ELN risk groups and may be affected by other co-existing genetic mutations.10,11 Previous study demonstrated that patients with DNA methylation regulatory gene mutations (DNMT3A, IDH1/2, TET2, SETBP1) was shorter in overall survival (OS) rate than patients without DNA methylation regulatory gene mutations in ELN favorable group, particularly in those with NPM1 mutant. 12 In addition, the prognosis of patients with single mutations and patients with multiple mutations are different. “Patients with isolated nucleophosmin 1 (NPM1)” have higher complete remission (CR) rates, and better relapse-free survival (RFS) and OS.13–15 However, patients with NPM1 and DNMT3A co-mutation significantly have worse OS and leukemia-free survival compared to isolated NPM1-mutated patients.16–18 It has been demonstrated that CEBPA mutations that are in-frame mutations affecting the basic leucine zipper region (bZIP) confer a favorable outcome, irrespective of their occurrence as biallelic (CEBPAbi) or single mutation (CEBPAsm). There is no definitive conclusion about the impact of co-mutations. Tarlock et al. found that the presence of a colony-stimulating factor 3 receptor (CSF3R) mutation in patients with CEBPA mutations is associated with a remarkably high relapse risk and poor event-free survival (EFS).6,19–22 Therefore, more prognostic markers need to be urgently explored to guide optimal treatments.10,11,20
The interleukin-7 receptor (IL7R), a heterodimer consisting of the specific IL7Rα chain (also known as CD127) and the common γ-chain (also known as CD132), are mainly expressed in hematopoietic cells, including T-cells, B cells, NK cells, and innate lymphoid cells. IL7R are required for the normal T-cell development and homeostasis of mature T-cells. IL7R mutational activation is one of the drivers of acute lymphoblastic leukemia (ALL).23–27 Gain-of-function mutations in IL-7R have been identified in 7%–10% of T-cell acute lymphoblastic leukemia (T-ALL) patients and approximately 2%–3% of B-cell acute lymphoblastic leukemia (B-ALL) patients.28–30 Most IL7R mutations are located in exon 6 in T-ALL and B-ALL patients. In addition, IL7R exon 5 mutations have also been found in a few patients with B-ALL.23,31 A correlation of IL7R mutations with prognosis in ALL has been sporadically reported. Alsadeq et al. demonstrated that the IL7R mutation was an adverse prognostic factor in pediatric B-cell precursor ALL with higher central nervous system infiltration and relapse. Besides, the IL7R mutation in the relapse of T-ALL was correlated with worse survival.32,33 However, the effects of the IL7R mutation on AML have rarely been reported. In adult AML, Kim et al. reported that IL7R exon6 mutation was around 1% in AML. 31 A case report showed that an IL7R mutation was found in a secondary AML patient with a dismal clinical course. 34 Therefore, we conducted a large retrospective study to explore IL7R mutations and their effects on AML patients.
Methods
Patients and data collection
This study examined patients with newly diagnosed AML between January 2017 and July 2020 at Nanfang Hospital. The clinical data cutoff date was August 31, 2023. Patients who met the following criteria were analyzed: (1) aged between 14 and 80 years, (2) diagnosed with de novo AML or secondary AML, and (3) NGS data available at diagnosis.
Cytogenetic and molecular analyses
Cytogenetic analyses were performed with Giemsa and reverse banding techniques and fluorescence in-situ hybridization (FISH). Conventional cytogenetic karyotyping was processed by standard G-banding or R-banding cytogenetic methods. Karyotypes were classified according to the International System for Human Cytogenetic Nomenclature (ISCN 2020). A FISH analysis is performed according to the standard protocol, using the following probes: PML/RARA(15Q22;17Q21.1), aml1/eto(8q22;21q22), MLL(11q23), and CBFβ (16q22). NGS was used for molecular analyses. A genomic panel of 167 gene targets was detected in bone marrow (BM) samples by NGS at the time of diagnosis (Supplemental Table S1). NGS was carried out on the Ion Torrent platform (in the target rate of 97%–99%, with an average depth of 1000×, the average is 94%–97%). The reference sequence was used in the human genome GRCh37.35,36 Risk groups were assigned using the 2022 ELN risk stratification scheme.
Treatment protocol
Generally, patients receive “3 + 7” standard induction therapy, and patients who are unfit for standard induction chemotherapy receive lower-intensity induction therapy, including VA (venetoclax, azacitidine) and D-CAG (decitabine, cytarabine, aclarubicin, granulocyte colony-stimulating factor).37–39 After achieving CR, patients received cytarabine-based consolidation chemotherapy and/or bridged to auto-SCT/allo-SCT (Stem Cell Transplantation) based on Minimal Residual Disease (MRD) status and donor availability. 40 Patients who did not achieve CR received salvage chemotherapy, including CAG (aclarubicin, cytarabine, granulocyte colony-stimulating factor), FLAG (fludarabine, cytarabine, granulocyte colony-stimulating factor), and VAH (venetoclax, azacitidine, homoharringtonine).41–43
Evaluation, definitions, and statistics
This study focused mainly on OS, EFS, disease-free survival (DFS), cumulative incidence of relapse (CIR), and non-relapse mortality (NRM). OS was measured from the date of diagnosis until death or censored at the last follow-up. EFS was defined as the time from the date of diagnosis until documented failure to achieve CRc, relapse after CRc, or death from any cause, whichever occurred first. CRc comprised CR and CRi (defined as all the criteria for CR, except for neutropenia or thrombocytopenia). Relapse was defined by morphologic evidence of the original hematologic disease in the peripheral blood, BM, or any extramedullary site. DFS was defined as the time from CR to relapse or death from any cause. NRM was defined as death from any cause not subsequent to relapse.4,6,20,44,45
Comparisons of continuous variables between groups were conducted using the Mann–Whitney U test, while comparisons of categorical variables between groups were performed using the Chi-square test or Fisher’s exact test. OS, EFS, and DFS were compared using the Kaplan–Meier analysis with the log-rank test. Hazard ratios (HRs) and 95% confidence interval (95% CI) were calculated using the Cox proportional hazards models. The CIR and NRM were adjusted for the competing risk analysis. Competing events were defined as follows: for relapse, NRM; for NRM, relapse. The Cox proportional hazards model was used for the analysis of risk factors for time-to-event variables. Only variables with a p-value less than 0.05 were included in the multivariable analysis. The correlations among various mutations were analyzed by the “ggcorrplot” package. Mutation status and frequency of genetic abnormalities were analyzed by the “oncoplot” package. A forest plot with HR and 95% CI was a simple and intuitive description of relative risks between IL7R mutation and concomitant genetics. All the statistical tests were two-tailed with a significance level of 0.05. Statistical analyses were conducted using GraphPad Prism 7, SPSS version 24.0, and R statistical software (version 4.1.1).
Results
Patient baseline characteristics and IL7R mutations
Total of 346 newly diagnosed AML patients from Nanfang Hospital from January 2017 to July 2020 were analyzed in this study. There were 324 de novo AML and 22 secondary AML in the cohort, including 18 patients progressing from myelodysplastic syndrome. There were 186 men and 160 women, with a median age of 38 years (range: 14–77). IL7R mutations were observed in 33 patients (9.5%). The median age at diagnosis in the IL7R MUT group was older than that in the IL7R WT group (p = 0.031). Except for age, other baseline characteristics were similar between the two groups (Table 1). There were 21.2% for ELN favorable, 36.4% for intermediate-risk, and 42.4% for adverse-risk among patients with IL7R mutation. We further explore the correlation between IL7R mutations and other gene mutations with mutation rates more than 5%. We found co-occurrence of IL7R mutations with DNA methyltransferase 1 (DNMT1; p = 0.007) and CSF3R (p = 0.039), but no other genetic mutations in co-occurrence of IL7R mutations (Figure 1 and Supplemental Table S2).
Baseline characteristics (n = 346).
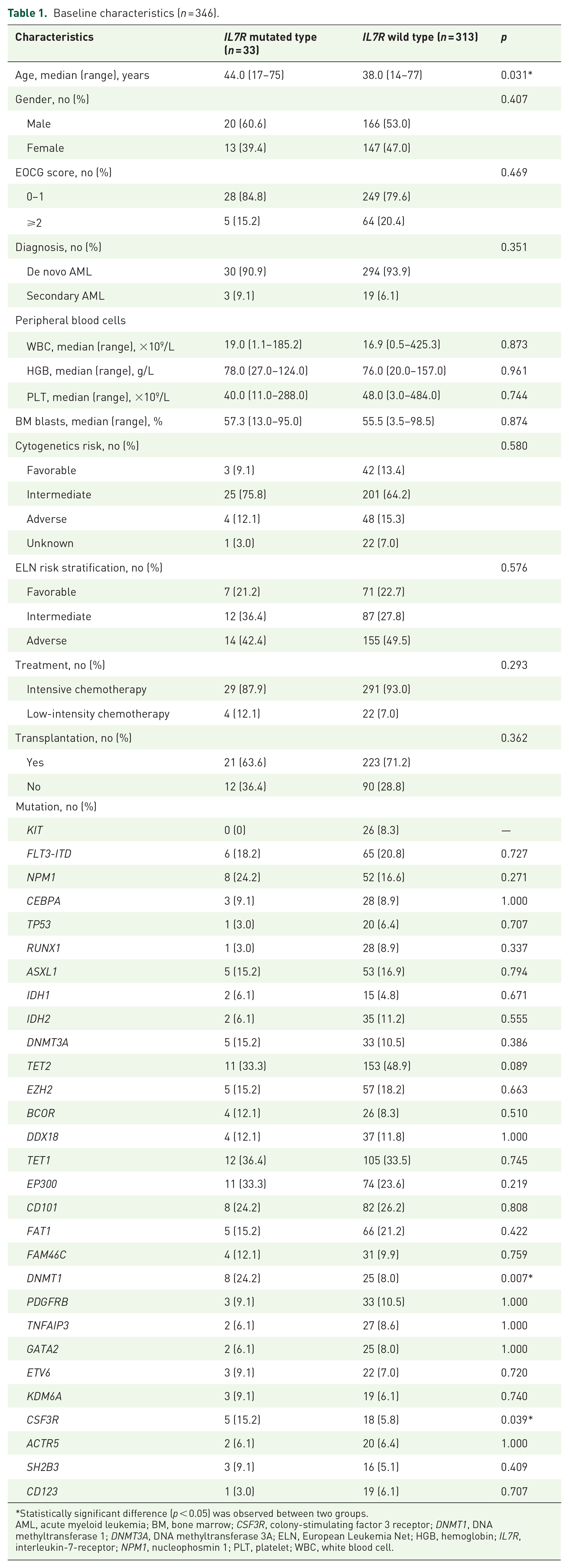
Statistically significant difference (p < 0.05) was observed between two groups.
AML, acute myeloid leukemia; BM, bone marrow; CSF3R, colony-stimulating factor 3 receptor; DNMT1, DNA methyltransferase 1; DNMT3A, DNA methyltransferase 3A; ELN, European Leukemia Net; HGB, hemoglobin; IL7R, interleukin-7-receptor; NPM1, nucleophosmin 1; PLT, platelet; WBC, white blood cell.

Correlation matrix for genetic mutations in AML patients (mutation frequency ⩾5%). Some co-occurrence mutations were negatively related, represented in blue, and others were positively related, represented in red. The darker the color, the higher the correlation was (p < 0.05).
Survival
With a median follow-up of 50.7 months (interquartile range (IQR) 17.3–62.2; 95% CI 46.5–54.3) the median OS was 24.9 months (IQR 7.9–62.0; 95% CI 9.0–60.5) in the IL7R MUT group and 50.6 months (IQR 21.8–64.1; 95% CI 46.9–54.0) in the IL7R WT group, corresponding to a 5-year OS of 51.5% (95% CI 37.0%–71.7%) for the IL7R MUT group versus 72.2% (95% CI 67.4%–77.3%) for the IL7R WT group (HR 2.0, 95% CI 1.2–3.5; p = 0.008, Figure 2a). The median EFS was 8.9 months (IQR 3.4–60.3; 95% CI 5.3–54.4) in the IL7R MUT group compared with 43.0 months (IQR 8.1–59.8; 95% CI 40.6–46.9) in the IL7R WT group, corresponding to a 5-year EFS of 36.1% (95% CI 23.2%–57.1%) for the IL7R MUT group versus 58.1% (95% CI 52.9%–63.8%) for the IL7R WT group (HR 1.9, 95% CI 1.2–3.1 p = 0.005, Figure 2b). The 5-year DFS was 75.1% (95% CI 59.6%–94.7%) in the IL7R MUT group and 73.5% (95% CI 68.3%–79.0%) in the IL7R WT group (HR 1.1, 95% CI 0.5–2.4 p = 0.885, Figure 2c). In the IL7R MUT and IL7R WT groups, the 5-year CIR was 25.7% (95% CI 10.8%–42.2%) and 25.3% (95% CI 20.2%–30.4%), respectively (p = 0.933, Figure 2d) and the 5-year NRM was 21.4% (95% CI 8.5%–38.2%) for the IL7R MUT group and 6.2% (95% CI 3.7%–9.5%) for the IL7R WT group (p = 0.004, Figure 2e). A multivariable analysis revealed that non-hematopoietic stem cell transplantation (HSCT), poorer prognostic stratification, and IL7R mutation were risk factors for OS and EFS. And poorer cytogenetic stratification, lower score of ECOG, and IL7R mutation were risk factors for NRM (Table 2).
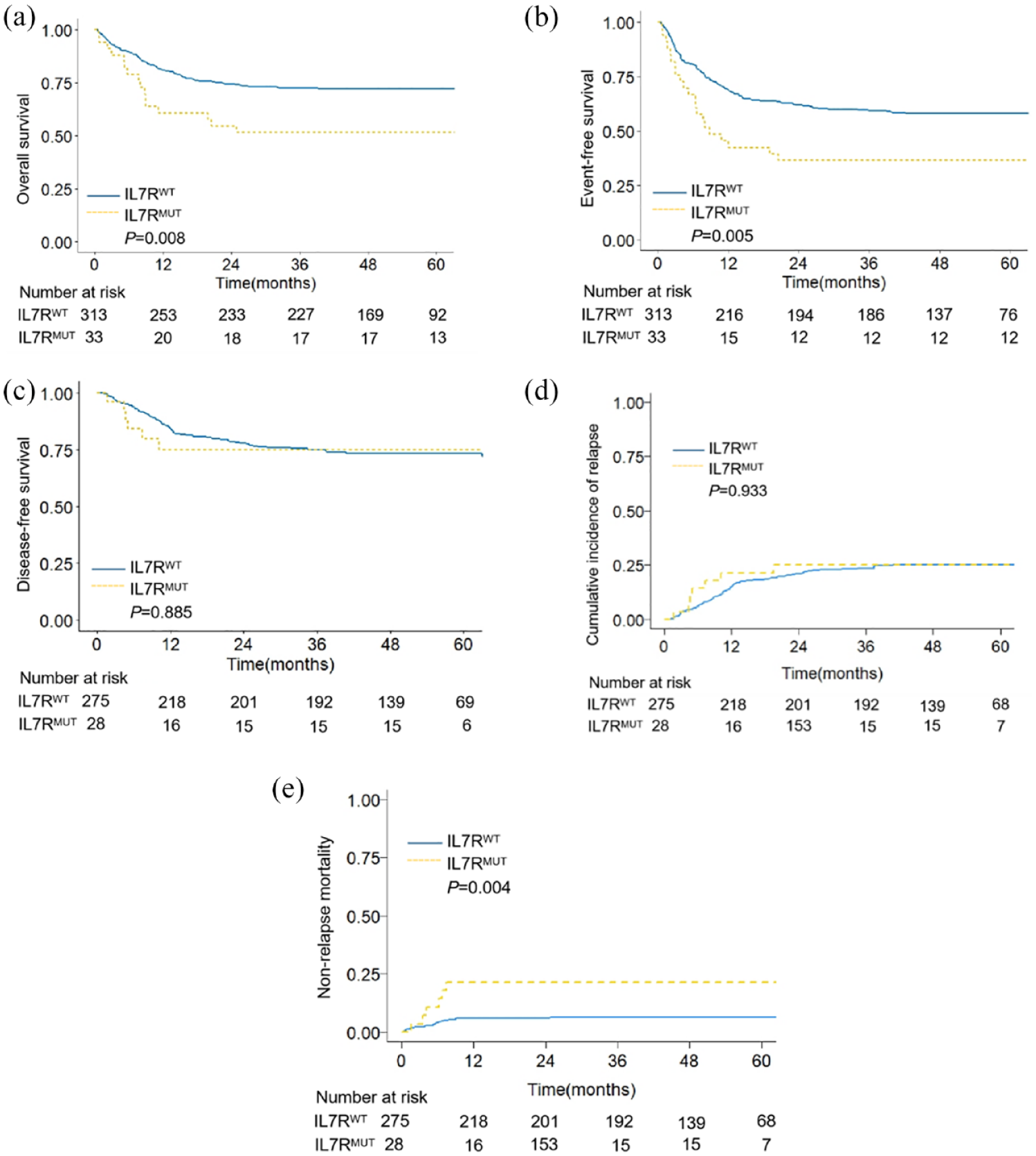
Survival between IL7R MUT and IL7R WT group among AML patients. (a) Overall survival, (b) event-free survival, (c) disease-free survival, (d) cumulative incidence of relapse, and (e) non-relapse mortality of the IL7R MUT group and IL7R WT group.
Uni- and multivariate analyses of overall survival, event-free survival, and non-relapse mortality in AML patients.
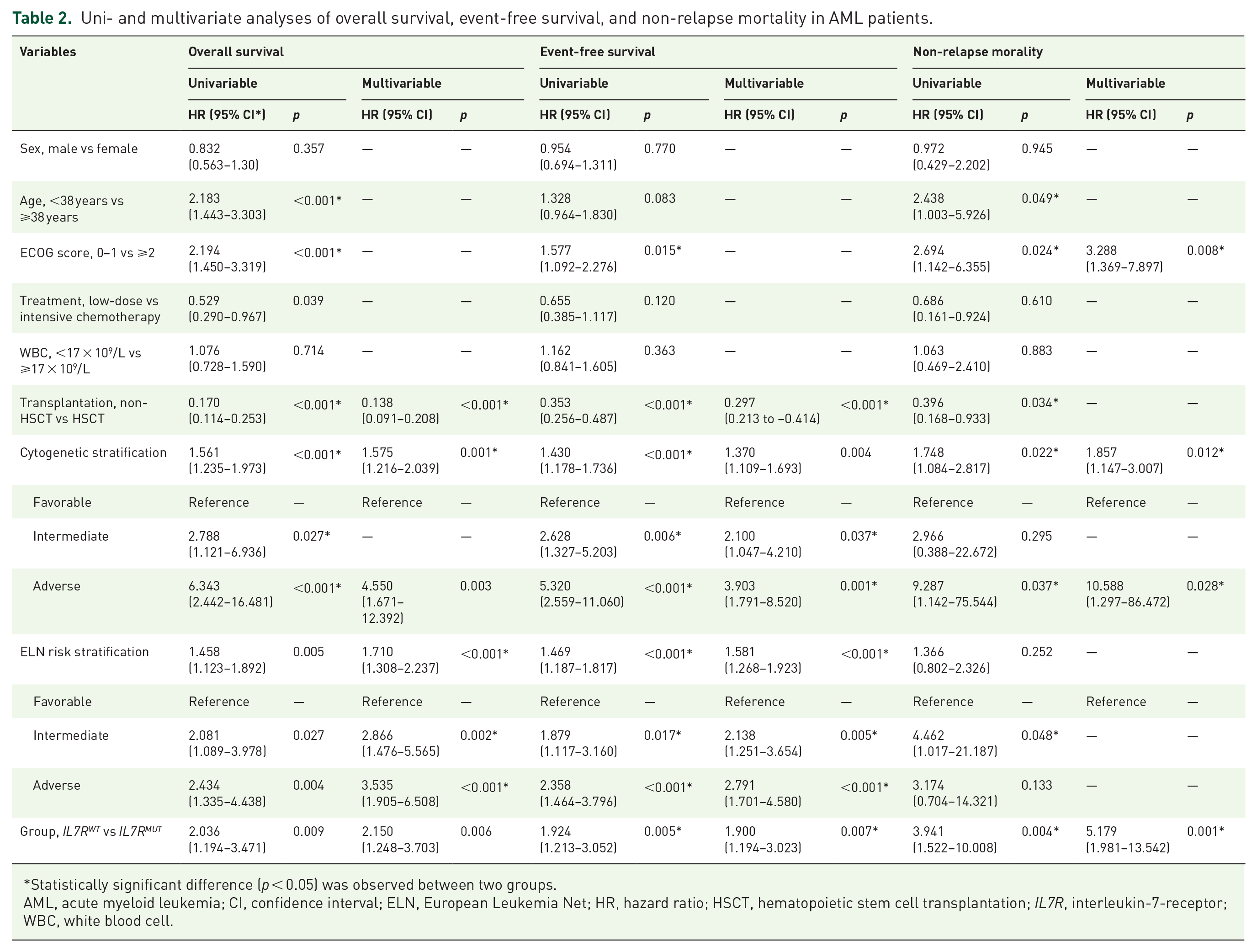
Statistically significant difference (p < 0.05) was observed between two groups.
AML, acute myeloid leukemia; CI, confidence interval; ELN, European Leukemia Net; HR, hazard ratio; HSCT, hematopoietic stem cell transplantation; IL7R, interleukin-7-receptor; WBC, white blood cell.
To explore the impact of age and treatment intensity on outcome, we do further subgroup analysis. It demonstrated that patients with IL7R mutation had poorer prognosis 5-year OS and EFS in the subgroup of age<60 and intensive treatment. There is no significance in 5-year OS, EFS in the subgroup of age ⩾60 and low-dose treatment (Supplemental Table S3).
Among 346 AML patients, 103 (29.8%) patients died—16 in the IL7R MUT group and 87 in the IL7R WT group. In the IL7R MUT group, the most common cause of death was infection (8 of 16 patients, 50.0%), which accounted for 37.5% for pneumonia cases and 43.8% for septicemia cases. In the IL7R WT group, the incidence of infection was 26.4%, including 20.7% of pneumonia and 17.2% of septicemia. Infection-related mortality in patients with IL7R mutation was higher than non-IL7R mutation (p = 0.003, Table 3).
Causes of death in AML patients (n = 103).

Statistically significant difference (p < 0.05) was observed between two groups.
AML, acute myeloid leukemia; GVHD, graft-versus-host-disease; IL7R, interleukin-7-receptor; TMA, thrombotic microangiopathy.
Effects of the IL7R mutation in AML patients receiving HSCT
Among 346 AML patients, 245 received HSCT. The baseline characteristics of the AML patients who underwent HSCT were similar between the IL7R MUT and IL7R WT groups (Supplemental Table S4). Among AML patients receiving HSCT, patients with IL7R mutations still had poorer outcomes than patients in the IL7R WT group. With a median follow-up of 49.6 months (IQR 35.8–56.6; 95% CI 47.6–51.6) for patients who underwent HSCT, the 5-year OS was 61.9% (95% CI 44.3%–86.6%) for the IL7R MUT group versus 85.3% (95% CI 80.7%–90.0%) for the IL7R WT group (HR 3.1, 95% CI 1.4–6.6; p = 0.003, Figure 3a). The 5-year EFS was 57.1% (95% CI 39.5%–82.8%) for the IL7R MUT group versus 72.6% (95% CI 67.0%–78.7%) for the IL7R WT group (HR 1.8, 95% CI 0.9–3.7; p = 0.083, Figure 3b). The 5-year DFS was 82.6% (95% CI 66.6%–100.0%) in the IL7R MUT group and 77.2% (95% CI 71.8%–83.1%) in the IL7R WT group (HR 1.732, 95% CI 0.3–2.7 p = 0.757, Figure 3c). In the IL7R MUT and IL7R WT groups, the 5-year CIR was 19.0% (95% CI 5.6%–38.4%) and 21.5% (95% CI 16.4%–27.2%), respectively (p = 0.790, Figure 3d). The 5-year NRM was 23.8% (95% CI 8.3%–43.6%) for the IL7R MUT group versus 5.8% (95% CI 3.2%–9.4%) for the IL7R WT group (p = 0.003, Figure 3e). A multivariable analysis showed that poorer prognostic stratification and IL7R mutation were risk factors for OS and NRM among patients who underwent HSCT. Poorer prognostic stratification and non-CR before HSCT were risk factors for OS and EFS. Besides, age >34 years was a risk factor for NRM (Supplemental Table S5).

Survival between IL7R MUT and IL7R WT group among AML patients undergoing HSCT. (a) Overall survival, (b) event-free survival, (c) disease-free survival, (d) cumulative incidence of relapse, and (e) non-relapse mortality of the IL7R MUT group and IL7R WT group.
Among the 244 AML patients who underwent HSCT, 41 patients died—8 in the IL7R MUT group and 33 in the IL7R WT group. In the IL7R MUT group, the most common cause of death was infection (37.5%) compared with that in the IL7R WT group (27.3%). Among patients receiving HSCT, there was also a trend of higher infection-related mortality in the IL7R MUT group than in the IL7R WT group (p = 0.072, Supplemental Table S6).
Among the 33 AML patients with IL7R mutations, 21 patients (21/33, 63.6%) received HSCT. Except for ages, other baseline characteristics were similar between the two groups (Supplemental Table S7). Among patients with IL7R mutations, 5-year OS rates were 61.9% (95% CI 44.3%–86.6%) for patients who underwent HSCT and 33.3% (95% CI 15.0%–74.2%) for patients who did not receive HSCT (p = 0.032), and the 5-year EFS rates were 38.1% (95% CI 22.1%–65.7%) and 33.3% (95% CI 15.0%–74.2%), respectively (p = 0.301, Figure 4 and Supplemental Table S7). HSCT might partially overcome the poor prognosis of patients with IL7R mutations.
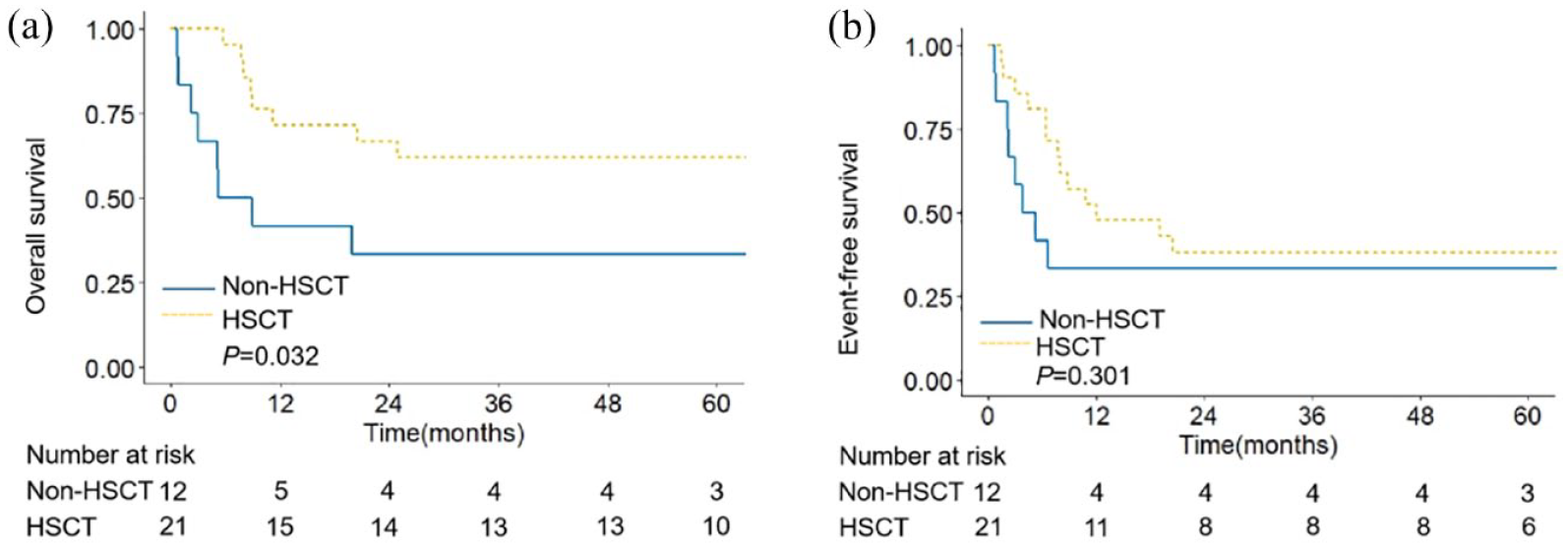
Survival between HSCT and non-HSCT group among AML patients with IL7R mutation. Overall survival (a) and event-free survival (b) analysis among 31 AML patients with IL7R mutation.
Effects of the IL7R mutation on concomitant genetics
We identified a total of 167 gene targets by NGS at the time of diagnosis. The detailed molecular mutation in which mutation rates were more than 5% is shown in Figure 5. For gene mutations, a total of 338 patients harbored at least one concomitant mutation. We further explored the effects of IL7R mutation on concomitant genetics in which the mutation rate was more than 8%. A forest plot showed that the IL7R mutation was a risk factor in patients with TET2 (p = 0.018) and DNMT3A mutation (p = 0.043, Figure 6). In TET2 and DNMT3A subgroups, the presence of IL7R mutation was associated with worse OS than in the IL7R WT patients (p < 0.05, Supplemental Figure 1). A tendency toward lower survival was observed in the NPM1 MUT /IL7R MUT group than in the NPM1 MUT /IL7R WT group (p = 0.073).

Mutation status and frequency of genetic abnormalities that detected in ⩾5% the patients. Mutations are listed on the left and frequency is listed on the right, with the percent of patients for each type of genetic abnormality, including activated signaling (green), DNA methylation (orange), chromatin modifiers (pink), myeloid transcription factors (blue), transcription (purple), and other (brown).

The effects of IL7R mutation in subgroups. Cox proportional hazard model for prognostic impact of IL7R mutation and concomitant genetics on overall survival in the cohort of 346 AML patients. The logarithm of HR is shown in the graph.
Discussion
Our study demonstrated that the IL7R mutation was an adverse prognostic factor for AML patients. Patients with IL7R mutations had significantly shorter OS and EFS and higher NRM than patients without IL7R mutations, even patients who underwent HSCT. The presence of IL7R co-mutation in the TET2 and DNMT3A subgroups was associated with decreased survival.
Somatic gain-of-function mutations in IL7R have been shown to act as oncogenes in T- and B-ALL.28,29 In our study, we analyzed 346 adult patients with newly diagnosed AML and identified mutations in IL7R in 9.5% (33/346). Approximately 10% of T-ALL patients are reported to have IL7R mutations, the poor prognosis might be due to the overactivation of the JAK/STAT and PI3K/AKT/mTOR pathways which are associated with glucocorticoid resistanc.23,29,46 Activation of the JAK/STAT pathway and PI3K/Akt/mTOR pathway led to the downregulation of p27Kip1 and the up-regulation of Bcl2, which promoted the proliferation and survival of T-ALL cells. IL7R mutation related to poor prognosis of ALL, Xiao et al. and Fu et al. showed that the IL7R mutation was associated with poor clinical outcomes in adult ALL patients.47–49 Richter-Pechańska et al. reported that pediatric T-ALL patients with IL7R mutations had lower EFS compared to IL7R wild-type patients. Li et al. showed that mutations in IL7R and its pathway-related genes such as JAK/RAS/AKT have lower RFS, which are associated with steroid resistance in pediatric T-ALL.32,46 However, the effect of IL7R mutation on AML pathogenesis and its related mechanisms have not been reported. In our study, we first demonstrated that IL7R mutation had a negative impact on survival among AML patients. The 5-year OS rate was 51.5% in the IL7R MUT group versus 72.2% in the IL7R WT group, corresponding to the 5-year EFS rate was 36.1% in the IL7R MUT group versus 58.1% in the IL7R WT group. The 5-year NRM for the IL7R MUT group was higher than IL7R WT group. The most common cause of death was infection. Infection-related mortality in patients with IL7R mutation was higher than in patients without IL7R mutation. The mechanism of IL7R mutation in infection remains unclear. Some polymorphisms of the IL7Rα were associated with immunodeficiency and inflammatory diseases.50–52 Ampuero et al. further reported that single nucleotide polymorphisms of IL7R would be related to the severity of adults with community-acquired pneumonia (CAP). 53 It had been reported that polyglutamylation and deglutamylation of IL-7Rα tightly controlled the development and effector functions of ILC3s, which promoted lymphoid organogenesis and potentiated immune responses against bacterial infection.54,55 Therefore, the mechanism of poor prognosis in IL7R mutation patients might be due to abnormal lymphocyte dysfunction and reduced anti-infection and anti-tumor immune function.
HSCT has been proven to be the most effective therapy for AML patients. However, among AML patients undergoing HSCT, patients with IL7R mutation still had poorer outcomes compared with patients in IL7R WT group. Our findings are in accordance with the previous study, showing that HSCT cannot improve OS in IL7-receptor pathway mutated (IL7Rp MUT ) T-ALL patients compared with IL7-receptor pathway non-mutated (IL7Rp WT ) T-ALL patients. 56 Patients with IL7R mutations who received HSCT had better OS than those who did not receive HSCT (4-year OS: 60.2% vs 31.2%, p = 0.028). We found that HSCT might partially overcome the poor prognosis of AML patients with IL7R mutation.
In our study, among TET2 MUT and DNMT3A MUT AML patients, patients with IL7R mutations had shorter OS than patients in the IL7R WT group. DNA methylation regulatory gene mutations such as those in DNMT3A, TET2, IDH1, and IDH2 have been shown to be associated with poor prognosis.12,16,57 It is reported that DNMT3A was identified to negatively regulate the SOCS5 expression levels and SOCS5 downregulation potentiates the expression of IL7R, JAK-STAT signal transduction, and leukemia progression. In our study, patients with TET2/IL7R or DNMT3A/IL7R co-mutation have a poorer prognosis. The reason might be due to aberrant DNA methylation (hypermethylation) indirectly leads to hyperactivation of JAK/STAT signaling.58–62
There is strong therapeutical potential to target the IL-7/IL-7R pathway in T-ALL. For example, the use of downstream signaling elements JAK inhibitors, Bcl-2 drug inhibitors, and the reducing agent n-acetylcysteine (NAC) have been shown to be effective in the treatment of T-ALL.63–65 These targeted drugs also might have the potential therapeutic value in AML, which will need to be further explored.
It has been reported that IL7R mutations are located in exon 5 and exon 6 in T-ALL and B-ALL. Limited by the depth and sensitivity of NGS technology in our study, we detected mutation sites of IL7R were only covered exon 5 and exon 6. In our study, all AML patients with IL7R mutations were located in exon 6 (c.731C>T; p.T244I). Besides, this is small sample size, monocentric, and retrospective study. Future large and multicenter prospective studies will be further explored.
Conclusion
Our study first demonstrates that the IL7R mutation is associated with an inferior prognosis for AML patients. Patients with IL7R mutations have poorer outcomes than those without IL7R mutation, even patients who have undergone HSCT. The presence of IL7R mutations was associated with higher NRM, shorter OS, and EFS than in AML patients without IL7R mutations.
Supplemental Material
sj-docx-1-tah-10.1177_20406207241279533 – Supplemental material for Prognostic impact of IL7R mutations on acute myeloid leukemia
Supplemental material, sj-docx-1-tah-10.1177_20406207241279533 for Prognostic impact of IL7R mutations on acute myeloid leukemia by Qiqi Tao, Qiaoyuan Wu, Yutong Xue, Changkun Chen, Ya Zhou, Ruoyang Shao, Haiyan Zhang, Hui Liu, Xiangzong Zeng, Lingling Zhou, Qifa Liu and Hua Jin in Therapeutic Advances in Hematology
Footnotes
Appendix
Acknowledgements
We thank the patients, their families, and their caregivers; coinvestigators, collaborators, and members of the study team involved in this study. We express our gratitude to all members of the Department of Hematology, Nanfang Hospital, Southern Medical University for their support.
Declarations
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
