Abstract
Tick-borne encephalitis is an important human arbovirus neuroinfection spread across the Northern Eurasia. Inhibitors of tick-borne encephalitis virus (TBEV) strain Absettarov, presumably targeting E protein n-octyl-β-
Keywords
Introduction
Tick-borne encephalitis (TBE) is an important human arbovirus neuroinfection spread across the Northern Eurasia.1,2 It is caused by tick-borne encephalitis virus (TBEV) from the Flavivirus genus. Despite the availability of several vaccines, TBE incidence is high among unvaccinated people; therefore, a significant threat exists for travelers going to popular destinations in Europe and Siberia.1,3 The risk of TBEV infection among unvaccinated travelers to highly endemic regions is estimated to be 1/10,000. 4 There are no developed, approved, and widely accepted methods of post-onset TBE therapy, and directly acting small molecule drugs are required as a therapeutic option and post-tick bite prophylaxis.
TBEV is divided into three subtypes (European, Eu; Siberian, Sib; Far-Eastern, FE), 5 and two phylogenetically separate groups (Baikalian and Buryat-Mongol). 6 Recently, a new Himalayan subtype was also proposed. 7 TBEV-Eu is the most common in the continental Europe. TBEV-Sib is prevalent on the Russian territory. 6 TBEV-FE circulates in China, Japan, Mongolia, and the Russian Far-East. 5 All TBEV subtypes co-circulate in Russia and in the Baltics.6,8 Subtypes are rather closely related genetically, and the variation of polyprotein sequences between subtypes is about 2–7%.5,6,9 Clinical manifestations vary from asymptomatic to encephalitis and death for all the TBEV subtypes. 1
Certain small molecule TBEV inhibitors were recently suggested and tested against Eu strains Neudoerfl and Hypr10,11 or Absettarov.12–14 We reported several series of potent TBEV reproduction inhibitors15,16 identified with the help of molecular docking against the n-octyl-β-
In the present work, we demonstrated an antiviral activity of 4-aminopyrimidine N-oxides against a panel of strains belonging to all main TBEV subtypes. Compounds varied in their antiviral activity against strains Absettarov and 256, which did not differ in E protein amino acid sequence. We observed variation of the ratio of infectious virions versus decoy E protein containing particles, including non-infectious and immature virus particles, E protein aggregates and membrane complexes, in the virus sample. These non-infection particles could influence observed activity values of small molecules targeting envelope proteins.
Materials and methods
Sequence analysis
E protein sequences were obtained from GenBank via Pubmed Protein interface (access date 26.10.2015) using search string “(tick-borne encephalitis) AND ((E protein) OR (envelope protein))” and lower sequence length limit of 240 amino acid residues. TBEV strain DV936k sequence (GenBank accession no. GU125722.1) was updated and uploaded to GenBank later (GenBank accession no. GU125722.2). Multiple sequence alignment was performed via Clustal Omega web service19,20 with default parameters (Supplementary alignment file). Sequences lacking residues 46 to 285 were excluded and substitutions in the pocket region compared to the TBEV strain Absettarov were analyzed using an in-house Python 3.5 script.
Cells and viruses
Porcine embryo kidney (PEK) cell line was maintained at 37°C in media 199 on Hanks’ balanced salt solution and Earle’s balanced salt solution (2:1, v:v, FSBSI “Chumakov FSC R&D IBP RAS”, Russia) supplemented with 5% fetal bovine serum (FBS, Invitrogen).
Vero cell line was maintained at 37°C in DMEM media with L-glutamine (FSBSI “Chumakov FSC R&D IBP RAS”, Russia) supplemented with 10% fetal bovine serum (FBS, Invitrogen).
TBEV strains used in the work are presented in Table 1.21,22 TBEV strains from laboratory collection were multiplied in PEK and/or Vero cells. Cultural fluids were collected 72 h post infection (cytopathic effect over 75% of cell monolayer), aliquoted, and stored at −70°C. Each strain sample was titrated by plaque assay in PEK cells at least three times to determine geometrical mean titer of infectious virions and by qPCR at least three times to determine the mean concentration of RNA-containing particles.
TBEV strains used in the study.

aM – passages in mouse brain (Mx – passages performed by strain authors before the viruses were obtained in the laboratory); V – passages in Vero cells; P – passages in PEK cells.
4-Aminopyrimidine N-oxides
The compounds were synthesized from commercially available starting materials by previously described methods16,23,24 and stored as solutions in DMSO at −20°C.
Plaque assay
Ten-fold dilutions of virus suspension in medium 199 made on Earle’s balanced salt solution (FSBSI “Chumakov FSC R&D IBP RAS”, Russia) were added to PEK cell monolayers in two replicates and incubated in a CO2-incubator for 1 h at 37°C. Then cells were overlaid with 1.26% methylcellulose (Sigma) prepared in media 199 on Hanks’ balanced salt solution and Earle’s balanced salt solution (2:1, v:v) containing 2% FBS. After incubation at 37°C in a CO2-incubator for six days, cells were fixed with 96% ethanol. Plaques were stained with 0.4% gentian violet. Plaques were counted and the virus titer was calculated as log10PFU/mL. Each strain was titrated in at least three biological replicates to determine the mean concentration of infectious virions.
Plaque reduction assay
Two-fold dilutions of studied compounds with concentrations starting from 50 µM were prepared in medium 199 made on Earle's balanced salt solution. Equal volumes of virus suspension, containing 20–35 PFU, were added to compound dilution (1:1, v:v). The control virus was added to the same sequential concentrations of DMSO, as in compounds dilutions. Virus-compound mixtures were incubated at 37°C in a CO2-incubator for 1 h and then added to the cells, seeded on 24-well plates, in two replicates and incubated at 37°C in a CO2-incubator for 1 h. Then the cells were overlaid with methylcellulose and incubated for six days, and plaques were stained as described for the plaque assay. EC50 values were calculated according to Reed-and-Muench method. 25 At least two biological replicates were performed to determine each compound activity.
ELISA
The content of E protein in virus samples was assessed by enzyme-linked immunosorbent assay (ELISA) using commercial kit D1154 VectorTBE-antigen (Vector-Best, Russia) according to the manufacturer’s instructions.
The standard curve was calculated for purified standard antigen prepared from TBEV strain Sofjin 26 (ST1, Figure S1), obtained by Dr. V. N. Lyapustin as described elsewhere. 27
To determine the amount of E protein in the standard ST1, two-fold dilutions of bovine serum albumin (BSA, Combithek) with mass range from 1 to 0.01 mg/mL were mixed with 3×Sample buffer (SDS-mercaptoethanol) and resolved in 12% SDS-PAGE at MiniPROTEAN cell (Bio-Rad). Gel was fixed in water solution containing 10% of acetic acid and 50% of ethanol, stained by Coomassie blue (Figure S2), and scanned. Optical density of protein bands was estimated in OneDscan (DSP Inc., USA). A calibration curve of bands’ optical density from protein concentration was built based on BSA dilutions, and was used to determine the amount of total E protein content in the standard sample ST1. The procedure was repeated in three independent experiments.
To obtain a standard calibration curve for E protein concentration from optical density of samples, series of two-fold ST1 dilutions was measured in ELISA with VectorTBE-antigen (Vector-Best, Russia) according to the manufacturer’s instructions (Figure S1).
The standard calibration curve was used to estimate the amount of E protein in the virus samples by their optical density measured in ELISA. Optical density of six two-fold dilutions of each sample was studied in at least two biological replicates.
The amount of E protein from infectious particles and genome-containing particles (GCP) was estimated assuming average molecular mass of TBEV E protein of 53 kDa and 180 protein molecules per viral particle.
qRT-PCR
qRT-PCR procedure was performed as described earlier28,29 with some changes. Sabin1 strain of poliovirus type 1 was used as an internal control and was added to the samples prior to RNA extraction. Reverse transcription was performed with M-MLV reverse transcriptase (Promega, USA) according to the manufacturer’s protocol with primers Pow-TBE-3′: 5′-
Statistical analysis
Mann–Whitney two-tailed U-test was applied to compare strain 256 sample’s characteristics to the other strain samples. The threshold of statistical significance was set at p ≤ 0.05. Analysis was performed with an in-house Python 3.5 script.
Results
Selection of TBEV strains
To investigate the spectrum of anti-TBEV activity of 4-aminopyrimidine N-oxides, we selected seven TBEV strains representing all three main TBEV subtypes from the laboratory collection. These strains were isolated in various regions of Russia and the Baltics and had a different passage history (Table 1). Strain sequencing and characterization have been presented previously.21,22
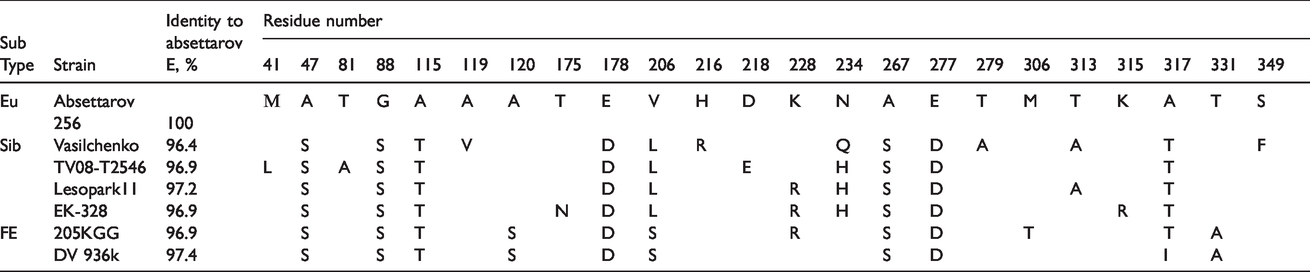
Our previous studies showed that 4-aminopyrimidine N-oxides would most likely interact with the ectodomain of the virus envelope protein E. Thus, we compared the protein E sequences of the selected TBEV strains. Variable amino acid positions are presented in Table 2 and in Figure S3. Strain 256 did not contain amino acid substitutions in the E protein compared to strain Absettarov, thus allowing direct comparison of the compounds’ activity against the strains with identical E proteins.
Comparison of the sE protein sequences of the TBEV strains used in activity spectrum study. All variable residues are shown.
Antiviral activity spectrum
The spectrum of anti-TBEV activity of nine 4-aminpyrimidine N-oxides, which had shown EC50 < 50 µM against strain Absettarov in our previous study in the focus reduction assay in PEK cells, 16 was assessed against selected strains in the plaque reduction assay in the same cells against 20–35 PFU. The EC50 values are shown in Table 3.
Anti-TBEV activity spectrum of 4-aminopyrimidine N-oxide
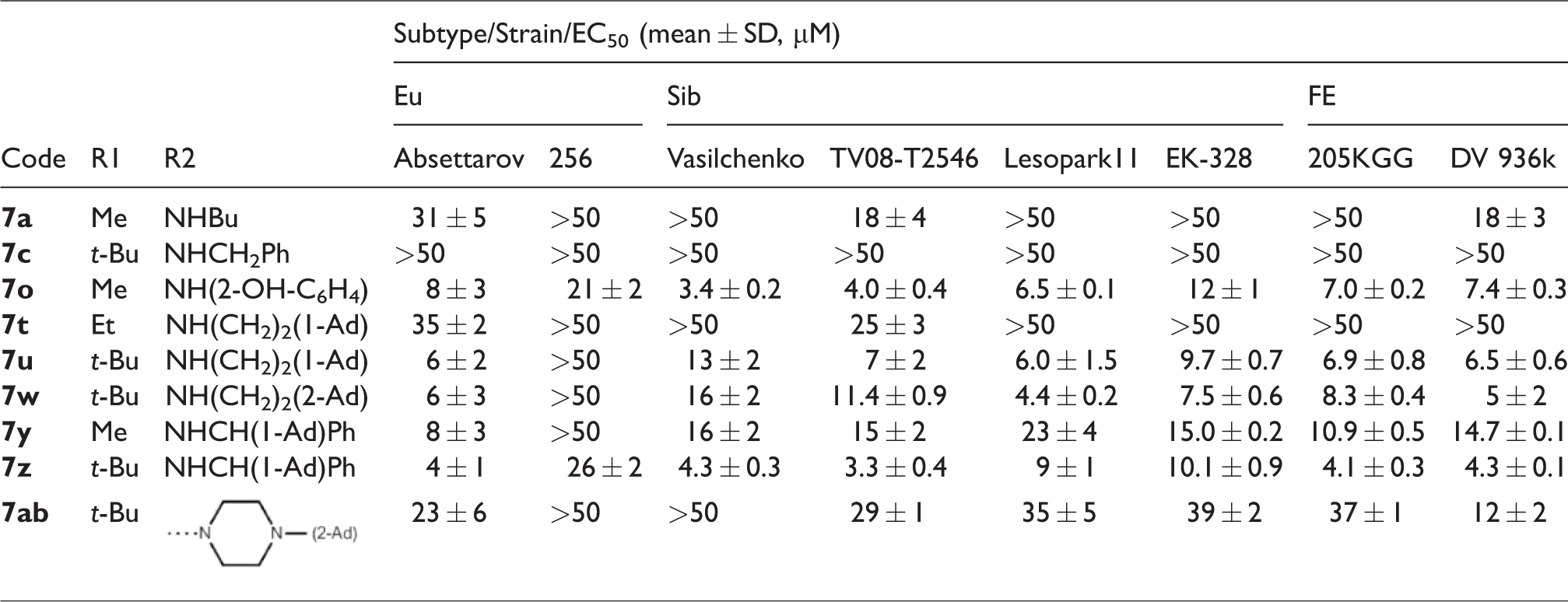
The majority of the studied compounds showed a broad spectrum of activity, inhibiting the reproduction of strains belonging to all main TBEV subtypes. Compound
All strains except 256 bore differences in the amino acid sequence of protein E; however, inhibitory activity of the most potent compounds against these strains was at the same level. Compounds
E protein sequence variation
Our analysis of the TBEV strain Absettarov envelope protein soluble ectodomain (sE) homology model 16 revealed that the following 34 residues should be considered as comprising the pocket: 46–55, 128–132, 140, 142, 199, 201, 204–206, 211, 212, 214, 273–275, 278–280, 282, 284, 285. To assess the variability of β-OG pocket, we analyzed full and partial TBEV E sequences from GenBank (Supplementary alignment file) containing residues 46 to 285 (E protein numbering), which comprised the whole pocket (Figure S4). The sequence of strain Absettarov E protein was used as a reference. Of 570 retrieved sequences, 168 (30%) did not contain amino acid substitutions in the pocket region, and six were identical to strain Absettarov E protein residues 1 to 395, comprising the whole soluble ectodomain sE. The remaining 402 sequences had no more than four substitutions in a limited number of positions at the pocket periphery (Figure 1).
(a) Distribution of amino acid substitutions in the β-OG pocket of TBEV E protein sequences obtained from GenBank in comparison with the one of strain Absettarov. *Combination also occurs in triple substitutions; (b) Location of the most frequently substituted residues in the E protein β-OG pocket (represented by surface) of TBEV strain Absettarov model together with binding mode of compound
The most common (258 sequences, 45%) was the double substitution A47S + V206S/L on the pocket periphery (Figure 1(a)). In rare cases, it was accompanied by another one, with the most common combination A47S + V206L + T279A met in 11 sequences (2%). The most common single substitution was N52S, observed in 68 sequences (12%). Only one sequence with four substitutions in the pocket was found (GenBank accession no. AIB53033).
Quantification of E protein in virus samples
We determined the concentration of infectious virions in plaque assay in PEK cells (Table 4) and calculated E protein amount associated with them (Table 4, Table S1). For the calculation of the total amount of E protein associated with GCP, which include mature, immature, partially mature, and defective virus particles, we used qRT-PCR (Table 4). The concentration of GCP for all strains was higher than the concentration of infectious virions measured in plaque assay. The highest GCP to PFU ratio was observed for strain 256, while the lowest – for strain Absettarov (Figure 2). Since these strains have identical E protein amino acid sequences, it can be assumed that an excessive amount of GCP could affect the binding of antiviral compounds to infectious virions of strain 256.
Quantitative characterization of TBEV strain samples. Values are presented as mean ± ½ 95% CI. *Corresponds to significant difference with strain 256 with p ≤ 0.05.
Log10 concentration of E protein in the strain samples – measured in ELISA and estimated from GCP or PFU concentrations, assuming average molecular mass of TBEV E protein of 53 kDa and 180 protein molecules per virus particle (Table S1). *Corresponds to significant difference with strain 256 with p ≤ 0.05 .
Virus samples may contain not only infectious and genome containing particles, but also E protein aggregates, membrane complexes and virus-like particles covered with E protein, but lacking RNA.31–34 We determined the total concentration of the E protein in our virus samples using a commercial ELISA kit. The main assumption was that the sensitivity of ELISA kit is comparable for virions and decoy E protein, and does not depend on the virus subtype.
Concentration of total measured E protein in all strain samples was comparable or lower than in strain 256, while concentration of infectious virions was higher than in strain 256 (Table 4). These data indicated that the amount of decoy E protein per infectious virion in strain 256 exceeded these amounts for the other strains.
Discussion
The main goal of our study was to determine the spectrum of antiviral activity of the 4-aminopyrimidine N-oxides, which previously showed high activity against TBEV strain Absettarov. Based on the results, the most active compounds showed good antiviral activity against all selected TBEV strains representing three main subtypes. It gave the prospect that they have a broad spectrum of antiviral activity and would effectively inhibit all strains of TBEV.
Initially 4-aminopyrimidine N-oxides were selected as a potential TBEV inhibitor based on a docking study of their ability to interact with E protein’s β-OG pocket (Figure S5). Based on time-of-addition assay, we also confirmed that the compounds would most likely interact with the E protein 16 and the soluble ectodomain is the only easily accessible component of it. 35
The amino acid composition of mosquito-borne flavivirus E protein β-OG pocket was shown to influence different aspects of the flavivirus life cycle, including pH requirements for fusion, virus in vivo properties, such as neurovirulence, neuroinvasiveness, etc.36–39 Therefore, we conducted a substitution analysis of the residues that should be considered as comprising the pocket for the strains selected in our work and obtained in GenBank in comparison with strain Absettarov’s ones. All the selected strains, as well the strains presented in the GenBank, did not have substitutions directly inside the pocket, indicating its high conservation. Substitutions appear in a very limited number of positions, which are located only on the periphery of the β-OG pocket (Figure 1).
Nevertheless, comparing the inhibitory effects of the compounds against strains 256 and Absettarov, both belonging to Eu subtype and expressing E proteins with the identical amino acid sequences, we observed substantial differences in their activity, which needed additional investigation. The possible explanations were that the inhibitors could have a target other than E protein or the replication kinetics of the strains may account for difference in inhibitory activity of the compounds. However, our time-of-addition assay 16 had indicated that compounds most likely interact with the E protein and affected fusion and early stages of virus reproduction cycle, thus we proposed a hypothesis that this differential inhibitory effect can be explained by difference in decoy E protein content in viral strain samples. Cells infected with flaviviruses can produce not only fully functional mature infectious virions, but also defective virions and virus-like particles lacking RNA.31–34 Virions also greatly vary in the degree of maturation (e.g. completeness of furin cleavage of prM into M and E protein conformational switch during the virion exocytosis) that may affect their infectivity. The presence of E protein aggregates and membrane-bound structures produced during cell infection or virion degradation upon storage is also anticipated. Virus-like particles without genome were shown to possess antigenic structure similar to infectious virions and thus to bind neutralizing antibodies. 40 Therefore, decoy E proteins in non-infectious virus particles and immature virions, E protein aggregates and membrane complexes, may in principle bind small molecules. The reproduction capability of different virus particles differs, and for a virus sample characterized by a high content of decoy E protein, molecules of the inhibitor could be wasted due to binding with it. Thus, EC50 values of TBEV reproduction inhibitors targeting E proteins may be influenced by the ratio between the amounts of E protein from infectious virions and E protein from decoy particles.
The ratio between the total measured concentration of E protein in the sample and estimated E protein concentration from infectious particles represents how many decoy E protein molecules may compete with target E proteins from infectious virions for binding to inhibitor (Figure 2). The greater this ratio, the higher is the probability of the inhibitor to bind E protein molecule that does not take part in the infection process. A high content of decoy E protein explains the low activity of 4-aminopyrimidine N-oxides against the strain 256 compared to the one for strain Absettarov: to inhibit formation of the same number of plaques, a greater amount of compound is required in the case of the strain 256 due to compounds binding to decoy E protein.
The content of decoy E protein most likely depends on the intrinsic properties of the strain (e.g. accuracy of the viral polymerase or properties of the viral proteins responsible for the virion assembly), the method of virus sample preparation (multiplicity of infection, cell line, degree of the cytopathic effect on the moment of virus harvesting), and storage conditions. Nevertheless, the extent to which these factors influence the ratio between infections virions and decoy particles, as well as the difference in the ability to bind small molecules between different decoy particles is a matter of further research. Apparently, the ratio between the total amount of E protein and the one from infectious virions could be a critical factor for in vitro experiments based on the infectivity measurement, influencing the observed activity values of small molecule antivirals targeting the envelope proteins.
Conclusions
On the example of 4-aminopyrimidine N-oxides, we demonstrated that small molecule compounds exhibit a broad range of anti-TBEV activity. Compound
Our results showed that the outcome of activity measurement of a small molecule targeting the envelope protein of a TBEV strain could be influenced by the ratio of the infectious virions and decoy E protein in the virus sample. This ratio varied between the samples of the studied strains and could misrepresent the results of infectivity measurement in vitro assays. Testing potential reproduction inhibitors against TBEV strains with high content of E protein out of infectious particles could be an important step in the study of the antiviral activity of compounds binding to the envelope proteins. This fact should be considered during the selection and characterization of virus preparation for assessment of the antiviral compounds and comparison of their activity.
Supplemental Material
sj-pdf-1-avc-10.1177_2040206620943462 - Supplemental material for Spectrum of antiviral activity of 4-aminopyrimidine N-oxides against a broad panel of tick-borne encephalitis virus strains
Supplemental material, sj-pdf-1-avc-10.1177_2040206620943462 for Spectrum of antiviral activity of 4-aminopyrimidine N-oxides against a broad panel of tick-borne encephalitis virus strains by Evgenia V Dueva, Ksenia K Tuchynskaya, Liubov I Kozlovskaya, Dmitry I Osolodkin, Kseniya N Sedenkova, Elena B Averina, Vladimir A Palyulin and Galina G Karganova in Antiviral Chemistry and Chemotherapy
Footnotes
Authors’ contributions
GGK, EVD, KKT, LIK, DIO designed the study; EVD performed sequence analysis, cell-based assays and experimental data interpretation and analysis; KKT performed protein and RNA content measurements; KNS, EBA provided the compounds; GGK, LIK, DIO, KNS, EBA and VAP supervised the study; EVD, DIO, LIK wrote the manuscript draft; all the authors provided feedback.
Acknowledgements
The authors thank Dr. Alexander Litov for qPCR standards preparation. Free academic license for OpenEye software was kindly provided by OpenEye Scientific Software Inc. Free academic license for Instant JChem, used for structure database management, was kindly provided by ChemAxon (Instant JChem 18.3.0, 2018, ChemAxon ![]() )).
)).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by Russian Foundation for Basic Research, grants no. 15–04-08365 and 18–03-00651. BSL-3 facilities, virus and cell collection were supported by the State research funding of FSBSI “Chumakov FSC R&D IBP RAS”.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
