Abstract
Background
The replacement of β,γ-pyrophosphate by β,γ-phosphonate moieties within the triphosphate chain of 5′-triphosphate nucleoside analogues was previously studied for various antiviral nucleoside analogues such as AZT and 2′,3′-dideoxynucleosides. Thus, it has been shown that these chemical modifications could preserve, in some cases, the terminating substrate properties of the triphosphate analogue for HIV-RT. Herein, we aimed to study such 5′-triphosphate mimics based on the scaffold of the well-known antiviral agent 9-[(2-phosphonomethoxy)ethyl]adenine (PMEA, Adefovir).
Methods
Synthesis involved coupling of a morpholidate derivative of PMEA with appropriate pyrophosphoryl analogues. The relative efficiencies of incorporation of the studied diphosphate phosphonates were measured using subtype B WT HIV-1 RT in an in vitro susceptibility assay, in comparison to the parent nucleotide analogue (PMEApp).
Results
Searching for nucleoside 5′-triphosphate mimics, we have synthesized and studied a series of diphosphate analogues of PMEA bearing non hydrolysable bonds between the and phosphorus atoms. We also examined their relative inhibitory capacity towards HIV-1 reverse transcriptase in comparison to the parent nucleotide analogue (PMEApp). Only one of them appeared as a weak inhibitor (IC50 = 403.0 ± 75.5 µM) and proved to be less effective than PMEApp (IC50 = 6.4 ± 0.8 µM).
Conclusion
PMEA diphosphoryl derivatives were designed as potential substrates and/or inhibitors of various viral polymerases. These modifications dramatically affect their ability to inhibit HIV-RT.
Keywords
Introduction
The acyclic nucleoside phosphonate 9-[2-(phosphomethoxy)ethyl]adenine (PMEA, Figure 1) exhibits a broad-spectrum activity against different types of DNA viruses and retroviruses.1,2 Its orally bioavailable form, the bis(pivaloyloxymethyl) prodrug (bis(POM)PMEA, Adefovir dipivoxil), has been approved for the treatment of chronic hepatitis B3 and other types of prodrugs are still under investigations.4–6 To achieve its inhibitory effect on viral synthesis, PMEA must be converted intracellularly to its active diphosphorylated metabolite, PMEApp (Figure 1). PMEApp has been described to interact as an alternative substrate and as a competitive inhibitor of both herpes simplex type 1 (HSV-1) DNA polymerase7,8 and reverse transcriptases.9–11 Variable inhibitory effects on human cellular DNA polymerases were observed, especially against DNA polymerase for which K
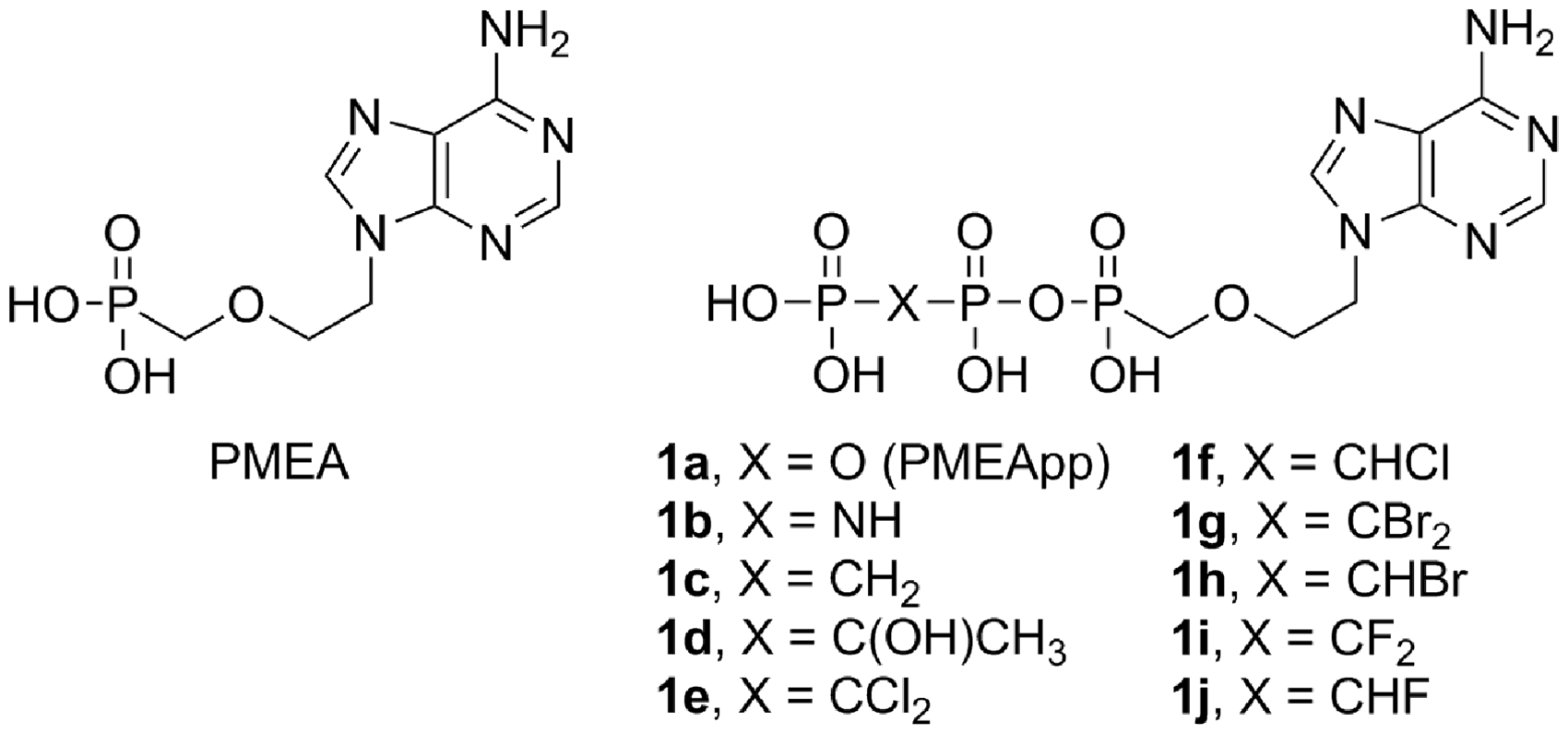
Chemical structures of PMEA, PMEApp 1a, and target diphosphate analogues 1b-j.
As part of a research program, we decided to synthesize new nucleoside 5′-triphosphate mimics based on the PMEA scaffold and incorporating chemical modifications of the Pβ–O–Pγ phosphoester bonds. Replacement of the anhydride oxygen with isosteric groups leading to non-hydrolysable bonds, the resulting analogues were designed as biological tools for the study of substrate properties of cellular and/or viral enzymatic systems, as well as new potential therapeutic agents.16,17 Based on previously published works on related topic, 16 requirements in the design of modified triphosphate analogues emerged: (i) the anhydride bond between β- and γ-phosphates, which is unaffected during the DNA biosynthesis, could be replaced with non-hydrolysable bond; (ii) similar modification could also be introduced between the 5′-position of sugar and the phosphorus atom; (iii) the anhydride bond between α- and β-P atoms should be preserved in order to provide the possibility of the mimetic to interact with targeted polymerase as substrate.
In this respect, PMEApp constitutes an attractive model to study chemical modifications on the pyrophosphoryl residue due to its phosphonate structure, characterized by a stable P–C bond (toward phosphohydrolase-hydrolysis) between acyclic nucleoside moiety and α-phosphorus atom, and its broad and high affinity for viral polymerases. Herein, we report the full accounts of the synthesis of compounds 1b-j and their study as terminating substrates in the DNA chain elongation catalyzed by human immunodeficiency virus (HIV) reverse transcriptase.
Experimental section
Material and methods
1H NMR (250 MHz) and 13C NMR (100 MHz) spectra were recorded with proton decoupling at ambient temperature. Chemical shifts (δ) are quoted in parts per million (ppm) referenced to the residual solvent peak chloroform (CDCl3) at 7.26 ppm and 77.0 ppm, deuterium oxide (D2O) at 4.63 ppm relative to tetramethylsilane (TMS). COSY experiments were performed in order to confirm proton assignments as well as 2D 1H-13C heteronuclear COSY for the attribution of 13C signals. 13P NMR spectra were recorded at ambient temperature at 100 MHz. Chemical shifts are reported relative to external phosphoric acid (H3PO4). 19F NMR spectra were recorded at ambient temperature at 235 MHz. Chemical shifts are reported relative to external trichlorofluoromethane (CFCl3). Coupling constants,
Chemistry
9-[(2-Phosphomethoxy)ethyl]-adenine (PMEA) and its phosp.
horomorpholidate derivative 2 were synthesized according to a published procedure. 18
The tributylammonium salts of pyrophosphate 3a and diphosphonic acids 3b-d were obtained fromtheir commercially available forms: tetra sodium pyrophosphate decahydrate, tetra sodium imidodiphosphonate, methanediphosphonic acid and 1-hydroxyethylidene diphosphonic acid, respectively. The halomethylidene diphosphonic acids 3e-j were obtained from their ethyl esters precursors following a usual way, 19 and stored as sodium forms after passage over a Dowex 50WX2 cation exchange resin column and freeze-drying. Tetraethyl methylenediphosphonate 4 was commercially available. The halomethylene diphosphonate esters 5e, 5g, 6f and 6h were prepared using literature methods. 20
Detailed description of experimental procedures and compound characterization are provided as supplementary data.
In vitro drug susceptibility assays with recombinant subtype B WT HIV-1 RT
The p66RTB gene construct allowing the bacterial expression of the wild-type (WT) HIV-1 RT was described elsewhere.21,22 The recombinant clade B WT HIV-1 RT was co-expressed with HIV-1 protease in
Standard RT activity was assayed using 250 µg/mL of activated calf thymus DNA (GE Healthcare). To determine IC50 values, reactions were performed with 10 nM enzyme and 5 µM each dNTP as a mixture (dATP, dCTP, dGTP, dTTP) containing 100 µCi/mmol of [3H]-labelled deoxythymidine 5'-triphosphate (Perkin Elmer), for 15 min with increasing amounts of phosphonate, compounds 1a (PMEApp as reference), 1c, 1e to 1j. Each aliquot was spotted in duplicate on DE81 ion-exchange paper discs.
Molecular modeling of HIV-1 RT in complex with diphosphate phosphonate analogues 1c, 1e, 1g, 1i
All models were based on the X-ray structure of the RT in complex with dsDNA and incoming PMPApp (PDB code 1T05). The UCSF Chimera software (PMID: 15264254) was used to replace the PMP – moiety by the PME – equivalent. Moreover, the oxygen of the β,γ bridge of the diphosphate phosphonate PMPApp was replaced by CH2 (compound 1c), CCl2 (compound 1e), CBr2 (compound 1g) and CF2 (compound 1i) groups.
Results
The synthesis of PMEA diphosphate 1a and its mimetics 1b-j was carried out according to a general procedure (Scheme 1),23,24 which requires preliminary preparation of the phosphoromorpholidate derivative 2 of PMEA. This was accomplished by usual reaction of PMEA with morpholine and
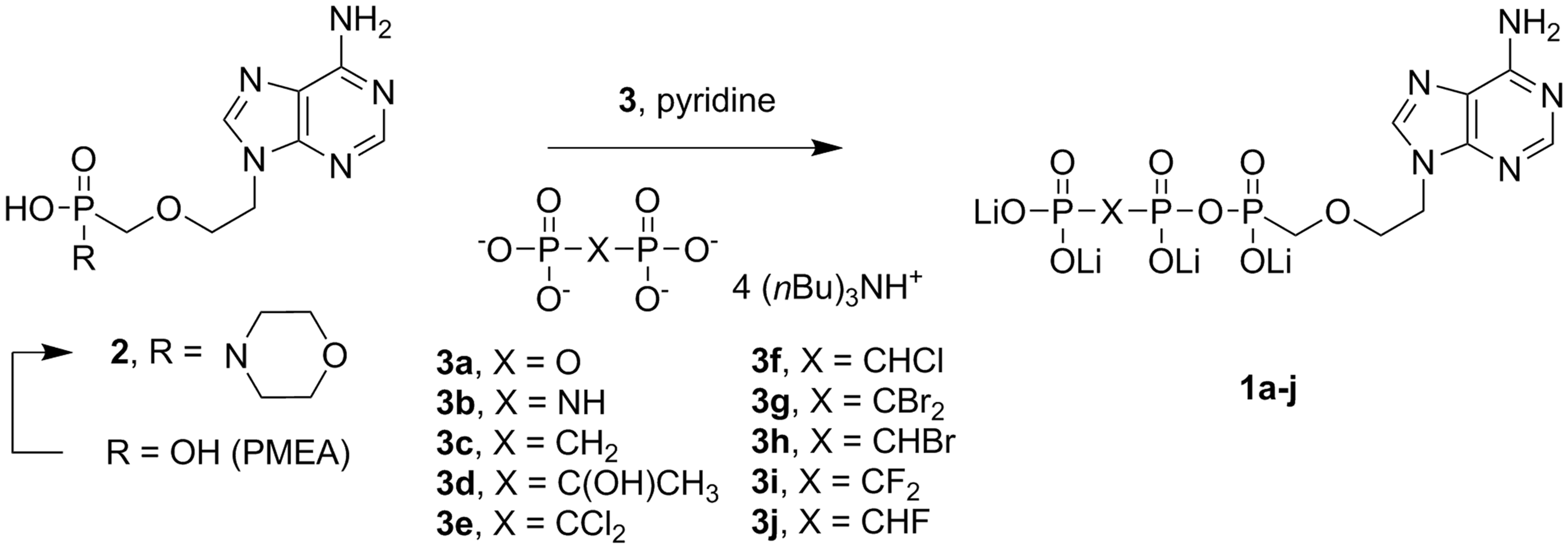
Final step in the synthesis of PMEApp 1a, and target diphosphate analogues 1b-j.
Moreover, in contrast to the dichloro- and dibromo-analogues (3e-h), nucleophilic dehalogenation of difluoromethylene-diphosphonates into the corresponding monofluoro esters by conventional methods28,29 was unsuccessful. In such conditions, P–C bond cleavage was observed resulting in the formation of dialkyl difluoromethylphosphonates. Thus, we decided to select the first approach (i.e. direct halogenation) leading in one step to a mixture of the mono- and difluoro- compounds through reaction of electrophilic fluorinating reagents with the carbanions of alkyl methylidenediphosphonates. Reported methodologies used perchloryl fluoride
19
or acetyl hypofluorite.
30
Herein,
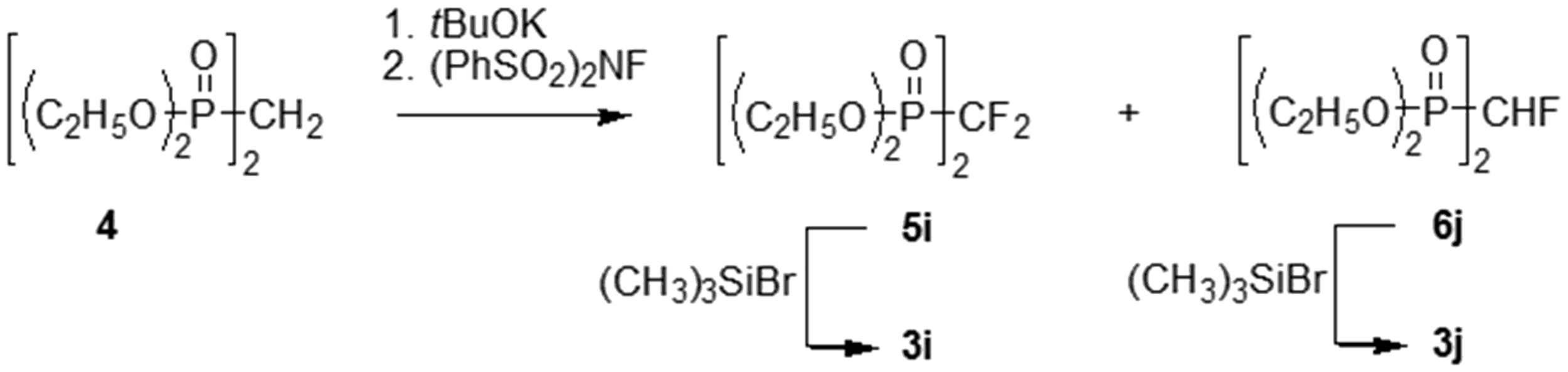
Synthesis of the mono- and difluorobisphosphonic acids 3i, j.
Crude reaction of the phosphoromorpholidate derivative of PMEA 2 with pyrophosphate 3a or the appropriate tributylammonium salts of the diphosphonic acids 3b-j was firstly purified by a Dowex 1X2 chromatography using a gradient of aqueous lithium chloride in 0.01 M hydrochloric acid. 18 Then, DEAE-Sephadex A25 chromatography gave PMEA diphosphate 1a and its mimetics 1b-j. The low yield obtained for derivative 1b (18%) was probably due to the chemical instability of the imido functionality 32 during purification step at acidic pH. Structures of the different mimetics of PMEA diphosphate were assigned on the basis of their NMR data (Tables 1 and 2), MS and UV spectra. Purity was checked by analytical high pressure liquid chromatography (HPLC) and high resolution mass spectra (HRMS).
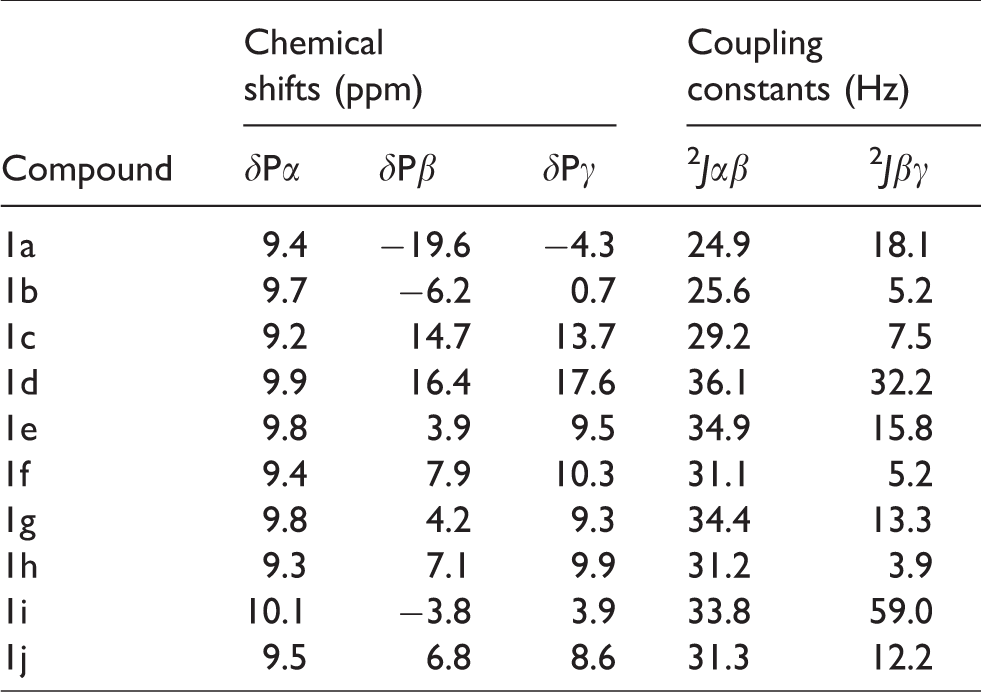
Selected 13P NMR data of the new PMEA diphosphate mimetics 1b-j.
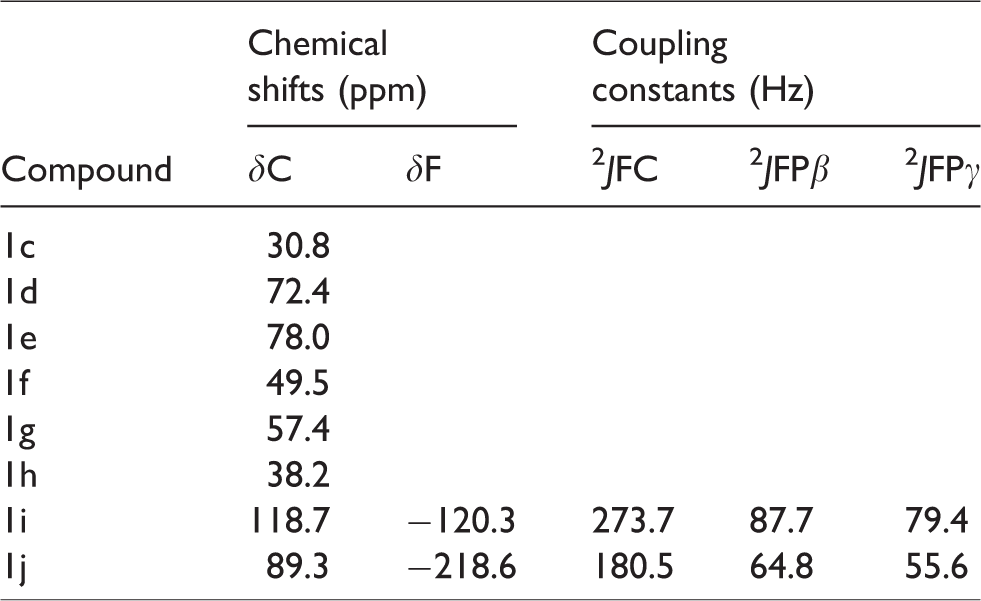
Selected 13C and 19F NMR data of the new PMEA diphosphate mimetics 1c-j.
To evaluate the inhibitory activity of diphosphate phosphonates 1a, 1c, 1e to 1j on the reverse transcriptase (RT) of HIV-1, their relative efficiencies of incorporation were measured using subtype B WT HIV-1 RT in an in vitro susceptibility assay. The calculated 50% inhibitory concentration (IC50) values obtained in this assay showed that PMEApp 1a is active (IC50 = 6.4 ± 0.8 µM), this value was in agreement with literature data, 33 whereas the diphosphate phosphonate analogues are truly less potent (IC50 >1000 µM or IC50 = 403.0 ± 75.5 µM for compound 1i).
Discussion
The synthesis of new mimics of PMEApp incorporating non hydrolysable bond between the β- and γ-P atoms has been carried out by reaction of the morpholidate derivative of PMEA 2 with the appropriate diphosphinic acids 3b-j (Scheme 1). The target diphosphate analogues 1b-j were isolated as lithium forms in 18–65% yields.
The 13P NMR spectra of the mimetic phosphonates 1b-j showed characteristic downfield shifts which permitted, from a straightforward comparison with literature data, the direct assignment of resonances for α-, β- and γ-phosphorus atoms (Table 1). The α-P resonance was relatively independent of the substituent nature between β and γ phosphorus atoms. Compared to the parent phosphate, the phosphorus–phosphorus coupling constant
2
The resonance of carbon atom between β-P and γ-P showed a normal dependence to the electronegativity of substituent (Table 2). The downfield shifts increased in the series δ(CF2) > δ(CHF) > δ(CCl2) > δ(COHCH3) > δ(CBr2) > δ(CHCl) > δ(CHBr) > δ(CH2). Finally, a single fluorine resonance was observed in 19F NMR spectra for both the difluoromethylene compound 1i (δ −120.3 ppm, dd) and the mixture of diastereoisomeric monofluoromethylene compounds 1j (δ −218.6 ppm, ddd) showing that the fluorine environments could be considered as magnetically equivalent. Coupling constants were different for the two phosphorus nuclei directly bonded to the fluorinated β,γ-methylene bridge.
2
The substrate properties of these PMEA diphosphoryl derivatives were comparatively studied, in cell free solutions, towards HIV-1 reverse transcriptase. Indeed, it has been previously shown in various nucleotide series that chemical modifications on 5′-phosphate residues could preserve, in some cases, the terminating substrate properties of the triphosphate analogue in the DNA biosynthesis catalyzed by different polymerases.17,38–40 The substitution of β,γ-pyrophosphate by β,γ-phosphonates was rather systematically studied for a series of antiviral nucleoside analogues (AZT and 2′,3′-dideoxynucleosides)41,42 including modification of the triphosphate chain at the - position and presents some similarity with the compounds under study. Thus, in the particular case of (
Several factors including size, polarity, and electronegativity may modulate the activity, even so a certain tolerance of the HIV-1 reverse transcriptase to the γ-P-substituents was demonstrated in literature. 42
To understand why diphosphate phosphonates analogues 1c, 1e, 1g and 1j are poor substrates of HIV-1 RT in comparison to PMEApp 1a, we performed modeling replacing the oxygen of the β,δ bridge of the diphosphate phosphonate by CH2, CCl2, CBr2 or CF2 groups, with respect to specific geometry and bond distances. According to Tuske et al.,43 amino acids R72 and K65, and D113 to a lesser extent, play a key role in the binding of the nucleotide in the active site and in their incorporation by HIV-1 RT (Figure 2(a)). If the substitution of oxygen atom of the, β,γ-pyrophosphate bond by a methylene group (CH2),
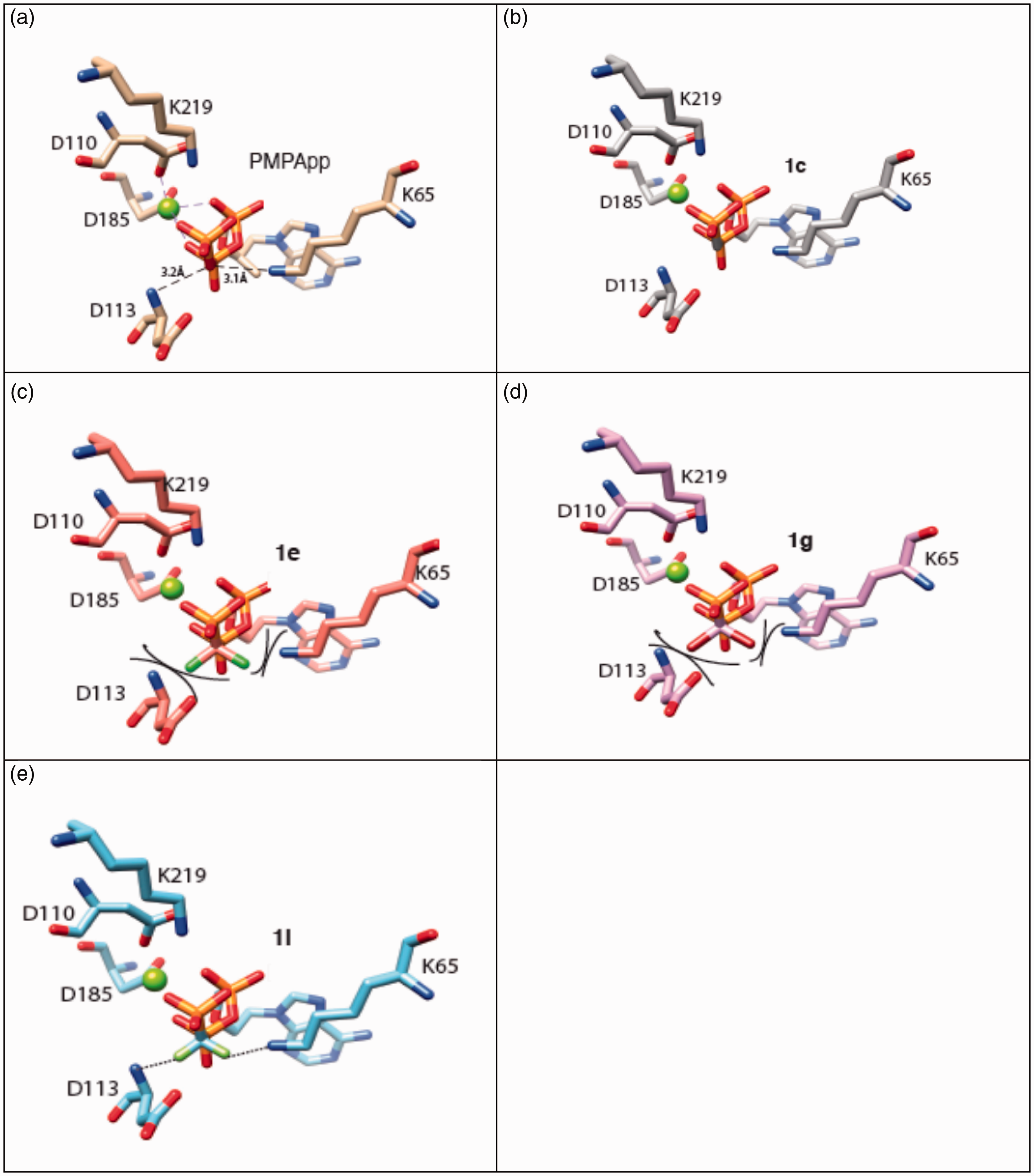
(a) Structure of HIV-1 RT active site in complex with PMPApp. (b), (c), (d) and (e) models for the putative positioning of acyclic diphosphate phosphonates 1c, 1e, 1g, 1i in the active site of HIV-1 RT. The atomic coordinates (PDB 1T05 – HIV-1 RT in complex with PMPApp) were used to visualize the complex HIV-1 RT- diphosphate phosphonate, after modeling, replacing PMP- moiety by PME- one and the oxygen of the, bridge of the diphosphate phosphonate by CH2, CCl2, CBr2 or CF2 groups, respecting specific geometry and bond distances. One magnesium ion is represented as green sphere. Amino acid R72 and the second magnesium ion are intentionally omitted for figure clarity. (a) Structure 1T05: distances between the O of the, bridge and amino acids K65 and D113 are mentioned in dotted line. (c) and (d) steric hindrance is mentioned in full line. (e) Interactions are mentioned in dotted line.
Another important remark is the difference that can be observed when comparing the crystallographic data from complexes of HIV-RT with purine or pyrimidine nucleotides. With purine nucleotides, as PMPApp, the amino acid R72 plays a crucial role in stacking the nucleobase, while only the amino acid R65 interacts with the O of the β,γ bridge. With pyrimidine nucleotides, their binding in the RT active site is slightly different, both amino acids R72 and K65 interact with the O of the α,β bridge and consequently the binding of the nucleotide is less sensible to the chemical modification of the bridge. This may explain why the modified phosphonate analogues of AZT are recognized and substrates for HIV-1 RT. 41 Thus, they are efficiently incorporated in the growing nucleic acid chain.
Conclusion
Nine diphosphate analogues of PMEA have been designed as isostere of the parent diphosphorylated form, PMEApp. The benefit of stable P–C bonds replacing scissile P–O–C linkage constitutes an attractive interest to evaluate the electronic and stereochemical requirements for binding to relevant proteins. It was observed that HIV-RT does not extend the DNA primer with the synthesized compounds. Within this acyclic series, the replacement of the β,γ bridge of the diphosphate phosphonate by CH2, CHCl, CCl2, CHBr, CBr2, CHF or CF2 groups has a drastic effect on the recognition by the HIV-RT.
Supplemental Material
SI Synthesis and Substrate Properties of PMEApp analogues -Supplemental material for Synthesis and substrate properties towards HIV-1 reverse transcriptase of new diphosphate analogues of 9-[(2-phosphonomethoxy)ethyl]adenine
Supplementary Material
Footnotes
Authors’ note
This article is dedicated to Dr. Gilles GOSSELIN on the occasion of his retirement.
Acknowledgments
We thank the “Fonds der Chemischen Industrie” for a post-doctoral fellowship (W. L.). We thank the “Agence Nationale de Recherche sur le Sida et les Hépatites Virales” for its financial support.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplementary material
Supplementary material (synthetic procedure and compounds characterization) is provided as a separate electronic file (PDF format) and was submitted along with the manuscript.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
