Abstract
Background
The novel phenanthridinone derivative HA-719 has recently been identified as a highly potent and selective inhibitor of hepatitis C virus replication. To elucidate its mechanism of inhibition, we have isolated and analyzed a clone of hepatitis C virus replicon cells resistant to HA-719.
Methods
To isolate HA-719-resistant replicon cells, Huh-7 cells containing subgenomic hepatitis C virus replicons (genotype 1b) with a luciferase reporter (LucNeo#2) were cultured in the presence of G418 and escalating concentrations of HA-719. After several passages, total RNA was extracted from the growing cells, and Huh-7 cells were transfected with the extracted RNA. Limiting dilution of the transfected cells was performed to obtain an HA-719-resistant clone.
Results
The 50% effective concentration (EC50) of HA-719 for hepatitis C virus replication was 0.058 ± 0.012 µM in LucNeo#2 cells. The replicon cells capable of growing in the presence of G418 and 3 µM HA-719 were obtained after 18 passages (72 days). The HA-719-resistant clone LucNeo719R showed 98.3-fold resistant to the compound (EC50 = 5.66 ± 0.92 µM), but the clone had no cross-resistance to telaprevir (NS3 inhibitor), daclatasvir (NS5A inhibitor), and VX-222 (NS5B inhibitor). The sequence analysis for the wild-type and LucNeo719R identified 3, 2 and 7 mutations in NS3/4 A, NS4B, and NS5A, respectively, but no mutations in NS5B.
Conclusion
None of the amino acid mutations in the resistant clone corresponds to those reported to confer drug-resistance to current anti-hepatitis C virus agents, suggesting that the target of HA-719 for hepatitis C virus inhibition differs from those of the existing agents.
Introduction
Hepatitis C virus (HCV) is a single stranded positive-sense RNA virus of Flaviviridae family. 1 HCV genome encodes at least 10 viral proteins categorized as structural proteins (C, E1, and E2) and non-structural proteins (p7, NS2, NS3, NS4A, NS4B, NS5A, and NS5B). HCV is a causative agent of chronic hepatitis that leads to liver cirrhosis and hepatocellular carcinoma. At present, 130–150 million people worldwide are chronically infected with this virus, and HCV infection is a serious public health problem.2–4
HCV is classified into six major types of genomes (genotypes 1‒6). The previous standard treatment for HCV-infected patients is the combination therapy with pegylated interferon-α (peg-IFN-α) and ribavirin (RBV). 5 However, this interferon-based therapy is not always effective for all patients. In addition, it has serious side effects, such as flu-like symptoms, anemia, loss of appetite, and depression. Several direct-acting antiviral agents (DAAs) have recently been approved for their clinical use. These include NS3 protease inhibitors (simeprevir, asnaprevir, vaniprevir, and paritaprevir), the NS5A inhibitors (daclatasvir (DCV), ledipasvir, and ombitasvir), the NS5B polymerase inhibitor (sofosbuvir) and their combination drugs.6,7 These DAAs are highly effective without peg-IFN-α and RBV. In fact, recent clinical trials with a once daily fixed-dose combination of sofosbuvir and ledipasvir revealed that the rates of sustained virological response were 97 to 99% in genotype 1-infected but previously untreated patients. 8
Alternatively, studies on antivirals targeting host cellular factors are also considered. The immunosuppressive drug cyclosporin A (CsA) has been reported to exert anti-HCV activity in vivo and in vitro via interaction with cyclophilin A (CypA).9–11 CypA is a crucial cellular cofactor for HCV replication and does display a peptidyl-prolyl cis-trans isomerase activity that catalyzes the cis-trans isomerization of the prolyl peptide bond preceding proline residues.11,12 Non-immunosuppressive derivatives of CsA, such as DEB025 and NIM811, have also been developed as cyclophilin inhibitors. 13 Other host cellular factors, including the liver-specific microRNA miR-122, phosphatidylinositol-4-kinase IIIα, silibinin and toll-like receptors, are also possible candidates for anti-HCV strategies. 5 The combination chemotherapies of such agents and DAAs may have potential for protecting HCV-infected patients from life-threatening complications.
We have previously reported the novel phenanthridinone derivative HA-719 (Figure 1) as a highly potent and selective inhibitor of HCV replication in vitro.
14
HA-719 exerted anti-HCV activity in dose-dependent fashion with a 50% effective concentration (EC50) of 0.063 ± 0.010 µM in genotype 1b subgenomic replicon cells. The compound did not show cytotoxic effects on the human hepatoma cell line Huh-7 at concentrations up to 40 µM. Although the molecular target of HA-719 remains unknown, HA-719 did not affect the enzymatic activity of NS3 protease or NS5B polymerase. In the present study, we have isolated a drug-resistant clone of HCV replicon cells and characterized its profiles to gain an insight into the inhibitory mechanism of HA-719.
Chemical structure of HA-719.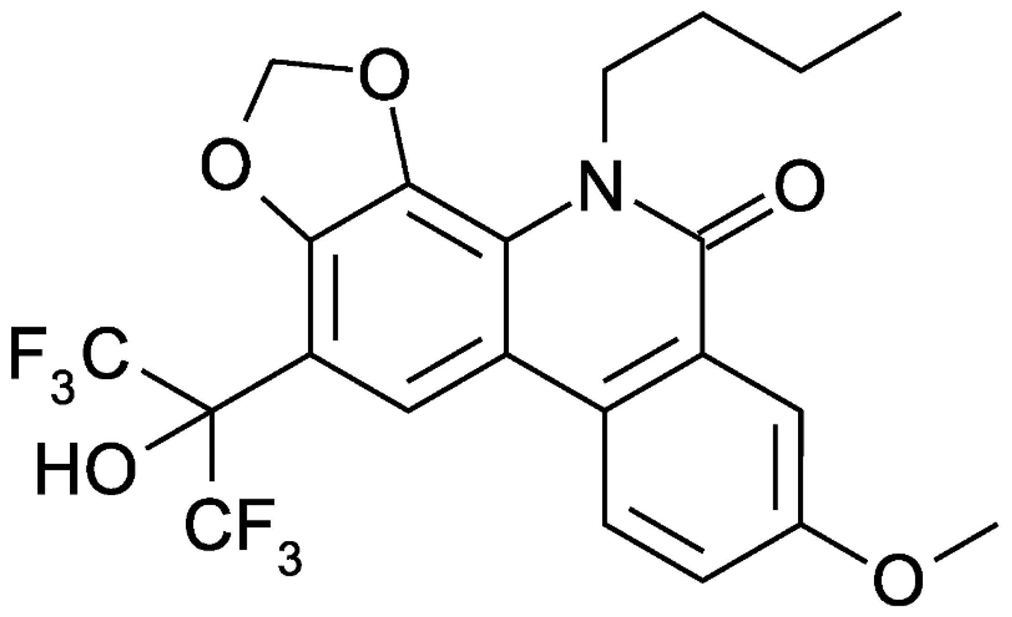
Methods
Compounds
HA-719 was synthesized by Dr. Hashimoto and his colleagues, according to the method previously described. 15 The NS3 protease inhibitor telaprevir (TLV), the NS5A inhibitor DCV, and the non-nucleoside NS5B polymerase inhibitor VX-222 were purchased from AdooQ BioScience (Irvine, CA).16–18 All compounds were dissolved in dimethyl sulfoxide at a concentration of 20 mM and stored at −20℃ until use.
Cells
Huh-7 cells containing self-replicating subgenomic HCV replicons (genotype 1b) with a luciferase reporter, LucNeo#2, were used for experiments. 19 The cells were cultured in Dulbecco’s modified Eagle medium with high glucose (Nacalai Tespue, Kyoto, Japan) supplemented with 10% heat-inactivated fetal bovine serum (Thermo Scientific, Waltham, MA), 100 U/ml penicillin G, 100 µg/ml streptomycin, and 1 mg/ml G418 (Nacalai Tespue).
Establishment of HA-719-resistant replicon cells
To induce viral resistance to HA-719, LucNeo#2 cells (8 × 104 cells/well) were cultured in a six-well plate and serially passaged in the presence of 1 mg/ml G418 and escalating concentrations of the compound. Fresh culture medium containing the compound was added to the cells every three to six days. The cells were subcultivated, whenever they reached 70 to 80% confluence. At the time of subcultivation, the drug concentration was increased, if the number of viable cells was sufficient. When the drug concentration reached far above its EC50 for HCV inhibition in LucNeo#2 cells, the cells were expanded, and total RNA was isolated from the cells. Huh-7 cells were transfected with the isolated RNA, and limiting dilution of the transfected cells was conducted to obtain their clones. After selection with G418, the anti-HCV activity of HA-719 was examined in the obtained clone LucNeo719R and compared to that in the wild-type LucNeo#2.
Colony formation assay
LucNeo#2 and LucNeo719R cells (1 × 105 cells/well) were cultured for three weeks in a six-well plate in the presence of 1 mg/ml G418 and various concentrations of HA-719. The culture medium was changed twice per week. After three weeks of cultivation, the cells were fixed with 99.5% ethanol and stained with 2% crystal violet (Wako, Osaka, Japan).
RNA quantification by real-time RT-PCR
LucNeo#2 and LucNeo719R cells (5 × 103 cells/well) were cultured in a 96-well plate in the absence of G418 and in the presence of various concentrations of test compounds. After incubation for three days at 37℃, the cells were washed once with phosphate-buffered saline and treated with lysis buffer containing deoxyribonuclease in TaqMan® Gene Expression Cell-to-CT™ Kit (Life Technologies, Waltham, MA). The lysate was subjected to reverse transcription real-time polymerase chain reaction (RT-PCR), according to the manufacturer’s instructions. The 5′-untranslated region of HCV RNA was quantified using the sense primer 5′-CGGGAGAGCCATAGTGG-3′, the antisense primer 5′-AGTACCACAAGGCCTTTCG-3′ and the fluorescence probe 5′-CTGCGGAACCGGTGAGTACAC-3′ (Life Technologies).
Sequence analysis
Total RNA was isolated from LucNeo#2 and LucNeo719R cells by RNeasy mini kit (Qiagen, Venlo, Netherlands), according to the manufacturer’s instructions. For RT-PCR, first-strand cDNA synthesis was performed by SuperScript III First-Strand Synthesis System kit (Life Technologies). PCR amplification of the non-structural protein region (from NS3 to NS5B) was performed with KOD-Plus-DNA polymerase (Toyobo, Osaka, Japan). The primer pair used for PCR amplification was 5′-ATCATCACCAGCCTCACAGGTCGGGACAAGAACC-3′ (forward; NS3 region) and 5′-CTTGGTCTTTACTGCCCAGTTGAAGAGGTACTTGC-3′ (reverse; NS5B region). The PCR conditions consisted of pre-incubation at 94℃, 3-min activation of the polymerase followed by 40 cycles of PCR (denaturation at 94℃ for 15 s, annealing at 60℃ for 45 s, and extension at 68℃ for 6 min) with final extension at 68℃ for 6 min. PCR products were sequenced by the primer walking DNA sequencing method.
GenBank accession number
GenBank accession number for the nonstructural protein region of LucNeo719R cells is LC034308.
Results
HA-719-resistant replicon cells
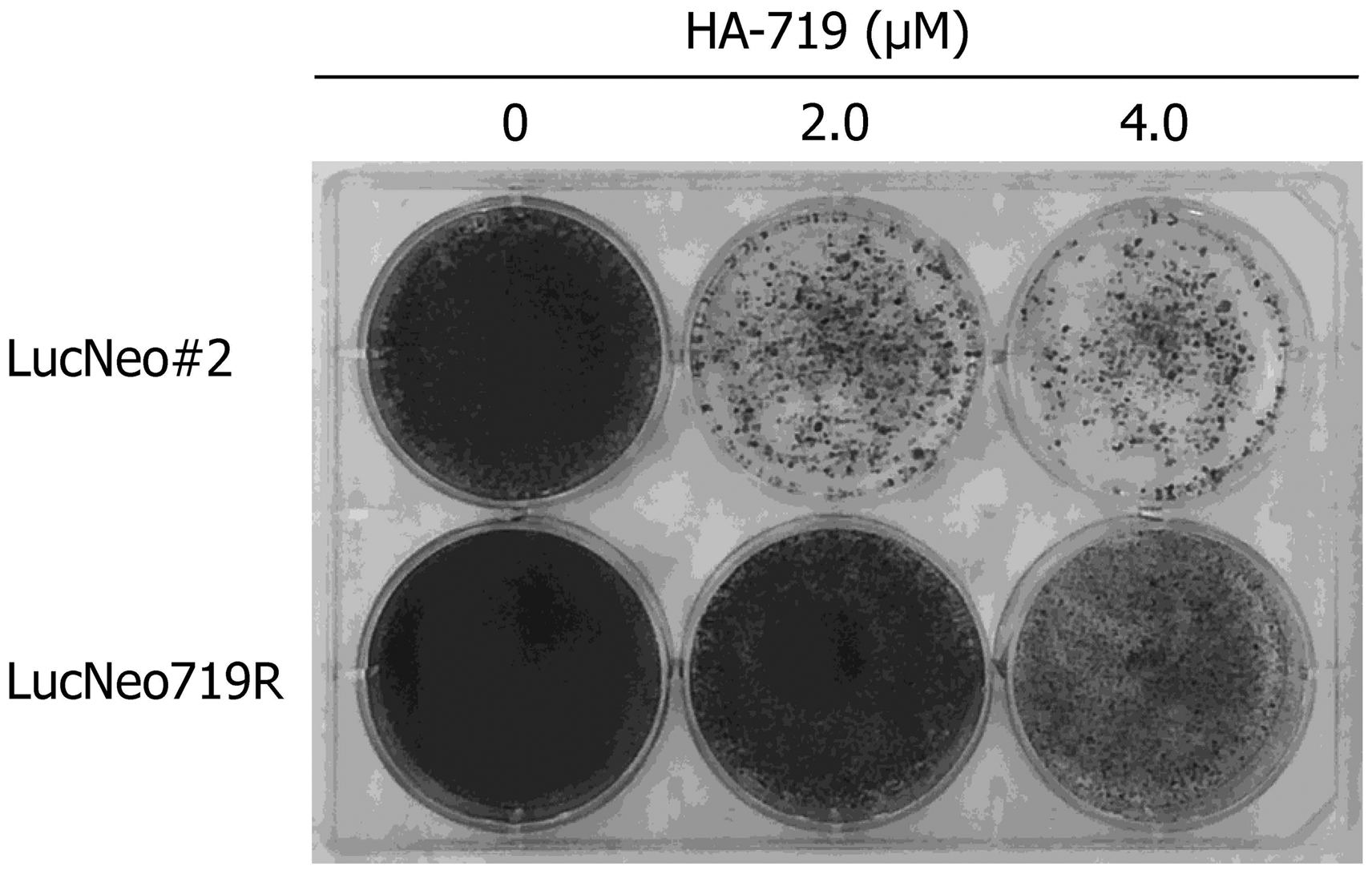
Induction of drug-resistant viruses often provides important information of the characters of compounds, including their mechanism of action. LucNeo#2 cells were cultured in the presence of 1 mg/ml G418 and HA-719 at a concentration close to its EC50 for HCV inhibition in LucNeo#2 cells. The concentration of HA-719 was increased gradually after each passage and could reach 3 µM after 72 days of cultivation (Figure 2(a)). After transfection of the resistant replicon RNA to Huh-7 cells and cloning of the transfected cells, the obtained cells harboring the replicon RNA (LucNeo719R) were examined for their susceptibility to HA-719. As shown in Figure 2(b), the HA-719 was much less inhibitory to HCV replication in LucNeo719R cells than in LucNeo#2 cells. The EC50 values were 9.5 and 0.050 µM in LucNeo719R and LucNeo#2 cells, respectively. In the colony formation assay, HA-719 could not significantly reduce the number of LucNeo719R colonies even at concentrations 40 to 80-fold higher than its EC50 in LucNeo#2 cells (Figure 3).
Establishment of HA-719-resistant-replicon cells. (a) LucNeo#2 cells (wild-type replicon cells) were cultured and serially passaged in the presence of G418 and escalating concentrations of HA-719. When the drug concentration was reached far above its EC50 for HCV inhibition in the wild-type replicon cells (arrow), total RNA was isolated from the cells. Huh-7 cells were transfected with the isolated RNA, and limiting dilution of the transfected cells was conducted to obtain a drug-resistant clone (LucNeo719R). (b) The activity of HA-719 was determined in LucNeo#2 cells (closed bars) and LucNeo719R cells (shadow bars) by real-time RT-PCR. Colony formation assay of LucNeo#2 and LucNeo719R. While under G418 selection pressure, the parental replicon cells LucNeo#2 and the established replicon cells LucNeo719R were treated with various concentrations of HA-719.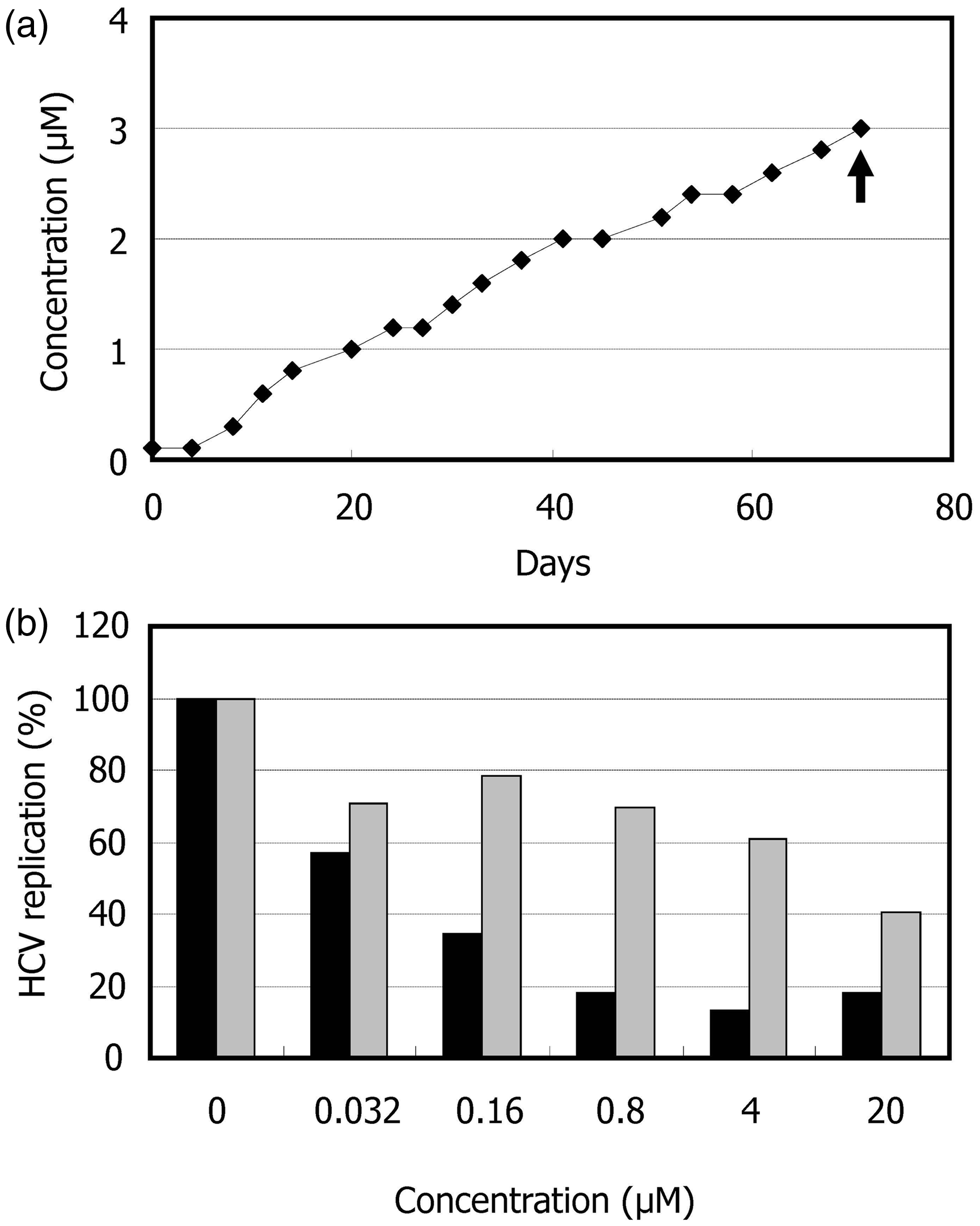
Anti-HCV activity of DAAs in LucNeo719R cells
Antiviral activity of HA-719 or HCV reference inhibitors.
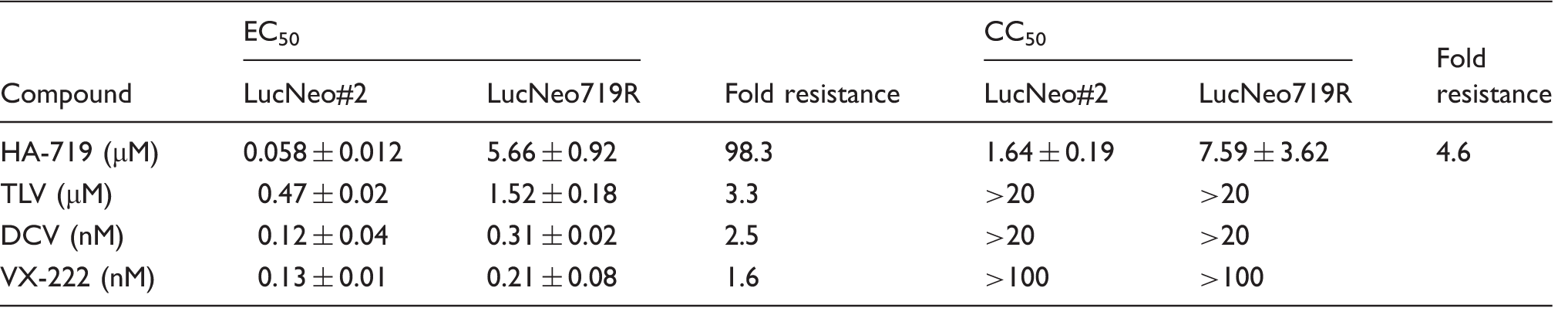
Note: All data represent means ± ranges for two independent experiments.
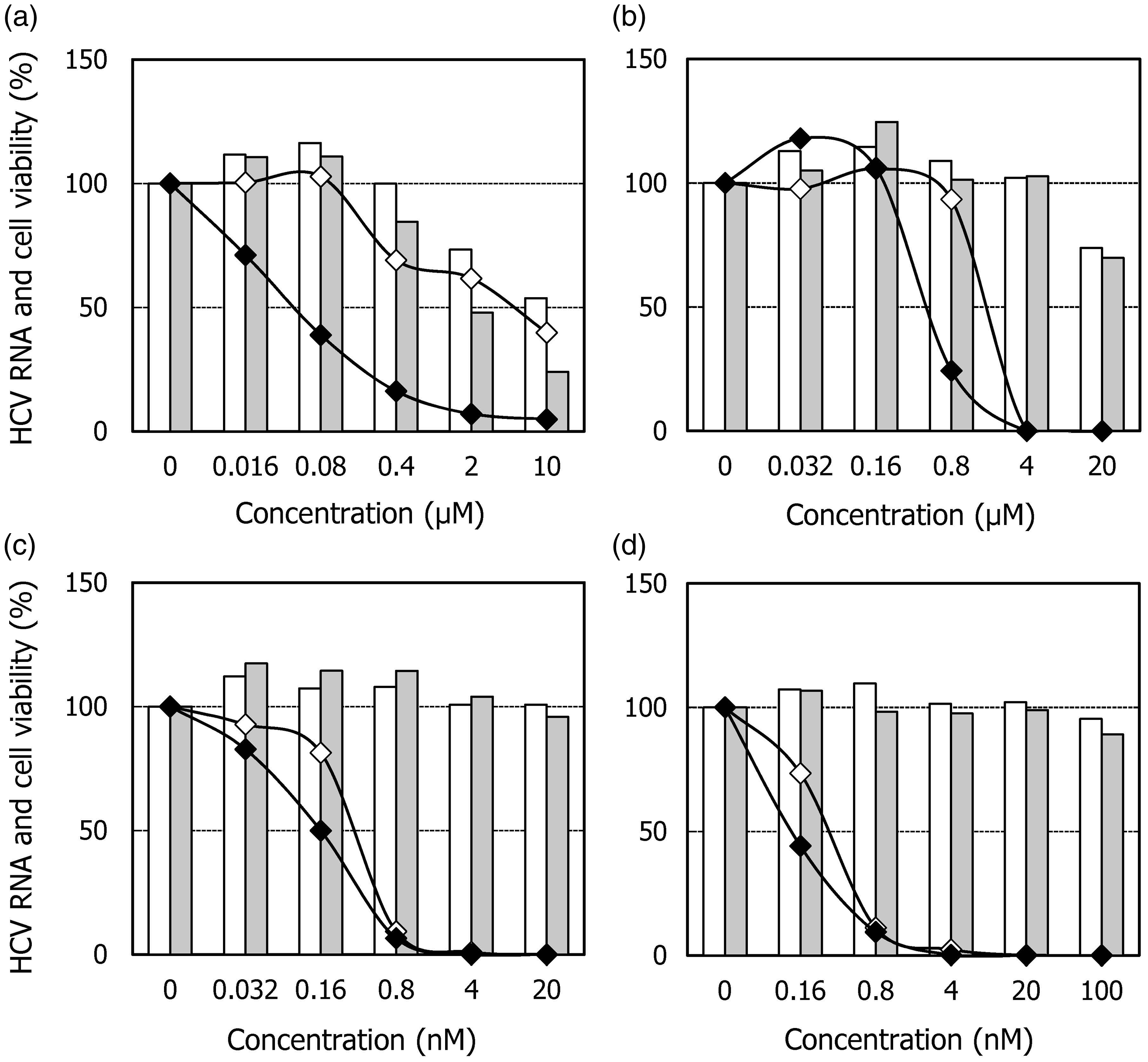
Effects of HA-719 or various DAAs on the replication of HCV RNA replicons in LucNeo#2 cells (closed diamonds) and LucNeo719R cells (open diamonds) and the proliferation of these cells (shadow bars and open bars, respectively). The cells were cultured in the presence of various concentrations of (a) HA-719, (b) TLV, (c) DCV, or (d) VX-222. After incubation for three days, the cells were subjected to real-time RT-PCR and tetrazolium dye methods to measure RNA copy number and viable cell number, respectively.
Mutations of resistant RNA replicon
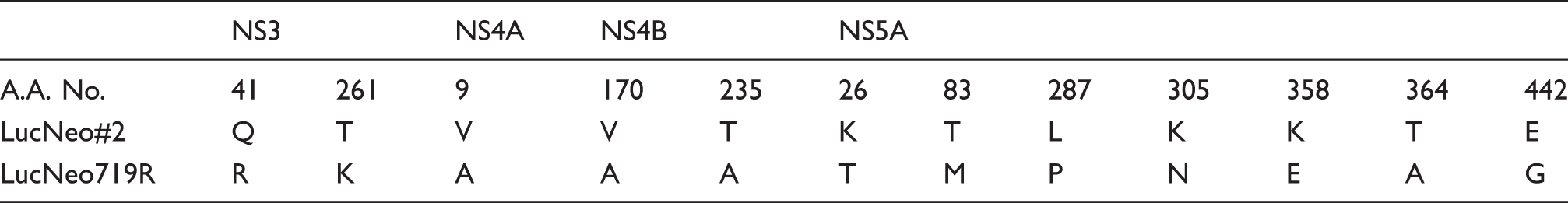
Amino acid mutations of nonstructural HCV proteins in LucNeo719R cells.
Discussion
To gain an insight into the antiviral mechanism of HA-719, we established the resistant clone of HCV replicon cells LucNeo719R and evaluated the anti-HCV activity of various DAAs in this clone. TLV (NS3 protease inhibitor) and VX-222 (non-nucleoside NS5B inhibitor) were equally active in LucNeo719R cells and its parental replicon cells LucNeo#2 (Figure 4(b) and (d), Table 1). Sequence analysis detected Q41R and T261K mutations in NS3 region but did not detect any mutations in NS5B region (Table 2). Q41R mutation was reported as a mutation induced in in vitro selection experiments with the NS3 protease inhibitor simeprevir. 20 However, the engineered Q41R mutation of a replicon construct had no significant impact on the activity of simeprevir. On the other hand, mutations at NS3 positions 43, 80, 155, 156, and 168 conferred various degrees of resistance to simeprevir. These mutations were not identified in the present study. Furthermore, the mutations that confer resistance to other NS3 protease inhibitors, such as TLV, boceprevir, BI-201335 and vaniprevir, were not observed. 21 In our previous report, HA-719 did not inhibit the enzymatic activity of NS3 protease and NS5B polymerase. 14 These results suggest that HA-719 exerts its anti-HCV activity through a mechanism other than NS3 protease and NS5B polymerase inhibition.
Cross-resistance of LucNeo719R cells to DCV (NS5A inhibitor) was not observed (Figure 4(c), Table 1). However, cross-resistance is not always observed among different NS5A inhibitors. For instance, the common mutation Q30E of genotype 1a HCV confers high-level resistance to DCV (∼7500 fold) but only 5‒50 fold resistance to several second generation NS5A inhibitors, including BMS-766, GS-5816, and MK-8742. 22 Therefore, it is still possible that the molecular target of HA-719 is NS5A, which needs to be excluded in further experiments. Sequence analysis identified many mutations in NS5A region. In general, most mutations that confer resistance to currently available NS5A inhibitors are located within the first 100 amino acids of NS5A (domain I). 23 By contrast, we detected many mutations in domain II (residues 250 to 342) and domain III (residues 356 to 447) in LucNeo719R cells (Table 2). CypA binds to NS5A domains II and III, and the mutations in NS5A domains II and III regions conferred drug resistance to CypA inhibitors.12,23–26 In addition, NS5A E442G mutation in LucNeo719R cells was previously identified as a mutation that conferred low-level resistance to CsA. 26 However, cross-resistance of LucNeo719R cells to CsA was not observed in our study (data not shown). These results suggest that the molecular target of HA-719 may not be CypA.
Oxidative stress is involved in HCV replication. For instance, exogenous H2O2 inhibited HCV replication in Huh-7 cells through elevation of cytosolic Ca2+.27,28 Lipid peroxidation products and arsenic trioxide, which produce reactive oxygen species (ROS) and deplete intracellular glutathione, similarly suppressed HCV replication.29,30 CsA and interferon-γ (IFN-γ) also inhibited HCV replication in a redox-sensitive manner. 31 Therefore, antioxidants, such as N-acetyl-L-cysteine (NAC) and vitamin E (VE), accelerated HCV replication or countered the inhibitory effects of ROS on HCV replication.27,28,30,31 Furthermore, NAC and VE canceled the anti-HCV activity of CsA.31,32 Although we also examined whether NAC and VE canceled the anti-HCV activity of HA-719 in LucNeo#2 cells, they did not weaken the anti-HCV activity of HA-719 (data not shown). These results suggest that HA-719 exerts anti-HCV activity through the mechanism other than oxidative stress.
In conclusion, our results suggested that the novel phenanthridinone derivative HA-719 exerts its anti-HCV activity through a mechanism not involved in NS3 protease, NS5B polymerase, or oxidative stress. NS5A itself or host cellular factors interacting with NS5A still remains a possible target for the inhibition of HCV by HA-719. This suggests its interaction with another allosteric site of NS5A, which may not be the target of existing NS5A inhibitors.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work described in this paper was partially supported by Grants-in-Aid for Scientific Research from the Science and Technology Incubation Program in Advanced Regions, Japan Science and Technology Agency (JST).
