Abstract
Purpose
We aimed to use machine learning (ML) algorithms with clinical, lab, and imaging data as input to predict various outcomes in traumatic brain injury (TBI) patients.
Methods
In this retrospective study, blood samples were analyzed for glial fibrillary acidic protein (GFAP) and ubiquitin C-terminal hydrolase L1 (UCH-L1). The non-contrast head CTs were reviewed by two neuroradiologists for TBI common data elements (CDE). Three outcomes were designed to predict: discharged or admitted for further management (prediction 1), deceased or not deceased (prediction 2), and admission only, prolonged stay, or neurosurgery performed (prediction 3). Five ML models were trained. SHapley Additive exPlanations (SHAP) analyses were used to assess the relative significance of variables.
Results
Four hundred forty patients were used to predict predictions 1 and 2, while 271 patients were used in prediction 3. Due to Prediction 3’s hospitalization requirement, deceased and discharged patients could not be utilized. The Random Forest model achieved an average accuracy of 1.00 for prediction 1 and an accuracy of 0.99 for prediction 2. The Random Forest model achieved a mean accuracy of 0.93 for prediction 3. Key features were extracranial injury, hemorrhage, UCH-L1 for prediction 1; The Glasgow Coma Scale, age, GFAP for prediction 2; and GFAP, subdural hemorrhage volume, and pneumocephalus for prediction 3, per SHAP analysis.
Conclusion
Combining clinical and laboratory parameters with non-contrast CT CDEs allowed our ML models to accurately predict the designed outcomes of TBI patients. GFAP and UCH-L1 were among the significant predictor variables, demonstrating the importance of these biomarkers.
Keywords
Introduction
According to the Global Burden of Disease Study 2016 and a subsequent 2018 study, the global incidence of traumatic brain injury (TBI) is estimated at 27–69 million cases each year.1,2 There were 223,135 TBI-related hospitalizations in 2019 and 64,362 TBI-related deaths in 2020 in the United States, indicating that this is a significant public health issue with devastating outcomes. 3 Considering the broad spectrum of its clinical manifestations and injury heterogeneity, as well as the high incidence and prevalence of TBI worldwide, prognostication has become increasingly important.
Research has been conducted to generate predictor variables, methods, and models for enhancing the precision of prediction of outcomes following a TBI, which aids in treatment decisions and the management of expectations.4–10 The Glasgow Coma Scale (GCS) has been used to promptly classify the severity of TBI for decades and it correlates with patient mortality and morbidity. 11 However, GCS is subject to interobserver variation and poorly correlates with mortality and morbidity at the favorable end of the spectrum. 11 In addition, imaging is crucial for identifying and prognosticating TBI patients. Imaging is necessary for TBI patients to identify injuries that may require immediate procedural intervention, that may benefit from early medical therapy or neurologic supervision, and to determine the prognosis of patients. 12 Especially non-contrast head computed tomography (CT), which quantifies neuro-parenchymal and bony injury, is vital for diagnosing, prognosis, and triaging TBI in the acute phase.11,12 Multiple scoring or classifying systems, including the Marshall, Rotterdam, Helsinki, and Neuroimaging Radiological Interpretation System (NIRIS) scores, are based solely on non-contrast CT.13–16 Moreover, a common data element (CDE) database for CT imaging was developed to facilitate the eventual systematic characterization of the natural history and prognostic factors in TBI. 17 In addition to these clinical and imaging tools, the diagnostic and prognostic capabilities of blood-based biomarkers, such as S100B, glial fibrillary acidic protein (GFAP), ubiquitin C-terminal hydrolase L1 (UCH-L1), Interleukin 10, and Amyloid β1-40, have been investigated.18–20
As the number of identified prognostic factors increases, physicians are required to manage more complex clinical, laboratory, and imaging data, necessitating the employment of more sophisticated analytical techniques. Deep learning and machine learning (ML)-based prediction models can employ this vast quantity of data to develop accurate prognosis models. While utilizing deep learning and ML models allows us to process vast amounts of data efficiently, interpretability issues make clinicians hesitant. 21 Especially deep learning models are referred to as “black boxes” frequently. 22 An interpretable ML-based predictive system incorporating clinical variables, blood biomarkers, and imaging biomarkers may improve the prognostic prediction, triage management and treatment strategy in TBI patients. Therefore, in the present study, we aimed to use ML algorithms using the clinical, lab, and imaging data as input to predict various outcomes in TBI patients while utilizing the SHapley Additive exPlanations (SHAP) approach to establish the interpretability of the models.
Materials and methods
Ethical considerations
The Institutional Review Board at Stanford University approved the study. The study complied with the Health Insurance Portability and Accountability Act. Patients or their legal authorized representatives were consented in the subacute phase of the trauma, after the initial workup was completed, and consent was obtained for being able to use their blood collected as part of the standard of care and for collecting their outcome.
Patient selection
In this retrospective cohort study, all consecutive patients transported to the Stanford Healthcare Emergency Department by ambulance or helicopter, for whom a trauma alert was initiated according to the established criteria 23 and who underwent non-contrast head CT scan due to TBI suspicion between November 2015 and April 2017 were evaluated for eligibility. The guide provided in the reference explains how and when the trauma alert was activated. The inclusion criteria were as follows: 1) Patients over the age of 18 transported by ambulance or helicopter with a trauma alert activated; 2) patients underwent a non-contrast head CT for suspected TBI; and 3) patients had blood biomarker results. Both penetrating and blunt trauma patients were included.
Data extraction
From electronic medical records, demographic and clinical information was extracted. Specifically, age, gender, the time elapsed between trauma and admission, GCS, mechanism of TBI, other major intracranial injury (OMII), for example, a stroke that would have caused the patient to get involved in a trauma, and other major extracranial injury (OMEI) at the time of trauma were collected. OMEI included fractures, cardiac diseases, pain and weakness, organ lacerations, operations, pneumothorax, and infections. The following follow-up data were also collected: disposition from the emergency department, disposition at discharge, duration of intensive care unit stay, and mortality data.
Our Institutional Review Board (IRB) approved the use of blood collected for clinical care but not utilized for standard-of-care clinical analysis within 48 h. We collected these blood samples just before discarding them per laboratory protocol. Using a sandwich enzyme-linked immunosorbent assay, samples were analyzed for GFAP and UCH-L1. All blood samples were collected, processed, and analyzed using the same procedures. The lower limit of quantification for GFAP was 3 pg/mL, and the limit of detection was not determined. The lower limit of quantification of UCH-L1 was 14 pg/mL, and the limit of detection was 6 pg/mL. Specimens with signal levels exceeding the quantification range were diluted and retested.
Three categories of outcomes are designed to predict: discharged or admitted for further management (prediction 1); in-hospital mortality (deceased or not deceased [prediction 2]); and course of hospital stay (admission only, prolonged stay, or neurosurgery performed [prediction 3]). Prolonged stay was defined as advanced care unit stay.
Imaging
The head CT was done within 30 min of admission. The non-contrast head CTs were reviewed by two experienced neuroradiologists, M.W. and B.J., with 25 and 12 years of experience in neuroradiology, respectively, for TBI CDEs developed by the National Institutes of Health. 17 The presence or absence of the following CDEs was documented: skull fracture; pneumocephalus; hemorrhage; parenchymal injuries; mass effect; herniation; or shift. The volumes of epidural hemorrhage, subdural hemorrhage, cerebral hematoma, and contusion were manually measured. We manually estimated the volume of these lesions using consecutive CT imaging slices. The volume attributed to each slice was calculated by multiplying the measured area by the slice thickness. We determined the total volume of the hemorrhage by summing the volume contributions of all the slices where the lesions were visible. Additionally, midline shift distance was recorded. A 5-mm cutoff was established for the diagnosis of midline shift.
Predictor variables
Predictor variables included age, gender, GCS, mechanism of TBI, the time elapsed between trauma and admission, OMII, OMEI, GFAP, UCH-L1, skull fracture, pneumocephalus, hemorrhage, parenchymal injuries, mass effect, herniation, or shift, and the volumes of epidural hemorrhage, subdural hemorrhage, cerebral hematoma, and contusion. CT CDEs and novel blood biomarkers were added to the traditional predictor variables in TBI, such as GCS, to evaluate their predictive value in ML models.
Machine learning models
All analyses were generated using Python (version 3.7). Before training the model, a correlation matrix of TBI features was conducted to determine the potential correlations between the different features. The cohort was randomly divided into two sets for the predictions of discharged or admitted (prediction 1) and deceased or not deceased (prediction 2): training set (75%, 330 patients) and testing set (25%, 110 patients). The cohort was randomly divided into a training set (80%, 215 cases) and a testing set (20%, 56 cases) for the prediction of admission only, prolonged stay, or neurosurgery performed (prediction 3). Several ML models, including XGBoost, Random Forest, decision tree, support vector machines, and logistic regression, were trained using the training sets to compare their performances. During training, cross-validation was employed to prevent model overfitting and enhance its robustness. While four folds were used for training, one-fold of the training set data was used for validation. The ML model with the best average performance across all cross-validation sets was utilized for further testing. Using a grid search strategy, the hyperparameter was tuned. We evaluated the relative significance of predictor factors using SHAP. SHAP values represent the significance and direction of associations between features and outcomes. The individual contribution of each feature to the model predictions can be visualized using a matrix of SHAP values. Thus, the function of each model feature was represented in a more comprehensible fashion.
Statistical analysis
Fisher’s exact test and Chi-square test were utilized to determine the differences between categorical data, such as gender. The Kruskal–Wallis and Wilcoxon rank sum tests were used to compare continuous data differences. Using Cohen’s Kappa coefficient with or without quadratic weighting, correlations between predictions and the ground truth were compared. All statistical analyses were conducted utilizing RStudio (version 4.1.0). The level of statistical significance was set at 0.05 for all analyses.
Results
The initial screening included 662 patients admitted to the emergency department with trauma alert and available non-contrast head CT scans. Eight patients under the age of 18, four with restricted records, and four with underlying brain pathologies unrelated to TBI, such as brain tumors, were excluded. One hundred eighty-two patients were excluded for lack of blood samples. Additionally, 24 patients were excluded due to the absence of GFAP and UCH-L1 test results. A total of 440 patients were finally included in the study. It is worth noting that our patient cohort was utilized in another study.
24
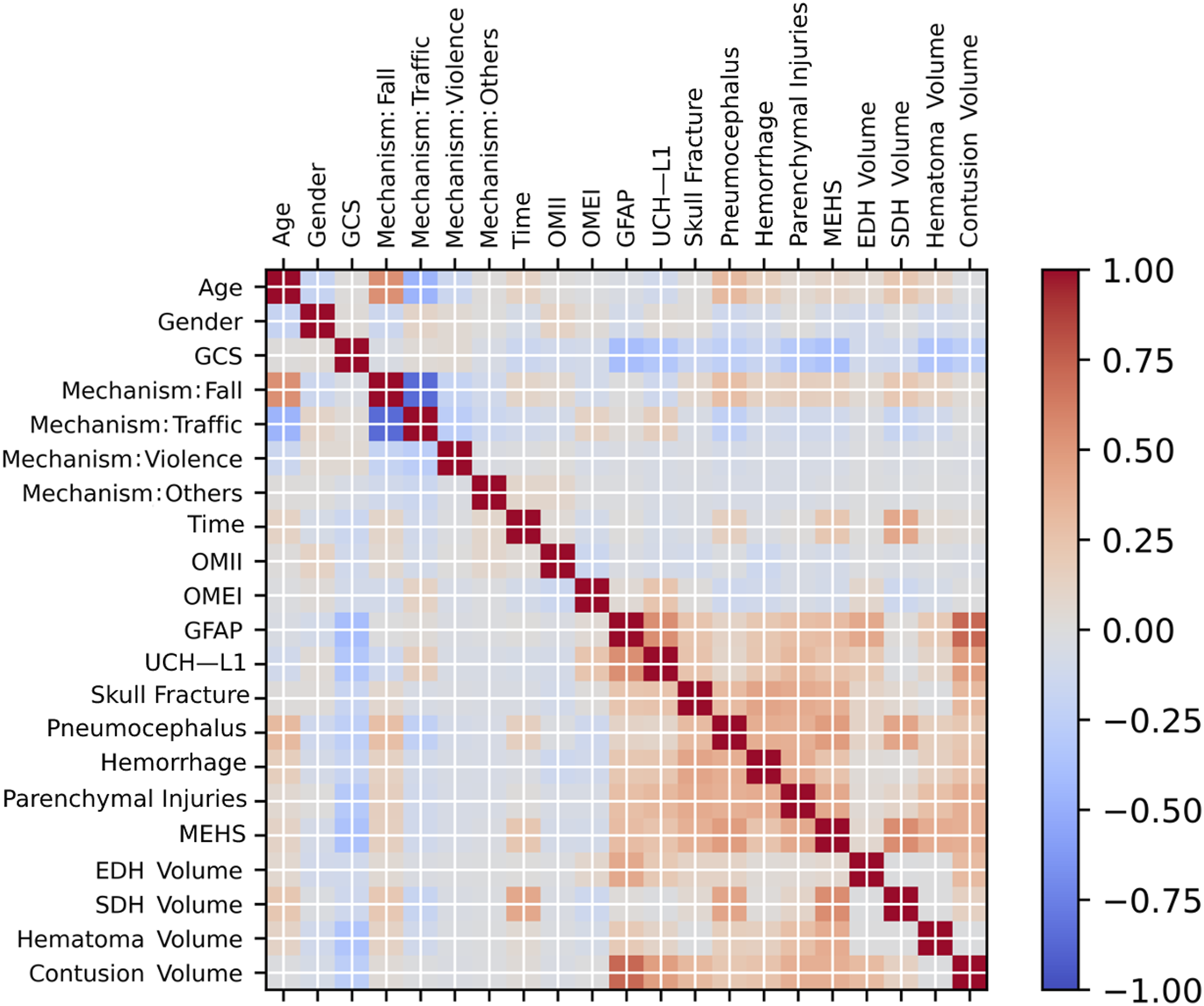
Four hundred forty patients were used to predict predictions 1 (discharged or admitted for further management) and 2 (deceased or not deceased). Two hundred seventy-one patients were utilized in the prediction of prediction 3 (admission only, prolonged stay, or neurosurgery performed). Figure 1 depicts the selection of patients. Figure 2 demonstrates the correlation matrix of TBI features. There were relatively high positive correlations between GFAP and the volume of contusions. Patientselection. Thecorrelationmatrixoftheincludedfeatures.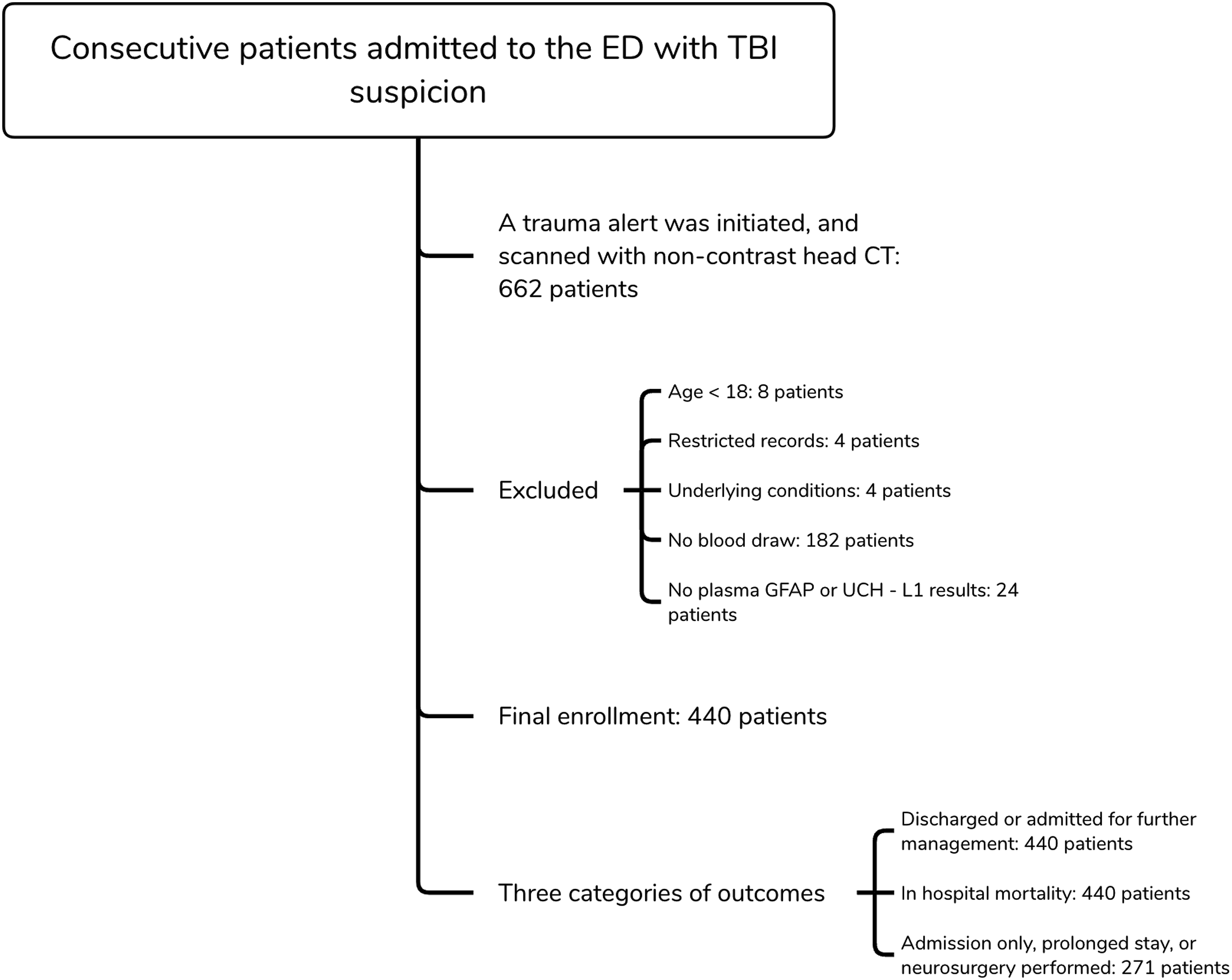
Prediction 1 and 2
Characteristics of the patient population used in predictions 1 and 2.
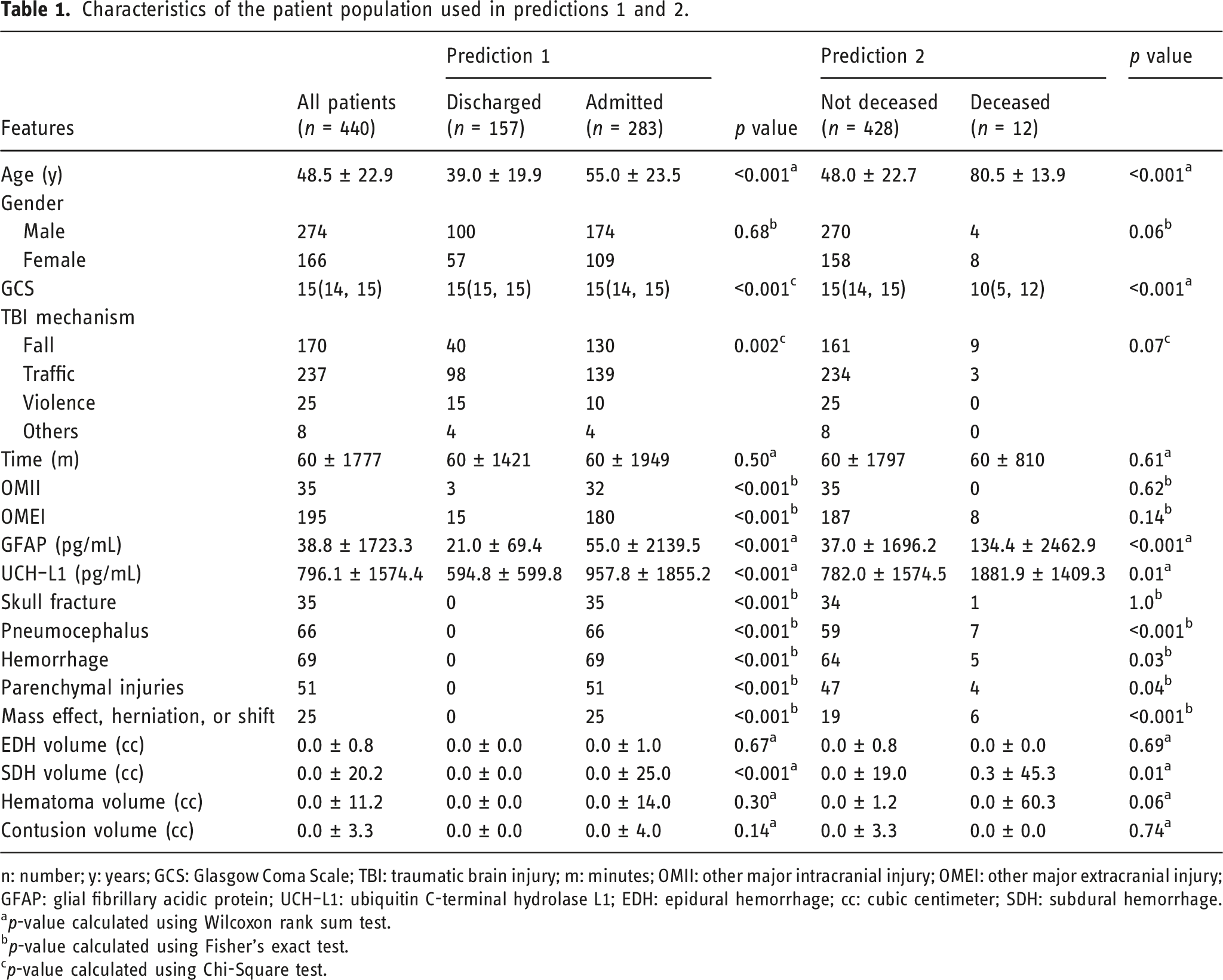
n: number; y: years; GCS: Glasgow Coma Scale; TBI: traumatic brain injury; m: minutes; OMII: other major intracranial injury; OMEI: other major extracranial injury; GFAP: glial fibrillary acidic protein; UCH–L1: ubiquitin C-terminal hydrolase L1; EDH: epidural hemorrhage; cc: cubic centimeter; SDH: subdural hemorrhage.
ap-value calculated using Wilcoxon rank sum test.
bp-value calculated using Fisher's exact test.
cp-value calculated using Chi-Square test.
During the training phase with cross-validation, Random Forest models achieved an average accuracy of 1.00 with a Kappa value of 0.99 for prediction 1 and an accuracy of 0.99 with a Kappa value of 0.82 for prediction 2. The average accuracy of XGBoost was 0.98 and 0.97 for predictions 1 and 2, respectively, making it the second-best performer. The average accuracies of the decision tree, support vector machines, and logistic regression models were below 0.90. Based on the initial results, only Random Forest models were used in the testing stage and analyzed with SHAP.
During the testing stage, the model for prediction 1 (discharged or admitted for further management) produced the most accurate results. The test set accuracy was 0.95, and the Kappa value was 0.88. In the testing stage, the accuracy of the model for prediction 2 (deceased or not deceased) was 0.98, with a Kappa value of 0.49.
According to the results of the SHAP analyses, the five most important features for prediction 1 were, in descending order: OMEI, hemorrhage, UCH-L1, OMII, and age. The top five features for prediction 2 were GCS, age, GFAP, UCH-L1, and mass effect, herniation, or shift, in descending order. Figure 3 depicts the bar plots and beeswarm plots for the results of SHAP analyses for predictions 1 and 2. (a) Bar and (b) beeswarm plots of the features for the result of SHAP analysis in prediction 1, and (c) bar and (d) beeswarm plots of the features for the result of SHAP analysis in prediction 2.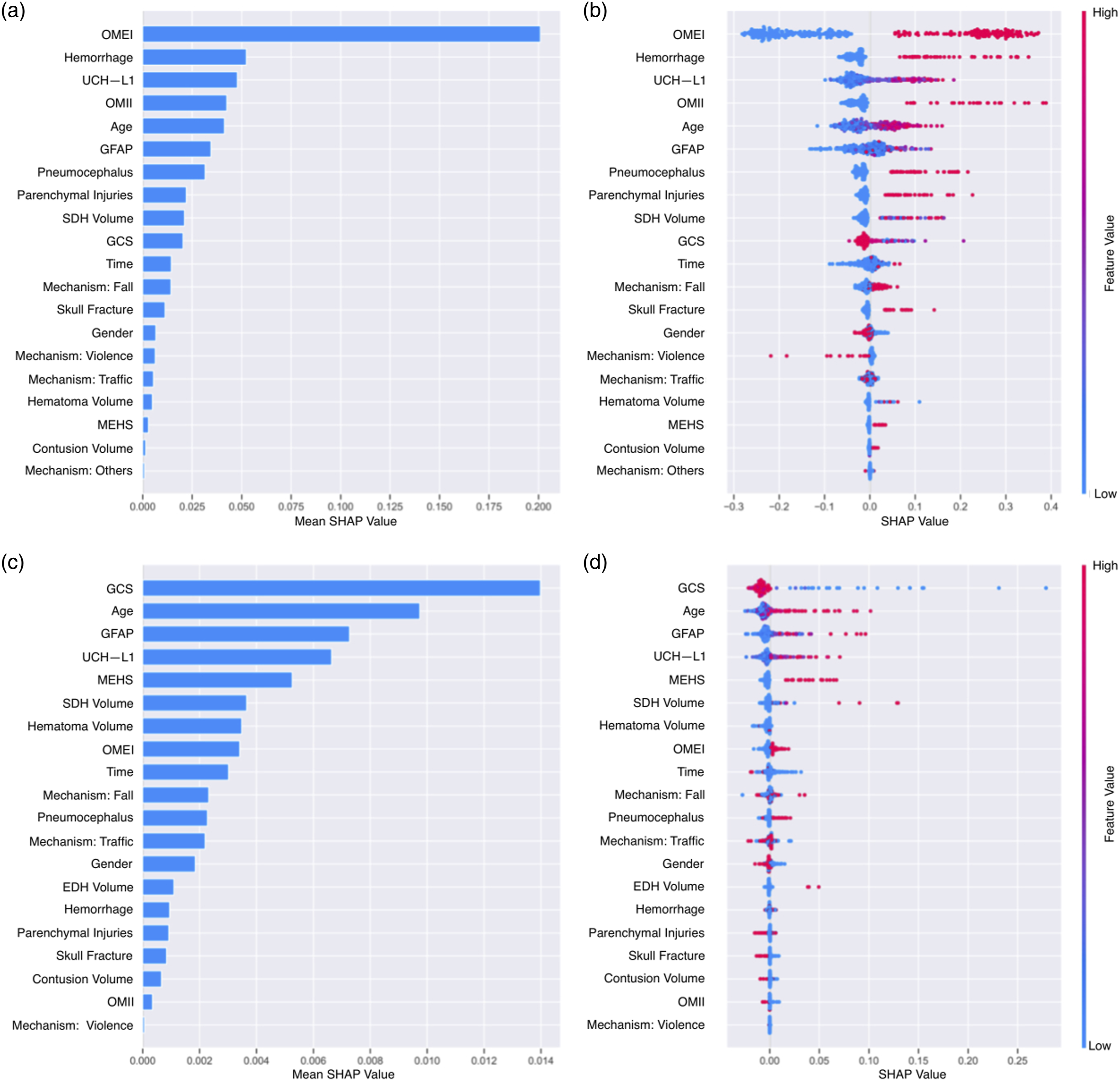
Prediction 3
Characteristics of the patient population used in prediction 3.
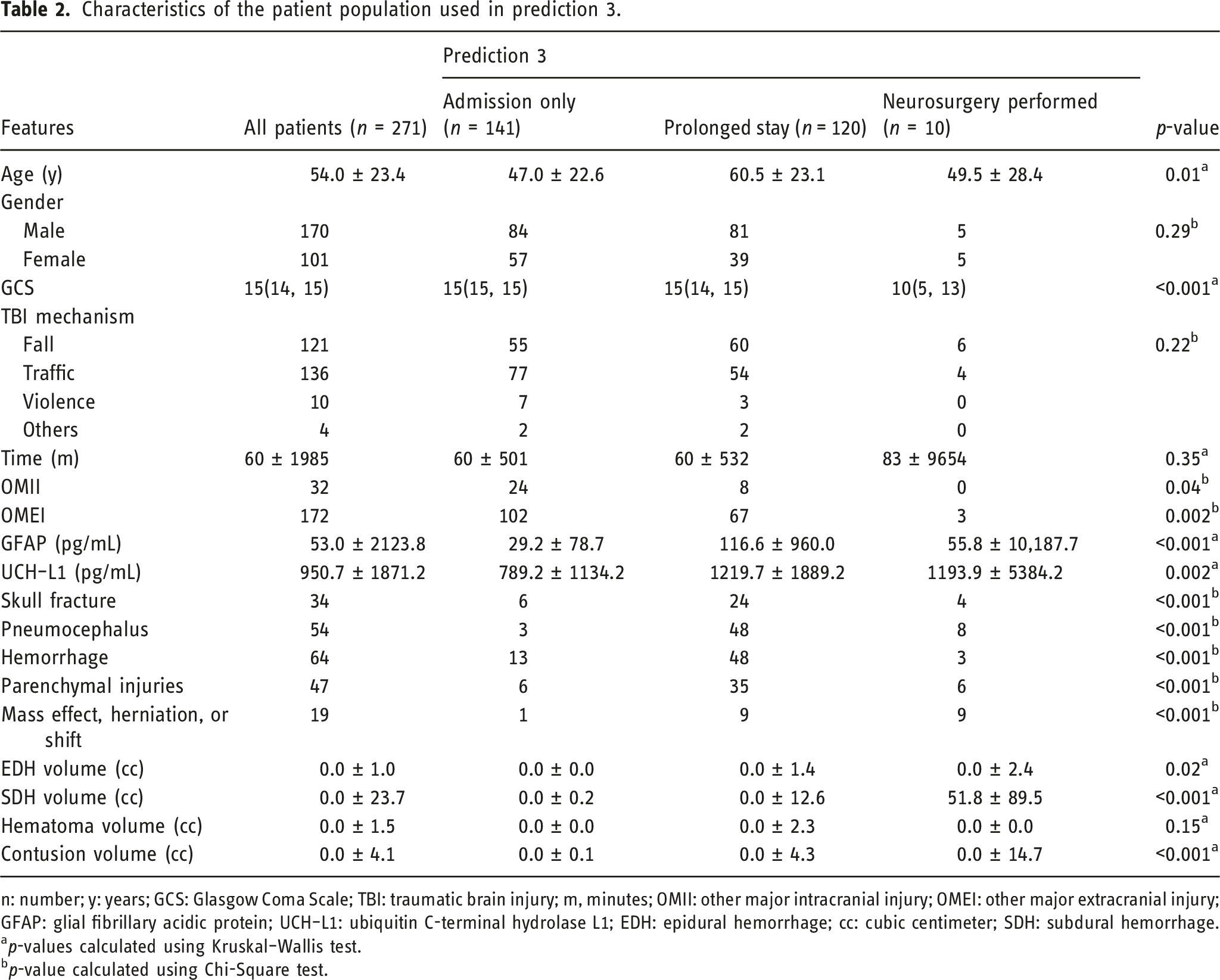
n: number; y: years; GCS: Glasgow Coma Scale; TBI: traumatic brain injury; m, minutes; OMII: other major intracranial injury; OMEI: other major extracranial injury; GFAP: glial fibrillary acidic protein; UCH–L1: ubiquitin C-terminal hydrolase L1; EDH: epidural hemorrhage; cc: cubic centimeter; SDH: subdural hemorrhage.
ap-values calculated using Kruskal–Wallis test.
bp-value calculated using Chi-Square test.
In the training phase with cross-validation, the Random Forest model achieved a mean accuracy of 0.93 and a Kappa value of 0.88 for prediction 3. XGBoost's average accuracy for prediction 3 was 0.92, making it the second-best model. The average accuracies of the decision tree, support vector machines, and logistic regression models were less than 0.90. Based on these results, only the Random Forest model was utilized and analyzed with SHAP during the testing phase. In the testing phase, the accuracy of the model for prediction 3 was 0.82, with a Kappa value of 0.72.
Since prediction 3 does not contain a binary outcome with three distinct outcomes, the significance of the features varies for each outcome. Overall, GFAP, subdural hemorrhage volume, pneumocephalus, UCH-L1, and hemorrhage were, in descending order, the five most significant features for prediction 3 based on the results of the SHAP analyses. When predicting admission only and prolonged stay, the top five most significant features did not change, except for the order of UCH-L1 and pneumocephalus in predicting prolonged stay. The top five predictors of “neurosurgery performed” were pneumocephalus, mass effect, herniation, or shift, GCS, GFAP, and subdural hemorrhage volume, in decreasing order, which differed from the overall results. Figure 4 depicts the bar plots and beeswarm plots for the results of SHAP analyses for prediction 3. (a) Bar plot of the features for the result of SHAP analysis in prediction 3. (b) Swarmplots for the results of SHAP analyses in predictions of the admission only, (c) prolonged stay, and (d) neurosurgery.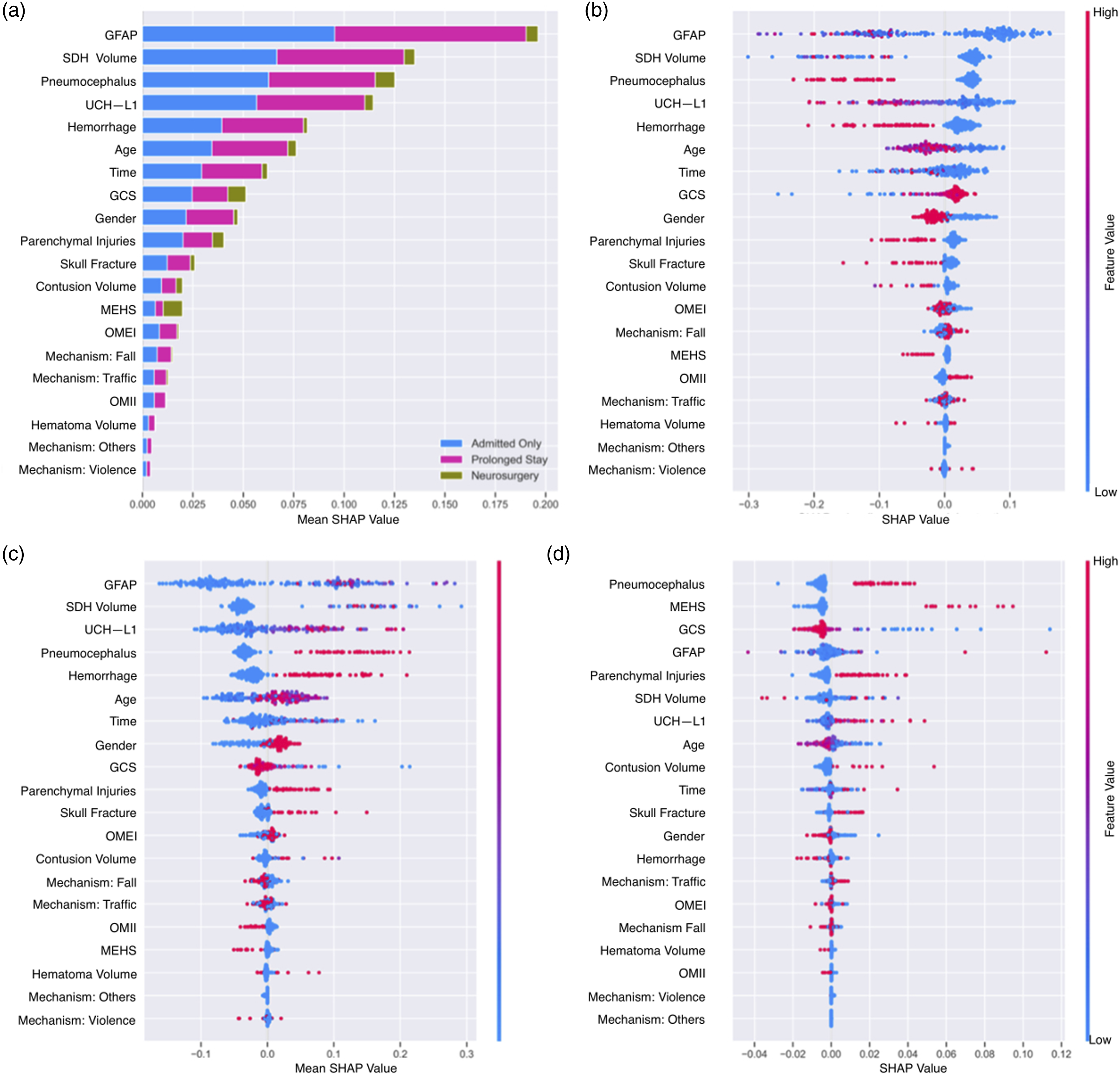
Performances of Random Forest models.
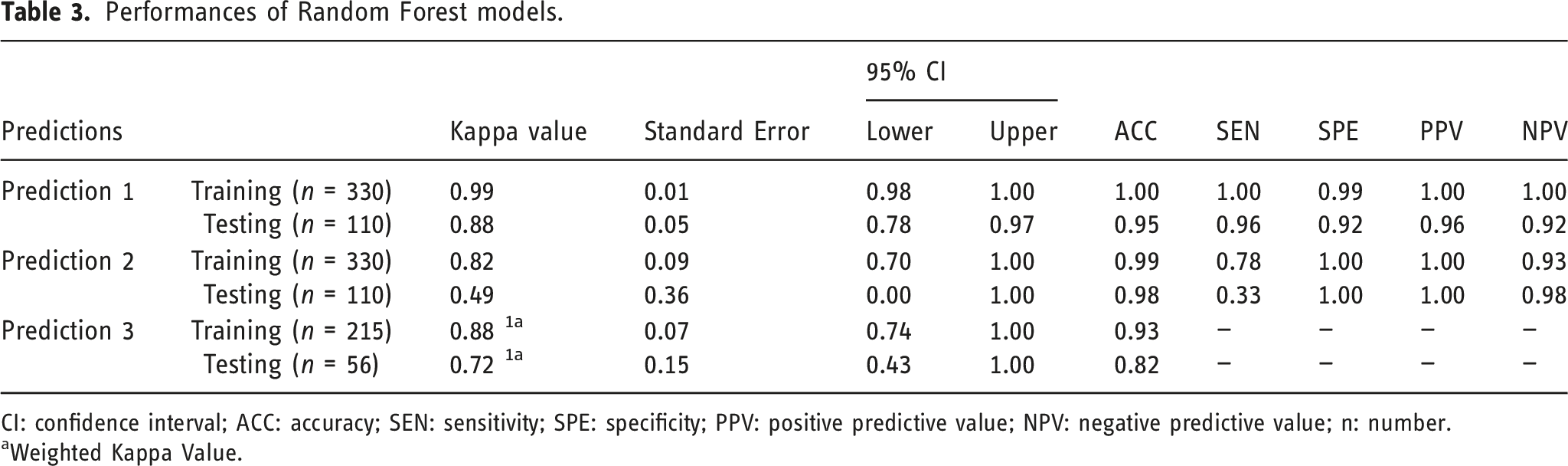
CI: confidence interval; ACC: accuracy; SEN: sensitivity; SPE: specificity; PPV: positive predictive value; NPV: negative predictive value; n: number.
aWeighted Kappa Value.
Discussion
This study presents a series of ML models that accurately predict the groups stratified based on the first decision at the emergency department (discharged or admitted for further management), mortality, and hospital course (admission only, prolonged stay, or neurosurgery performed) in TBI patients. We have chosen these outcomes to predict in order to assess the value of CT CDEs and novel blood biomarkers in real-world clinical scenarios, with the goal of improving the prediction of TBI patient prognosis. The most successful model was Random Forest for prediction 1 (discharged or admitted for further management), prediction 2 (deceased or not deceased), and prediction 3 (admission only, prolonged stay, or neurosurgery performed) with accuracies of 0.95, 0.98, and 0.82 in the test sets, respectively.
Predicting TBI outcomes with prognostic models, deep learning, or ML is not a novel concept. In the context of the International Mission for Prognosis and Clinical Trials in TBI (IMPACT) model, our study investigates the potential of ML algorithms by incorporating clinical, laboratory, and imaging data to predict outcomes. While the IMPACT model is well-established and focuses on clinical, imaging, and demographic factors, our study aims to supplement it by incorporating blood biomarkers and other additional predictors. Furthermore, there are studies describing models to predict the functional outcome or The Glasgow Outcome Scale-Extended,5,25–29 and more recent studies using images as inputs.8,30,31 Furthermore, there are studies in the literature similar to ours that describe models for predicting in-hospital mortality,29,32–35 early mortality,36–38 discharge position,39,40 need for hospital admission, 6 emergency neurosurgery, 41 and length of hospital stay. 4 In addition to contributing to the body of knowledge by describing the efficacy of incorporating ML into patient care to predict multiple outcomes simultaneously in TBI patients, this study is unique since it has used blood biomarkers such as GFAP and UCH-L1, and non-contrast CT CDEs as input variables. Although other studies have utilized CT data as input variables, the CDEs employed in this study make it different. CDEs were established to promote the use of similar nomenclature and criteria in defining intracranial injuries across all imaging examinations; thus, we believe their use is crucial.
In our study, the model for prediction 1 (discharged or admitted for further management) produced a test set accuracy of 0.95, a sensitivity of 0.96, a specificity of 0.92, a positive predictive value (PPV) of 0.96, and a negative predictive value (NPV) of 0.92. Similarly, Marincowitz et al. predicted the need for hospital admission, which was defined as a deterioration measure intended to encompass the need for the hospital admission. 6 In this well-written study, their model had an accuracy of 0.32, a sensitivity of 0.99, a specificity of 0.07, a PPV of 0.29, and an NPV of 0.94. In their study, the five most predictive factors were injury severity on CT (Modified Marshall Criteria), GCS, number of injuries, the hospital admitted to, and subdural hemorrhage. Similarly, our study revealed that OMEI (similar to number of injuries), OMII (similar to number of injuries), and hemorrhage were among the top five most predictive factors for admission or discharge. Consequently, even though the designed outcomes were not identical, our results align with theirs, supporting our findings. In addition, UCH-L1 was among the top five most predictive factors in our study, indicating the importance of the biomarker.
Our model for prediction 2 (in-hospital mortality) yielded a test set accuracy of 0.98, a sensitivity of 0.33, a specificity of 1.00, a PPV of 1.00, and an NPV of 0.98. In the testing set, only three of 110 patients were deceased, which may be the primary reason for the low sensitivity. The best-performing model of Matsuo et al. in the test set showed a sensitivity of 0.88, a specificity of 0.88, and an accuracy of 0.89 in predicting in-hospital mortality. 29 Furthermore, the best-performing model of Abujaber et al. in the test set yielded an accuracy of 0.96, a sensitivity of 0.73, a specificity of 0.99, a PPV of 0.88, and an NPV of 0.97. 32 Moreover, the best-performing model of Hsu et al. in the test set yielded an accuracy of 0.93, a PPV (precision) of 0.93, and a sensitivity (recall) of 0.93. 33 Finally, the best-performing model of Rau et al. was artificial neural network-based and yielded an accuracy of 0.92, a sensitivity of 0.84, and a specificity of 0.93 in the test set. 34 While it is not ideal to directly compare models based on these results, our model produced comparable outcomes, except for sensitivity, due to the small number of deceased patients in our test set. As in our study, GCS was the most or second most significant factor in three of these studies.29,33,34 UCH-L1 and GFAP were among our study’s top five most predictive factors, again indicating these biomarkers' significance.
Our model for prediction 3 (course of hospital stay [admission only, prolonged stay, or neurosurgery performed]) yielded a test set accuracy of 0.82. Similarly, Moyer et al. predicted the need for emergency neurosurgery within the 24 h following the admission. 41 However, our model's outcome was not binary, with three possible options. Pneumocephalus, mass effect, herniation or shift, GCS, and parenchymal injuries were among the top five variables in predicting the need for neurosurgery, highlighting the significance of imaging. GFAP was also among the top five most predictive factors for the need for neurosurgery.
Bazarian et al. demonstrated the high sensitivity and NPV of the UCH-L1 and GFAP tests for predicting the absence of intracranial lesions on head CT scans. 18 While this supports its potential significance in excluding the requirement for a CT scan in TBI patients in emergency departments, our research also indicates that their level can be utilized to predict TBI patients' hospital course. Furthermore, other studies have utilized these biomarkers to predict the prognosis of TBI patients. Korley et al. demonstrated that day-of-injury plasma concentrations of GFAP and UCH-L1 have good to excellent predictive value for death and unfavorable outcomes, particularly in patients with a GCS score of 3 to 12. 42 Moreover, Helmrich et al. showed that serum biomarkers, particularly UCH-L1, provide incremental prognostic value for functional outcome prediction after TBI when combined with established prognostic models. 43 There are other studies highlighting the diagnostic and prognostic potential of protein biomarkers, specifically GFAP and UCH-L1, in TBI patients.44,45 It has been shown that GFAP and UCH-L1 can aid in detecting intracranial lesions, predicting unfavorable outcomes, and guiding therapeutic interventions, according to meta-analyses and longitudinal studies.44,46 Moreover, evaluating multiple biomarkers with distinct cellular origins can improve outcome prediction models, highlighting the importance of incorporating these biomarkers into evaluations of TBI patients. 45 Additionally, numerous studies have proposed cutoff values by illustrating a correlation between elevated GFAP and UCH-L1 levels with TBI diagnosis or prognosis. Papa et al. demonstrated that, utilizing a UCH-L1 cutoff level of 0.09 ng/mL for detecting intracranial lesions on CT, the classification performance yielded a sensitivity of 100% and a specificity of 21%. 47 Moreover, the classification performance for predicting the necessity of neurosurgical intervention, using a UCH-L1 cutoff level of 0.21 ng/mL, achieved a sensitivity of 100% and a specificity of 57%. 47 Furthermore, Mondello et al. demonstrated that an analysis of the glial-neuronal ratio, defined as the ratio of GFAP concentration (ng/mL) to UCH-L1 concentration (ng/mL), for predicting focal mass lesions with a cutoff value of >1.43, resulted in a specificity of 83% and a sensitivity of 60%. 48
Our study has several limitations. First, it shares the inherent constraints of retrospective studies. Additionally, since our findings were sourced from a single institution, it is crucial to validate these results externally within a broader and more diverse patient population. Such validation ensures the findings’ consistency and generalizability across varied settings and populations. We also did not account for comorbidities that could impact patient recovery. Furthermore, an imbalance in training data distribution might cause learning algorithms to underperform on the minority class. 49 Meanwhile, imbalances in the test data can lead to misleading conclusions with certain metrics. 49 Therefore, it is especially noteworthy to approach the findings with caution due to the class imbalances observed in Predictions 2 and 3.
Conclusion
ML might be helpful in accurately predicting the hospital course of TBI patients by combining clinical and laboratory parameters with non-contrast head CT CDEs. Blood biomarkers like GFAP and UCH-L1 were among the significant variables for prediction, demonstrating the originality of our study. ML models have the potential to enhance prognostic classification.
Supplemental Material
Supplemental Material - Enhancing hospital course and outcome prediction in patients with traumatic brain injury: A machine learning study
Supplementary Material for Enhancing hospital course and outcome prediction in patients with traumatic brain injury: A machine learning study by Guangming Zhu, Burak B Ozkara, Hui Chen, Bo Zhou, Bin Jiang, Victoria Y Ding and Max Wintermark in The Neuroradiology Journal.
Footnotes
Author contributions
Conceptualization, G.Z. and M.W. Methodology, G.Z. and M.W. Software, G.Z. and B.O. Validation, B.O., H.C., B.Z., B.J., V.D., and M.W. Formal analysis, G.Z. Investigation, G.Z. and M.W. Resources, M.W. Data Curation, H.C., B.Z., B.J., and V.D. Writing - Original Draft, G.Z. and B.O. Writing - Review & Editing, M.W. Visualization, B.B.O. and G.Z. Supervision, M.W. Project administration, M.W.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
