Abstract
Objective
Extrachromosomal circular DNA (eccDNA) has been shown to be involved in several physiological and pathological processes including immunity, inflammation, aging, and tumor. However, the expression of eccDNA in cartilage has not been reported until now. In this study, we aimed to investigate the landscape of eccDNA in articular cartilage and analyze the potential roles in osteoarthritis (OA).
Methods
The samples of articular cartilage were obtained from total knee arthroplasty (TKA) donors with OA. The mitochondrial DNA (mtDNAs) and the linear DNAs from chondrocytes of articular cartilage were removed. Then the eccDNAs were enriched for cir-DNA sequencing. After quality control evaluation, we systematically revealed the identified eccDNA data including size distribution, the size range, and sequence pattern. Moreover, we explored and discussed the potential roles of eccDNA in OA via motif analysis and Gene Ontology (GO)/Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis.
Results
The chondrocytes from OA cartilage contained an abundance of eccDNAs, which was termed as OC-eccDNAs (OA cartilage-derived eccDNA). The characteristics of OC-eccDNAs were tissue-specific, including the distribution, the size range, and sequence pattern. Moreover, the functional analysis indicated that eccDNA may be involved in the homeostasis maintenance of chondrocytes and participated in the process of OA.
Conclusions
Our data first showed the landscape of eccDNA in articular cartilage and preliminarily indicated the potential roles of eccDNA in OA.
Introduction
Osteoarthritis (OA) is a common degenerative disease of the musculoskeletal system. The total number of patients with OA disease all over the world has exceeded 250 million, and the number of patients is still rising rapidly and continuously. 1 With the aging of our society, the incidence rate of OA is increasing, which will become the most important factor leading to the dysfunctions of the elderly and seriously affect the quality of life of the patients. 2 At present, the clinical therapeutic effect of OA is still not ideal. Therefore, it is urgent to strengthen the relevant research so as to provide new theoretical support for the prevention and treatment of OA.
OA is a total joint disease involving cartilage, subchondral bone, and synovium. 3 Among them, cartilage injury is the core pathological change of OA and the basic element of OA disease diagnosis. A variety of biological factors such as aging, inflammation, and mechanical stimulation will destroy the homeostasis of chondrocyte and then aggravate OA progression. 4 The articular cartilage consists of chondrocytes and extracellular matrix, which mainly depends on the infiltration of joint fluid to maintain nutrition. As there is no distribution of blood vessels, nerves, and lymphatic vessels, the repair and regeneration ability of articular cartilage after injury is very poor. 5 Until now, there are no effective measures to protect OA chondrocytes from degeneration. More studies on the homeostasis maintenance of chondrocytes in the process of OA needed to provide a new potential target for this disease.
Extrachromosomal circular DNA (eccDNA) refers to the circular DNA derived from genomic DNA but free of chromosomes. Recent studies have shown that eccDNA is extensively expressed in different tissues and may be involved in varied physiological and pathological processes including those of immune system, inflammation, aging, and tumors.6-9 However, the expression of eccDNA in cartilage tissue and its pathological significance in OA are still unknown. Here, we extracted eccDNA from total knee arthroplasty (TKA) donors and first investigated the characteristics of eccDNA in articular cartilage using cir-DNA sequencing, and discussed the potential roles of eccDNA in OA.
Methods
Human Cartilage Tissue Collection and Isolation of Chondrocytes
Human cartilage was obtained from OA patients (n = 10, aged 69.3 ± 6.53 years, 4 male and 6 female patients). Subjects with tumors, diabetes, or other severe diseases in the last 5 years were excluded. Human articular cartilage explants were cultured according to the previously described methods. 10 Briefly, freshly obtained germ-free human cartilage was washed with phosphophate buffer saline (PBS) and cut into 4-mm-diameter blocks by the bistoury. Subsequently, the bocks were digested in high-glucose dulbecco’s modified eagle medium (DMEM) supplemented with 0.2% type II collagenase for 6 hours. Then the digestion suspension was collected and centrifuged at 400 g for 5 minutes following the filtration with a 40-μm cell strainer. Finally, the sediment (primary chondrocyte) was re-suspended with high-glucose DMEM containing 10% fetal bovine serum (FBS) and cultured in a 5% CO2 incubator.
Circle-Seq
The method of Circle-Seq chondrocyte eccDNA was optimized based on the description of Henrik Devitt Møller et al., including multiple steps as follows: (1) Total DNA was extracted using a QIAamp DNA Mini Kit (Qiagen, Santa Clarita, USA); (2) then, total DNA was alkaline treated to separate chromosomal DNA, lipids, and protein by rapid DNA denaturing-renaturing, followed by column chromatography on an ion exchange membrane column (Plasmid Mini AX; A&A Biotechnology, Pomerania, Poland); (3) the remaining linear DNA was removed by exonuclease (Plasmid-Safe ATP-dependent DNase, Epicentre, San Diego, USA), assisted by rare-cutting endonuclease MssI (only support Homo sapiens) that digested mitochondrial circular DNA (mtDNA, 16 kb) and made additional accessible DNA ends for exonuclease; (4) EccDNA-enriched samples were used as a template for phi29 polymerase reactions (REPLI-g Midi Kit) amplifying DNA; (5) amplified circular DNA was cleaned (AMPure XP beads) and sheared by sonication (Bioruptor) to an average fragment size of 200-300 bp; (6) libraries for next-generation sequencing were prepared using the NEBNext Ultra DNA Library Kit for Illumina (San Diego, USA) according to the manufacturer’s protocol (New England Biolabs, Ipswich, USA) and subjected to sequencing on Illumina NovaSeq 6000 using PE150, collecting up to 80 million paired-end reads/samples.
EccDNA Mapped and Identified Method for Circle-Seq Data
The sequencing reads from Circle-Seq were aligned to reference genome sequences (GRChg38) using the BWA program (Burrows-Wheeler-Alignment Tool, https://github.com/lh3/bwa) to dig eccDNA originated from the chromosome. Subsequently, the candidate reads were sorted, labeled, and ranked by Circle-Map to detect and identify eccDNA, which refers to the practices of Henrik Devitt Møller.
Motif Analysis of Junction
The HOMER’s findMotifsGenome.pl tool was used for Motif analysis. The input file is the peak file and the genome fasta file. The 200bp DNA sequence around the strat and the end of junction is extracted according to the peak file, and the sequence is compared with the Motif database, including HOMER and JASPAR to obtain the Motif.
Enrichment Analysis
The “Cluster Profiler” R package was utilized to conduct Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses based on the eccDNA-related genes. P values were adjusted with the Benjamini-Hochberg (BH) method.
Results
Cir-DNA Sequencing Revealed the Landscape of eccDNA in Articular Cartilage from OA Patients
First, we harvested articular cartilage (n = 10) from OA patients after TKA (Suppl. Fig. S1A) and collected eccDNA of chondrocytes for cir-DNA sequencing ( Fig. 1A and Suppl. Fig. S1B). The quality of cir-sequencing output reads was evaluated as excellent (Suppl. Fig. S1C and S1D). Then we used BWA software to perform genomic alignment of reads with GRch38. For mapped reads, we utilized the Circle-Map software for eccDNA identification and detected 39,342 eccDNA (detected eccDNA) ( Fig. 1A ). Next, we developed a filter standard of split reads >1 to identify the detected eccDNA, and finally 15,121 eccDNA (identified eccDNA) were obtained ( Fig. 1B ). Subsequently, we analyzed the size of the identified eccDNA in Figure 1B . We found that the size distribution of eccDNA in chondrocytes was between 0.1 kb and 10 kb, which was similar to the size distribution of eccDNA in muscle tissue but was distinct from that of serum (<1 kb) and tumor cells (1 kb-1 mb) ( Fig. 1C ). As to the distribution of eccDNA on chromosomes, the results uncovered that chondrocyte eccDNA was distributed in all other regions except for Y chromosome and the short arm of chromosome 13, 14, and 15, which was similar to that in Hela cells, but a bit different from that of glioma cells and yeast cells ( Fig. 1D ). In addition, previous studies have demonstrated that eccDNA may derive from repetitive sequences on the genome. 11 Indeed, we found that most of the reads corresponding to eccDNA of chondrocytes matched the repetitive sequences of the genome, including long interspersed nuclear elements (LINEs, 19.2%), short interspersed nuclear elements (SINEs_Alu, 18.8%), long terminal repeat (LTR, 11.3%), and a small number of telomere sequences ( Fig. 1E ), which was partly the same with muscle tissue, but different from plasma. 12 Excluding the overlapping parts, 8,915 genes were annotated, which included mRNA (6,468), long noncoding RNA (1,075), multipletype (697), pseudogene (219), miRNA (1), and others (455) ( Fig. 1F ). These data indicate that articular cartilage contains an abundance of eccDNA, whose size range and chromosome origin may be relatively specific.
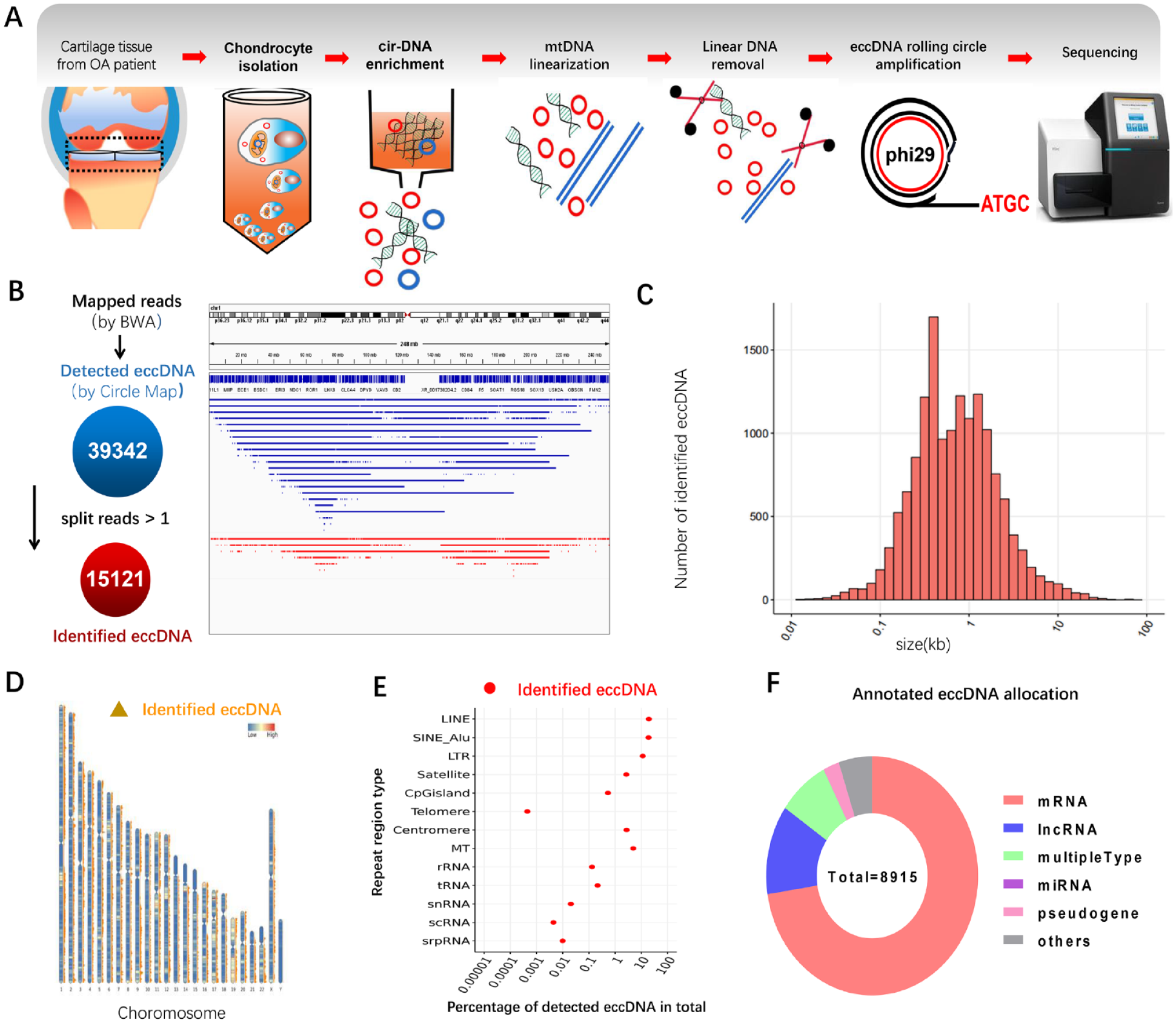
The landscape of extrachromosomal circular DNA in articular cartilage. (
OA-Related Transcription Factors Involved in eccDNA Cyclization
Previous studies have suggested that the junction point of eccDNA and its nearby area played an important role in eccDNA formation. Therefore, we separately extracted the 200-bp sequence from upstream and downstream of the junction point start site or the stop site, combing with the HOMER software to blast the extracted sequence with the known motif and predict the de novo motif ( Fig. 2A ). To explore the role of the transcription factor in eccDNA formation and effects, we performed transcription factor-based enrichment analysis on mapped known motifs and de novo motifs, respectively, and extracted the TOP20 (based on P value) collection of the two outputs ( Fig. 2B , Suppl. Table S1 and S2). Interestingly, most of the transcription factors with a high P value were highly related to the development and degeneration of cartilage, such as MYB, STAT3, NKX, NRF2, ZNF215, and FOX. Moreover, some transcription factors including KLF4, NRF2, STAT3, and NF-kB have been proved to be the key pathogenic factors in OA ( Fig. 2B ). These results indicate that OA-related transcription factors may be involved in the formation of eccDNA in chondrocytes.
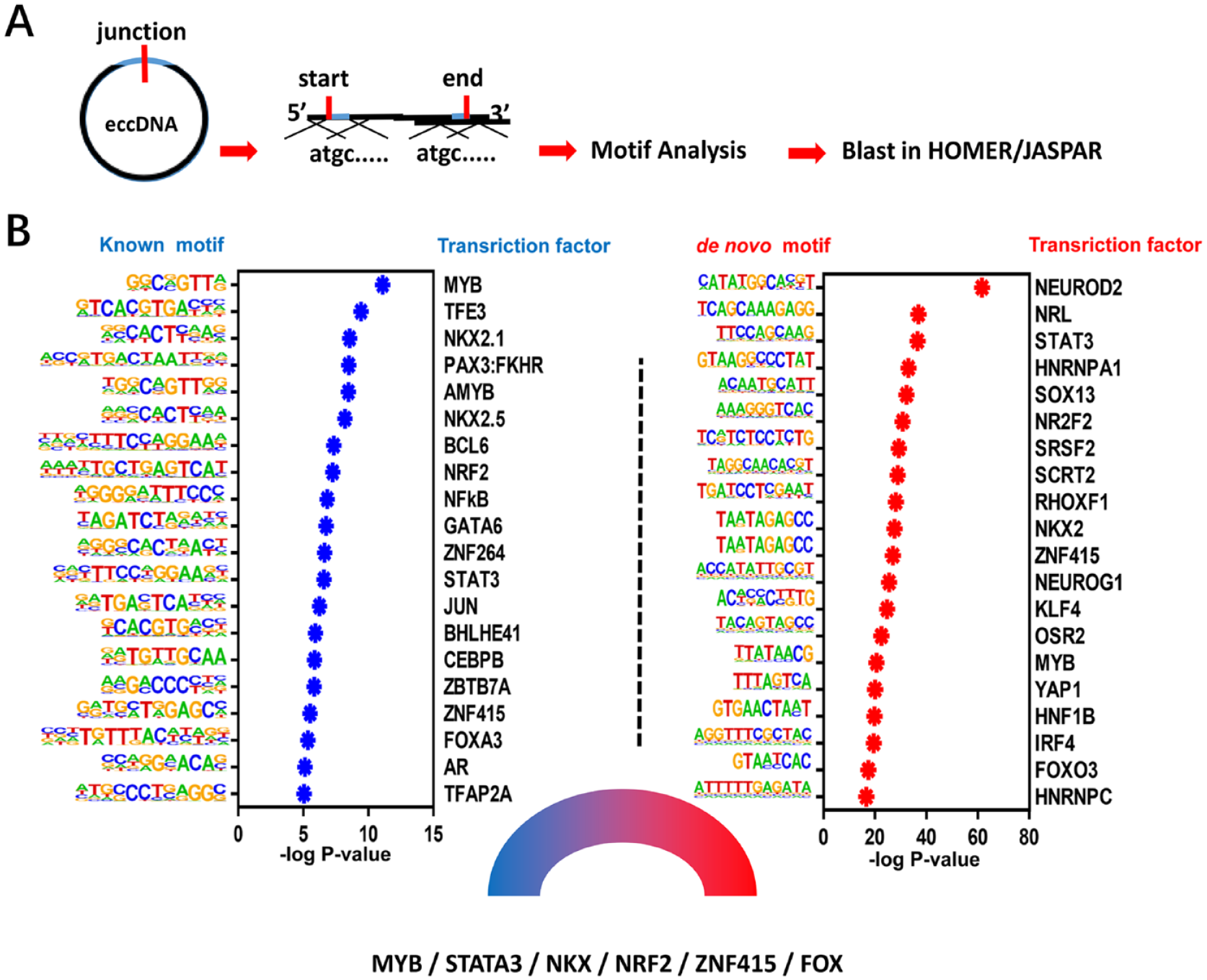
The potential transcription factors that participate in the formation of eccDNA. (
The GO/KEGG Pathway Analysis for the eccDNA in OA
Next, we performed GO and KEGG analyses of the aforementioned genes (
Fig. 3A
and
B
). The data suggest that eccDNA may be involved in the pathophysiological regulation of degenerated chondrocytes in morphology, cell polarization, cell-extracellular matrix (ECM) interaction, and so on. From the top enriched terms, we found that abundant genes were involved in anabolism and catabolism, senescence, and inflammation of chondrocyte, such as Aggrecan (ACAN), matrix metalloproteinases (MMPs), a disintegrin and metalloproteinases with thromnospondin motifs (ADAMTSs), interleukin-1β (IL-1β), and so on (
Fig. 3C
), indicating that eccDNA may participate in the regulation of cartilage homeostasis. Moreover, GO and KEGG analysis both revealed that the target genes of eccDNA were significantly enriched in nervous system-related pathways, such as axon guidance and synapse organization, suggesting a potential role of eccDNA in joint pain of OA. Besides, we sorted the eccDNA according to circle score to analyze the genes annotated by TOP20 eccDNA and found that these genes are mainly related to the formation of nerve synapses (CNTN4, CABLES), cell stress response (SGK1), chromatin stability and DNA damage repair (NT5DC1, PDS5A, STK11), protein degradation and reparation (FBXW7, PCMT1, UBXN8), transcription inhibition (CBFA2T2, NCOR2), and cell adhesion and skeletal remodeling (MAGI1, ARHGEF26) (
Fig. 3C
and
D
). Furthermore, certain eccDNAs that potentially targeted OA-related genes from
Figure 3C
and
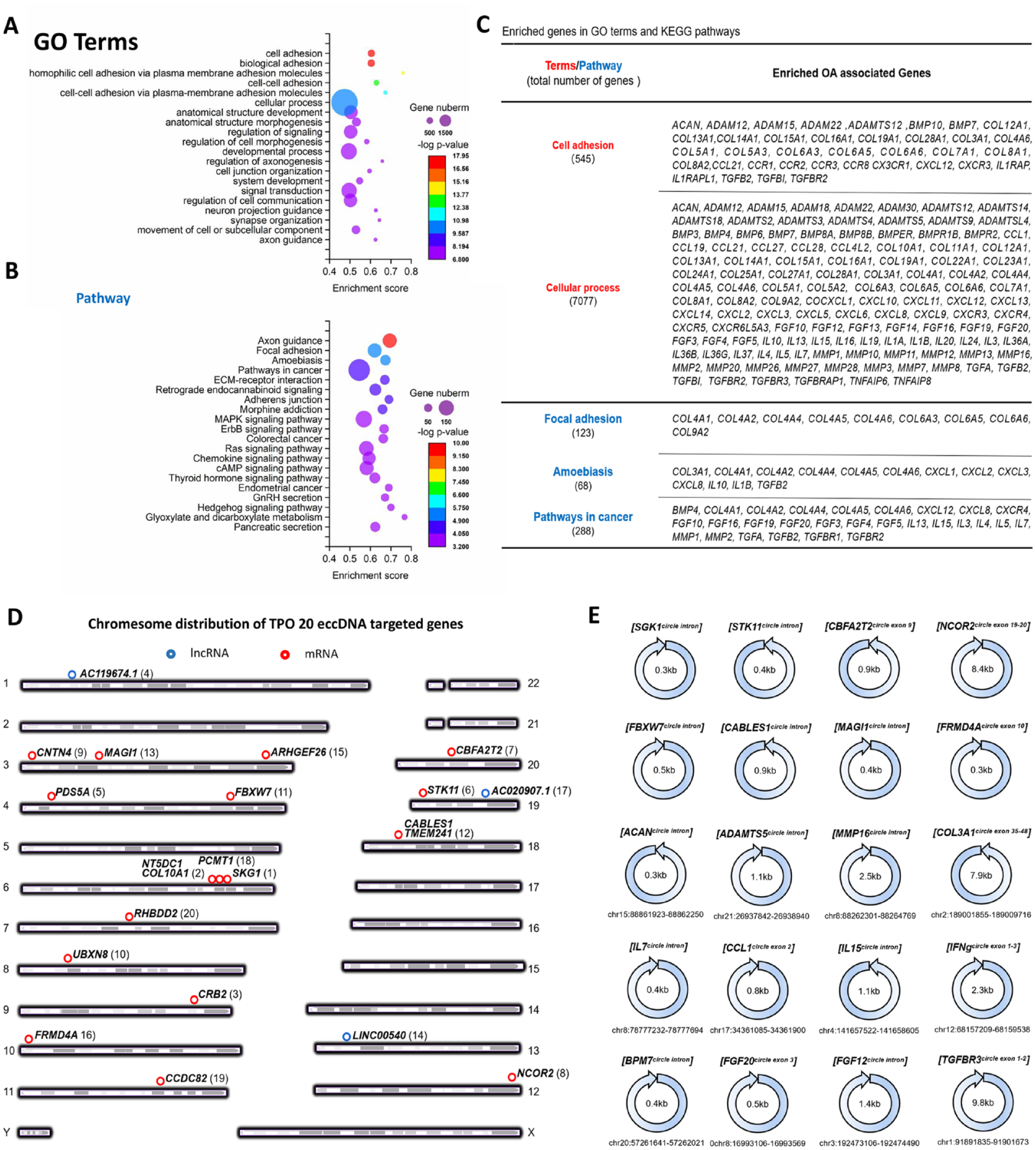
The pathway and target gene analysis for eccDNA suggested the potential functions in OA. (
Discussion
In recent years, more and more studies have shown that eccDNA is closely involved in the occurrence and development of several diseases such as cancer.9,13 Turner et al.9,13 reported that eccDNA was found in nearly half of human cancers, and its frequency varied by tumor type. They also revealed that driver oncogenes in eccDNA were amplified commonly, which contributed to the transcription of oncogenes out of chromosomes.14,15 Wu et al. 7 investigated the structure of eccDNA by integrating ultrastructural imaging, long-range optical mapping, and computational analysis of whole-genome sequencing, and their data suggested that oncogenes encoded on eccDNA increased copy number with high transcription levels. In addition, eccDNA displays significantly enhanced chromatin accessibility than it is typical of chromosomes and has more ultra-long-range interactions with active chromatin, 7 indicating it might possess powerful transcriptional capacity potentially. Moreover, the studies performed by Zou et al. 16 and Jiang et al. 17 suggested that eccDNA might act as a pool of micro-RNA or its precursor in tumor progress. Koche et al. also found that cancer-causing lesions can increase the formation of functional eccDNA in neuroblastoma, which may play an important role in tumor progression. 18 Apart from tumor, eccDNA has also been related to the aging process.8,19,20 The accumulation of copper protein 1 (CUP1) eccDNA has unique site specificity in the copper-induced aging model of yeast. 20 Furthermore, the heterogeneity of circular DNA that offers flexibility in adaptation is significantly diminished with age. 8 As aging is the main cause of OA, it is speculated that eccDNA may play an important role in the maintenance of chondrocyte homeostasis and the process of OA disease. Just like tumor cells, gene transcription on eccDNA (not on chromosomal DNA) may be actived in aged chondrocytes and be involved in the OA process. As eccDNA comes from genome, it may regulate chondrocyte phenotypes by epigenetic means, such as affecting the replication and transcription of chromosomal DNA. In addition, as micro-RNA play a importance role in chondrocyte homeostasis,21,22 eccDNA that annotated as micro-RNA might be involved in cartilage destruction through sustaining micro-RNA supply. More studies are needed to investigate the roles and mechanisms of eccDNA on chondrocyte.
Synovium is an important part of the joint sliding system, which is composed of loose connective tissue. Its main functions include secreting synovial fluid, reducing joint surface friction, preventing joint adhesion, providing nutrients for articular cartilage, absorbing and swallowing various metabolites and cell fragments in the joint cavity, and so on. When the joint is subjected to stress injury (such as degeneration, trauma, infection, and rheumatism), the synovium will be stimulated to produce inflammatory reaction and participate in the repair of articular cartilage injury. It is found that chronic and low-grade inflammation is the main feature of OA synovitis, which is closely involved in the formation of OA cartilage damage repair microenvironment. Therefore, trying to regulate the state of synovitis to reshape the microenvironment of OA joint, and then promote the damage and repair of chondrocytes, is expected to become a feasible strategy for clinical OA treatment. However, the formation mechanism of OA synovitis has not been very clear until now. Previous studies have suggested that the fragments from dead chondrocytes and degraded cartilage matrix can stimulate synovial tissue to produce inflammatory response and mediate the formation of chronic synovitis in the process of OA. 23 In addition, our group reported that the exosome-like vesicles from OA chondrocytes with inflammatory stimulation could increase the mature IL-1β production of synovial macrophages and aggravated synovitis in OA. 10 Recently, Wang et al. revealed that eccDNAs are apoptotic products with high innate immunostimulatory activity. Their data indicated that apoptosis inducers can increase eccDNA generation, and eccDNAs can function as potent innate immunostimulants in a manner that is dependent on eccDNA circularity and the cytosolic DNA sensor Sting. 6 Therefore, we deduced that eccDNA from the apoptotic or senescent chondrocytes may participate in the formation of OA chronic synovitis via stimulating synovial inflammatory cells.
This study first mapped out the landscape of eccDNA in articular cartilage and preliminarily indicated the potential roles of eccDNA in OA, which may open up a new vision on the roles and mechanisms of eccDNA in joint diseases. Based on the biological characteristics of eccDNA, we speculate that it may play a potential role in cartilage-related diseases, such as OA. But, it is a pity that the sample size in this study was relatively limited, and validation of bioinformatic results was absent. So, the roles and mechanisms of eccDNA in joint diseases need further research. In addition, the formation mechanisms of eccDNA in chondrocytes and its changes in the pathological process are also worthy of further attention.
Supplemental Material
sj-docx-1-car-10.1177_19476035231205690 – Supplemental material for Cir-DNA Sequencing Revealed the Landscape of Extrachromosomal Circular DNA in Articular Cartilage and the Potential Roles in Osteoarthritis
Supplemental material, sj-docx-1-car-10.1177_19476035231205690 for Cir-DNA Sequencing Revealed the Landscape of Extrachromosomal Circular DNA in Articular Cartilage and the Potential Roles in Osteoarthritis by Wei Xiang, Tongyi Zhang, Song Li, Yunquan Gong, Xiaoqing Luo, Jing Yuan, Yaran Wu, Xiaojing Yan, Yan Xiong, Jiqin Lian, Guangyu Zhao, Changyue Gao, Liang Kuang and Zhenhong Ni in CARTILAGE
Supplemental Material
sj-pdf-4-car-10.1177_19476035231205690 – Supplemental material for Cir-DNA Sequencing Revealed the Landscape of Extrachromosomal Circular DNA in Articular Cartilage and the Potential Roles in Osteoarthritis
Supplemental material, sj-pdf-4-car-10.1177_19476035231205690 for Cir-DNA Sequencing Revealed the Landscape of Extrachromosomal Circular DNA in Articular Cartilage and the Potential Roles in Osteoarthritis by Wei Xiang, Tongyi Zhang, Song Li, Yunquan Gong, Xiaoqing Luo, Jing Yuan, Yaran Wu, Xiaojing Yan, Yan Xiong, Jiqin Lian, Guangyu Zhao, Changyue Gao, Liang Kuang and Zhenhong Ni in CARTILAGE
Supplemental Material
sj-xls-2-car-10.1177_19476035231205690 – Supplemental material for Cir-DNA Sequencing Revealed the Landscape of Extrachromosomal Circular DNA in Articular Cartilage and the Potential Roles in Osteoarthritis
Supplemental material, sj-xls-2-car-10.1177_19476035231205690 for Cir-DNA Sequencing Revealed the Landscape of Extrachromosomal Circular DNA in Articular Cartilage and the Potential Roles in Osteoarthritis by Wei Xiang, Tongyi Zhang, Song Li, Yunquan Gong, Xiaoqing Luo, Jing Yuan, Yaran Wu, Xiaojing Yan, Yan Xiong, Jiqin Lian, Guangyu Zhao, Changyue Gao, Liang Kuang and Zhenhong Ni in CARTILAGE
Supplemental Material
sj-xls-3-car-10.1177_19476035231205690 – Supplemental material for Cir-DNA Sequencing Revealed the Landscape of Extrachromosomal Circular DNA in Articular Cartilage and the Potential Roles in Osteoarthritis
Supplemental material, sj-xls-3-car-10.1177_19476035231205690 for Cir-DNA Sequencing Revealed the Landscape of Extrachromosomal Circular DNA in Articular Cartilage and the Potential Roles in Osteoarthritis by Wei Xiang, Tongyi Zhang, Song Li, Yunquan Gong, Xiaoqing Luo, Jing Yuan, Yaran Wu, Xiaojing Yan, Yan Xiong, Jiqin Lian, Guangyu Zhao, Changyue Gao, Liang Kuang and Zhenhong Ni in CARTILAGE
Footnotes
Acknowledgments and Funding
The authors are grateful to Ms. JiaLi for excellent technical assistance. The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (no. 81871817; 82372495; 82002360; 81772330; 82202770) and the Natural Science Foundation of Chongqing (no. CSTB2022NSCQ-MSX1267; CSTB2022NSCQ-MSX0863).
Author Contributions
Z.N., L.K., and C.G. conceived and designed the experiments. WX, TZ, YG, YW, J.Y., and S.L. performed experiments. Y.X., J.L., and X.L. provided expert advice. All authors analyzed the data. W.X., L.K., X.L., and Z.N. wrote the paper.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
All procedures and protocols were approved by the ethics committee with informed consent obtained (2018128) and carried out in accordance with standard operative procedures.
Data Sharing
The authors declare that all the data will be shared online after the paper is published.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
