Abstract
Purpose
Chondrocytes, isolated from articular cartilage, are routinely utilized in cell-based therapeutics for the treatment of cartilage pathologies. However, restoration of the biological tissue faces hindrance due to the formation of primarily fibrocartilaginous repair tissue. Chondroprogenitors have been reported to display superiority in terms of their chondrogenic potential and lesser proclivity for hypertrophy. In line with our recent results, comparing chondroprogenitors and chondrocytes, we undertook isolation of progenitors from the general pool of chondrocytes, based on surface marker expression, namely, CD166, CD34, and CD146, to eliminate off-target differentiation and generate cells of stronger chondrogenic potential. This study aimed to compare chondrocytes, chondroprogenitors, CD34−CD166+CD146+ sorted chondrocytes, and CD34−CD166+CD146− sorted chondrocytes.
Methods
Chondrocytes obtained from 3 human osteoarthritic knee joints were subjected to sorting, to isolate CD166+ and CD34− subsets, and then were further sorted to obtain CD146+ and CD146− cells. Chondrocytes and fibronectin adhesion-derived chondroprogenitors served as controls. Assessment parameters included reverse transcriptase polymerase chain reaction for markers of chondrogenesis and hypertrophy, trilineage differentiation, and total GAG/DNA content.
Results
Based on gene expression analysis, CD34−CD166+CD146+ sorted chondrocytes and chondroprogenitors displayed comparability and significantly higher chondrogenesis with a lower tendency for hypertrophy when compared to chondrocytes and CD34−CD166+CD146− sorted chondrocytes. The findings were also reiterated in multilineage potential differentiation with the 146+ subset and chondroprogenitors displaying lower calcification and chondroprogenitors displaying higher total GAG/DNA content compared to chondrocytes and 146− cells.
Conclusion
This unique progenitor-like population based on CD34−CD166+CD146+ sorting from chondrocytes exhibits efficient potential for cartilage repair and merits further evaluation for its therapeutic application.
Introduction
Articular cartilage serves to resist compression while providing a frictionless surface for load bearing and joint movement. 1 Hyaline cartilage consists primarily of a collagenous extracellular matrix with interspersed sparse populations of chondrocytes and their progenitors. 2 The relative avascular nature limits intrinsic repair and regeneration. Chronic inflammatory states and traumatic injury cause irreversible cartilage degeneration, and this progression can lead to the development of osteoarthritis. Cell-based therapy as a treatment strategy commonly employs bone marrow–derived mesenchymal stem cells (BM-MSCs) and chondrocytes for repair, with reports exhibiting regeneration with functional improvement. 3 However, histological analysis of the regenerated tissue shows the presence of mixed fibrohyaline and fibrocartilage, which eventually results in inferior biomechanical properties.4-7 To remedy this, the search for alternative strategies with better chondrogenic ability and lower hypertrophic inclination is still underway.
Articular cartilage–derived chondroprogenitors offer promise for efficient cartilage regeneration as they exhibit a commitment toward chondrogenic lineage with lower expression of hypertrophy markers.8-10 To optimize the use of chondroprogenitors, numerous techniques for isolation based on migratory ability, selective adhesion to fibronectin, and specific cell surface marker profile have been studied. 11 The fibronectin differential adhesion assay, a commonly employed technique for isolation of chondroprogenitors, has repeatedly yielded cells demonstrating superiority for chondrogenesis. Some characteristics of the chondroprogenitors include differential adhesion to fibronectin,2,12 differential expression of integrins and Notch proteins,13,14 and high replicative potential, with the ability to form large number of colonies even with low seeding density.15,16 They are also likened to mesenchymal stem cells (MSCs) since they are plastic adherent, express classical MSC markers, do not express hematopoietic markers, and display multilineage potential.17-19 Furthermore, several studies assessing the effects of different culture conditions and interventions such as physioxia 20 and specific growth factors21-23 on chondroprogenitors report its conduciveness for chondrogenesis. Chondroprogenitors, when compared to BM-MSCs and chondrocytes, report a higher expression of chondrogenic genes such as aggrecan, collagen type II, SOX-9, and PRG418,24 and a lower expression of hypertrophic markers such as RUNX2 and collagen type X,8,10,25 indicative of superior potential for neocartilage formation.
Concerning surface markers indicating enhanced chondrogenic profile, several reports have suggested the use of CD166 and CD146 as markers of high chondrogenicity while assessing induced pluripotent stem cells (iPSCs) and MSCs.26-28 Chondroprogenitors obtained using these surface markers displayed decreased heterogeneity, increased proliferation, with robust extracellular matrix production, increased GAG/DNA content, and expression of chondrogenic genes. Specific cell markers can also help distinguish chondroprogenitors from the available pool of chondrocytes, notably following culture expansion when phenotypic drift following release from cartilage makes a comparison between the 2 cell groups difficult.29,30 Additionally, a recent study comparing human chondroprogenitors and chondrocytes demonstrated that progenitors with increased time in culture exhibited significantly lower CD34, higher CD166, and CD146 levels. 19 Contrarily, a few studies have also suggested that eliminating CD146 positive subsets may improve chondrogenic ability.31-33
To optimize chondroprogenitor use in cell-based therapy, our study aimed to see whether a constellation of reported CD markers would help identify progenitors with the highest chondrogenic ability and least tendency for hypertrophy.
Methods and Materials
Tissue Source and Study Design
Adult articular cartilage was obtained from 3 patients (age: 60 ± 8 years, 1 female and 2 males) undergoing total knee replacement surgery for the management of osteoarthritis (OA). The study was conducted after obtaining approval from the institutional review board, and written informed consent was obtained from each patient as per the ethical regulation of our institution. Osteoarthritic joints with a radiological Kellgren-Lawrence score of grade 4 was included. Joints with infectious, inflammatory, and posttraumatic causes were excluded. Cartilage shavings were primarily obtained from non–weight bearing areas with preserved full-depth cartilage. To release chondrocytes, articular cartilage slices were minced and subjected to subsequent overnight enzymatic digestion. Chondroprogenitors were isolated by subjecting the released chondrocytes to fibronectin differential adhesion assay.2,9 Chondrocytes and chondroprogenitors were then characterized for positive and negative MSC markers using fluorescence-activated cell sorting (FACS). In addition, chondrocytes were subjected to flow cytometric associated cell sorting to isolate CD34 negative (CD34−) and CD166 positive (CD166+) subsets, which were further sorted to procure both CD146 positive (CD146+) and negative (CD146−) cells. The unsorted chondrocytes and fibronectin adhesion–derived chondroprogenitors were used as controls. The aforementioned 4 categories, chondrocytes, chondroprogenitors, CD34−CD166+CD146+ sorted chondrocytes, CD34−CD166+CD146− sorted chondrocytes, were subjected to FACS, trilineage differentiation, and RT-PCR (reverse transcriptase polymerase chain reaction) analysis for markers of chondrogenesis and hypertrophy. To differentiate based on their chondrogenic potential, the pellets subjected for chondrogenic differentiation were also evaluated for total GAG/DNA accumulation (
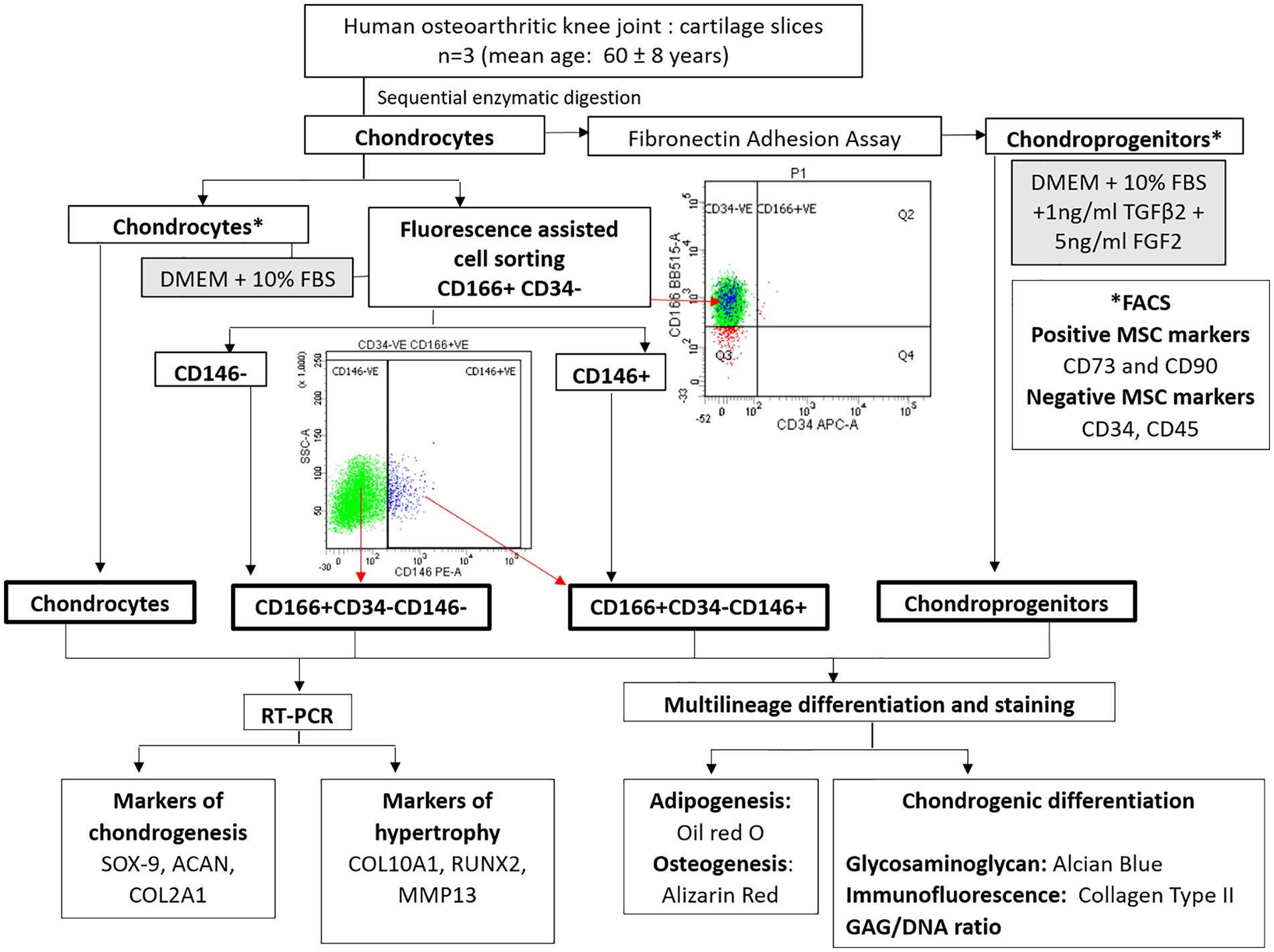
Study algorithm depicting the 4 groups used for comparison and their evaluation parameters. The 4 groups were chondrocytes, chondroprogenitors, CD166+/CD34−/CD146+ chondrocytes, and CD166+/CD34−/CD146− chondrocytes. CD = cluster of differentiation; SOX-9 = sex-determining region Y-box 9; ACAN = aggrecan; COL = collagen; RUNX2 = runt-related transcription factor-2; MMP-13 = matrix metalloproteinase-13.
Cell Isolation and Culture
Chondrocytes
Cartilage slices from OA knee joints were washed in PBS (phosphate-buffered saline) and minced to 1 to 2 mm. They were then subjected to sequential enzymatic digestion with 12 IU pronase (Mat No.: 10165913103, Roche, Switzerland) for 3 hours and 100 IU collagenase type II (Cat No.: LS004176, Worthington, Lakewood, WA) for 12 hours in a 37 °C shaking water bath. The released chondrocytes were filtered using a 40-µm cell strainer and counted using Invitrogen countess based on trypan blue dye exclusion. The obtained fresh cell isolate was loaded at a seeding density of 10,000 cells/cm2 and expanded using DMEM-F12 Ham (D8900, Sigma-Aldrich, 1:1 mixture), supplemented with 0.1 mM ascorbic acid, 10 mM HEPES, and 10% fetal bovine serum (FBS; Cat No.: 10270106, GIBCO, Thermo Fischer Scientific, Waltham, MA). Under standard culture conditions (humidified chamber with 5% CO2), adherent cells were expanded to sub-confluence up to passage 2 for further analysis.
Chondroprogenitors
To isolate chondroprogenitors, chondrocytes at a loading density of 400 cells/cm2 were plated onto precoated fibronectin (10 µg/mL) surfaces (Cat No.: F1141, Sigma Aldrich) as described by Nelson et al. 9 Following 20-minute incubation, nonadherent cells were removed, and the remaining adherent cells were maintained in culture for 2 weeks to achieve minimum population doublings of 5 per clone. The enriched clones were transferred to T-25 flasks and expanded to passage 2 for further analysis. Expansion medium consisted of DMEM-F12 with 10% FBS, 0.1 mM ascorbic acid, 10 mM HEPES, recombinant human transforming growth factor-β2 (1 ng/mL), and fibroblast growth factor-2 (5 ng/mL).
Fluorescence-Activated Cell Sorting
Positive and Negative MSC Markers
Phenotypic analysis of chondrocytes and chondroprogenitors was performed using BD FACS Celesta flow cytometer and analyzed using BD FACS Diva v 5.0.2 software. In brief, trypsinized cells were washed and incubated with each antibody as per the manufacturer’s instructions. The list of antibodies against the conjugated mouse, anti-human surface antigens included (a) positive MSC markers: CD 73-PE (Cat No.: 550257) and CD90-PE (Cat No.: 555596) and (b) negative MSC markers: CD 34-PE (Cat No.: 348057) and CD45-FITC (Cat No.: 555482). Unstained controls were performed for each conjugate.
CD166/CD34/CD146 Sorted Chondrocytes
Passage 2 chondrocytes were subjected to fluorescence-activated cell sorting (BD FACS ARIA III) to initially obtain CD166+/CD34− cells, which were further sorted to obtain CD146+ and CD146− subsets. The conjugates used were BB515 mouse anti-human CD166 Clone 3A6 (Cat No.: 565461), PC mouse anti-human CD34 Clone 581 (Cat No.: 560940) and PE, mouse, anti-human, CD146 Clone P1H12 (Cat No.: 550315). Before sorting, both IgG and unstained controls were run. The sorted cells (CD166+/CD34−/CD146+ and CD166+/CD34−/CD146−) were also expanded with the same culture medium as used for chondrocytes. Following sub-confluence, the cells were trypsinized and subjected to further analysis.
Reverse Transcriptase-Polymerase Chain Reaction
Total RNA was isolated from chondrocytes, chondroprogenitors, and sorted chondrocyte subsets using RNeasy Mini Kit (Qiagen, Cat No.: 74104, Germantown, MD). The concentration and absorbance (A260/A280) were determined using a Nanodrop spectrophotometer. A total of 280 ng of RNA was reverse transcribed to cDNA using Bio First-Strand synthesis system (Takara). Quantitative RT-PCR (7 ng of cDNA/10 µL reaction) was performed using Low Rox SYBR Master Mix Dttp Blue (Takyon, Cat No.: UF-LSMT-B0701, Eurogentec, Belgium). All experiments from 3 biological samples (n = 3) were performed in duplicate. The housekeeping gene (Ct), glyceraldehyde 3-phosphate dehydrogenase (GAPDH), was used to normalize relative expression for all target genes (ΔCt). The results were compared to the chondroprogenitor group, and 2^-ΔΔCt was determined as a result. The expression of genes associated with chondrogenesis, such as SOX-9, ACAN, and COL2A1, and genes associated with hypertrophy, such as RUNX2, MMP13, and COL10A1, were studied. The specific primers for the 6 genes, and the reference gene are listed in Supplemental Table S1.
Trilineage Differentiation
StemPro (Thermo Fischer, Cat Nos.: A1007201, A1007001, and A1007101) differentiation kits were used to induce osteogenic, adipogenic, and chondrogenic lineage differentiation. Cells were seeded at a density of 5,000 cells/cm2 in 12-well culture dishes and expanded to 60% confluence before inducing osteogenic and adipogenic differentiation. Cells grown in standard expansion medium over the same time frame served as controls. A total of 0.5 × 106 cells were placed into Eppendorf tubes, centrifuged at 400g for 12 minutes, and left undisturbed for 48 hours for 3-dimensional pellet cultures. The pellets were then treated with StemPro chondrogenic medium. For 3 weeks, a medium change was performed once every 3 days.
Postdifferentiation Staining
To validate osteogenic differentiation, the control and test cells were fixed in 70% ethanol for 1 hour at 4 °C, washed, and stained for 15 minutes with Alizarin red S solution (2 g in 100 mL, Sigma Aldrich) to confirm the presence of calcium deposition. Similarly, to validate adipogenic differentiation, control and test cells were fixed in 10% buffered formalin for 1 hour at 4 °C, washed, and stained with Oil red O (Cat No.: O0625, Sigma Aldrich) solution for 30 minutes to validate the presence of lipid droplet accumulation. The chondrogenic differentiated pellets were subjected to fixation using 4% paraformaldehyde for 15 minutes and paraffin embedded. To test glycosaminoglycan deposition, sections were stained with Alcian blue (pH 2.5, (Cat No.: J60122, Alfa Aesar) and neutral red (counterstain). To assess the presence of collagen type II, pellets were subjected to antigen retrieval using pronase plus hyaluronidase, overnight incubation with a primary mouse monoclonal anti-collagen type II antibody (DSHB Hybridoma Product II-II6B3, 5 g/mL), followed by secondary antibody IgG (H + L), goat anti-mouse, Alexa Fluor 594 (Invitrogen, Catalogue No.: A11032, 1:100 dilution) for 1 hour, and counterstained using DAPI (10 µg/mL, D9542, Sigma-Aldrich). Alizarin red S and Oil red O images were acquired using Leica DMIL, Alcian blue using Olympus bx43f, and collagen type II using fv1000 Olympus laser scanning confocal microscopes.
Biochemical Analysis: Total GAG/DNA Content
Pellets after chondrogenic differentiation were digested with Papain (125 µg/mL) at 65 °C overnight and used for GAG and DNA measurement. DNA concentrations were determined with Quant-iT Picogreen dsDNA reagent using Lambda DNA as a standard. Fluorescence intensity (Ex: 480 nm, Em: 520 nm) was measured with a SpectraMax i3x Reader (Norwalk, CT). The amount of GAG was determined by the dimethyl methylene blue dye method, using chondroitin 6-sulfate as the standard (Chondrex, Inc, Cat No.: 6022). Absorbance was measured with an Elisa reader at 525 nM. All experiments from 3 biological samples (n = 3) were performed in duplicate. GAG values were normalized to the DNA content.
Statistical Analysis
SPSS version 21.0 was used for data processing, and IGOR Pro Version 5.0.4.8 was used for graphical presentation (Wave metrics Inc.). Gene expression and GAG/DNA content were compared between groups using a one-way ANOVA with post hoc LSD correction. Numerical values were expressed as mean ± standard error mean. A P value of less than 0.05 was considered significant.
Results
Cell Morphology
Chondroprogenitors grew clonally following fibronectin adhesion assay and displayed spindle-shaped morphology on the further expansion (
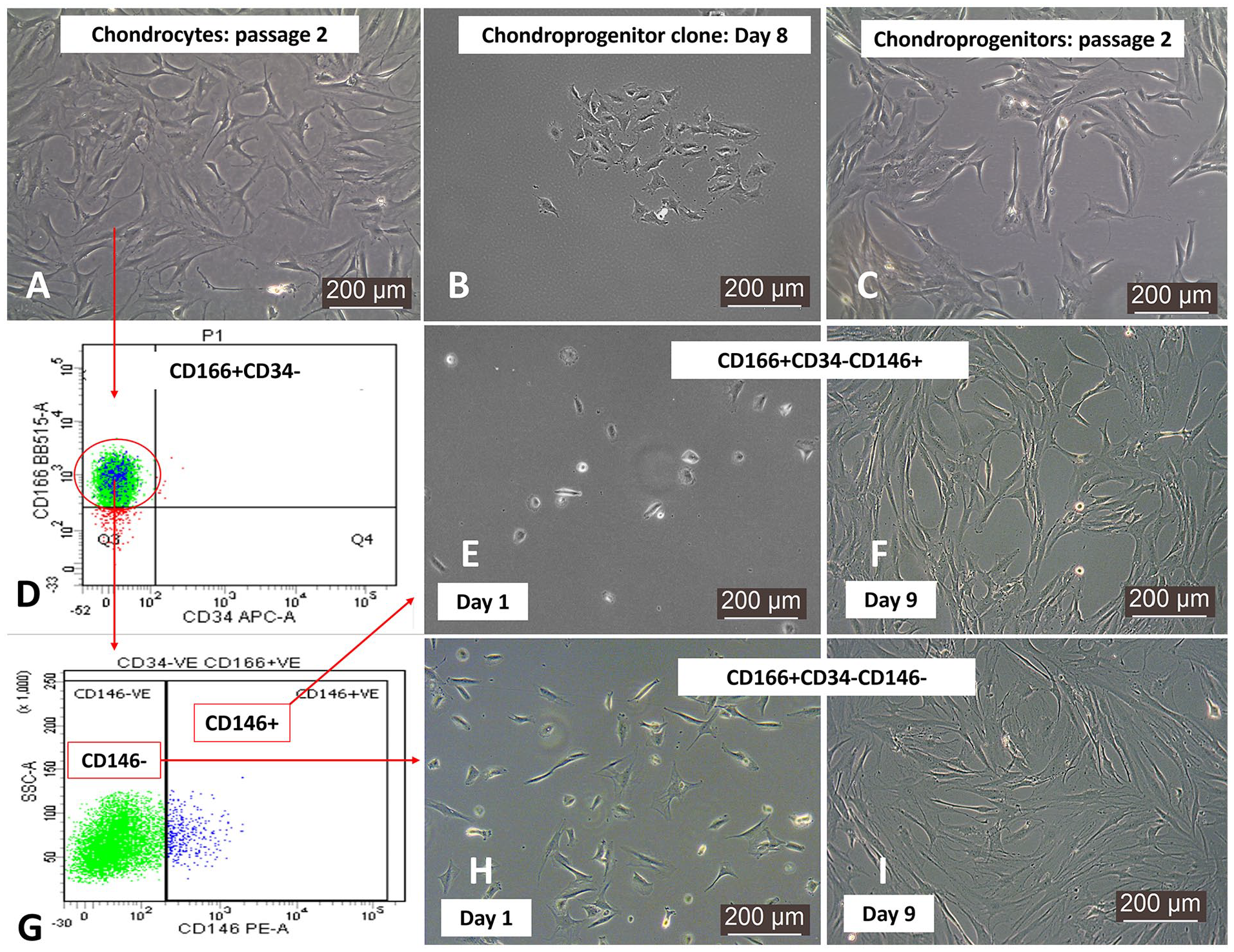
Representative phase contrast images of the different cell groups. (
FACS
When positive MSC markers were compared, both chondrocytes and chondroprogenitors showed high expression of CD73 and CD90. Both showed low expression of CD34 and CD45, with no significant difference observed between them (
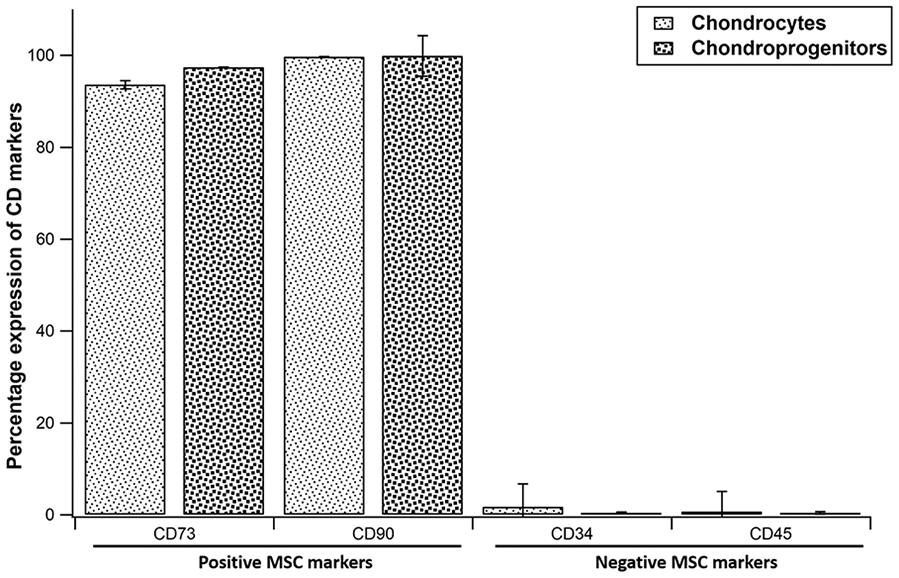
Percentage expression of positive and negative MSC markers on chondrocytes and chondroprogenitors. Data expressed as mean ± standard deviation.
RT-PCR
Chondroprogenitors exhibited significantly higher chondrogenic genes SOX-9 (P = 0.043) and COL2A1 (P = 0.005) than sorted 146− subsets. When compared to chondrocytes, progenitors showed significantly higher SOX-9 (P = 0.016) with lower expression of the hypertrophy marker MMP-13 (P < 0.001). In comparison to the sorted subsets (146+/146−), chondrocytes showed significantly higher expression of hypertrophy genes, COL10A1 and MMP-13 (P < 0.01). Comparison between the 2 sorted subsets revealed that CD146+ cells showed significantly higher levels of both ACAN (P = 0.025) and COL2A1 (P = 0.001) when compared to the CD146− cells (
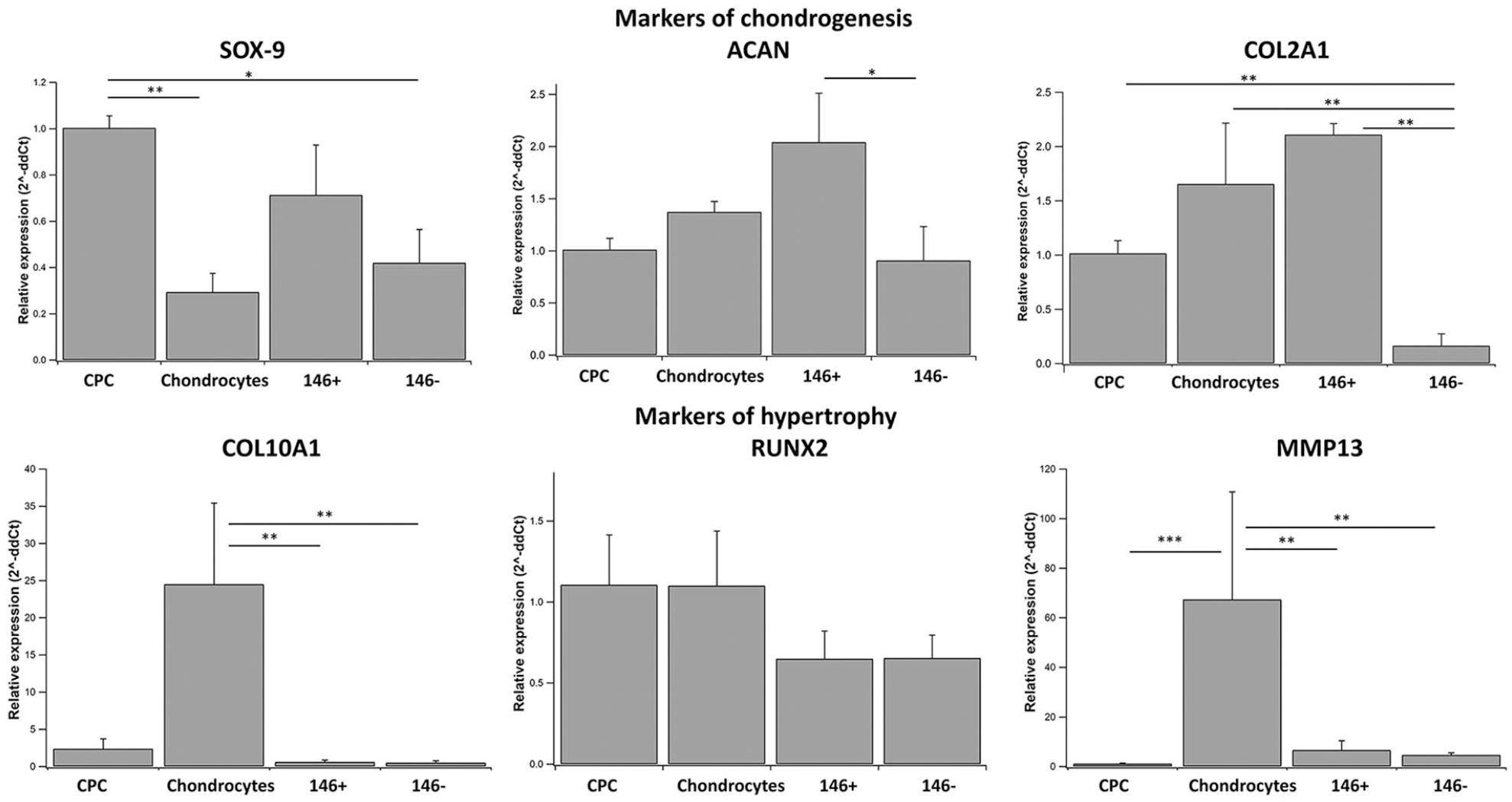
Relative expression of SOX-9, ACAN, COL2A1, COL10A1, RUNX2, and MMP13 across different study groups, when compared to the CPC group. 2^-ΔΔCt values are expressed as mean ± standard error mean (*P < 0.05, **P < 0.01, and ***P < 0.001), n = 3, each sample performed in duplicate. CPC = chondroprogenitors.
Trilineage Potential and GAG/DNA Content
All the study groups displayed a capacity for multilineage differentiation (
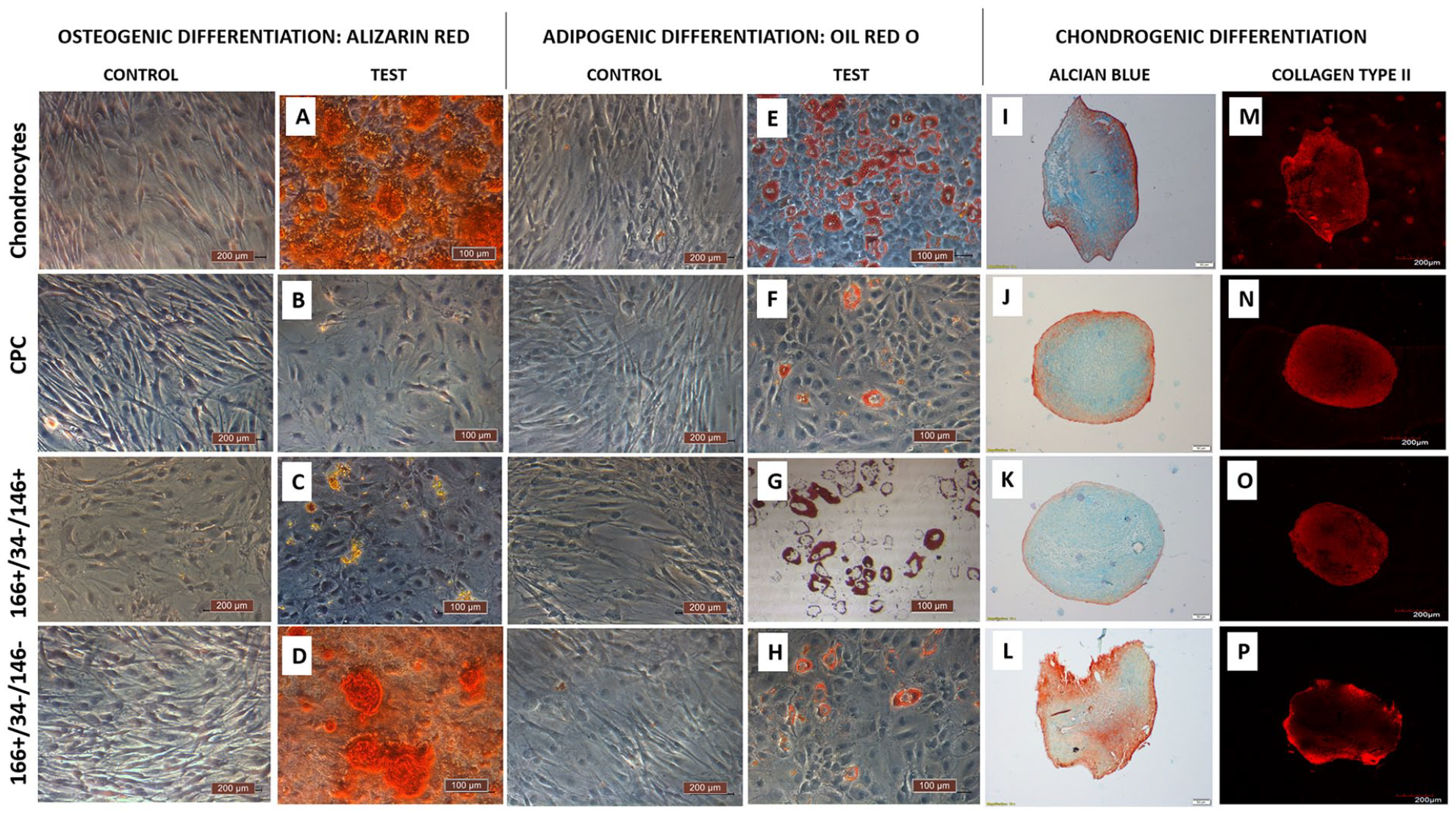
Trilineage differentiation of the different cell groups. Representative microscopic images of Alizarin red (
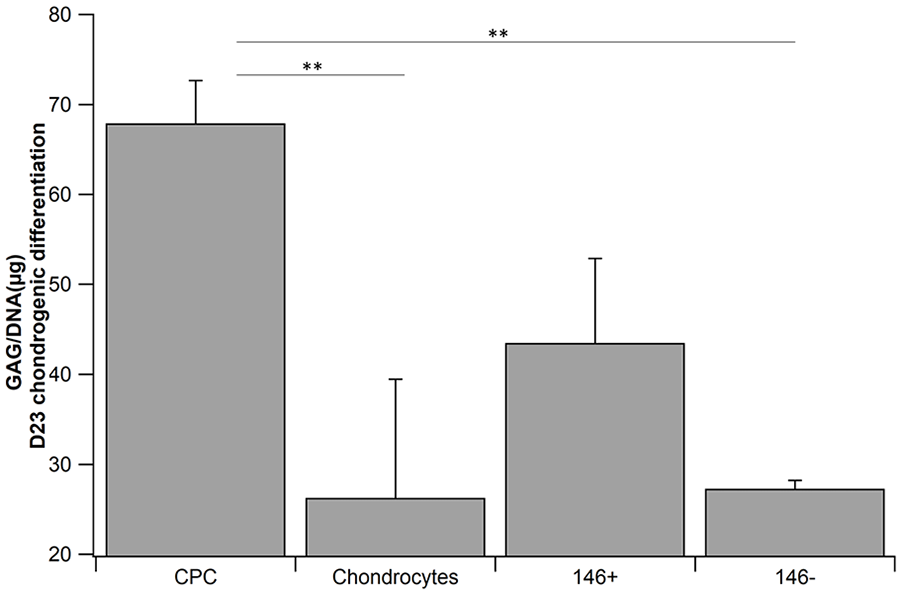
Estimation of GAG/DNA content in the chondrogenically differentiated pellet from CPC: chondroprogenitors, chondrocytes, CD166+/CD34−/CD146+ chondrocytes, and CD166+/CD34−/CD146− chondrocytes. Values are expressed as mean ± standard error mean (**P < 0.01), n = 3, each sample performed in duplicate. GAG = glycosaminoglycan.
Discussion
The therapeutic applications of chondrocyte use for cartilage regeneration are limited by the type of resultant repair, which comprises hyaline and fibrocartilaginous tissue. The phenotypic heterogeneity of chondrocytes has compelled the search for alternatives with superior chondrogenic potential. The emergence of the use of its progenitors has shown promise as they display higher chondrogenic capability with a reduced tendency for hypertrophy. Compared to commonly employed cells like chondrocytes and BM-MSCs, in vitro experiments show that fibronectin adhesion–derived chondroprogenitors demonstrate superiority for cartilage regeneration.8,10,34 However, the isolation of chondroprogenitors requires additional manipulation such as subjection to fibronectin, laminin adhesion, or based on its migratory potential.9,35,36 Another recommended method for isolation of chondroprogenitors includes subjecting the isolated chondrocytes to cell sorting based on cell surface marker expression.26,28 In line with our recent results comparing human chondroprogenitors to chondrocytes, we show that progenitors with time in culture demonstrate significantly lower expression of CD34 and higher expression of CD166 and CD146. 19 This was in accordance with previous reports by Adkar et al., reporting identification of a unique chondroprogenitor population with high chondrogenic potential based on sorting of human iPSCs using CD146 and CD166. 37 On the contrary, few studies report that elimination of CD146 positive subsets exhibits better chondrogenesis.31,33 Depletion of the CD34 population subset was considered based on our results and another report, which used this as a method to prevent contamination of MSCs with other cell types.19,38
Our experiment focused on comparing sorted chondrocytes, namely, CD166+/CD34−/CD146+ and CD166+/CD34−/CD146− cells with unsorted chondrocytes and fibronectin adhesion assay derived chondroprogenitors to assess whether inclusion or removal of CD146 subpopulation enhanced chondrogenesis, additionally assessing its comparability to the standard method for progenitor isolation.
Our findings demonstrated that chondroprogenitors had significantly higher expression of the chondrogenic gene SOX-9 when compared to chondrocytes and 146− subsets with lower expression of the hypertrophy gene MMP-13 compared to chondrocytes. This finding was revalidated by the total GAG/DNA content, which also showed the superiority of chondroprogenitors to chondrocytes and 146− subsets. This noteworthy finding was in accordance with a previous report by Khan et al. that showed higher expression of SOX-9 in chondroprogenitors when compared to chondrocytes, thus reiterating their potential for the regeneration of hyaline-like cartilage. 15 When chondrocytes were compared to sorted subsets, they displayed significantly higher expression of hypertrophic markers, namely, COL10A1 and MMP13, inferring that selection of prospective progenitors based on CD166, CD34, and CD146, from the heterogeneous population, does help deplete subpopulations with higher terminal differentiation potential. However, chondrocytes showed higher levels of COL2A1 than the CD146− subset. This finding may be attributed to the time point along the differentiation pathway from where the cells were procured. In the case of CD146, a protein that has been described as both a positive and negative chondrogenesis marker in several studies, our findings based on the comparison of the 146+ and 146− population showed that 146+ chondrocytes displayed significantly higher levels of both ACAN and COL2A1. These results were also reflected in multilineage differentiation analysis where chondroprogenitors and 146+ chondrocytes demonstrated a lower tendency for calcified matrix accumulation with 146− chondrocytes displaying the lowest GAG and collagen type II deposition. Thus, the findings in this study agree with previous literature reporting the use of CD166 and CD146 as markers of high chondrogenicity.26-28
Ozbey et al. report the presence of chondroprogenitors expressing putative stem cell markers with retained multilineage potential even at the deeper zones of cartilage. 17 Similarly, Nelson et al. report the presence of a viable pool of chondroprogenitors within osteoarthritic joints. 9 Since the availability of nondiseased joints is limited, chondroprogenitors were isolated from non–weight bearing areas of osteoarthritic joints. This showed the behavior of chondroprogenitors in patients with osteoarthritis, providing useful information while considering autologous chondroprogenitor cell therapy for these patients. However, the proposed strategy for isolating chondroprogenitors from chondrocytes would merit further evaluation using cells obtained from nondiseased joints and revalidation using in vivo study models. Studies have shown that chondrocytes and iPSCs display heterogeneity with off-target differentiation during the chondrogenic differentiation process.39,40 A recent single cell gene sequencing study revealed that iPSCs differentiated into other cell lineages during chondrogenic induction. Thus, future direction would be to include in-depth evaluation of progenitors using transcriptomic analysis which could provide valuable information regarding their chondrogenic potential.
This study showed that 146+ subsets show superiority to chondrocytes and 146− cells in terms of enhanced chondrogenesis with characteristics similar to chondroprogenitors. In terms of hypertrophy, the highest tendency was observed with chondrocytes. In conclusion, we have identified a unique progenitor population from chondrocytes that express CD166 and CD146, albeit CD34 negative, which show strong chondrogenic potential and lower hypertrophy comparable to chondroprogenitors. The findings of this first report will contribute to further investigations in the field of cartilage tissue engineering and its therapeutic application for joint diseases such as OA and osteochondral defects.
Supplemental Material
sj-pdf-1-car-10.1177_19476035211042412 – Supplemental material for Prospective Isolation and Characterization of Chondroprogenitors from Human Chondrocytes Based on CD166/CD34/CD146 Surface Markers
Supplemental material, sj-pdf-1-car-10.1177_19476035211042412 for Prospective Isolation and Characterization of Chondroprogenitors from Human Chondrocytes Based on CD166/CD34/CD146 Surface Markers by Elizabeth Vinod, Kawin Padmaja, Abel Livingston, Jithu Varghese James, Soosai Manickam Amirtham, Solomon Sathishkumar, Boopalan Ramasamy, Grace Rebekah, Alfred Job Daniel and Upasana Kachroo in CARTILAGE
Supplemental Material
sj-tif-1-car-10.1177_19476035211042412 – Supplemental material for Prospective Isolation and Characterization of Chondroprogenitors from Human Chondrocytes Based on CD166/CD34/CD146 Surface Markers
Supplemental material, sj-tif-1-car-10.1177_19476035211042412 for Prospective Isolation and Characterization of Chondroprogenitors from Human Chondrocytes Based on CD166/CD34/CD146 Surface Markers by Elizabeth Vinod, Kawin Padmaja, Abel Livingston, Jithu Varghese James, Soosai Manickam Amirtham, Solomon Sathishkumar, Boopalan Ramasamy, Grace Rebekah, Alfred Job Daniel and Upasana Kachroo in CARTILAGE
Footnotes
Author’s Note
Boopalan Ramasamy is also affiliated with Faculty of Health and Medical Sciences, The University of Adelaide, Australia.
Acknowledgments and Funding
The authors would like to acknowledge Ms. Immani Job and Mr. Ashok Kumar for technical support and the Centre for Stem Cell Research (A unit of inStem Bengaluru), Christian Medical College, Vellore, for infrastructural support. The research work presented in this article is a part of a proposed PhD thesis of the Tamil Nadu Dr. M.G.R. Medical University. The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was funded by the institutional fluid research grant (IRB Min No 11586), Christian Medical College, Vellore, and the Department of Biotechnology (BT/PR32777/MED/31/415/2019), Government of India.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
The study was conducted after obtaining approval from the institutional review board (IRB Min no: 11586).
Informed Consent
Written informed consent was obtained from each patient as per the ethical regulation of our institution.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
