Abstract
Objective
We aimed to analyze deoxycytidine-deoxyguanosine dinucleotide (CpGs) methylation profiles in DIO2, GPX3, and TXNRD1 promoter regions in osteoarthritis (OA) and Kashin-Beck disease (KBD) patients.
Methods
Blood samples were collected from 16 primary OA patients and corresponding 16 healthy individuals and analyzed for methylations in the CpGs of DIO2, GPX3, and TXNRD1 promoter regions using MALDI-TOF-MS. The methylation profiles of these regions were then compared between OA and KBD patients.
Results
DIO2-1_CpG_2 and DIO2-1_CpG_3 methylations were significantly lower in OA than KBD patients (P < 0.05). A similar trend was observed for GPX3-1_CpG_4, GPX3-1_CpG_7, GPX3-1_CpG_8.9.10, GPX3-1_CpG_13.14.15 and GPX3-1_CpG_16 (P < 0.05) as well as TXNRD1-1_CpG_1 and TXNRD1-1_CpG_2 methylation between OA and KBD patients (P < 0.05). However, there was no difference in methylation levels of other CpGs between the 2 groups (P > 0.05).
Conclusion
OA and KBD patients display distinct methylation profiles in the CpG sites of DIO2, GPX3, and TXNRD1 promoter regions. These findings provide a strong background and new perspective for future studies on mechanisms underlying epigenetic regulation of selenoprotein genes associated with OA and KBD diseases.
Osteoarthritis (OA), a degenerative joint disorder, is one of the leading causes of disability around the globe. The disease affects almost all joints, but more than 80% of OA occurs in the knees.1,2 Apart from disability, OA increase in the risk of developing cardiovascular diseases. 3 In addition, OA negatively affects mental well-being. In particular, the disease condition is associated with depression, suicidal ideation, memory loss, insomnia, and bipolar disorder.4-7 Current studies are focused on understanding the pathogenesis of OA with a view of rationalizing treatment approaches. However, the mechanism underlying OA pathogenesis remains to be validated.
Kashin-Beck disease (KBD) is an endemic OA, mainly prevalent in China, North Korea, and Russia.8,9 Generally, KBD manifests with pain, swelling, and stiffness of the joints as well as flexion of finger joints. Patients with this conditions find it very difficult to work or to take care of themselves. 10 By the end of 2017, there were 535,878 first-degree KBD patients in China. Among these, 12,730 were children ≤13 years old. 11 Exploring the etiology, pathogenesis, as well as biomarkers for early diagnosis of KBD is critical in guiding management strategies of the disease.
Primary OA and KBD share not only similar clinical characteristics but also the associated pathologic articular cartilage disorders, including chondrocyte apoptosis, inflammation, and cartilage degeneration manifest in both disease conditions.4,12,13 Nonetheless, OA and KBD exhibit distinct epidemiological characteristics. Whereas OA is globally distributed, KBD is only endemic in some regions. As search, discerning marked differences and similarities between OA and KBD will promote early diagnosis and effective treatment of the 2 diseases.
In recent years, increasing evidence shows that both OA and KBD are caused by epigenetic DNA modification. For instance, methylation of selenoprotein genes has been implicated in the development of OA and KBD.14,15 As such, selenoproteins are important targets in understanding arthritis diseases. Meanwhile, iodothyronine deiodinase (IDs), GPXs, and TrxRs family of proteins are key members of selenoproteins. These proteins participate in regeneration of skeletal muscles. Indeed, research shows that DIO2 is central to KBD progression in children. 16 On the other hand, GPX3 protects against oxidative-related damage of human chondrocytes 14 . Similar to GPX3, TXRND1 protects against oxidative-related cell damages.17-19 Increasing evidence shows that the cross-talk between inflammation and epigenetic regulators contribute to the development and/or progression of OA. 20 Nevertheless, methylation profiles of selenoprotein genes in OA and KBD patients remain unknown. Therefore, we evaluated the epigenetic changes in DIO2, GPX3, and TXRND1 genes in OA and KBD patients.
CpGs methylations in DIO2, GPX3, and TXRND1 promoter regions of OA and KBD patients were assessed using the high-throughput, cheap, and highly accurate matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI-TOF MS) technique, 21 DNA methylation differences between KBD and OA will deepen our understanding of the epigenetic mechanism underlying modification of selenoprotein genes in OA and KBD patients.
Materials and Methods
Participant Selection
We collected 32 blood samples from 16 primary OA patients and 16 healthy subjects. All study participants were Han Chinese, drawn from all ages and either gender ( Table 1 ). Clinical examinations and radiographic test were performed on each participant. Primary OA diosmosis was based on the proposed guidelines by the Diagnostic and Therapeutic Criteria Committee of the American Rheumatism Association (ACR). KBD was diagnosed based on the clinical diagnosis criteria of China (WS/T 207-2010). Patients with underlying genetic bone-, cartilage-, and arthritis-related diseases as well as other skeletal disorders were not included in the study.
The Basic Characteristics of the Subjects for MALDI-TOF MS Experiment.
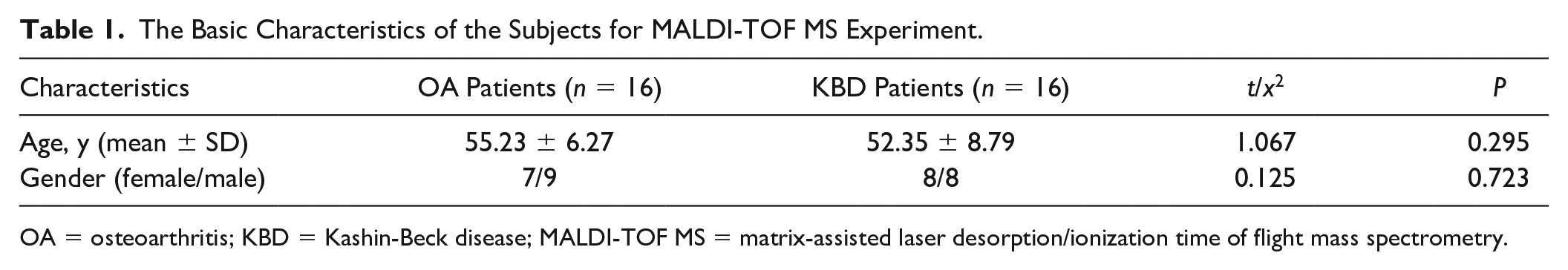

OA = osteoarthritis; KBD = Kashin-Beck disease; MALDI-TOF MS = matrix-assisted laser desorption/ionization time of flight mass spectrometry.
Collection of Blood Specimens and DNA Extraction
Protocol for this study was approved by the Human Ethics Committee of Xi’an Jiaotong University, People’s Republic of China. All participants consented to the study in writing. Briefly, 3 mL of blood from all subjects was first collected in biochemical anticoagulation test tubes. Genomic DNA was then extracted from the blood samples based on the QIAamp DNA Blood Mini Kit (QIAGEN, Germany), following the manufacture’s protocol. Efficiency of extraction was validated by agarose gel electrophoresis.
Primer Design and Synthesis
Primers for the methylated CpG sites in the DIO2, GPX3, and TXNRD1 promoter regions were designed using the Agena software (http://www.epidesigner.com/index.html). Methylation patterns in CpG sites were determined based on the sequence of the three target genes. The size of the target fragments ranged between 200 and 600 bp for all the genes. DIO2, GPX3, and TXNRD1 fragments contain 4, 25, and 13 CpGs, respectively. Each forward primer was conjugated with a 10-mer tag. Also, reverse primers were conjugated with a T7 promoter tag to adjust for the differences in melting temperature. The properties of primers used in this experiment including sequences of the target fragments and CpGs are highlighted in Tables 2 and 3 , respectively. The primers were synthesized by Liuhe Huada Gene Technology Co., Ltd. (Beijing, China).
The Information of Methylation Primers of DIO2, GPX3, TXNRD1 Used in This Study.
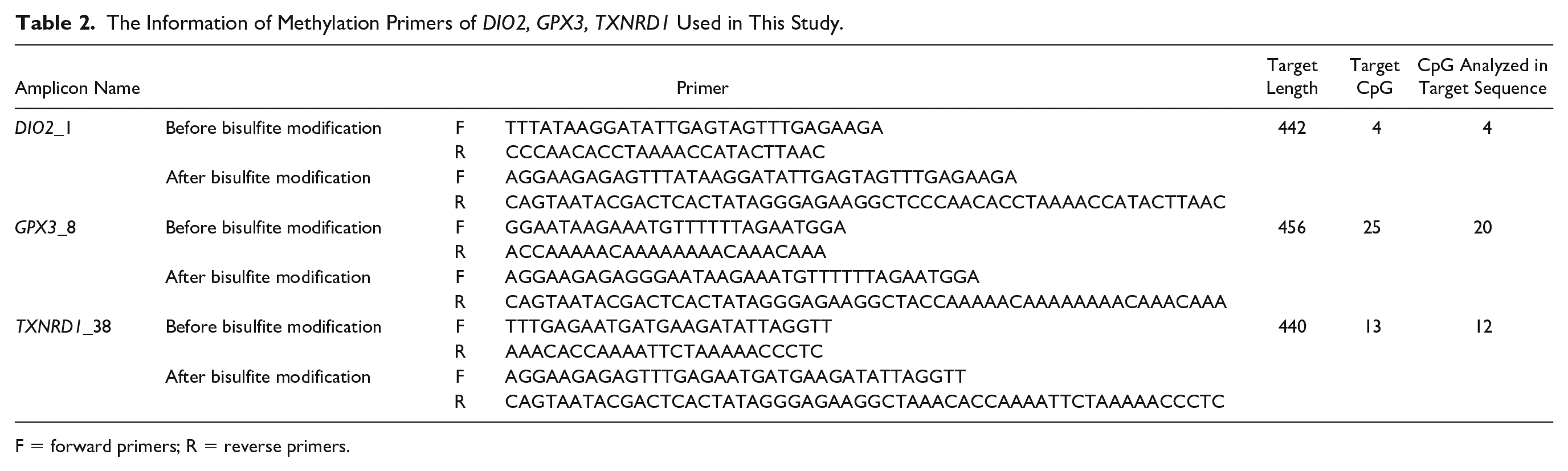

F = forward primers; R = reverse primers.
The CpGs in the Target Sequences.

Methylation Assessment
Briefly, 200 ng of genomic DNA of each participant was treated with bisulfite based on the EZ-96 DNA methylation kit (Zymo Research, Irvine, CA, USA), according to the manufacturer’s instructions. Methylation of DIO2, GPX3, and TXNRD1 was then quantitatively analyzed using the Agena MassARRAY platform (CapitalBio Corporation, Beijing, China). The platform composes of a MALDI-TOF mass spectrometer and an RNA base-specific cleavage (Mass CLEAVE) module. The methylations were detected using the Spectro CHIP (Agena Bioscience, California, USA) and Mass ARRAY Compact System (Agena Bioscience, California, USA) on the Agena MassARRAY platform. The data imported and analyzed using the EpiTYPER software version 1.0 (Agena Bioscience, San Diego, CA, USA).
Statistical Analysis
Continuous variables were expressed as means ± standard deviations (SD). Differences between OA and KBD patients were analyzed using chi-square (χ2) test and Student t test for 2 independent samples. All statistical analyses were performed using SPSS V. 23.0 (IBM Corp Armonk, NY, USA). Statistical significance was set at P < 0.05.
Results
Baseline Characteristics of the Study Population
There was no significant difference in demographic characteristics of the 16 OA and 16 KBD patients evaluated in this study between the 2 groups with regard to age and gender (P > 0.05) (Table 1).
DIO2 Methylation
Methylation profile of the 4 CpGs in DIO2 promoter region is shown in Figure 1 . We found methylation levels of DIO2-1_CpG_2 and DIO2-1_CpG_3 were significantly lower in OA than KBD patients (P < 0.05). However, there was no significant difference in the methylation level of DIO2-1_CpG_1 and DIO2-1_CpG_4 between the 2 groups of patients (P > 0.05) ( Fig. 2 ).
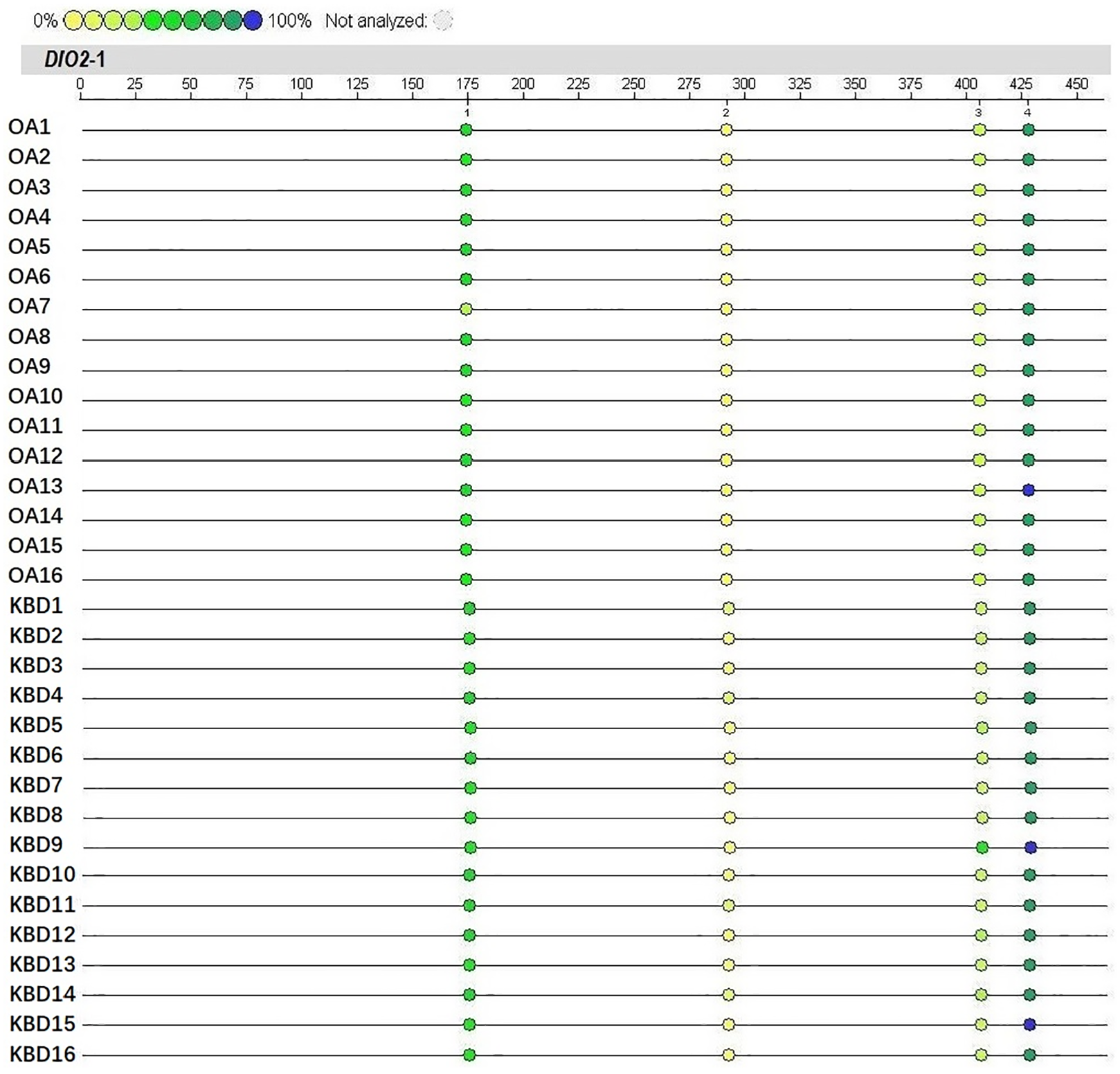
Methylation levels of CpGs in the promoter region of DIO2 gene in patients with osteoarthritis (OA) and Kashin-Beck disease (KBD). The color intensity is directly proportional to the methylation level. The numbers 1, 2, 3, and 4 represent CpGs DIO2-1_CpG_1, DIO2-1_CpG_2, DIO2-1_CpG_3, and DIO2-1_CpG_4, respectively.
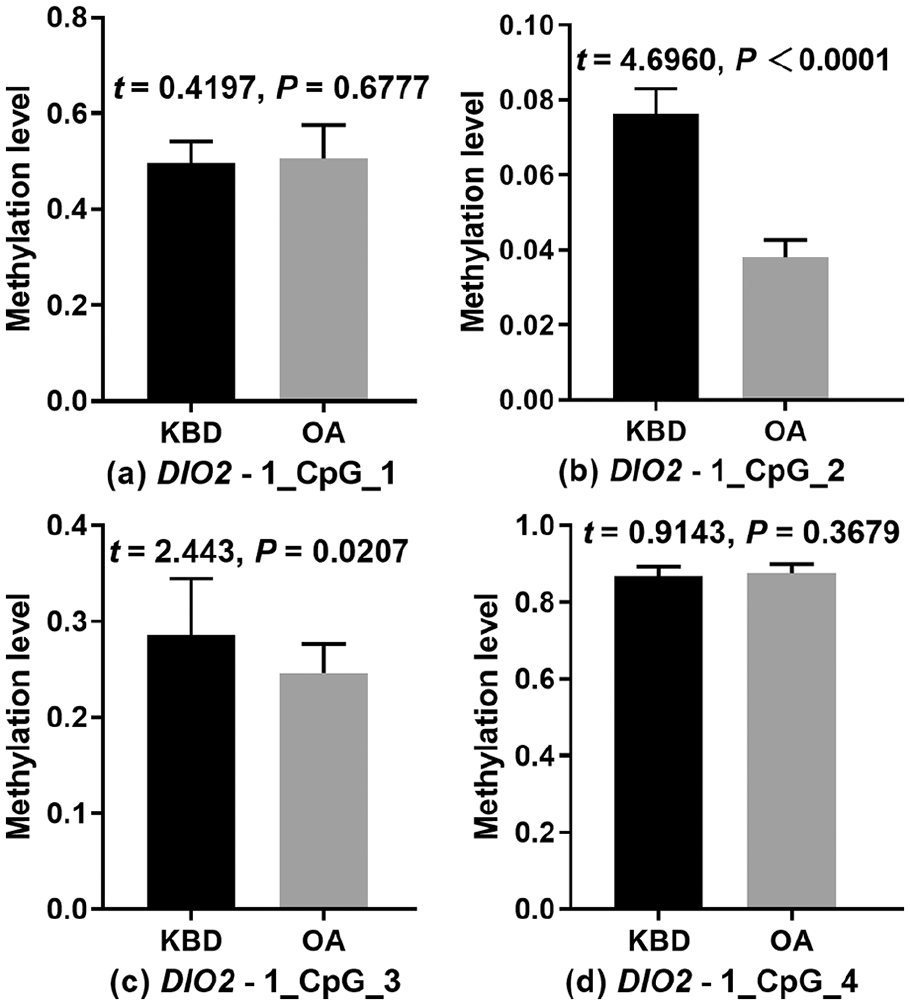
Methylation levels of CpG sites in DIO2 promoter region in osteoarthritis (OA) and Kashin-Beck disease (KBD) patients.
GPX3 Methylation
Methylation profile of 14 CpGs (CpG Units) in GPX3 promoter region is summarized in Figure 3 . Methylated GPX3-1_CpG_4, GPX3-1_CpG_7, GPX3-1_CpG_8.9.10, GPX3-1_CpG_13.14.15, and GPX3-1_CpG_16 were significantly lower in OA than KBD patients (P < 0.05). Conversely, there was no difference in the levels of methylated GPX3-1_CpG_3, GPX3-1_CpG_5.6, GPX3-1_CpG_11, GPX3-1_CpG_17.18, GPX3-1_CpG_20, GPX3-1_CpG_21, GPX3-1_CpG_22, GPX3-1_CpG_23, GPX3-1_CpG_24 between the groups (P > 0.05) ( Fig. 4 ). However, the methylation patterns of GPX3-1_CpG_1, GPX3-1_CpG_2, GPX3-1_CpG_12, GPX3-1_CpG_19, and GPX3-1_CpG_25 were not assessed because they were either large or small, beyond the detection range of the Mass ARRAY platform.
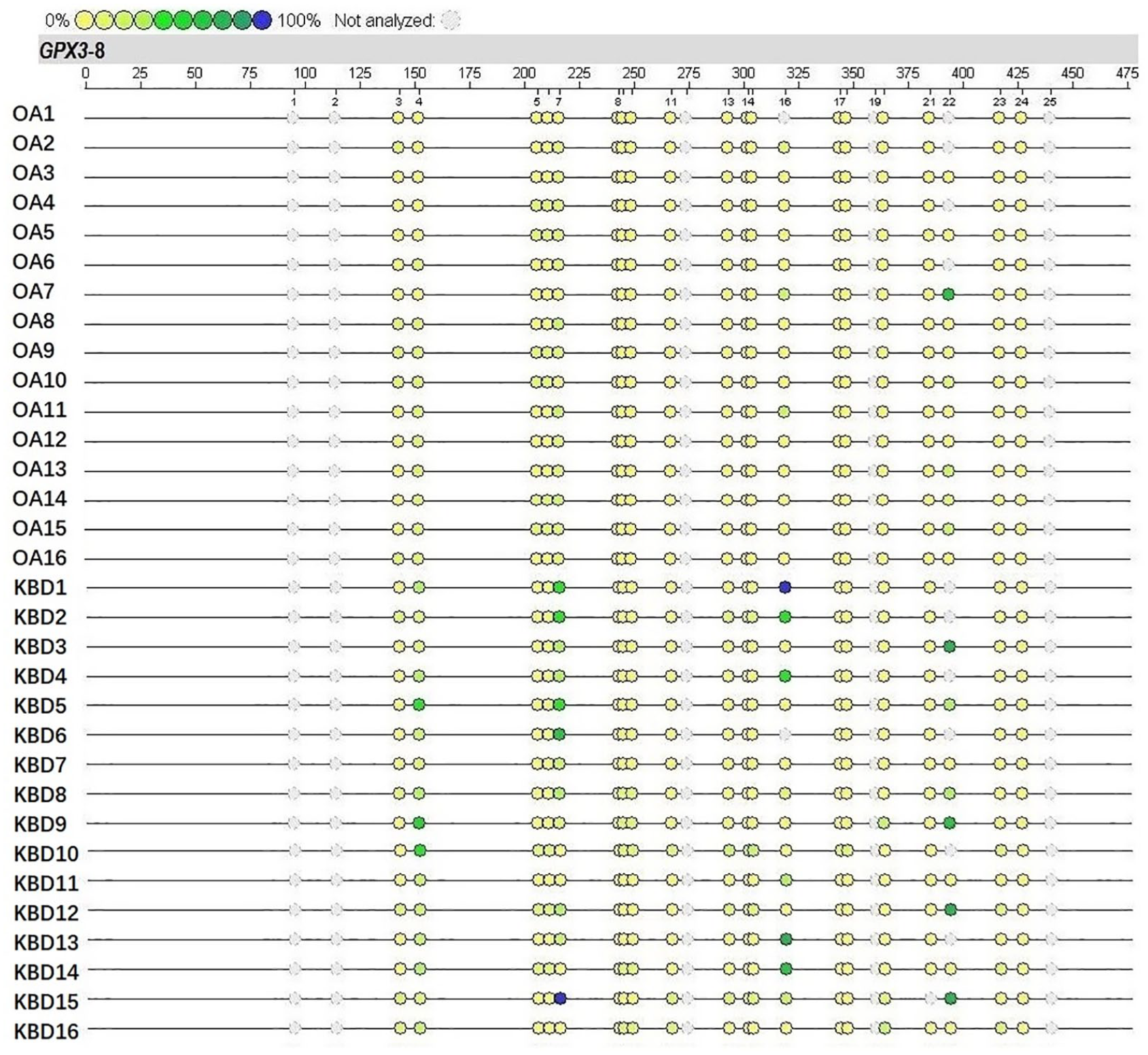
Methylation levels of CpGs in the promoter region of GPX3 gene in osteoarthritis (OA) and Kashin-Beck disease (KBD) patients. The color intensity of the dots is directly proportional to the methylation level. The numbers 1 to 25 in the figure represent CpGs GPX3-8_CpG_1 - GPX3-8_CpG_25.
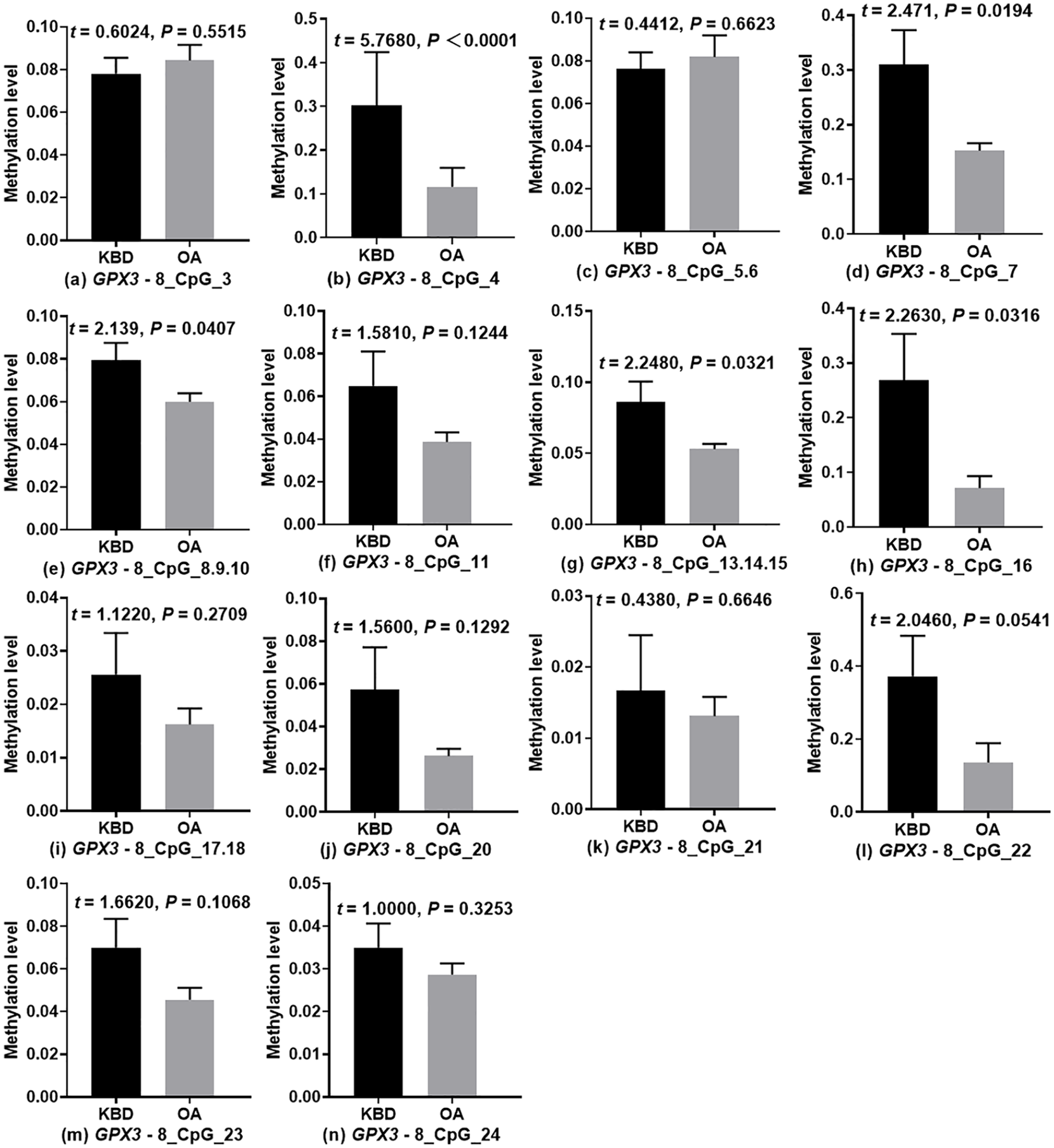
Methylation levels of CpGs in the promoter region of GPX3 in osteoarthritis (OA) and Kashin-Beck disease (KBD) patients.
TXNRD1 Methylation
Methylation profiles for 9 CpGs (CpG Units) in the TXNRD1 promoter region are shown in Figure 5 . Methylated TXNRD1-38_CpG_1 and TXNRD1-38_CpG_2 were significantly lower in OA than KBD patients (P <0.05). On the other hand, there no difference in the levels of methylated TXNRD1-38_CpG_3, TXNRD1-38_CpG_5.6.7, TXNRD1-38_CpG_8.9, TXNRD1-38_CpG_10, TXNRD1-38_CpG_11, TXNRD1-38_CpG_12, and TXNRD1-38_CpG_13 between the 2 groups (P > 0.05) ( Fig. 6 ). Methylation of TXNRD1-38_CpG_4 was not detectable because it is too small, beyond the detection range of the Mass ARRAY platform.
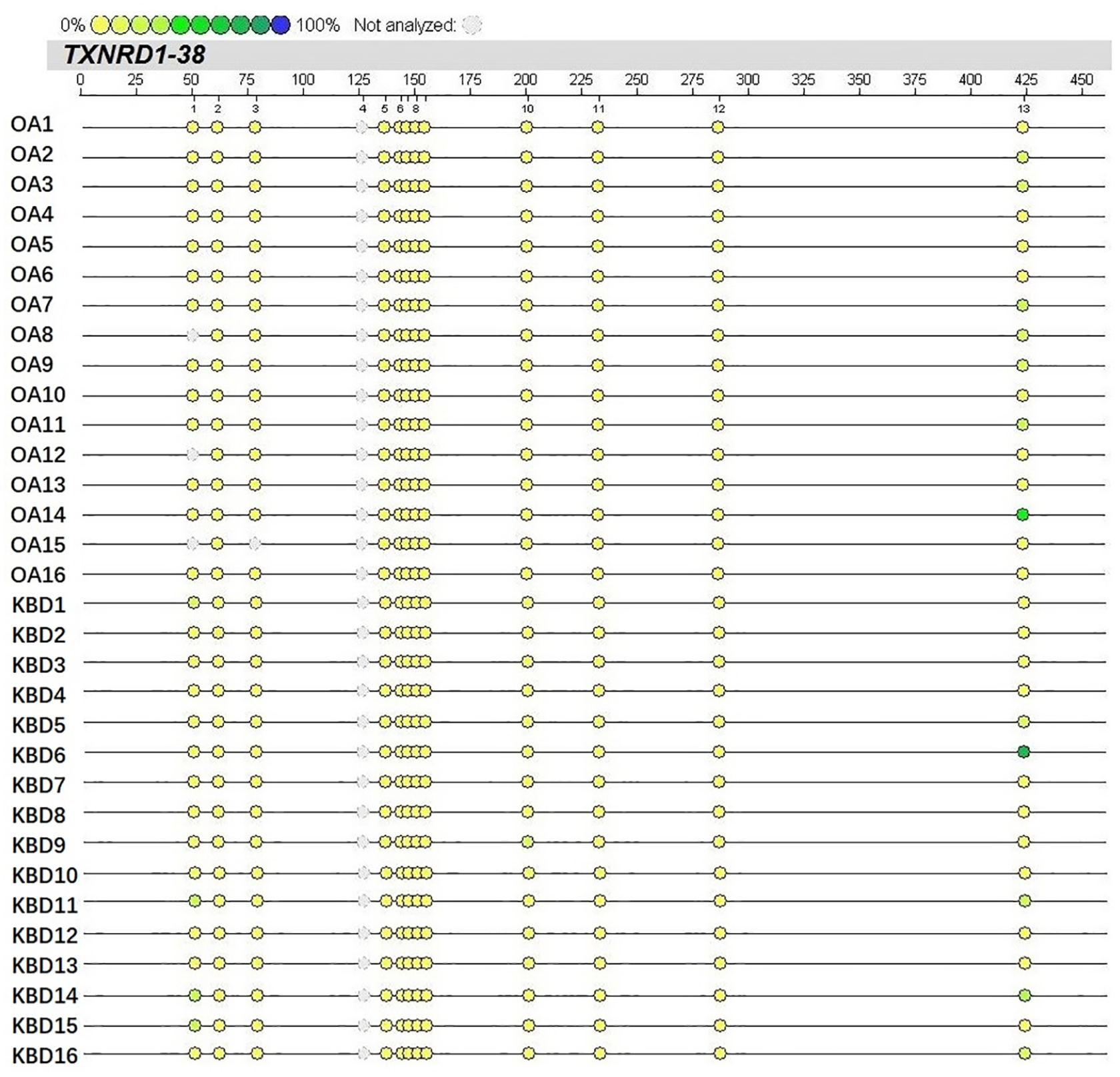
Methylation levels of CpGs in the promoter region of the TXNRD1 gene in osteoarthritis (OA) and Kashin-Beck disease (KBD) patients. The color intensity of the dots is directly proportional to the methylation level. The numbers 1 to 13 represent CpGs TXNRD1-38_CpG_1- to TXNRD1-38_CpG_13.
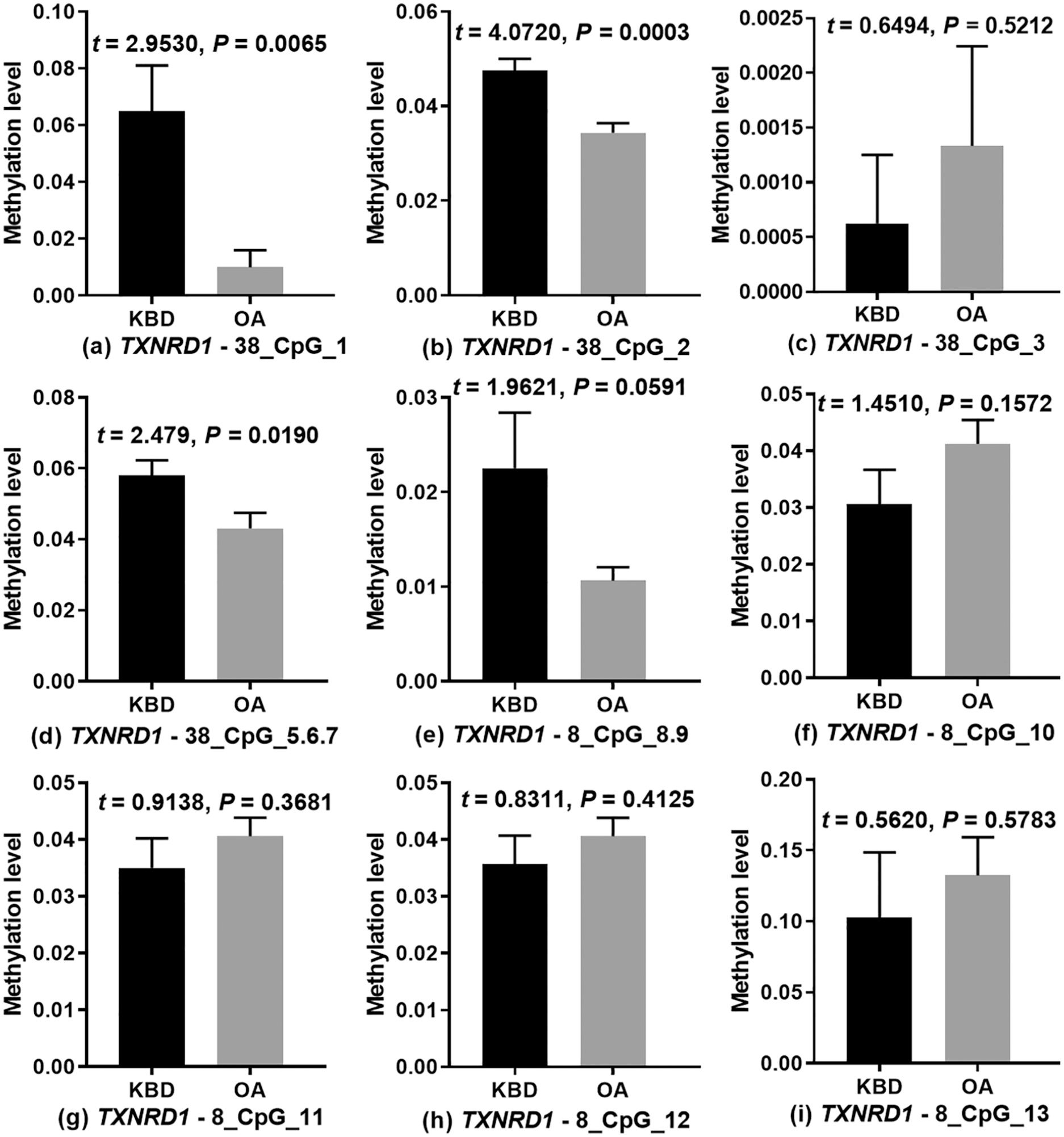
Methylation levels of CpGs in the promoter region of TXNRD1 in osteoarthritis (OA) and Kashin-Beck disease (KBD) patients.
Discussion
Herein, we explored the methylation patterns of selenoprotein genes in patients with OA and KBD using the MALDI-TOF MS technique. Overall, 42 CpG sites in the promoter region of three selenoprotein genes (DIO2, GPX3, TXRND1) were analyzed. We found significant pathogenetic differences between OA and KBD patients. Most previous OA research focused on the pathogenesis, management, and treatment modules of the disease.2,15,22-24 Clinical evidence shows that symptom management does not sufficiently ameliorate OA. Research on mechanism underlying OA development is particularly important in guiding the development of better treatment therapeutic strategies, which will improve the quality of life of affected patients. On the other hand, KBD is characterized by joint deformation. In extreme cases, the effected individual becomes disabled. Though critical, early diagnosis and prevention of KBD remain a persistent challenge. Numerous research studies have shown that OA and KBD patients exhibit oxidative damage and apoptosis of cartilage chondrocytes.25-30 Accordingly, reducing the levels of reactive oxygen species (ROS) in chondrocytes has been reported to effectively delay and prevent oxidative related damage to the cells.31,32
Selenium (Se) is a natural mineral widely distributed in the environment. 33 It is an essential trace element in human and animals, easily obtained from food. 34 In the body, selenium is distributed in various tissues, organs, and body fluids, with the highest concentration found in the kidneys. Besides the known toxic effects, there has been a keen interest in selenium owing to its nutritional value. 35 In one study, it was found that selenium deficiency is closely associated with Keshan disease (KSD) and KBD.36,37 Given that several inorganic selenium antioxidant compounds have been implicated in the oxidation of the selenium, it is particularly necessary to explore the biological functions of the mineral. Meanwhile, selenium compounds prevent oxidative DNA damage.31,36 Oxidative stress also causes degeneration of the extracellular matrix, which exacerbate apoptosis and necrosis of KBD and OA articular chondrocytes.31,38,39 These findings demonstrate the protective antioxidant properties of selenium on apoptosis and necrosis of chondrocytes.
To date, 25 selenoproteins have been identified and isolated in humans. Functionally, selenoprotein (selenocysteine) mainly regulates various biological functions such as oxidation, inflammation, and apoptosis.17,40,41 The iodothyronine deiodinase (ID) family participates in maintaining metabolic balance of thyroid hormones in the body, as well as regulates regeneration of skeletal tissues.17,40,41 Several studies show that DIO2 mRNA is overexpressed in the articular cartilage of OA patients.22,42 Meanwhile, DIO2 has been implicated in KBD progression in children. 16
Recent studies show that, GPXs protects against oxidative damage by degrades ROS. For instance, GPX3 exerts an anti-inflammatory biological effect against activated H2O2-mediated lipoxygenase. Selenium up-regulates the level expression of GPX3 mRNA following oxidative-related damage of human chondrocytes. 14
On the other hand, most research on the biological function of TrxRs mainly focuses on removing excess free radicals in the cytoplasm and mitochondria to protect cells against oxidative stress.29,43 In related studies, it has been found that low Se level in the body increases production of ROS and activates the Nrf2 signaling pathway. Together, they enhance the antioxidant capacity of the body.17-19 This underlines the role of selenoproteins in genetic regulation of OA and KBD diseases.
Recent studies have revealed significant difference in the methylation patterns of DIO2, GPX3, and TXNRD1 promoter regions between OA and KBD patients. In this study, we found there were significantly fewer methylated CpGs sites in the DIO2, GPX3, and TXNRD1 regions in OA than KBD patients. Although the 2 diseases exhibit several similarities across many characteristics, the above methylation differences suggest of distinct epigenetic modification of selenoprotein genes toward OA and KBD disease development. Even so, the exact mechanism underlying epigenetic modification of selenoprotein genes in the development of OA and KBD remain to be elucidated. Nevertheless, findings of this study provide a strong background that will guide future research perspectives on OA and KBD.
Overall, compared with individual with KBD, OA patients exhibit higher methylations in the DIO2, GPX3, and TXNRD1 promoter regions. This implies that KBD and OA are caused by distinct epigenetic modifications. These changes disrupt the expression of selenoproteins, modulating anti-oxidation and anti-chondrocyte apoptosis. In general, findings of this study provide new insights into epigenetic mechanism underlying repressed expression of selenoprotein in OA and KBD disease conditions.
Supplemental Material
sj-pdf-1-car-10.1177_1947603520988165 – Supplemental material for Patients with Osteoarthritis and Kashin-Beck Disease Display Distinct CpG Methylation Profiles in the DIO2, GPX3, and TXRND1 Promoter Regions
Supplemental material, sj-pdf-1-car-10.1177_1947603520988165 for Patients with Osteoarthritis and Kashin-Beck Disease Display Distinct CpG Methylation Profiles in the DIO2, GPX3, and TXRND1 Promoter Regions by Rongqiang Zhang, Hao Guo, Xiaoli Yang, Dandan Zhang, Di Zhang, Qiang Li, Chen Wang, Xuena Yang and Yongmin Xiong in CARTILAGE
Footnotes
Author Contributions
Study design and conception: Yongmin Xiong, Rongqiang Zhang. Data collection: Hao Guo, Rongqiang Zhang, Xiaoli Yang, Di Zhang, Dandan Zhang, Qiang Li, Chen Wang, Xuena Yang. Data analysis and interpretation: Hao Guo, Rongqiang Zhang, Yongmin Xiong. All authors read, revised and approved submission of the final draft.
Acknowledgments and Funding
We appreciate Chao Lu (Honghui Hospital Affiliated to Xi’an Jiaotong University) for assisting in collecting clinical materials. The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported financially by the National Natural Science Foundation of China (81773372, 81573104), Key Research and Development Program in Shaanxi Province (2020SF-076), Research Project from Health Commission of Shaanxi Provincial Government (2018A017) and the Education Department of Shaanxi Provincial Government (19JS015).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Protocol for this study was approved by the Human Ethics Committee of Xi’an Jiaotong University, People’s Republic of China.
Informed Consent
All participants consented to the study in writing.
Trial Registration
Not applicable.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
