Abstract
NKX3.1 is a tumor suppressor down-regulated in early prostate cancers. A SNP (rs2228013), which represents a polymorphic
Introduction
Adenocarcinoma of the prostate, like many malignancies, initiates in epithelial cells that acquire the precursor or gatekeeper mutations required for development of the malignant phenotype. In the majority of early prostate cancers a region of 8p21.2 is lost resulting in loss of the homeobox gene
Loss of NKX3.1 expression is a very early event in prostate carcinogenesis. Gene targeting studies in mice showed that
Although no somatic mutations of
Growth suppression by NKX3.1 is affected, in part, by inducing expression of insulin-like growth factor binding protein-3 (IGFBP-3), a known growth suppressor protein and down-regulator of insulin-like growth factor-I (IGF-I) activity. IGF-I is a peptide growth factor that regulates cell growth, differentiation, and apoptosis by binding to the IGF receptor-I (IGFR-I).
10
IGFs are present in abundance in the circulation. Circulating IGF-I is bound mainly to IGFBP-3, one of the most abundant serum proteins.
11
Although IGFBP-3 can inhibit the interaction of IGF-I with its receptor at the cellular level, serum IGFBP-3 also serves to stabilize circulating IGF-I.
12
Because of the effects of IGF-I on cell growth, survival, and apoptosis, the influence of both serum IGF-I and IGFBP-3 concentrations on cancer risk has been studied by a number of investigators.
12
Serum IGF-I levels are associated with an elevated risk of prostate cancer in a variety of studies13-17 that have been confirmed by meta analyses,18,19 Here we show that NKX3.1(R52C) and a protein engineered for loss of the serine 48 phosphorylation site NKX3.1(S48A) do not activate expression of IGFBP-3. Consistent with this loss of function, we hypothesize that the presence of the variant NKX3.1 protein may interact with circulating serum IGF-I to affect prostate cancer risk. Determination of
Results
Effect of NKX3.1(R52C) on IGFBP-3 expression and IGF-IR activation
Amino acid 52 affected by rs2228013 is an arginine, located in a consensus motif that is a site for phosphorylation at serine 48. Replacement of arginine 52 with cysteine decreases phosphorylation at serine 48 by 70%.
20
Thus, a missense mutation at serine 48 potentially generates a protein with analogous, but more absolute, loss of serine 48 phosphorylation compared to NXK3.1(R52C). Expression of NKX3.1(R52C) in PC-3 cells induced substantially less
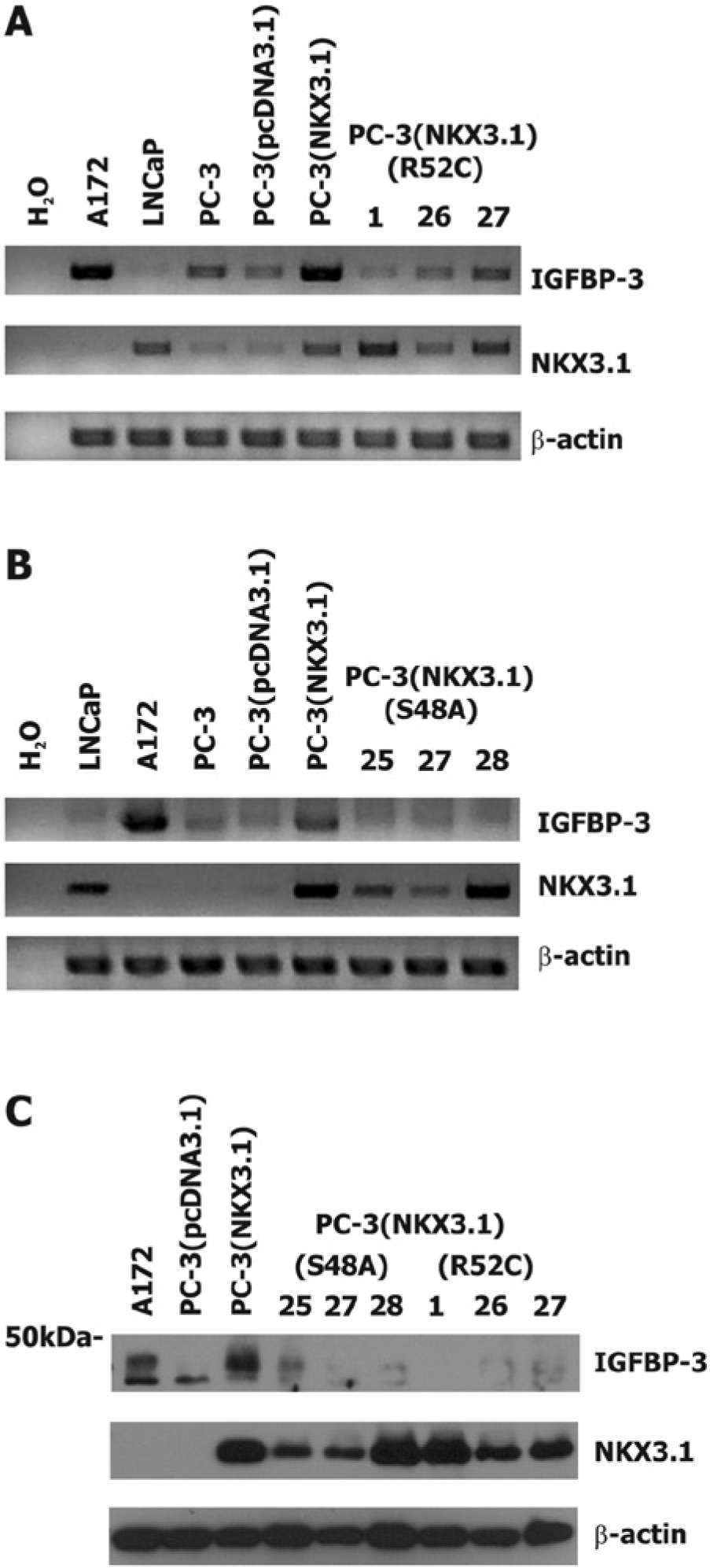
The effect of NKX3.1 variant proteins on IGFBP-3 expression in PC-3 cell clones. (
NKX3.1 expression attenuated IGFR-I activation in PC-3 cells via induced expression of IGFBP-3. The effect of NKX3.1 on IGFR-I activation was not seen when Long R-IGF-I, an IGFR-I ligand that does not bind to IGFBP-3 was used, or when cells were pretreated with
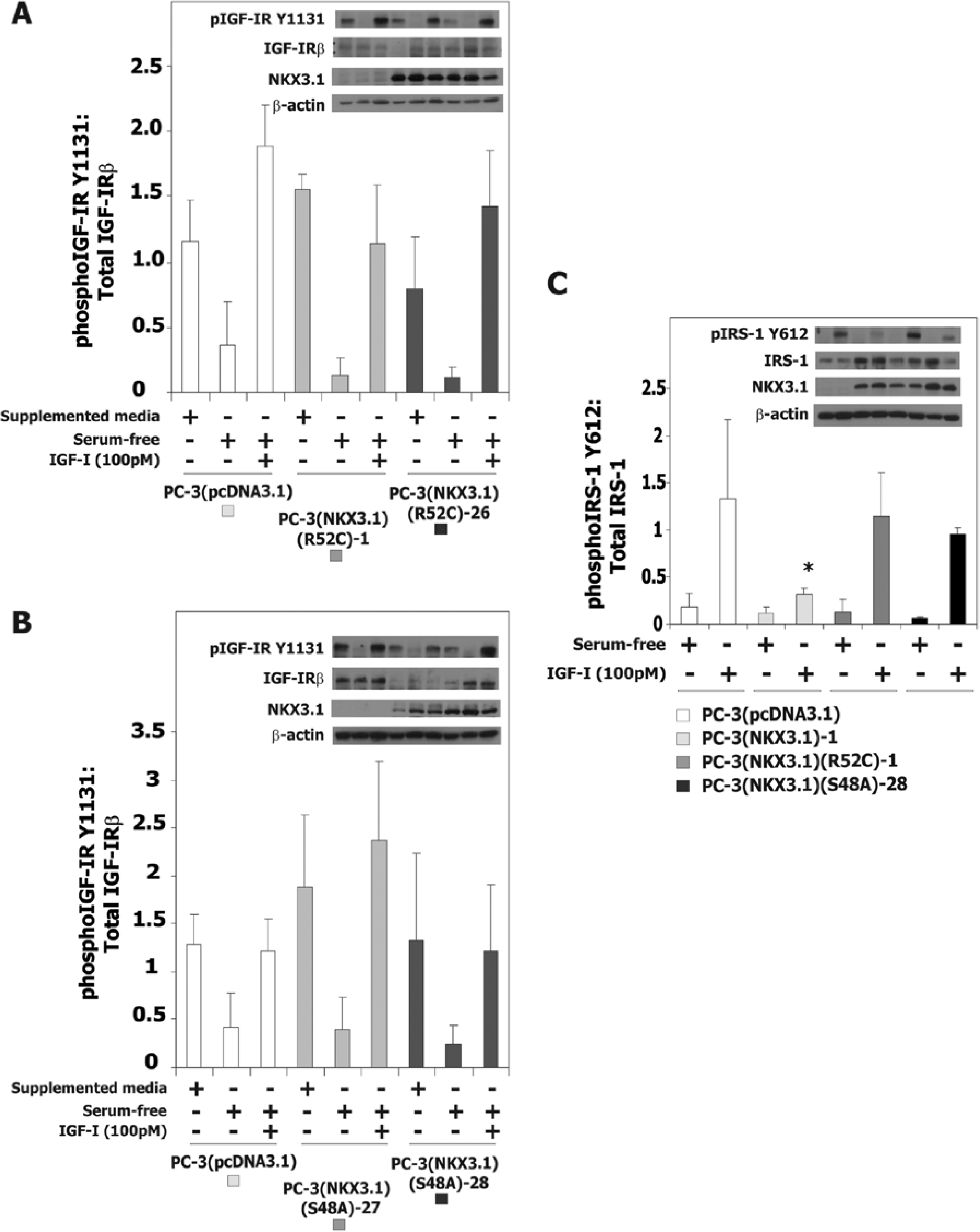
The effect of NKX3.1 variant proteins on IGF-IR activation in PC-3 cells. (
NKX3.1 suppresses cell proliferation in culture, an effect that is abrogated by siRNA to
Effect of NKX3.1 Expression on Cell Proliferation.
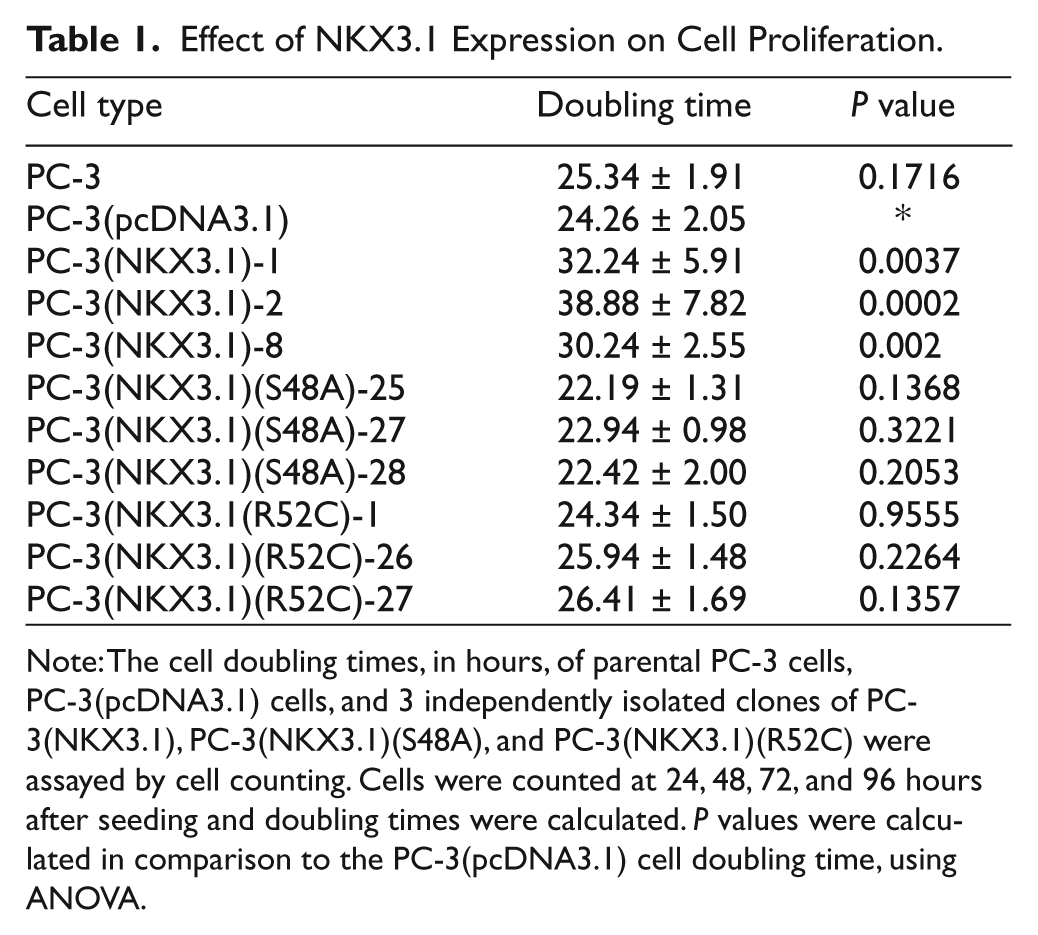
Note: The cell doubling times, in hours, of parental PC-3 cells, PC-3(pcDNA3.1) cells, and 3 independently isolated clones of PC-3(NKX3.1), PC-3(NKX3.1)(S48A), and PC-3(NKX3.1)(R52C) were assayed by cell counting. Cells were counted at 24, 48, 72, and 96 hours after seeding and doubling times were calculated.
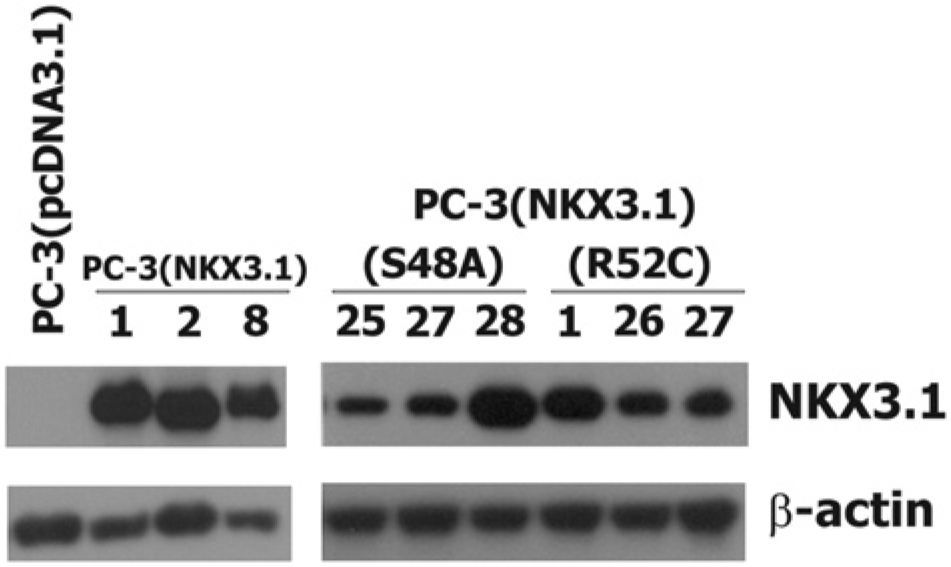
Expression of NKX3.1 in derivative PC-3 cell clones. Panels from western blots of β-actin and NKX3.1 are shown from exponentially growing cultured cells from which equal amounts of total protein were loaded onto a gel. Numbers indicate identities of individual independently derived clones. A cell extract from PC-3 cells transfected with the empty vector is shown at the far left as a negative control.
NKX3.1(C154T) as a risk factor
Previously we had shown that among 1,253 cases and controls in the Physicians’ Health Study (PHS)
Odds Ratios of Prostate Cancer by
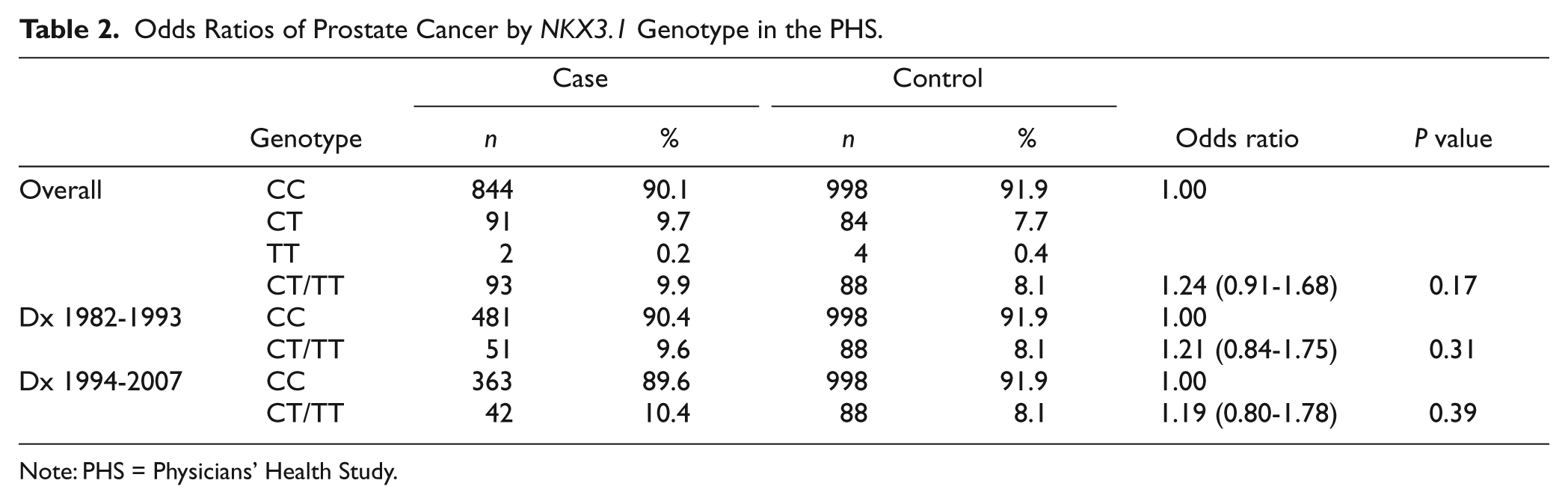
Note: PHS = Physicians’ Health Study.
Odds Ratios of Prostate Cancer by
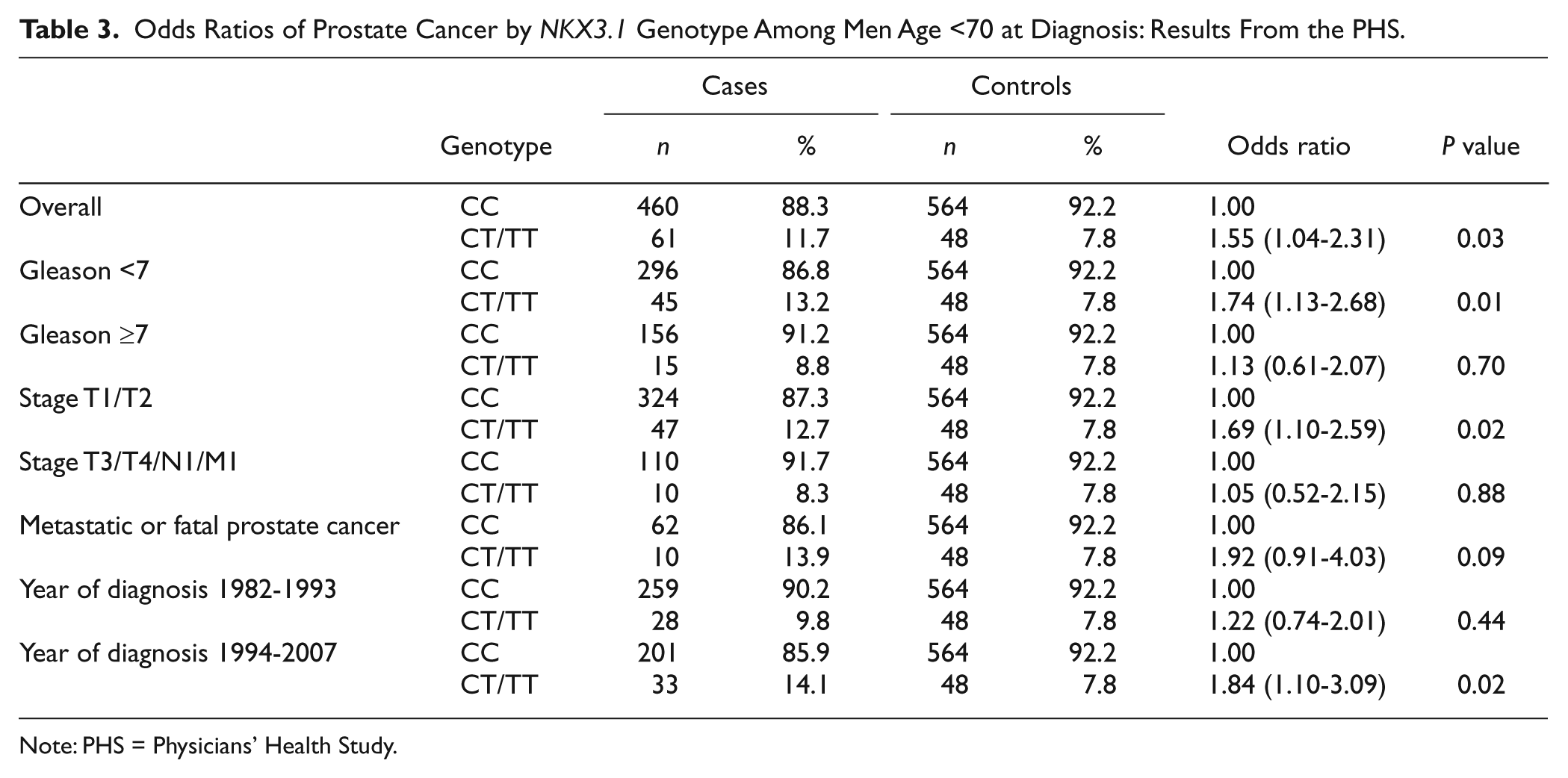
Note: PHS = Physicians’ Health Study.
Analysis of NKX3.1 genotype and serum IGF-I
Since NKX3.1 regulates local IGFBP-3 expression in prostate and IGFBP-3 attenuates IGF-I signaling, we asked whether
Odds Ratios of Prostate Cancer by
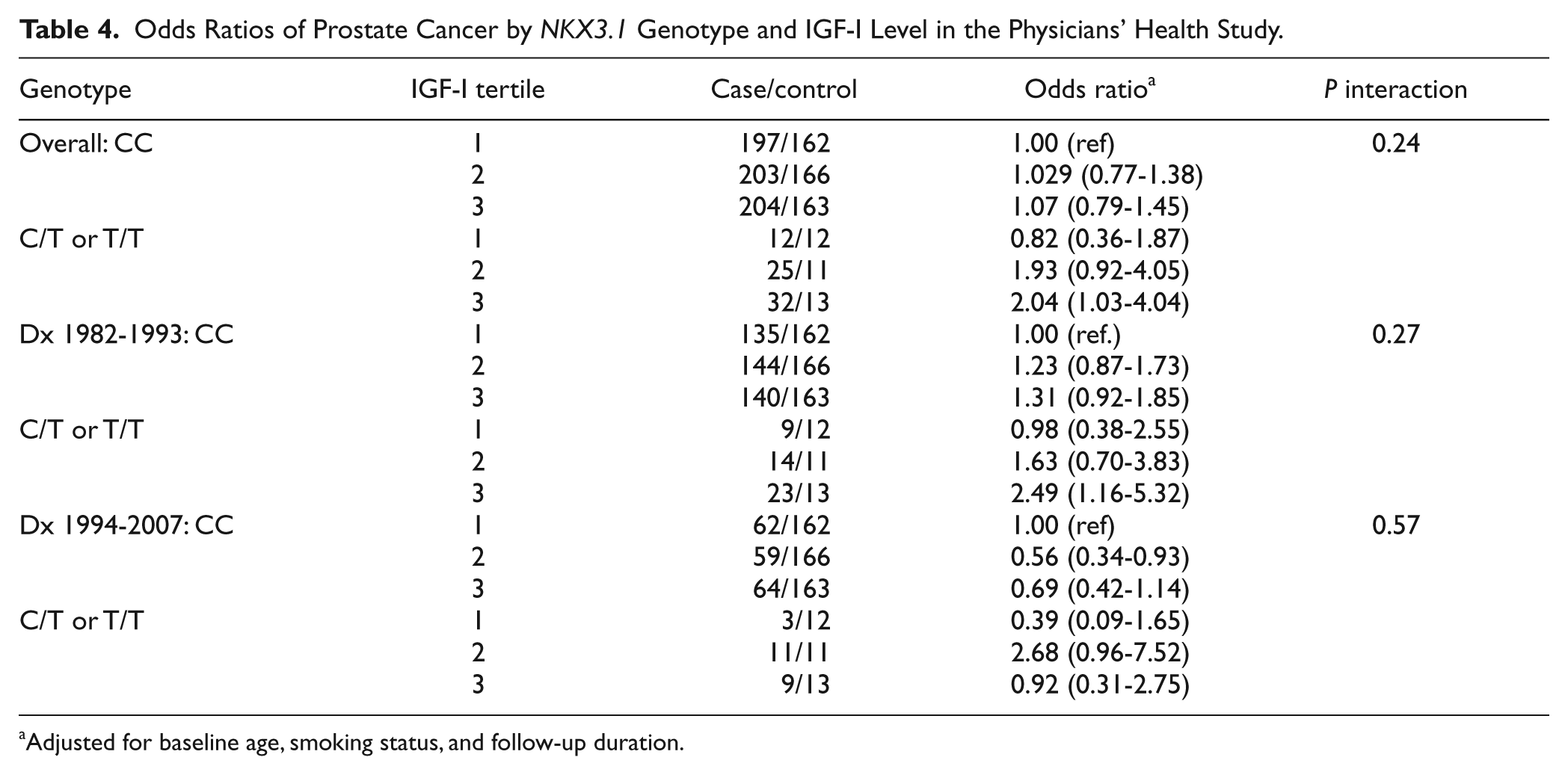
Adjusted for baseline age, smoking status, and follow-up duration.
We analyzed
Characteristics of the Two Study Populations.
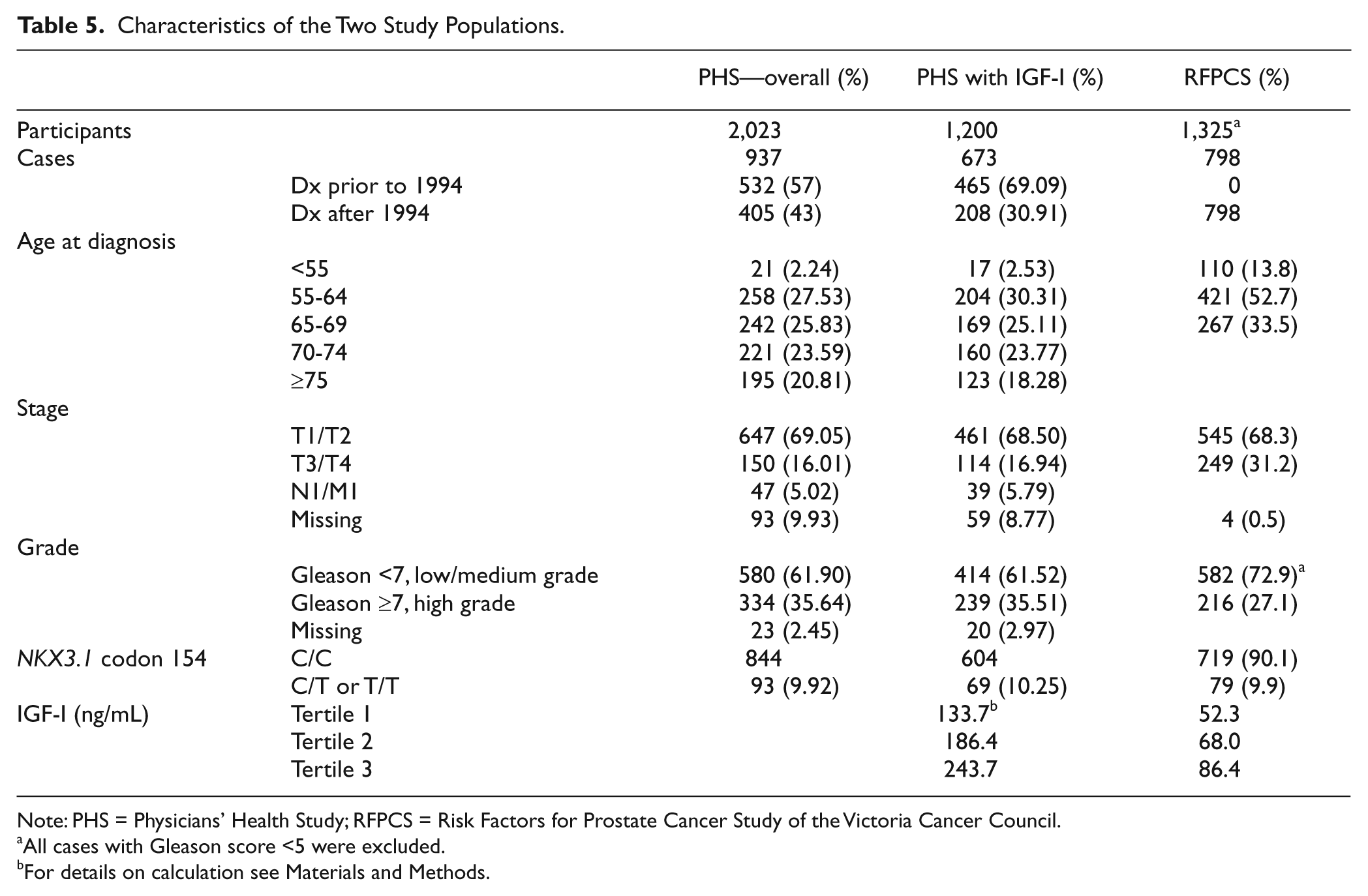
Note: PHS = Physicians’ Health Study; RFPCS = Risk Factors for Prostate Cancer Study of the Victoria Cancer Council.
All cases with Gleason score <5 were excluded.
For details on calculation see Materials and Methods.
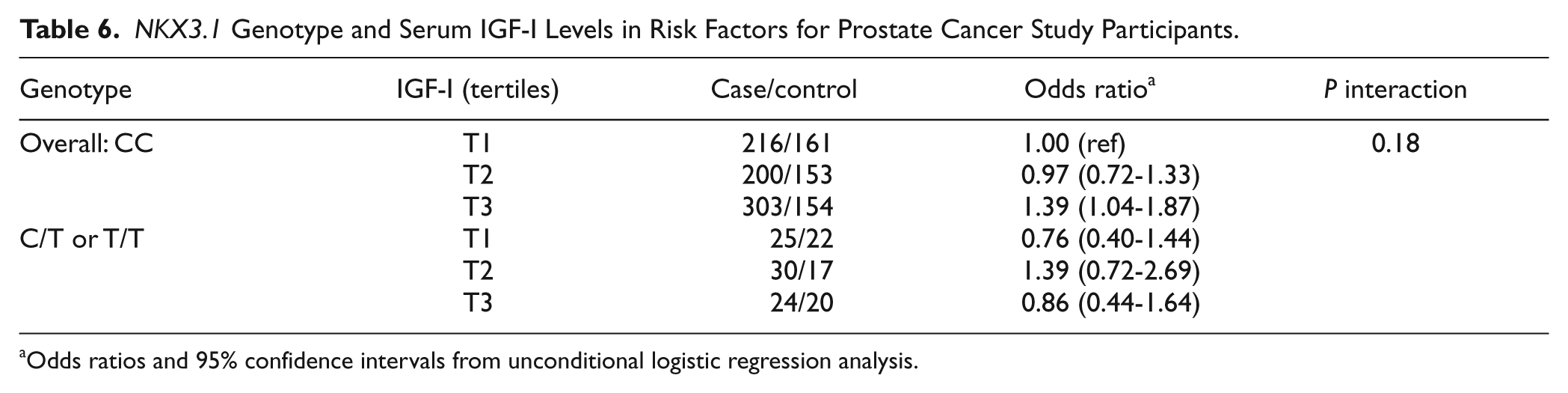
Odds ratios and 95% confidence intervals from unconditional logistic regression analysis.
Serum IGF-I Levels and Prostate Cancer Risk in Risk Factors for Prostate Cancer Study Participants.
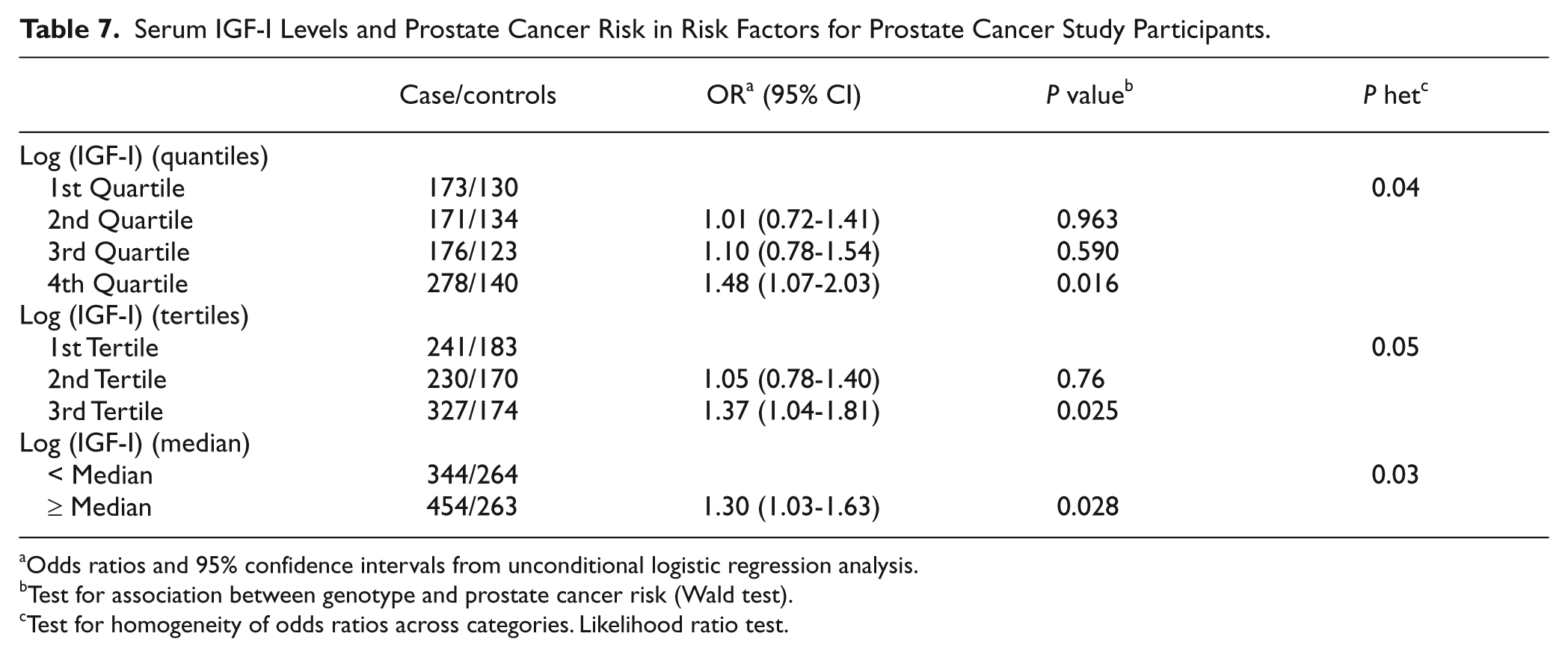
Odds ratios and 95% confidence intervals from unconditional logistic regression analysis.
Test for association between genotype and prostate cancer risk (Wald test).
Test for homogeneity of odds ratios across categories. Likelihood ratio test.
Discussion
NKX3.1 is important for prostate epithelial cell development, growth control, and differentiation.4,22 Murine
NKX3.1 exerts a broad range of effects on prostate epithelial cells. When NKX3.1 is down-regulated in the course of prostate carcinogenesis,
3
those effects vary with gene copy number as methylation due to
Previously we showed that both in cultured cells and
In the data we present here, the presence of a variant
Differences in the association between
The PHS and RFPCS also differed in the timing of blood collection that was used for serum IGF-I determinations. The PHS group was a nested case-control study, with prospectively collected samples. IGF-I levels were determined on samples collected before the diagnosis of cancer. In contrast, for the RFPCS case-control study, the blood was collected after diagnosis and perhaps after treatment had been initiated. Thus, in the case-control setting, one is always concerned that the disease, or treatment, or both could affect the blood levels.
Almost all prostate cancer initiation is accompanied by decreased expression of NKX3.1 protein. We do not know whether there is a threshold of NKX3.1 expression that predisposes cells to malignant transformation or whether the risk of malignant transformation is related to the level of NKX3.1. Consistent with the role of NKX3.1 as a tumor suppressor, a T164A missense mutation in the NKX3.1 homeodomain that reduced DNA binding by 95% was found to cosegregate with early prostate cancer in a family.
9
Phenotypic effects resulting from reduced levels of NKX3.1 protein may reflect the fact that other members of the NK protein family are also haploinsufficient. For example, both missense and truncation mutations that cause loss of NKX2.5 protein are autosomal dominant determinants of congenital cardiac abnormalities.37,38 Our data and work with gene targeted mice suggest that tumor suppression by NKX3.1 correlates with protein expression levels and that optimal function of the protein requires native expression from 2 intact
Materials and Methods
Cell culture and reagents
The prostate cancer cell lines PC-3 and LNCaP and the A172 human glioblastoma cell line were obtained from the American Type Culture Collection (Rockville, MD). PC-3 and A172 cell lines were grown as previously described. 21
Plasmids and transfection full length NKX3.1, NKX3.1 (R52C), and NKX3.1(S48A) were expressed from constructs in the mammalian expression vector, pcDNA3.1 (Invitrogen, Carlsbad, CA) as previously described. 20 Transient and stable transfections were carried out as described earlier. 21
Western blot analysis
Immunoblotting was done as previously described. 21 Antibody reagents were as follows: β-actin (Sigma, St. Louis, MO) 1:10,000; NKX3.1 21:2,000; IGFBP-3 (sc-9028, Santa Cruz Biotechnology, Santa Cruz, CA) 1:8,000 at 4°C overnight, followed by 3 washes in phosphate-buffered saline. Horseradish peroxidase conjugated goat-anti-rabbit and goat-anti-mouse (ImmunoPure antibodies, Pierce Biotechnology, Rockford, IL) secondary antibodies in 1% milk or 1% bovine serum albumin were applied for 1 hour at room temperature. Signal detection was performed with Super-Signal West Pico Chemiluminescent Substrate (Pierce Biotechnology).
Reverse transcriptase PCR analysis
Total RNA was extracted using the RNeasy Mini Kit (Qiagen, Venlo, the Netherlands) and cells were homogenized using the Qiashredder (Qiagen) method. A total of 125 to 250 ng of RNA was added to the RT-PCR master mix from One-step RT-PCR kit (Qiagen) (includes 5× buffer, DNTPs, and Taq polymerase). The following primers were used in the RT-PCR reactions: β-Actin (Fwd 5′-GGC CAC GGC TGC TTC-3′ and Rev 5′-GTT GGC GTA CAG GTC TTT GC-3′); NKX3.1 (Fwd 5′-GCC GCA CGA GCA GCC AGA GAC A-3′ and Rev 5′-TTC AGG GCC GGC AAA GAG GAG TG-3′); IGFBP-3 (Fwd 5′-CGC CAG CTC CAG GAA ATG-3′ and Rev 5′-GCA TGC CCT TTC TTG ATG ATG-3′); IGFBP-4 (Fwd 5′-TTA GCC CAA GAG GTC TGA GC-3′ and Rev 5′-CTG TGC TTC AAG TCT TCC TTT G-3′); Lamin A/C (Fwd 5′-AAC TTC AGG ATG AGA TGC TGC G-3′ and Rev 5′-GTC CAG AAG CTC CTG GTA CTC GT-3′). RT-PCR was performed in a Techne Techgene PCR machine; 30 minutes at 50°; 15 minutes at 94°; 22 to 30 cycles of 30 seconds to 1 minute at 94°, 30 seconds to 1 minute at melting temperatures of 55° to 65°, and 30 seconds to 1 minute at 72°; followed by 15 minutes at 72°. Samples were stored on ice and mixed with 10× Blue Juice gel loading buffer (Invitrogen) and run on a 1.5% agarose gel containing 0.1 µg/mL ethidium bromide in TAE buffer. Gels were imaged on a luminometer and recorded using a Kodak 1D digital camera.
IGF-IR activation and signaling
IGF-IR signaling was assayed as described previously.
20
Western blot analysis was completed as described above using anti-IGF-IR (#3027, Cell Signaling, Beverly, MA), anti-phospho-IGF-I Receptor (Tyr1131) (#3021, Cell Signaling), anti-IRS-1 (06-248, Upstate, Placid Lake, NY), anti-phospho-IRS-1(Y612) (44-816G, Biosource, Carlsbad, CA) primary antibodies. Bands were quantified by Scion Imager software and
Cell proliferation assay
PC-3 and derivative cell lines were seeded in triplicate in 96-well plates at a concentration of 4,000 cells per well in IMEM containing 10% fetal bovine serum (PC-3) or 10% fetal bovine serum plus 1.2 mg/mL G418 (PC-3(pcDNA3.1) and PC-3(NKX3.1)) and incubated for 24 hours at 37°C. At 24, 48, 72, and 96 hours after seeding, wells were trypsinized, suspended in IMEM, and immediately counted in a Beckman Coulter Z1 cell counter. Doubling times were calculated using Microsoft Excel, and
Study cohorts
Physicians’ Health Study: The participants of the Physicians’ Health Study (PHS) who comprised the prostate cancer cases and controls have been previously described in articles describing the effect of serum IGF-I 22 and NKX3.1 genotype 23 on prostate cancer risk. The current analysis includes 937 cases and 1,086 controls, using a prospective nested case-control design. Of the cases, 580 had Gleason score <7, 334 had Gleason score ≥7, and 23 cases had unknown Gleason scores. Among the cases, 647 were T1/T2 and 197 were locally advanced or metastatic at presentation, with the remaining 93 cases of unknown stage. Median age of cases at diagnosis was 70 years.
Risk factors for prostate cancer study
The Risk Factors for Prostate Cancer Study (RFPCS) is a population-based case-control study of prostate cancer in Australia. Prior approval of the study protocol was obtained from all relevant hospital and cancer registry human research ethics committees in Melbourne and Perth. Eligible cases comprised male residents of Melbourne and Perth diagnosed between 1994 and 1997 and recorded in the population-based cancer registries with a histopathology confirmed diagnosis of adenocarcinoma of the prostate (International Classification of Diseases, 9th revision, rubric 185), excluding well-differentiated tumors (defined as low grade, i.e., those with Gleason scores <5). Cases had to be <70 years of age at diagnosis. Controls were randomly selected from men on the current state electoral rolls and were frequency matched to the predicted age distribution of the cases in a ratio of one control per case. Potential controls were matched against the cancer registries at the time of recruitment to exclude men with a known history of prostate cancer. Controls were identified and interviewed contemporaneously with the cases over the period 1994 to 1997. The cohort comprised >95% of Caucasians. The lack of racial diversity is not critical to this analysis since the frequency of the NKX3.1 polymorphism does not vary between races. 23 A total of 798 cases and 527 controls were analyzed. Of the total cases, 582 were described as low-to-moderate grade and 216 as high grade (Gleason score >7 or poorly differentiated or undifferentiated tumors). Among the cases, 545 were T1/T2 and 249 were locally advanced or metastatic at presentation, with the remaining 4 cases of unknown stage. Median age for cases and controls was 62 years.
Serum IGF-I assay
Serum IGF-I levels were determined by ELISA with reagents from Beckman Coulter (DSL, Webster, TX). All assays were performed in the laboratory of one of the authors (MJP). Reliability of laboratory assays of plasma levels are always checked before measuring the real samples. The mean intrapair coefficients of variation for blinded duplicate quality control samples were 2.6% for IGF-I, and the long-term intraperson correlation coefficient for these biomarkers was 0.66 for IGF-I (3 years apart). 14 Cases and controls were separated by tertile of serum IGF-I levels. The median values for the IGF-I levels in the tertiles from the controls in each study are reported. For the PHS, the IGF-I measurements were made at 4 different times when batches of 92, 257, 73, and 105 control samples were assayed. Batch-specific cutoff points were used for tertiles, and overall median values are reported as the mean of the 3 median values. Inclusion of the IGF-I values for the cases as well as controls changed the overall means by less than 1% for each tertile.
Statistical analysis
For the PHS, the baseline characteristics and information at prostate cancer diagnosis for all the 937 cases and 1,086 controls and for the subgroup of 673 cases and 527 controls with both the genotype data and plasma IGF-I levels are presented in Table 5. Using Pearson’s goodness-of-fit test, the NKX3.1 SNP does not violated Hardy-Weinberg equilibrium (
We first conducted case-control analysis assessing the overall association of the NKX3.1 polymorphism with risk of developing prostate cancer among all the 937 cases and 1,086 controls. We also evaluated the associations separately by Gleason grade (<7 vs. ≥7), by clinical stage (localized T1/T2 vs. advanced T3/T4/N1/M1), by fatal prostate cancer as outcome, and by year of the cancer diagnose (1982-1994 vs. 1997-2007) (Table 2). These analyses were then repeated by median age at diagnosis (<70 vs. ≥70 years). Finally, we assessed interaction between baseline plasma IGF-I levels and the NKX3.1 polymorphism (Table 4). We then assessed this interaction in the RFPC study using the same strategy. All statistics were calculated using SAS (version 9.1.3; SAS Institute Inc, Cary, NC), with a 2-sided significance level of 0.05.
Footnotes
Acknowledgements
We thank Dallas English and John Hopper, principal investigators of the Risk Factors for Prostate Cancer Study, for their contributions to this work.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by NIEHS Grant ES09888 to EPG and by DOD Grants W81XWH-07-1-0263 to EPG and PC-05-0590 to EM. PHS was supported by NIH Grants CA42182, CA90598 and DOD Grants PC050569, CA097193. The RFPC study was supported by grants from the Australian National Health and Medical Research Council (#504700 and #504702) and by infrastructure from the Cancer Council Victoria.
