Abstract
In bacteria, replication is a carefully orchestrated event that unfolds the same way for each bacterium and each cell division. The process of DNA replication in bacteria optimizes cell growth and coordinates high levels of simultaneous replication and transcription. In metazoans, the organization of replication is more enigmatic. The lack of a specific sequence that defines origins of replication has, until recently, severely limited our ability to define the organizing principles of DNA replication. This question is of particular importance as emerging data suggest that replication stress is an important contributor to inherited genetic damage and the genomic instability in tumors. We consider here the replication program in several different organisms including recent genome-wide analyses of replication origins in humans. We review recent studies on the role of cytosine methylation in replication origins, the role of transcriptional looping and gene gating in DNA replication, and the role of chromatin’s 3-dimensional structure in DNA replication. We use these new findings to consider several questions surrounding DNA replication in metazoans: How are origins selected? What is the relationship between replication and transcription? How do checkpoints inhibit origin firing? Why are there early and late firing origins? We then discuss whether oncogenes promote cancer through a role in DNA replication and whether errors in DNA replication are important contributors to the genomic alterations and gene fusion events observed in cancer. We conclude with some important areas for future experimentation.
Introduction to DNA Replication
Making another strand of DNA that is exactly the same as the existing strand can be argued to be the single most important job of a cell. Unwinding, replicating, and rewinding 3 billion base pairs must be an incredibly challenging task. The cell accomplishes this by creating a series of sites at which replication will be initiated and by opening the double-stranded DNA at those sites to form a small bubble. DNA is replicated outward from these sites in both directions by replication “forks.” Once these forks are formed, they may encounter obstacles and roadblocks: DNA adducts, the presence of other enzymes, or a lack of dNTPs. A complex machinery exists to stabilize replication forks, to help prevent their collapse and failure, and to prevent them from firing under conditions in which they are likely to fail. Fork failure has been associated with genetic changes, and recent data indicate that DNA replication may be an important contributor to the genetic alterations observed in humans. We might imagine that there would be a strong drive to organize DNA replication in a way that optimizes the cell’s ability to faithfully create a replica of its DNA.
DNA replication is initiated when a
The activity of S-phase kinases and Cdc7/Dbf4 is tightly regulated in order to ensure that DNA replication only occurs during S phase and that each portion of chromosomes is replicated only once per cell cycle. Changes in cyclin-dependent kinase activity, 10 proteasome-mediated destruction of Cdt1, 11 and a Cdt1 inhibitory protein called geminin prevent the assembly of new prereplication complexes until mitosis, when prereplication complexes are permitted to reassemble. By restricting the formation of the prereplication complexes to only a short window before S phase, the cell can ensure that origins fire only once during the cell cycle. 12 When the cells enter the next S phase, only existing complexes are permitted to fire.13,14 After an origin fires or when it is passively replicated by a replication fork from a neighboring origin, the origin loses its initiation proteins. 15 Thus, origins that have already fired are prevented from reloading a prereplication complex prior to the next cell division. 16 Through these mechanisms, the cell prevents origins from firing more than once per cycle, 17 which could result in re-replication, DNA damage, or cell death.
Even if DNA replication is restricted to one round in a cell cycle, it is still a process that is fraught with challenges. Replication forks must navigate challenging DNA structures such as triplet repeats, palindromic sequences, G-quartets, telomeric repeats, DNA adducts, and tRNA genes, all of which create problems that cause the replication forks to stall. 18 They must also replicate genes that are actively being transcribed, which may result in a collision with an RNA polymerase. There may be low levels of dNTPs or proteins important for the creation of the replisome. 19 As a result of all of these challenges, replication forks are often in danger. In budding yeast, forks stall approximately once per 10 kb. 20 When DNA polymerases encounter barriers, one possibility is that the blockage is removed and the polymerase can continue. 21 Other possible outcomes can result in errors in the passage of genetic information, for instance, single-stranded DNA and DNA breaks can be formed, which can lead to single-stranded 3′ tails from broken forks annealing with microhomology to single-stranded DNA nearby. 22 Indeed, a pattern of mutations characterized by microhomologies is induced by treatment with the inhibitor of ribonucleotide reductase hydroxyurea or the DNA polymerase inhibitor aphidicolin.23,24
When replication problems arise, the cell can activate checkpoint mechanisms that integrate information about the presence of DNA damage and the replication status of the cell and provide the cell with multiple different types of assistance. RPA-bound, single-stranded DNA accumulates at stalled forks and acts as a signal to checkpoint kinases. 25 The checkpoint kinases ATR/Mec1 and ATM/Tel1 stabilize stalled forks 19 and prevent their collapse.19,26-28 In yeast with mutations in the ATR pathway, stalled replication forks rapidly collapse into a reversed-fork conformation.19,27 These structures recombine very easily and can lead to genome rearrangements including inverted repeats19,27,29 and translocations.30-35 In an analysis comparing the most important contributors to cell death in yeast, checkpoint activation of stalled replication forks was found to be central to cell viability, while other pathways such as the regulation of mitosis, gene expression, and origin firing only contributed modestly. 36
Interest in replication fork–mediated genomic changes has increased recently because of deep sequencing studies of human genomes.22,28,34,37,38 The complex genome rearrangements discovered in some patients with genetic disorders suggest elaborate replication fork failure.22,28,34,37,38 One disorder that has been investigated is Pelizaeus-Merzbacher disease an X-linked myelination disorder often caused by nonrecurrent duplications of the proteolipid protein 1 gene. Analysis of this gene in patients revealed complex patterns of duplications.22,34 The types of alterations included multiple copy number changes, deletions, inverted duplications, triplications, insertion of short sequences at break points that were templated by nearby genomic regions, and microhomology at the break-point junctions. Based on comparisons with experimental observations from bacteria, yeast, and human studies,22,39 the changes were considered consistent with a replication-based model for their generation based on fork stalling and template switching. 40 Further studies of the dystrophin gene and other disease-causing genes revealed a similar pattern with deletions and duplications that were interpreted as likely resulting from stalled and dislodged replication forks that re-engaged at a different template based on a limited region of homology.41-43
Because of the critical importance of performing the complex act of DNA replication, we anticipate that the cell has evolved an optimized and efficient strategy to achieve high fidelity transmission of genetic information. Yet, we still do not fully appreciate the most important principles for the organization of DNA replication in eukaryotes. The last year has seen an explosion of information. High-throughput methods for identifying origins have become possible, and several reports have emerged describing the systematic identification of a large number of origins in multiple species. Information on the timing of DNA replication and on the 3-dimensional organization of chromatin has provided insights into the dynamics of origin firing and its relationship with the spatial organization of chromatin. Other studies have highlighted the role of new pathways in preparing origins for replication, in organizing 3-dimensional replication structure, and in creating replication checkpoints.
We review the relevant literature on DNA replication in different species, focusing on recent advances. We then synthesize this information to discuss the question: what are the most central problems that a cell faces when it replicates its DNA, and has it organized the selection of origins for firing and the timing of origin firing in order to minimize these problems? We then consider how these findings relate to the formation of tumors and whether they can explain the types of mutations observed in tumors. Finally, we articulate what we believe will be the exciting areas of research that will advance this field.
Replication in Different Organisms
Replication in bacteria
In a bacterial cell cycle, chromosome replication starts at a single genomic locus, at a well-defined sequence, the origin. The replication origin in
DNA replication in budding yeast Saccharomyces cerevisiae
Budding yeast such as
Since DNA is not loose within the nucleus but rather tightly wound around octamers of histones and further organized into 30-nm fibers of condensed nucleosomes, accessing a single strand of DNA for replication requires the ability to open and unravel chromatin and gain access to an individual strand. Depleting nucleosomes nearby the potential origin may facilitate formation of a prereplication complex. RNA polymerases face similar challenges as DNA polymerases58,59 and binding sites for transcription factors that regulate the accessibility of DNA to RNA polymerases can also facilitate replication initiation.
59
For instance, the B3 element of the ARS1 replication origin is a recognition site for the transcription factor Abf1. Replacing the Abf1 recognition site with the recognition site for a different transcription factor, GAL4, allowed the origin to retain its capacity to nucleate DNA replication.
60
Further, mutations that disrupt silent chromatin at telomeres in budding yeast activate a telomeric origin that is typically silent.
61
Thus, in
Replication in fission yeast
In the fission yeast
Replication in embryonic frogs and flies
Frog and fly embryos use a radically different replication model from the approach used by bacterium. Early in the development of both frog and fly embryos, when rapid chromosomal replication is needed to keep up with the fast pace of cell division, DNA replication initiates at many sites throughout the genome all at the same time.70,71 The regions of the chromosomes that are used to initiate DNA replication are not characterized by the presence of any particular sequence, and instead, origins are formed and fire from seemingly random sequences present at short, regular intervals. Any plasmid DNA injected into a
In both
To more directly test the relationship between origins of replication and transcription factor binding sites, an inducible transcription template was introduced into
Replication in metazoans
In mature cells of higher eukaryotes, there are specific sequences that identify the positions at which replication initiates. For instance, if the origin of replication in the human β-globin gene locus along with 8 kb of surrounding DNA is transferred to a different site in the genome, it can still direct site-specific initiation of replication. However, replication origins in higher eukaryotes, unlike in bacteria or budding yeast, do not share a consensus sequence, even though the proteins that are recruited to origins in higher eukaryotes are similar to those in bacteria and yeast.6,45 Unfortunately, the types of functional assays that successfully identified origins of replication in yeast based on their ability to promote the replication of bacterial plasmids, have not been successful in mammalian cells.45,80 However, enough origins have been identified that patterns have emerged. Some of the specific sequences enriched near origins in fission yeast are also important for higher eukaryotes as well. For instance,
In higher eukaryotes, origins are not utilized with the same efficiency; some initiate replication in almost every cell cycle, while others rarely fire and instead are passively duplicated by a replication fork initiated at neighboring origins. In
Many of the known replication origins map to intergenic regions close to promoters in higher eukaryotes,5,91-95 and highly expressed genes tend to cluster near origins of replication.47,48 Further, transcriptionally active regions in mammalian cells tend to replicate early in S phase.
95
Origins that fire early tend to have more open, accessible chromatin,95-96 acetylated histone lysines,
97
and high levels of activating histone dimethylation and trimethylation on H3K4.98-100 Modulation of histone acetylation and chromatin conformation patterns is sufficient to affect DNA replication programs.101-104 Origins of replication are also associated with nearby unmethylated CpG islands and promoters.93,105,106 Origins at unmethylated CpG islands replicate earlier than those at methylated CpG islands.
107
Origins activated late in S phase are associated with nontranscribed, heterochromatic regions.95,105,108,109 Eventually, all of the genome must be replicated, so origins in heterochromatic regions do ultimately fire, but they tend to fire late in S phase.
110
In humans as well as in
One example of an origin of replication that demonstrates the association between transcription and replication in mammalian cells is the β-globin locus, a site that is strongly transcriptionally induced in erythroid cells that synthesize large amounts of hemoglobin.112,113 In the human erythroleukemic cell line K562, expression of the β-globin locus is strongly dependent on a locus control region (LCR) >20 kb upstream of the β-globin gene that contains binding sites for transcription factors including those of the Maf and bZip family proteins. Although the LCR is located kilobases away, it is required for replication initiation because deletion of a region containing the LCR abolishes replication initiation from the β-globin origin. 114
The association between origins of replication and transcription start sites could reflect a functionally important role for transcription factors in activating replication origins. In viruses, yeast,
The oncogenic transcription factor c-Myc has also been implicated in the control of replication origin firing, 128 possibly through its role in remodeling chromatin. 129 Myc forms heterodimers with members of the Max family of proteins and binds to the E-box DNA sequence CACGTG. Myc-bound E-boxes can then associate with other proteins to remodel chromatin. 130 In addition to its role in promoting transcription, c-Myc was found to co-immunoprecipitate with protein components of the ORC, suggesting a role in replication. Myc overexpression resulted in DNA damage during S phase, 128 which may reflect a contribution of excessive origin firing. Further, c-Myc overexpression in primary human fibroblasts accelerated S phase, while c-Myc–deficient fibroblasts exhibited a prolonged S phase. 131
A recent paper closely investigated the role of c-Myc in regulating the firing of the lamin B origin, 132 one of the well-mapped human replication origins (Fig. 1). 133 The lamin B origin contains an E-box to which c-Myc and its partner Max bind in early G1 phase. The histone methyltransferase mixed lineage leukemia 1 (MLL1) was recruited by c-Myc to the lamin B origin and added methyl groups to lysine 4 of histone H3 in nearby nucleosomes. After MLL1 and c-Myc were released from the origin, the H3K4me3 modification persisted and served to recruit the histone acetylase HBO1, which acetylated lysines on histone H4. Hyperacetylation resulted in lower nucleosome occupancy, which facilitated the loading of MCM proteins at the origin.
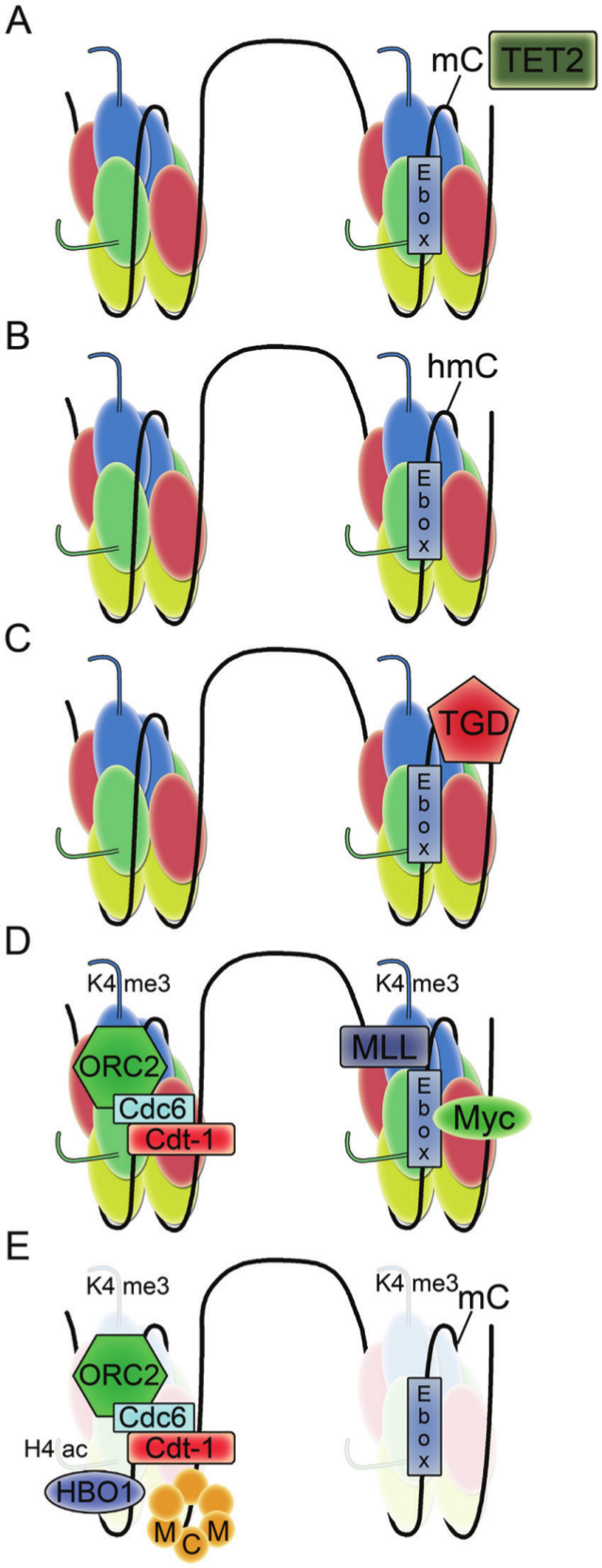
Model for the initiation of DNA replication at the lamin B2 origin. The TET2 enzyme converts methylated cytosine to hydroxymethylated cytosine. The hydroxymethylated cytosine is excised by thymine DNA glycosylase. The E-box with unmodified cytosine is recognized by Myc as origin proteins ORC2, Cdc6, and Cdt1 are bound. Myc recruits MLL, which modifies histone H3 on lysine 4 to the trimethyl form. The histone H3K4me3 mark promotes the binding of HBO1 acetylase and facilitates histone H4 hyperacetylation. The resulting hyperacetylation favors nucleosome remodeling that facilitates the loading of MCM proteins.
The authors also discovered that loading of the c-Myc protein to the origin was controlled by demethylation of CpG sites in the E-box. 132 During the transition from G0 to G1, CpG sites were first converted to hydroxy-CpG by Ten-Eleven Translocation (TET) enzymes and then repaired by thymine DNA glycosylase-mediated base excision repair, which removed the modified nucleotides. Removal of the modifications prepared the CpGs for Myc binding and the rest of the remodeling required for MCM complex loading. The findings support a model in which cytosine demethylation, Myc recruitment, and histone methylation-acetylation crosstalk result in nucleosome remodeling, replication complex recruitment, and origin licensing. It will be interesting to determine whether this model is confirmed and extended to other origins. For the DHFR origin, comparison of the active and inactive X chromosomes suggests that the retention of methylation at the CpG regions is not important for origin function.107,111 Further, mouse embryonic stem cells with genetic defects in CpG methylation did not exhibit differences in origin firing for many single copy loci. 134
Global origin mapping
Without a functional assay for sequences capable of initiating replication, 45 the main approach to understanding DNA replication in mammalian cells until recently had been to identify and investigate individual positions at which DNA replication initiates. With this strategy, about 30 origins were identified.86,135 In the past few years, with the widespread adoption of high-throughput sequencing technologies, there has been an explosion of information as hundreds of origins have been identified and mapped. The ability to systematically define human origins has made it possible to consider anew the properties of replication origins in humans and the important principles of DNA replication.
One of the first such studies involved mapping replication origins based on the sequence composition of the surrounding bases. The leading and lagging strands of replicated DNA display different rates for each of the possible base pair substitutions.136-138 In most species, the leading strand is richer in G relative to C and, to a lesser degree, richer in T relative to A. 139 The most widely held view is that the cause of this skew is deamination of C to T. 140 The rate of cytosine deamination is 140-fold higher in single-stranded than double-stranded DNA, 141 and the leading strand spends more time single-stranded than the lagging strand during DNA replication. In bacterial chromosomes, where there is a single origin and a single terminus, and each nucleotide is always either leading or lagging, a strong compositional bias is observed.139,142 Drawing GC skews, defined as (G – C)/(G + C), in sliding windows has become a standard method to identify the origin and terminus of replication in bacteria as the sign of the TA and GC skews changes abruptly when crossing replication origins and termination sites.143,144 In yeast chromosomes, there are many autonomously replicating sequences, only some of which are used in each replication cycle, so a particular single-stranded sequence may be replicated as the leading strand in some cell cycles and the lagging strand in others. At the very end of yeast chromosomes, however, there is only one leading/lagging option, and in these sequences, the same asymmetry present in bacterial chromosomes is observed. 145
Analysis of nucleotide composition asymmetry around experimentally determined origins in mammals revealed that in 6 out of 9 cases, the skew displayed an abrupt sign switch at the origin similar to the pattern in prokaryotes. 146 This observation led to the computational prediction of around 1,000 putative origins representing about 27% of the genome.146,147 Within 50 kbp of putative origins, the mean density of genes that are transcribed in the same direction as they are replicated was 8.2 times greater than genes transcribed and replicated in opposite directions. 147 These findings would suggest a strong concordance between directionality of transcription and replication in mammals but, as described further below, have been reanalyzed.
Experimental analyses have also been performed to systematically capture and map large numbers of replication origins. One approach has been to isolate transitory, RNA-DNA short nascent strand molecules created at the beginning of DNA replication based on their protection from exonuclease treatment and to hybridize them to microarrays.
105
Applying this approach with microarrays that cover 1% of the genome resulted in a dataset of 283 origins of replication with interorigin distances that ranged from 1 kb to 500 kb. Origin sequences were more evolutionarily conserved than expected by chance, and half of them mapped within or near CpG islands.
105
Most of the origins overlapped transcriptional regulatory elements, and in this study, a significant correlation was observed between origins and c-Jun and c-Fos binding sites. The position of origins correlated strongly with DNAse-hypersensitive sites, acetylated histone H4, and H3K4 dimethyl and trimethyl marks. The findings were consistent with the replication machinery preferentially recognizing open chromatin structures near promoters of actively transcribed genes, although approximately 30% of origins did not overlap with open chromatin marks. Similar findings were reported in other studies of origins identified by nascent strands99,148 or with a protocol in which restriction fragments containing origins are trapped in gelling agarose based on their partially circular nature and sequenced.
149
While the findings of enrichment near CpG islands and transcription start sites were consistent among these studies, unfortunately, the concordance between the actual sites identified as origins was low when comparing
Topological constraints, gene gating, and higher order chromatin structure
In addition to recent information on the mapping of origins genome-wide, there have also been a recent series of papers describing DNA replication with respect to the 3-dimensional topology of chromatin. All DNA-related processes including both replication and transcription generate torsional energy that can result in negatively or positively supercoiled DNA when the double helix underwinds or overwinds.
151
When 2 forks converge during replication termination or when 1 fork clashes with a transcription bubble,152,153 especially if the helix is anchored and cannot rotate, the expectation is that there will be positive supercoiling ahead of the replication fork or transcription bubble and negative supercoiling behind it (Fig. 2A).
152
If the torsional stress that is created by the encounter cannot simply diffuse through the chromosome by untwisting it, then type I and type II topoisomerases can make single- or double-stranded breaks, respectively, and catalyze strand passage reactions, thus changing the linkage of DNA molecules.151,153 Both
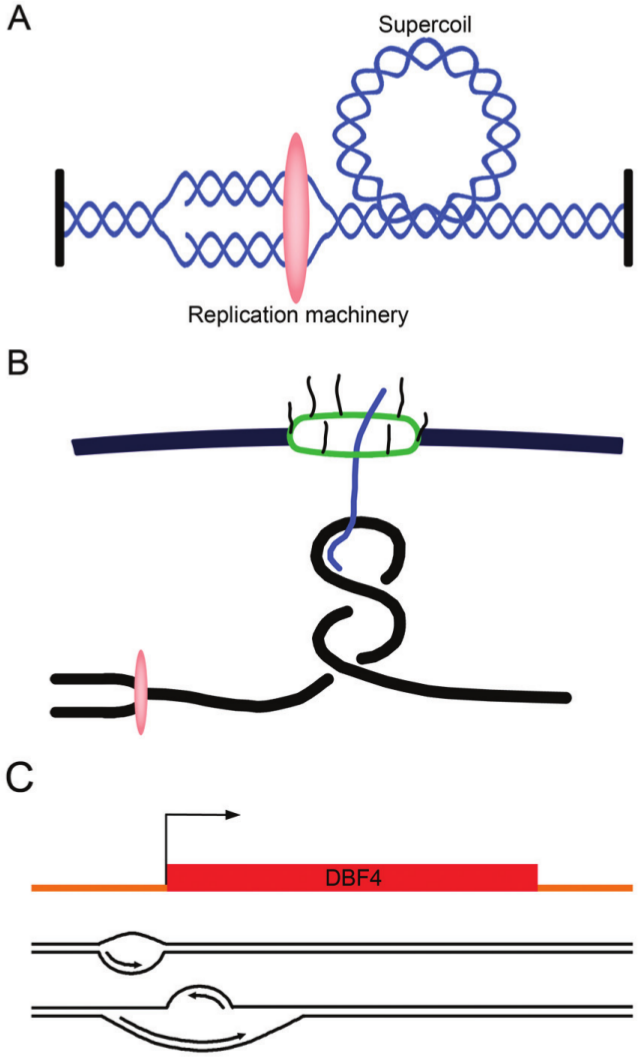
Topological problems and a potential solution associated with DNA replication. (
Recent studies have also shown that RNA polymerase II–transcribed units are organized into loops, 155 which may facilitate RNA polymerase recycling and promote repetitive rounds of transcription of the same gene. 155 Through gene gating, these loops are coupled to the nuclear pore complex, which allows for a coupling of transcription with mRNA export (Fig. 2B). The proteins that mediate this interaction are the THO/TREX and TREX-2 complexes and the nucleoporins.156-159 Gene gating might serve the purpose of decreasing the likelihood that the nascent RNA becomes tangled in the RNA polymerase bubble.160,161 However, by attaching the transcribed DNA to a fixed structure such as the nuclear pore, there may be an exacerbation of topological problems associated with transcription, especially when the replication fork arrives.154,156
The challenge faced by the replication fork when it encounters these gene loops is demonstrated by the finding that transcribed genes represent the most abundant sites of replication fork pausing in the yeast genome, and replisome pausing at transcribed genes is independent of the gene’s orientation with respect to the replication fork.
162
During S phase,
While the selection of individual origins suggests a randomness to the replication process in higher eukaryotes, in fact, the replication of eukaryotic chromosomes does exhibit a temporal and spatial organization within the nucleus. The characteristic timing and location of specific chromosomal regions are thought to reflect a higher order structure of the genome within the nucleus.167-169 One manifestation of the effects of a higher order structure of chromatin on replication is the presence of replication foci that are the nexus of multiple origins. Their position in the nucleus as well as temporal order of activation are inherited throughout cell cycles.170-173 Possibly as a result of this higher order structure, even though different individual origins may be selected to fire, there is a reproducible replication timing for broad regions of the genome.82,83,173-176 When followed for as many as 15 cell cycles, labeled foci that represent replication factories do not mix, separate, or change in shape.170,173 Moreover, replicon clusters that fire at different times during S phase occupy different subnuclear compartments. 170
High-resolution replication timing profiles in mouse embryonic stem cells revealed multimegabase, coordinately replicated regions of the genome that are separated into distinct nuclear regions. 176 The regions that act as boundaries were consistent between several embryonic stem cell lines and induced pluripotent stem cells. Upon differentiation to neural precursor cells, approximately 20% of the genome transitioned to different domains, with the predominant pattern being a consolidation into fewer, larger replication units in the differentiated cells. These studies and others have led to a model in which distant genomic regions of similar replication timing come together to form replication factories in which DNA is replicated in multiple regions simultaneously (Fig. 3).177-179
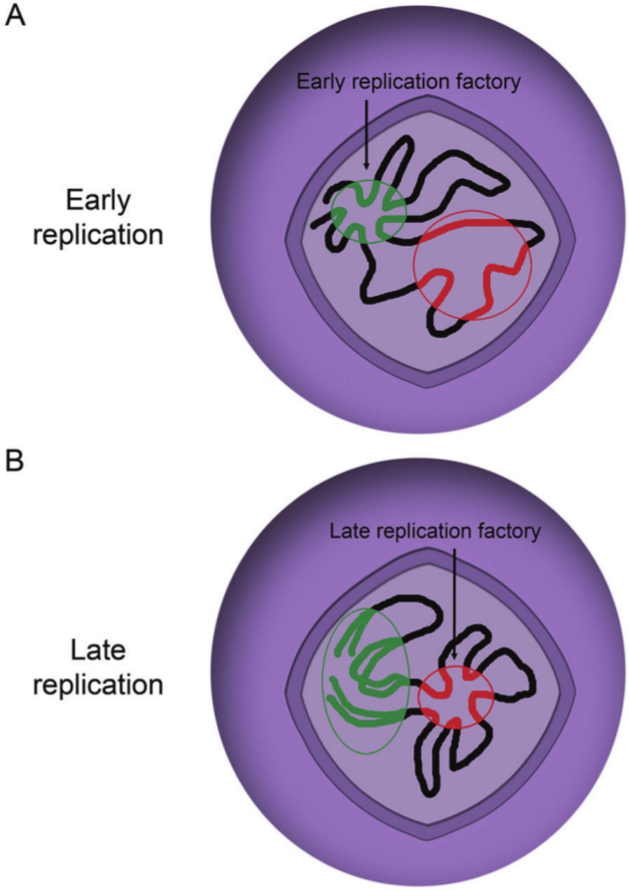
Model for the organization of genomic DNA with respect to DNA replication. Replication origins in open chromatin are grouped together into a replication factory early in S phase. Later in S phase, a different set of origins are clustered together to form a replication factory.
Proteins that regulate the formation of these replication factories have recently been identified. In mice, the absence of the nuclear matrix–localized Rif1 protein resulted in changes in the temporal order of origin firing as well as the physical definition of chromosome domains, 180 implicating Rif1 as part of the machinery that establishes the accessibility of different origin clusters for replication factors. In yeast, mutations in the forkhead transcription factors resulted in extensive changes in the timing of origin firing, with early firing origins consistently firing later than normal and late firing origins shifting to an earlier firing time. 181 The role of the forkhead proteins in transcription was unrelated to this phenotype, which more likely reflects the newly discovered ability of forkhead transcription factors to control the clustering of early origins and their association with the key initiation factor Cdc45 in G1. 181
Understanding the Organization of DNA Replication
How does the cell organize the firing of its origins?
Bacteria exhibit a very strict replication model in which a single origin is fired at a specific time, while origin sequences in embryonic flies and frogs vary from cycle to cycle, do not share consensus motifs, but are regularly spaced. Higher eukaryotes use a different approach from either of these. There are specific sequences that function consistently as origins in higher eukaryotes, but only a fraction fire with each cell cycle. The simplest model would be one in which origins are selected at random for firing. After all, since S phase is distinct from the phase in which origins are licensed, perhaps the location of origins of replication does not matter. 45 However, if potential origins were distributed randomly along the genome, one expects a geometric (exponential) distribution of separations. 182 This would result in some very large interorigin distances, which could raise problems. If the gap is too large, the cell might not have time to replicate that region, and the region might be unreplicated at mitosis or delay the length of S phase. 88
Recent studies employing DNA combing have suggested 2 different models to explain how the cell avoids this problem (Fig. 4). In DNA combing, sites of DNA replication are labeled at 2 different times with 2 different nucleotides and visualized with fluorescent antibodies. The data provide quantitative information on replication rates. Guilbaud and colleagues 183 performed DNA combing studies and found that DNAse-hypersensitive sites and CpG islands were more abundant at early firing locations and steadily decrease as the replication forks head toward late firing regions. Replication fork velocity did not change during S phase, but the global fork density increased over the course of S phase, which was interpreted to indicate that the efficiency of firing increases through S phase (Fig. 4A).88,184 This may reflect changes in DNA supercoiling in front of the fork, other proteins being recruited to the later origins, possibly because they have been released by the early firing forks, or increased S-phase kinase activity.185-187 Combining the DNA combing data with chromosome conformation capture that provides information on long-range chromatin interactions 177 revealed the presence of U-shaped domains that correspond to blocks of enriched interaction. Thus, the genome is segmented into replication timing domains that correspond to spatially compartmentalized chromatin units. These domains are insulated on either side by 2 boundaries of open, accessible, actively transcribed chromatin that are also enriched in the insulator binding protein CTCF. The authors proposed a model for U-shaped chromatin domains in which replication would first initiate in efficient zones with an open chromatin structure on the outside of the domain and then work progressively and with increasing efficiency toward origins at the base of the U with more compact chromatin.183,188 This “domino” model may help to explain why replication progresses much faster than the known speed of a single fork, how late firing origins that are in regions of inaccessible chromatin can initiate, and why adjacent origins fire synchronously. Further, under this model, even if there are long stretches of chromosomes that are origin-free, replicating them would not take excessively long. The increasing density of origin firing with S-phase progression predicted by this model has been observed in budding yeast and frog embryo extracts.62,71,184,189
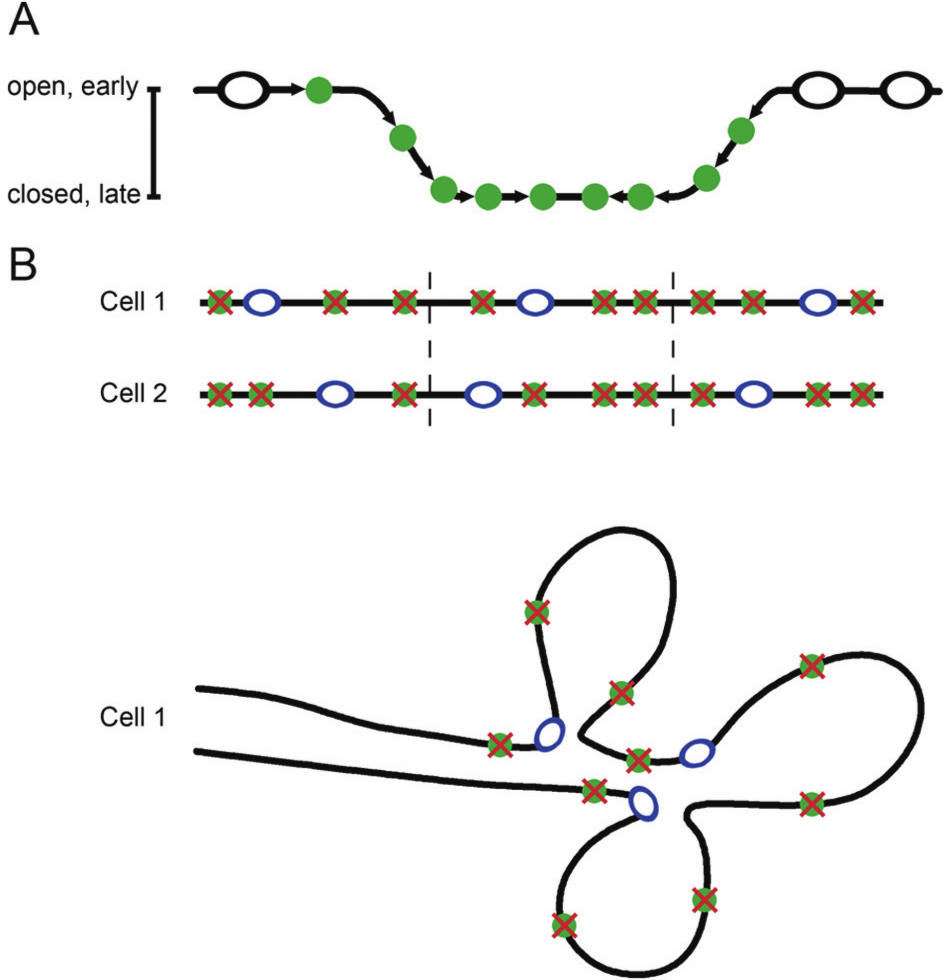
Two possible models to explain the lack of long gaps between replication origins. (
Cayrou and colleagues
148
also performed DNA combing experiments and arrived at a somewhat different model for DNA replication. They compared their DNA combing data in
Is the cell coordinating transcription and replication?
One of the clear messages from recent genome-wide analyses of origins in humans is an association with transcription start sites. As described above, in multiple species, early firing origins tend to have an open chromatin configuration and to be located near transcription start sites.5,91-94 In general, a tissue-specific gene replicates earlier in S phase in cells in which it is expressed than in cells in which it is transcriptionally silent.
191
It is important to note, however, that in at least some studies, a relationship between replication timing and transcription during early S phase was not observed.
192
The association between replication and transcription does not require transcription
However, even if transcription is not required
Another possibility is that the genome has been organized to minimize collisions between DNA and RNA polymerases. Because both forks advance and progression of the transcription bubble generates positive supercoiling, a head-on collision between replication and transcription machinery is expected to cause severe topological impediments and fork pausing.199-201 Fork stalling in response to head-on collisions with the RNA polymerase has been observed using electron microscopy to monitor fork progression after inverting the direction of ribosomal operons, 202 with 2-dimensional gel electrophoresis of replication fork intermediates, 203 and microarrays.204,205 Most studies indicate that replication is slowed more substantially as a result of head-on collisions with transcription units that oppose the fork compared to co-directional encounters,199,206 although co-oriented collisions can also result in RNA polymerase stalling.162,199,207,208 Replication fork arrest as a result of collisions with transcription complexes can lead to double-stranded breaks, DNA damage response, mutagenesis, and chromosomal deletions. 204
Prokaryotes have only one replication origin, and they have organized their genomes in a way that has been interpreted as minimizing collisions between replication forks and transcription bubbles. For
As described above, the most recent analysis suggests that human cells have not achieved co-directionality between transcription and replication. Necsulea and colleagues
150
analyzed experimental and skew predictions of origins. While for the computationally predicted origins there was a strong leading strand bias, for the experimental data, the association between the direction of transcription and replication was essentially random. Further, the leading strand fraction was similar for highly and lowly expressed genes and was not different for genes expressed in S phase. There are several possible explanations. One possibility is that head-on collisions in eukaryotes are not as consequential as might be expected based on findings in bacteria. Perhaps the head-on collisions studied most closely in bacteria are particularly disastrous. The severity of replication fork arrest due to a head-on collision is correlated with the level of gene expression, and in some studies, only heavily transcribed genes significantly impeded the progression of the replication fork when inverted.
204
Bacterial rRNAs, the most intensively studied genes, have very high transcription rates. Also, in bacterial rRNAs, there may be clustering of multiple RNA polymerases that work together,
215
so a collision with this multi-RNA polymerase complex may lead to particularly ruinous consequences. On the other hand, data in
Even in bacteria, whether the main goal of genome organization is eliminating head-on collisions is being reconsidered.
142
The frequency of co-directional genes is approximately 75% in
Another possibility is that eukaryotes have evolved other mechanisms to prevent head-on collisions. Yeast have evolved a clever solution in which they ensure that certain highly transcribed genes are replicated only in a single direction by erecting a nucleic acid–protein barrier to forks entering from the 3′ direction opposite to the direction of transcription.
219
For ribosomal DNA repeats in yeast, a replication fork barrier that consists of a
Another possibility is that in eukaryotes, the presence of a transcription loop is problematic, and the directionality of the replication machinery is not as important. In support of this model, the regions of the genome with the highest number of pause sites in
A final intriguing possibility is that the genome is, in fact, organized to minimize head-on collisions, but instead of solving the problem through the orientation of genes, higher eukaryotes resolve it in the way they activate their origins. Evidence for this model comes from a report on asymmetric bidirectional replication at the DBF4 origin. 227 Using 1-way PCR-based primer extension, the DBF4 origin was found to contain 2 initiation zones, one on the sense strand and one on the antisense strand, separated by approximately 400 bp that include the transcription start site (Fig. 2C). DBF4 replication starts from initiation zone I, which has more open chromatin, and then proceeds in the sense direction, that is, the direction of DBF4 transcription, toward initiation zone II. Replication of the antisense strand from initiation zone II began after the replication on the sense strand had reached or passed through this initiation zone. ORC binds both initiation zones, both have DNase I–hypersensitive regions, and replication of both strands proceeds as though it is a leading strand. Combining the asymmetric replication model with the fact that origins are often at the beginning of genes implies that the replication could be specifically oriented to follow the direction of transcription, with a subsequent origin firing to replicate the opposite strand.
It remains to be determined whether this asymmetric bidirectional replication model is more widely used. In a follow-up study, the same authors used 1-way PCR-based mapping to monitor the lamin B origin and concluded that it likely also contains 2 short initiation zones with a 40-bp noninitiation zone in between.
228
Further, when Cayrou and colleagues
148
used nascent strand purification to identify origins in
By using the same transcription factors that facilitate both transcription and origin firing, it would be possible for the cell to flexibly coordinate these processes. If the cell takes on a new fate or differentiates, it can both induce the expression of a particular gene, for instance, a hemoglobin or immunoglobulin component, and at the same time ensure that the portion of the genome encoding this newly critical gene will be replicated early. The reprogramming of cells when they take on a different fate, both in terms of replication and 3-dimensional chromatin structure, may help to explain why there is so little correlation between, for example, the origins found in an ovarian cancer cell line and a lymphoblastoid cell line. 80 This would be consistent with findings that cells from different tissue take on different 3-dimensional chromatin structures. 232 It would also be consistent with findings of significant overlap in origin firing between 2 similar types of cells, which would be expected to have much more similar higher order chromatin structures. 232
What causes origin interference?
In addition to the selection of a specific origin, there is likely also a method to deselect origins. Indeed, if there is a selection of one and only one origin within a replicon, there is likely a method of origin interference, that is, a mechanism whereby the selection of one origin for firing inhibits the selection of nearby origins. 233 In budding and fission yeast, when 2 origins are located close together, it is rare that both origins are active in the same cell during S phase.57,60,233 Origin interference is unlikely to be related to ORC binding or prereplication complex assembly as these factors bind efficiently to all origins. Certainly, as the replication fork from the selected origin passes, it will inactivate nearby forks. 234 Thus, potential origins that are not as efficient will be inactivated as a consequence of the selection of a more efficient origin nearby.
In addition to this passive mechanism, the ATM and ATR checkpoint kinases also play an active role in the unfolding of S phase, even in unstressed cells.235-239 ATR–/– mice are embryonic lethal, and cells from the mouse are not viable, 240 supporting an important role for ATR without checkpoint activation. The ATM and ATR kinases downregulate the Cdk2 and Cdc7 kinases and thereby slow down the rate of DNA replication by blocking origin firing. 241 Inhibiting ATM and ATR kinases, for instance, by adding caffeine or neutralizing antibodies, increases the replication rate. 241 The likely activator of ATM and ATR is RPA-bound, single-stranded DNA, which would be expected to be high near an actively firing origin. With this mechanism, the cell can inhibit replication forks near a replication zone. The role of ATR as an inhibitor of S-phase progression could also explain its role in halting DNA replication when there are replication problems. The role of ATM and ATR in limiting origin firing in the context of DNA damage can be viewed from this perspective as an extension of its role in origin interference in unstressed cells. DNA damage or stalled forks would result in more extensive single-stranded DNA and thus a stronger checkpoint response. 236
Why have late origins?
The role of the ATM and ATR proteins in limiting origin firing raises the question: why have early and late origins? Does the existence of both early and late replication origins allow the cell to use the early origins as sensors for replication conditions and then adjust the rate of replication for late origins based on the results of the early firings using the checkpoint response? Or alternatively, would firing all available origins at the same time lead to a depletion of replication factors, fork-stabilizing proteins, or dNTPs? Would it result in too many replication fork convergences for the cell to accommodate? Consistent with these theories, there have been suggestions of a checkpoint that limits the total number of replication forks at any time during S phase. 226 However, any hypotheses about the need for both early and late origins would have to incorporate the fact that embryonic frog and fly cells do manage to faithfully replicate their genomes with a pattern that involves firing many origins simultaneously, at least for a few cell cycles.
One current model is there are proteins or protein complexes that are rate limiting for origin firing. 242 Indeed, substrates of the kinases that activate S phase have been reported to be rate limiting for origin firing in budding yeast. 242 The formation of replicon foci could reflect the need to bring early origins in close contact with each other to benefit from the availability of these factors. The distance between these origins is consistent with replication initiation sites being defined by an optimal loop size that correlates with the intrinsic stiffness of DNA. 182 An origin exclusion zone would be created as those origins that are physically too far from the limiting replication reagents. In this case, the physical position of the origin, in addition to ATR/ATM activation, would serve to limit the activity of origins near those that were selected. As S phase progresses, the limiting factors may be released or increase in abundance, allowing more origin firing. 185 Origins fired later in S phase could reuse rate-limiting proteins released by the early firing origins. In addition, the organization of DNA replication in higher order substructures could allow the cell to compartmentalize important proteins. 45 Certain types of topoisomerases, chromatin remodelers, or histone chaperones might be concentrated in specific nuclear subcompartments. 45 This might be consistent with theories that replication organization facilitates the propagation of chromatin states during DNA synthesis.110,243
Does the Organization of the Genome for Replication Contribute to Carcinogenesis?
There is substantial evidence that inappropriate organization of DNA replication may be central to the process of tumorigenesis. Some of the data to support this perspective are based on the demonstration that tumors exhibit a replication-induced DNA damage response. Early stages of tumorigenesis are consistently associated with the engagement of the DNA damage checkpoint response in multiple tissues.244,245 Analysis of precancerous and cancerous lesions from human patients revealed foci of DNA repair proteins, suggesting the presence of DNA double-stranded breaks not found in normal tissues.244-246
According to this model, early in tumor progression, cell proliferation and transformation are inhibited by senescence. Analysis of colon and bladder precancerous lesions shows that senescence markers coincide with DNA damage response markers. 247 Di Micco and colleagues, 248 for instance, showed that senescence triggered by the expression of an activated oncogene H-RasV12 in normal human cells is a consequence of a robust DNA damage response. When Ras was expressed in DNA damage response–deficient cells, the cells continued to proliferate rather than senesce. With DNA combing, they found that oncogene activation led to an increased number of active replicons and an increased asymmetry in the progression between right and left forks emanating from the same origin in Ras-expressing cells. Such discontinuous fork advancement might reflect increased fork instability or extensive fork pausing, possibly resulting from excessive origin firing. An outstanding question for the field is whether the activation of oncogenic factors mediates increased origin firing and a DNA damage response through an increase in the levels of replication proteins like CDC6 or whether oncogene activation stimulates origin firing more directly through epigenetic changes.
Several other lines of evidence are consistent with oncogenes affecting DNA replication. The oncogenic transcription factors c-Myc,
128
E2F,
115
c-Jun, and c-Fos
105
have been associated not only with transcription but also with origins of replication and with promoting DNA replication. Indeed, the transcription factors most closely associated with oncogenic transformation are also those that are enriched near origins of replication. In both precancerous lesions and cancers, oncogene activation induces the stalling and collapse of DNA replication forks, which in turn leads to the formation of DNA double-stranded breaks.
249
Activation of oncogenes and more generally of growth signaling pathways induces a loss of heterozygosity and genomic instability in mammalian cells cultured
The types of mutations observed in human cancers are consistent with the types of mutations that occur as a result of DNA replication errors. Double-stranded breaks resulting from excessive origin firing could contribute to the genomic instability that characterizes most human cancers. This has been tested by monitoring fragile sites at which replication forks preferentially collapse. 256 These loci are prone to the formation of microdeletions and gross chromosomal rearrangements and thus represent an indicator of the presence of replication stress. In both human precancerous lesions and in a human skin xenograft hyperplasia model, loss of heterozygosity was associated with common fragile sites, indicating that the lesions likely experience DNA replication stress.244,245 In oral precancerous lesions, loss of heterozygosity at the common fragile site Fra3B was a better predictor of progression to cancer than the other markers investigated. 257
As described above for genetic disorders, the types of mutations observed in cancers are also indicative of DNA damage response. Recently, data from deep sequencing of tumors have resulted in a new term “chromothripsis” to define instability in 1% to 3% of cancers, resulting in a highly complex pattern of genomic rearrangements with multiple copy number variants. 258 This type of mutagenic event is consistent with the expected errors from fork failure and template switching. 28 Multiple copy number and structural changes consistent with microhomology-mediated, break-induced replication are thus common to both cancers and genetic disorders.40,258,259
The relationship between the organization of DNA for replication and somatic copy number alterations, a hallmark of cancer, has been investigated directly. 198 Analysis of thousands of cancer samples including 26 cancer types revealed that the 2 boundaries of copy number alterations tend to be close to each other in the nucleus and replicate at a similar time. In fact, long-range interaction and replication timing data were sufficient to identify copy number variations. The authors concluded that the spatial proximity of regions replicating at the same time is an important contributor to the mutations observed in cancer.198,260
It is possible that a similar model will prove to be explanatory for the recurrent chromosomal translocations that are hallmarks of many cancers. Translocations, like copy number variants, tend to join genetic loci that are in close spatial proximity.232,261 As an example, BCR and ABL are not only in close nuclear proximity but are also replicated at a similar time in S phase, which might help to explain their frequent fusion to form the BCR-ABL oncogene that drives leukemogenesis. 198 A recent high-throughput, genome-wide translocation sequencing study 262 revealed that double-stranded, break-induced translocations to the IgH or c-Myc loci are much more likely to occur at sites that are either close to those loci or about 300 to 600 bp on the sense side of active transcription start sites. Microhomologies characterized the translocations. The correlation with sites that are origins of replication is striking and suggests that a 3-dimensional clustering of early origins into replicons might provide an opportunity for the formation of translocations when double-stranded breaks formed by the DNA polymerase at these genomic loci are clustered together in S phase. Thus, the association between cancer and fragile site mutations, the examples of “shattered” chromosomes that might reflect DNA replication failure, the association between DNA replication and the formation of copy number alterations, and the possibility that common translocations are a reflection of the organization of chromatin around DNA replication all suggest that DNA replication is a central driver of the genetic component of tumorigenesis.
Finally, the recent discovery that the TET enzymes are important for preparing CpG sequences in E-boxes for c-Myc binding and origin firing 132 could have repercussions for our interpretation of the role of mutations in isocitrate dehydrogenases in cancer. Gain-of-function mutations in isocitrate dehydrogenase enzymes IDH1 and IDH2 are frequently observed in tumors including acute myelogenous leukemia (AML) and glioblastoma. 263 The mutant versions of these enzymes have a neomorphic ability to generate 2-hydroxyglutarate (2-HG), and high levels of this metabolite have been identified in the serum of patients with IDH1 or IDH2 mutations. 264 2-HG can inhibit the activity of TET enzymes, which use α-ketoglutarate as an oxygen donor. By inhibiting TET enzymes, the IDH1/2 mutations limit the removal of cytosine methyl groups. Indeed, AML patients with IDH1/2 mutations have hypermethylated DNA, and the hypermethylation preferentially targets promoter regions and CpG islands neighboring transcription start sites. 265 2-HG can also inhibit the activity of other enzymes that rely on α-ketoglutarate as an oxygen donor, including histone demethylases, and consistent with this hypothesis, tumors with IDH1/2 mutations contain hypermethylated histones. 266 Further, mutations in the TET enzymes themselves have also been identified in AML, and these are found in a distinct set of tumors from those with IDH1/2 mutations, indicating that inhibition of TET enzymes is a common pathway to tumorigenesis. 267 The prevailing model is that altered CpG methylation and histone methylation in these patients are causative for tumor growth, and certainly, this may be the most predominant effect. The new findings relating TET enzymes to the preparation of origins for firing 132 raise the possibility that IDH mutations affect replication as well as transcription. IDH mutations would be expected to result in an accumulation of methylated CpG dinucleotides, thus inhibiting c-Myc from binding to its E-box at specific replication origins. It is interesting to note that IDH mutations are associated with a good prognosis in gliomas, 268 glioblastomas, 269 and acute myeloid leukemias. 270 There are a number of reasons that these tumors could have a favorable prognosis, including that they tend to have normal karyotypes, 271 they have altered methylation patterns, and they are associated with metabolic changes. It will be interesting to determine whether origin firing is impaired in patients with tumors with IDH mutations and whether this limits the tumor’s ability to grow.
Anticipated Future Directions
Recent studies have highlighted the importance of DNA replication for genetic diseases and cancer. We anticipate several emerging areas for the field of DNA replication. Application of methods for systematically identifying large numbers of origins of replication combined with methods to assess higher order chromatin structure will permit us to better understand the relationship between chromatin structure, origin selection, and replication kinetics. Our hope is that it will be possible to perform such studies in samples isolated directly from tissues. Such studies will likely help us to define the most important principles of DNA replication in higher eukaryotes: for example, what controls origin selection, origin interference, and the orderly timing of S phase. We anticipate that these technologies will allow us to better understand and classify the types of S phases in different cells, in different differentiation states, in response to different stimuli and in pathological states.
We also anticipate that more careful studies of the biophysical properties of replication forks, transcription loops, and nuclear gating will be forthcoming. Such studies might illuminate the biomechanical importance of the mechanisms that relieve the torsional stress associated with DNA replication such as topoisomerases, helicases, and checkpoint proteins. They might also provide more information on the topological state of DNA when it is replicated, transcribed, and subjected to both at the same time.
Further detailed analyses of the replication patterns at individual origins and of the factors required for DNA replication will also advance the field by providing more mechanistic insights and by potentially implicating other important pathways in the control of replication. It will also be important to test further some of the hypotheses put forward here such as whether the presence of a transcription factor binding site is an important regulator of origin firing, whether most promoters fire from a single origin or use bivalent origins, whether CpGs at myc-binding origins are demethylated with each cell division,272-274 and whether IDH mutations affect DNA replication as well as transcription.
Another question that we anticipate will be addressed is about the organization of genetic information into genomes. Issues surrounding DNA replication can explain many of the properties of bacterial genomes including the positioning of genes, the orientation of genes, nucleotide composition, and the locus-specific mutation rate. We wonder whether there are important rules of replication in higher eukaryotes that will have explanatory power for the locations of genes, their orientation and spacing, as well as the nucleotide composition and evolution rate that will be discovered.
Our expectation is that the field will also provide information on origin selection, origin firing, and origin dynamics in tumors. Correlating information about replication dynamics, 3-dimensional chromatin structure and the types of genetic events that drive tumorigenesis will allow us to better understand the molecular basis for this disease. For example, it will also be important to assess whether transcription factors or chromatin remodelers promote tumorigenesis through their roles in DNA replication in addition to their roles in the activation of cyclin-dependent kinases and transcription. From this perspective, it is especially interesting that mutations in the Rif1 protein that organizes S-phase progression in mice have been found in breast cancer patients.275,276 Many anticancer treatments involve drugs that interfere with DNA replication. A better understanding of the role of checkpoints and replication forks might help us to design better anticancer treatments.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received the following financial support for the research, authorship, and/or publication of this article: National Institute of General Medical Sciences (NIGMS) Center of Excellence grant P50 GM071508 (David Botstein, PI), National Institutes of Health (NIH) Oncology: Molecular Basis of Cancer grant 2T32 CA009538 (James Broach, PI), NIH/NIGMS grant 1R01 GM081686, and NIH/NIGMS grant 1R01 GM086465.
