Abstract
The cell cycle ensures genome maintenance by coordinating the processes of DNA replication and chromosome segregation. Of particular importance is the irreversible transition from the G1 phase of the cell cycle to S phase. This transition marks the switch from preparing chromosomes for replication (“origin licensing”) to active DNA synthesis (“origin firing”). Ubiquitin-mediated proteolysis is essential for restricting DNA replication to only once per cell cycle and is the major mechanism regulating the G1 to S phase transition. Although some changes in protein levels are attributable to regulated mRNA abundance, protein degradation elicits very rapid changes in protein abundance and is critical for the sharp and irreversible transition from one cell cycle stage to the next. Not surprisingly, regulation of the G1-to-S phase transition is perturbed in most cancer cells, and deregulation of key molecular events in G1 and S phase drives not only cell proliferation but also genome instability. In this review we focus on the mechanisms by which E3 ubiquitin ligases control the irreversible transition from G1 to S phase in mammalian cells.
E3 Ubiquitin Ligases Control Cell Cycle Transitions
Changes in protein abundance are critical for transitions in the cell division cycle. Cell cycle changes at both the protein and transcript levels have been analyzed globally and reveal that the proteome is even more dynamic than the transcriptome. 1 Such findings highlight the importance of ubiquitin-mediated protein degradation for properly-regulated cell cycle progression. Ubiquitination control is lost or perturbed in many cancers through the amplification or inactivation of key E3 ubiquitin ligases.2-5 These perturbations often impact the critical transition from G1 phase to S phase of the cell cycle. Replication licensing occurs during the G1 phase of the cell cycle at thousands of sites throughout the human genome known as origins of replication. These origins are the genomic loci where DNA replication is initiated during S phase. Replication licensing is accomplished by the assembly of prereplication complexes or “preRCs.” Licensing is complete once MCM complexes have been loaded onto origin DNA by ORC, Cdc6, and Cdt1 (Figure 1). Replication origins are first recognized and bound by the origin recognition complex (ORC) that remains associated with origins throughout the cell cycle. ORC is composed of constitutively expressed Orc2-6 and cell cycle regulated Orc1 (reviewed in DePamphilis 6 ). Cdc6 is then recruited to origins and interacts directly with ORC. Cdt1 interacts with the MCM complex in the nucleus during G1 through direct binding to the Mcm6 subunit.7-10 Cdt1 and MCM complexes are then recruited to origins through direct interaction of Cdt1 with both Cdc6 9 and Orc6. 11 Cdt1 is absolutely required for MCM recruitment but lacks any enzymatic activity. It is through the ATPase activity of ORC and Cdc6 that MCM helicases are finally loaded onto origin DNA.
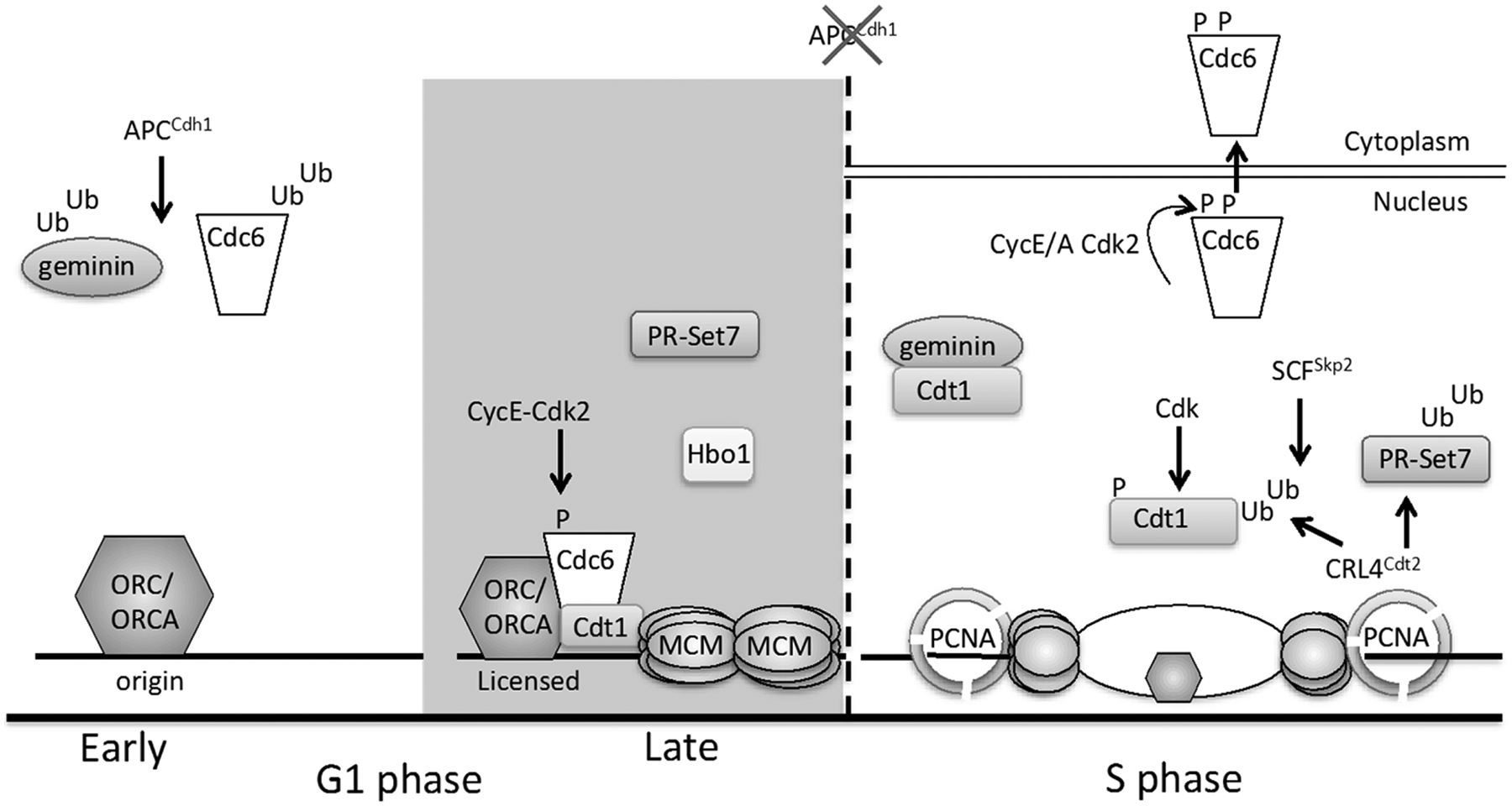
DNA replication origin licensing control in G1 and S phase. Origins are licensed by the DNA loading of MCM complexes in late G1. During early G1, MCM loading is blocked through the APCCdh1-mediated destruction of Cdc6. Cdc6 binding to APCCdh1 is blocked in late G1 by Cyclin E/Cdk2-mediated phosphorylation (Ser54), whereas the licensing inhibitor geminin is ubiquitinated by APCCdh1 throughout G1 phase. In late G1, MCM complexes are loaded at origins by the combined action of ORC/ORCA, Cdc6, and Cdt1 with contributions from the PR-Set7 and Hbo1 enzymes that may function through modification of histones at origins and/or nonhistone proteins. Once cells enter S phase, Cdc6 is phosphorylated at Ser106 phosphorylation, which promotes Cdc6 nuclear export. Three overlapping mechanisms suppress Cdt1 activity during S phase: (1) geminin accumulates and prevents Cdt1 from binding MCM. (2) Cdt1 is phosphorylated by Cdk2, and this phosphorylation induces Cdt1 ubiquitination by SCFSkp2. (3) Cdt1 and PR-Set7 both associate with DNA-loaded PCNA at replication forks, and PCNA interaction triggers ubiquitination by CRL4Cdt2.
Once MCM complexes have been loaded, the origins are licensed for replication, although MCM is not yet active as a DNA helicase in G1 because activation requires phosphorylation events mediated by S phase protein kinases. After MCM loading, however, ORC, Cdc6, and Cdt1 are no longer required for DNA synthesis in S phase.12,13 Origin licensing is tightly regulated to prevent MCM loading outside of G1 phase because new licensing during S and G2 phases can lead to rereplication that ultimately promotes genome instability. The consequences of rereplication include gene amplification and aneuploidy that drive oncogenesis. 14 Therefore, it is imperative that cells prevent rereplication. Origin licensing is restricted to G1 primarily through ubiquitin-mediated proteolysis of licensing components and cell cycle regulators.
Ubiquitin-mediated proteolysis by the 26S proteasome is initiated through polyubiquitination of substrate proteins. Polyubiquitination is accomplished through the activities of E1, E2, and E3 enzymes, and substrate specificity is determined by the E3 ubiquitin ligases (reviewed in Wilkinson
15
and Pickart
16
). The 3 most important E3 ubiquitin ligase complexes that regulate the G1/S transition in mammalian cells are the anaphase promoting complex/cyclosome (APC/C), Skp1-Cul1-F-box protein complex (SCF), and Cul4-RING-E3 Ligase 4 (CRL4) (Figure 2). These complexes are members of the largest single class of RING E3 ubiquitin ligases, the cullin-RING ligase (CRL) superfamily (reviewed in Jackson & Xiong
17
and Zimmerman
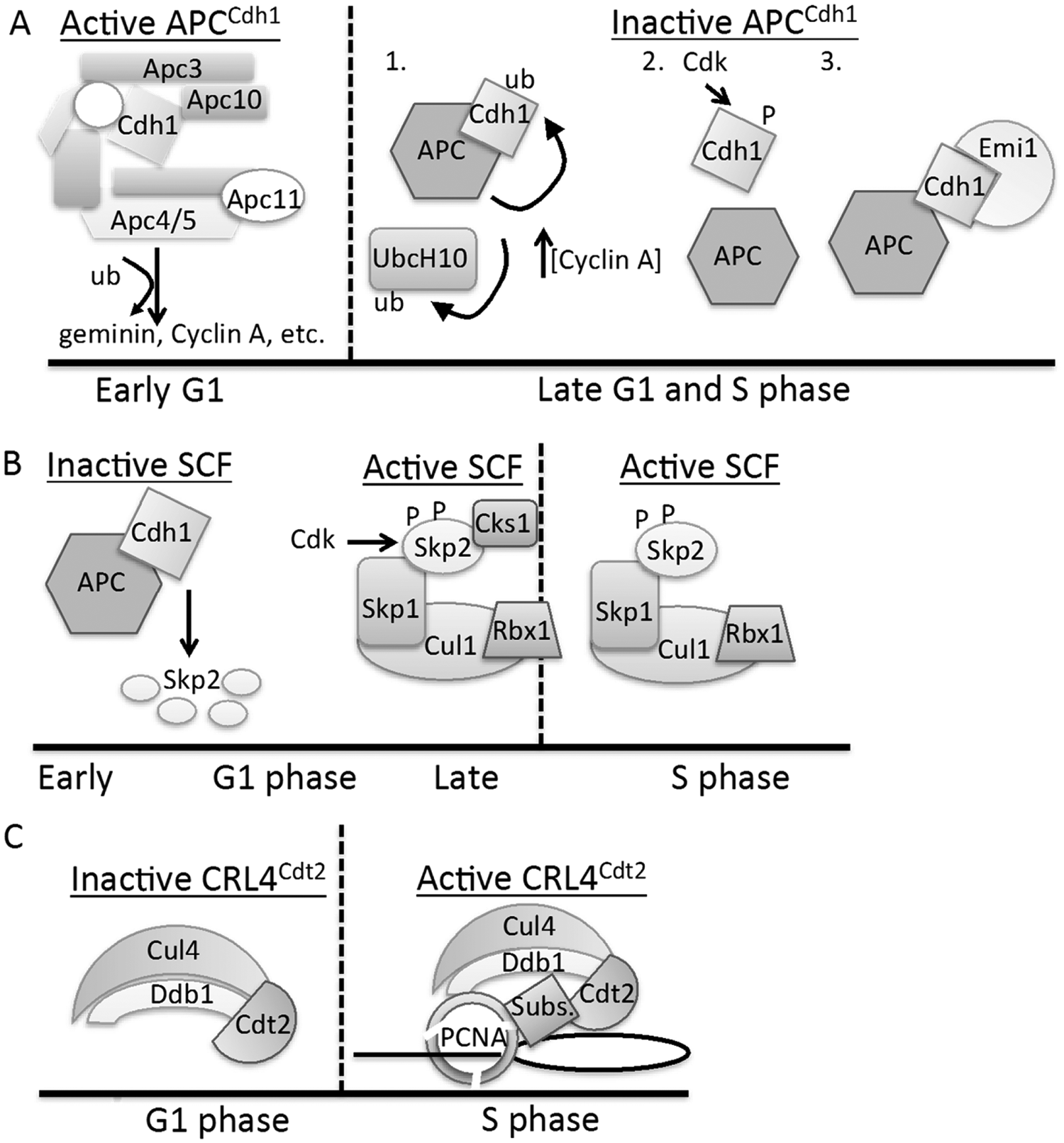
Three E3 ubiquitin ligases, APCCdh1, SCFSkp2, and CRL4Cdt2, at the G1/S transition.
The APC/C complex is active not only during the second half of mitosis (from anaphase on) but also during G1. APC/C contains the cullin protein Apc2, the Apc11 RING protein, either the Cdh1 (during G1) or Cdc20 (during G2/M) adaptor protein, and 11 other proteins whose exact functions remain unknown (reviewed in Nakayama & Nakayama
19
). The adaptor proteins mediate substrate interactions although some core APC/C proteins (Apc10 and Apc5) can also interact with substrates.20-23 Nearly all APC/C targets contain at least 1 peptide sequence known as a “destruction motif” or “degron” that mediates interaction with the adapter proteins.24-27 The 2 most common motifs are the destruction box (D-box) and the KEN-box, although there are several others that have been more recently identified (reviewed in McLean
Our focus here is on APCCdh1 since it functions primarily during telophase and the G1 phase of the cell cycle as opposed to APCCdc20 that functions in anaphase. Cdh1 protein levels remain constant throughout the cell cycle, but Cdh1 activity is regulated by phosphorylation.30-33 During early to mid-G1, Cdh1 is dephosphorylated and APCCdh1 is active, but as cells reach the G1/S transition, Cdh1 is phosphorylated by Cyclin-dependent kinases (CDKs) and this phosphorylation disrupts its interaction with the APC/C. Additionally, E2F transcription factor activity during late G1 induces accumulation of the APC/C pseudosubstrate, Emi1, which binds Cdh1 in S phase and thereby competitively inhibits APCCdh1 activity.34-36 Moreover, the two E2 enzymes that interact with the APC/C, UbcH10 and UbcH5, are degraded during S phase, and this downregulation further suppresses APC/C activity.37-39 Although loss of Cdh1 does not prevent cell cycle progression, it does result in a shorter G1 phase with impaired origin licensing followed by a prolonged and defective S phase with accumulation of DNA damage.40-42
In contrast to APC/C, the SCFSkp2 ubiquitin ligase complex becomes active later at the G1/S transition and remains active throughout S phase. The SCF complex is composed of only 4 subunits: Cul1 (cullin scaffold), Roc1 (RING-domain protein), Skp1 (adaptor protein), and one of many F-box protein substrate adaptors. Skp1 associates with many different F-box proteins that confer substrate specificity. During G1, APCCdh1 inhibits SCFSkp2 activity through ubiquitination and degradation of Skp2; this degradation is dependent on the N-terminal D-box of Skp2.43,44 Later in G1, however, Skp2 is phosphorylated at Ser64/72 by Cyclin E-Cdk2, and Skp2 continues to be phosphorylated, but by Cyclin A-Cdk2 in S phase. Phosphorylation at Ser64, and to a lesser extent at Ser72, prevents Skp2 ubiquitination by APCCdh1.43-45 Once active, SCFSkp2 promotes S phase progression and contributes to the inhibition of origin licensing during S phase (described below).
There are 69 F-box proteins encoded in the human genome, but substrates have been elucidated for fewer than 20%.
46
For substrates that have been identified, substrate phosphorylation is typically essential for direct interaction with the F-box protein and thus the SCF enzyme. Phosphorylation creates a negatively charged site on the substrate that docks with a positively charged surface on the F-box protein (e.g., Skp2).47,48 A notable exception to this requirement is p21, which does not require phosphorylation for interaction with Skp2. For efficient ubiquitination of the p21 and p27 CDK inhibitors, the SCFSkp2 complex instead requires the cofactor Cks1.49-51 Cks1 performs a docking function to facilitate the interaction between Cyclin-CDKs and their substrates. In addition, Cks1 promotes interactions between Cdk2-bound, phosphorylated CKIs and Skp2.
47
There are two CKS (Cyclin-dependent kinase subunit) paralogs in mammals, Cks1 and Cks2, which tightly associate with Cyclin-CDK complexes, compete for Skp2 binding, and are essential for cell viability.52,53 Only Cks1 can promote substrate interaction with Skp2.
52
It is clear that SCFSkp2 requires Cks1 association for a subset of its substrates, but the p21 and p27 CDK inhibitors are the only ones identified in this group thus far; others may yet be discovered (reviewed in Krishnan
The most recently characterized E3 ubiquitin ligase required for proper cell cycle progression is CRL4Cdt2. This complex is composed of Cul4, Roc1/2, an adaptor protein DDB1, and a substrate receptor or DCAF (DDB1- and Cul4-associated factor) that interacts directly with DDB1. There are at least 20 known DCAFs encoded in the human genome (reviewed in Havens & Walter
55
). We focus primarily on the CRL4Cdt2 complex in this review because it has the highest relevance for the G1/S transition. The DCAF Cdt2 (Cdc10-dependent transcript 2) is unique among DCAFs in that its interaction with substrates is dependent on the substrate first interacting with DNA-loaded PCNA. PCNA is a processivity factor for DNA polymerase δ that is loaded at replication forks during S phase and during the DNA synthesis steps of DNA repair. Proteins that interact with PCNA contain a motif known as a PCNA-interacting protein box (PIP box). CRL4Cdt2 substrates contain a specialized PIP box known as the PIP degron. The PIP degron differs from the canonical PIP box in 2 ways. First, a TD motif is present within the PIP box that strengthens the interaction with PCNA.56-59 Second, the PIP degron contains a conserved basic residue 4 amino acids downstream from the PIP box that is required for interaction with Cdt2.56,59,60 All
DNA Replication Origin Licensing Regulation before the G1/S Transition
Geminin and Cdt1
During S phase and G2, Cdt1 is bound by its inhibitor, geminin.62,63 Structural studies of mouse geminin and Cdt1 found that a dimer of geminin binds to Cdt1. 64 The human geminin-Cdt1 complex was also crystallized, and interestingly this study identified not only the trimeric form but also that the complex could dimerize to form a heterohexamer. Further analysis suggested that the trimeric geminin/Cdt1 complex may be permissive for origin licensing, whereas the heterohexameric complex is inhibitory for origin licensing due to the burial of Cdt1 residues required for licensing.65,66 More recently, Mcm9 was shown to interact with Cdt1, and this interaction prevents formation of the inhibitory geminin/Cdt1 heterohexamer to promote origin licensing. 67 The molecular mechanism controlling the apparent switch between licensing-competent and licensing-incompetent Cdt1 is still not fully understood (reviewed in Caillat & Perrakis 68 ). During G1, APCCdh1 targets geminin for degradation, allowing Cdt1 to recruit MCM complexes to origins. Geminin interacts with the Cdh1 adaptor protein through a D-box at its N-terminus. 69 Overexpression of a D-box mutant, nondegradable form of geminin in primary fibroblasts blocks chromatin-association of MCM complexes and inhibits origin licensing. 70 Thus, low levels of geminin are required for efficient licensing during late G1.
In addition to geminin degradation, another Cdt1 regulatory mechanism in G1 has been suggested. Cdt1 was shown to interact with APCCdh1 in a proteomics study, and
Cdc6
APCCdh1 targets not only geminin but also Cdc6 for degradation during early G1.
74
Cdh1 binding to Cdc6 is dependent on both a KEN-box and a D-box found at the N-terminus of Cdc6. APCCdh1 prevents Cdc6 accumulation until late in G1, when Cdc6 is phosphorylated at Ser54 by Cyclin E-Cdk2. This phosphorylation protects Cdc6 from APCCdh1-mediated degradation by disrupting the interaction between Cdc6 and Cdh1.
75
Additionally, APCCdh1 activity is inhibited late in G1 (as previously described) to allow cell cycle progression into S phase. Importantly, Cdc6 is stabilized, but geminin is not. Thus, in late G1, ORC is already bound to origins, Cdt1 is actively recruiting MCM complexes to origins, and now that Cdc6 is present, MCM loading can be completed. Cdc6, Cdt1, ORC, and MCM complexes are the essential replication licensing proteins and, together with nucleoplasmin, are fully sufficient for licensing
PR-Set7 (Set8)
The requirement for nucleoplasmin in the reconstituted licensing reaction with purified proteins has been attributed to a need to decondense chromatin. This notion underscores the importance of chromatin structure near origins in controlling the efficiency of both origin licensing in G1 and origin firing in S phase. Recently, the methyltransferase PR-Set7 (also known as Set8) has emerged as a new critical player in origin licensing. PR-Set7 is the sole enzyme responsible for monomethylation of histone H4 at lysine 20 (H4K20me1), and both PR-Set7 and H4K20me1 are tightly cell cycle- regulated. Importantly, H4K20me1 is enriched at origins of replication in
As with many origin licensing factors, PR-Set7 abundance is regulated both transcriptionally and posttranslationally.
The absence of the PR-Set7 methyltransferase can only result in corresponding changes in histone methylation if histone H4K20 monomethylation is simultaneously converted either to unmethylated K20 or to di- or trimethylated H4K20 (or both). H4K20me1 could promote origin licensing in G1 by serving as the template for di- and trimethylation by both Suv4-20h1 and 2 methyltransferases.85-87 Interestingly, the ORCA protein can specifically bind to trimethylated H4K20.73,88 The decrease in H4K20me1 during S phase could thus be due in part to the transition to these higher methylation states, although these marks have not yet been specifically examined at replication origins. Loss of both Suv4-20h1 and Suv4-20h2 causes defects in S phase entry, but the effects are less severe than loss of PR-Set7, suggesting additional roles for PR-Set7 in the G1/S transition.78,86 One such role could be the methylation of an as yet unidentified nonhistone protein required for progression from G1 to S phase. Currently, the only nonhistone substrate of PR-Set7 that has been identified is p53. 89 However, it is unlikely that p53 methylation is fully responsible for the replication defects caused by PR-Set7 depletion because these phenotypes are not entirely p53 dependent, and p53 null cells are viable.77,90,91 The mechanism by which PR-Set7 promotes cell cycle progression is therefore still unclear.
E2F and the mixed-lineage leukemia methyltransferase
The proteins responsible for origin licensing are clearly regulated by ubiquitin-mediated proteolysis, but they are also the products of transcriptionally regulated genes. The transcription factors controlling these genes are themselves also regulated by ubiquitin-mediated proteolysis. The most important class of transcription factors involved in progression from G1 to S phase is the E2F family. E2F proteins play key roles in cell cycle progression. They are often categorized as activator (E2F1-3) or repressor (E2F4-8) E2Fs, although members of either group can both activate and repress gene transcription depending on the circumstances (reviewed in Wong
Tight control over E2F activity is required to prevent aberrant DNA replication. Specifically, overexpression of
Recent studies have described an MLL-E2F axis that promotes transcription of E2F target genes, particularly
Cyclin-CDK Regulation before the G1/S Transition
Cyclin D
Cyclin D1 is the first cyclin expressed during G1 in response to mitogenic signals and peaks immediately before the G1-S transition. It interacts with both Cdk4 and Cdk6 to form active Cyclin-CDK complexes that phosphorylate Rb to relieve repression of E2F-dependent genes, including
Cyclin E
CKIs: p21 and p27
High CDK activity is required for progression from G1 to S phase, and one critical CDK-mediated event is the recruitment of MCM activating subunits to stimulate origin unwinding. Destruction of the CKIs p21 and p27 prior to the G1/S transition is required for full CDK activity (Figure 3). P21 and p27 are regulated posttranslationally, in part, through ubiquitination by the SCFSkp2 E3 ubiquitin ligase complex. This SCFSkp2-mediated ubiquitination requires that p21 and p27 be in a complex with Cyclin-CDK.125,126 Phosphorylation of p27 by Lyn or BCR/ABL kinases at Tyr88 ejects the inhibitory domain of p27 from the Cdk2 active site.127,128 The now-active Cyclin E-Cdk2 can then phosphorylate p27 at Thr187 to stimulate its interaction with the SCFSkp2-Cks1 E3 ubiquitin ligase.47,126 Cyclin E-Cdk2 also phosphorylates p21 at Ser130 to stimulate its interaction with SCFSkp2-Cks1.51,129 Although p21 and p27 both require association with Cks1 and Cyclin A/E-Cdk2 for proper SCFSkp2-mediated degradation, they differ in that phosphorylation of p21 is not strictly required for proteolytic degradation as it is for p27. In fact, Ser130 phosphorylation of p21 by mitogen-activated protein kinases (MAPKs) has been shown to stabilize p21. 130 Although it has not been tested, it is possible that Ser130 phosphorylation strengthens the interaction between p21 and Cyclin A/E-Cdk2 that is absolutely required for proper degradation of p21 by SCFSkp2-Cks1.
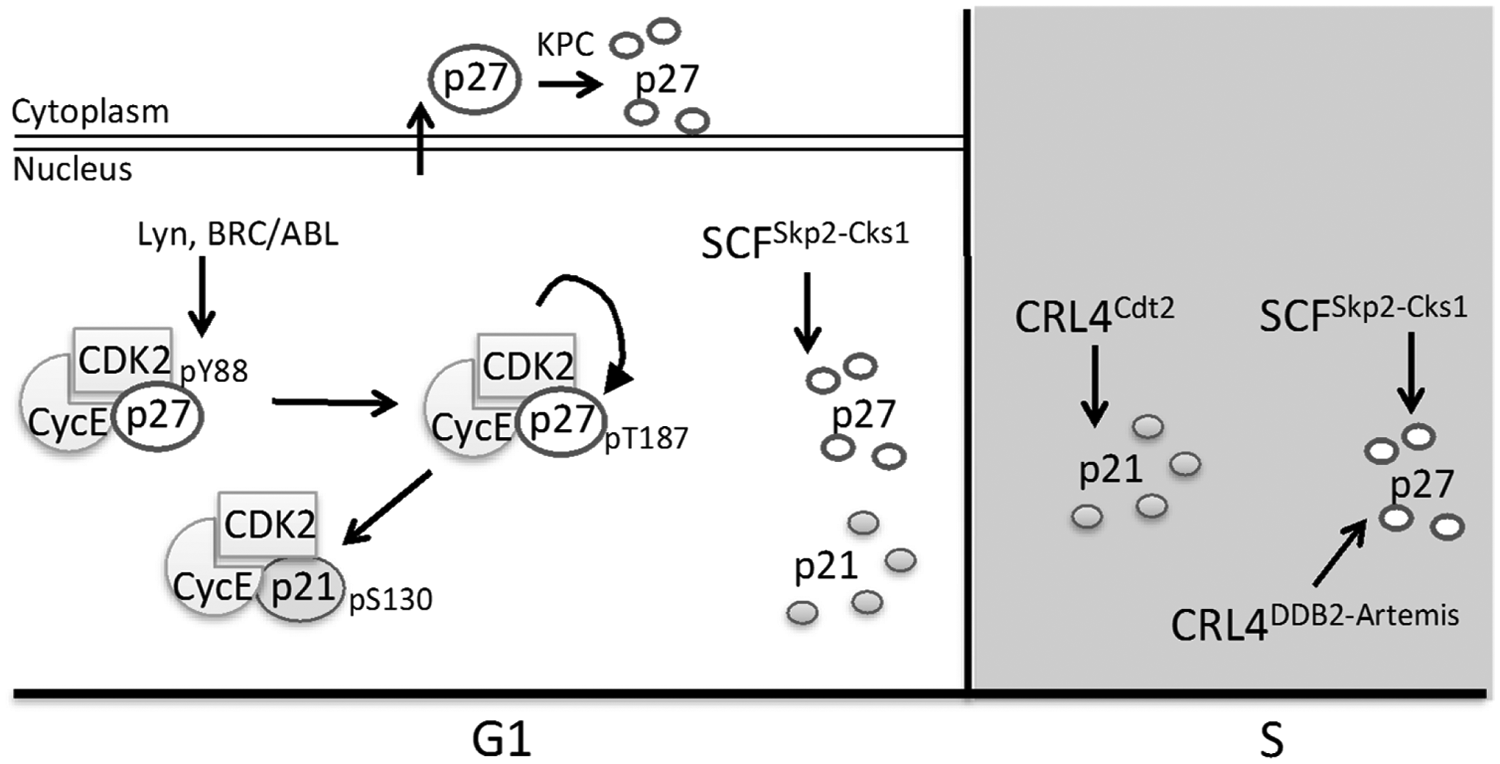
CDK inhibitor control in G1 and S phase. During G1, CDK activity is lowest due to high levels of p21 and p27. Lyn and BRC/ABL kinases phosphorylate p27 in late G1, relieving its inhibition of Cyclin E-Cdk2 without disrupting CDK association. The active Cyclin E-Cdk2 then phosphorylates both p27 and p21, promoting their interaction with and ubiquitination by SCFSkp2-Cks1. High CDK activity is maintained during S phase through ubiquitination of p27 and p21 by SCFSkp2-Cks1, p27 ubiquitination by CRL4DDB2-Artemis, and p21 ubiquitination by CRL4Cdt2.
As previously mentioned, Cks1 is required for degradation of both p21 and p27 by SCFSkp2. Cks1 promotes the stable interaction between Skp2 and phosphorylated p27.49,50,131 Cks1 also promotes the phosphorylation-independent interaction between p21 and Skp2.
51
Knockout of
In addition to changes in stability, free p27 (not bound to Cyclin-Cdk2) is exported from the nucleus during G1 through interaction with Cyclin D2-Cdk4/ 6. Additionally, p27 phosphorylation by PI3K/Akt and Ras signaling pathways (induced by growth factors) promotes its cytoplasmic relocalization.133-135 Cytoplasmic p27 can be targeted by the Kip1 ubiquitylation-promoting complex (KPC).136-138 Although KPC inactivation has little effect on cell cycle progression, when combined with Skp2 downregulation cells cannot enter S phase from quiescence. 136
Summary of G1 events
By the end of G1, origins have been licensed and are ready for replication initiation during S phase. Once MCM complexes have been loaded at origins, the MCM loading factors Cdc6, Cdt1, and ORC are no longer required for cell cycle progression. APCCdh1 is inactive and has allowed Skp2 to accumulate and interact with the SCF E3 ubiquitin ligase. Another consequence of inactive APCCdh1 is accumulation of geminin that prevents additional MCM loading at S phase onset through direct Cdt1 inhibition. Finally, Cyclin E-Cdk2 is active and promotes the transition into S phase. At this point, the business of G1 phase is complete and cells are poised to initiate DNA replication through phosphorylation-mediated recruitment and activation of MCM helicase activators, DNA polymerases (α, δ), and processivity factors (PCNA, RFC, etc.).
DNA Replication Origin Licensing Regulation after the G1/S Transition
As cells enter S phase, it is imperative that origin licensing is inhibited. Once a given origin has fired and bidirectional replication forks have already been established, relicensing of that same origin could be followed by a refiring or “rereplication” event. 139 Rereplication is an aberrant phenomenon (unlike endoreduplication), and rereplication ultimately results in double-strand DNA breaks.140,141 Most recently, rereplication has also been established as a direct cause of gene amplification.14,142 The induction of both DNA damage and gene amplification drives genome instability and may be one underlying cause of oncogenesis. In normal cells, however, several overlapping mechanisms suppress rereplication, and these are discussed below.
Geminin and Cdt1
Arguably the most important mechanism to prevent rereplication during S phase is the simultaneous degradation and inhibition of Cdt1. Cdt1 is a substrate of both CRL4Cdt2 and SCFSkp2 during S phase. 143 CRL4Cdt2 targets Cdt1 for degradation only when Cdt1 is bound to PCNA loaded onto DNA at replication forks. Since PCNA is DNA loaded both in S phase and after DNA damage, Cdt1 ubiquitination by CRL4Cdt2 can be induced outside of S phase by exogenous DNA damage.144-147 Cdt1 contains a well-conserved PIP degron at its N-terminus through which it interacts with both DNA-loaded PCNA and Cdt2.59,148-150 Overproduction of Cdt1 or expression of Cdt1 lacking a PIP degron promotes rereplication in many experimental systems.148,149,151-155 Removal of Cdt1 from chromatin after ubiquitination by CRL4Cdt2 is partially dependent on the activity of the p97 ATPase and its cofactor UFD1 (which possesses a ubiquitin-binding domain). Depletion of p97 in HeLa cells resulted in stabilization of Cdt1 in S phase and after DNA damage. 156
During S phase, SCFSkp2 can also ubiquitinate Cdt1 to promote its degradation. Cdt1 interaction with Skp2 is dependent upon a Cy-motif that mediates interaction with and phosphorylation by Cyclin E/A-Cdk2. 143 Cdk2 can phosphorylate human Cdt1 at Thr29 to stimulate ubiquitination by SCFSkp2. 153 Interestingly, the same site was recently shown to be phosphorylated by JNK after cellular stress, although JNK-mediated phosphorylation had no effect on Skp2 binding or Cdt1 stability. 157 It is not yet clear how Skp2 binding and Cdt1 degradation were blocked when Thr29 was phosphorylated by JNK, but several other JNK-mediated phosphorylation sites in Cdt1 that block targeting by CRL4Cdt2 likely have a much stronger effect on overall Cdt1 stability than the T29 site. 158 For instance, five phosphorylation sites in the C-terminal one third of human Cdt1 were recently mapped that confer resistance to CRL4Cdt2 despite being quite distant from the PCNA and Cdt2 binding sites at the N-terminus of Cdt1. 158 Both the JNK and p38 stress MAP kinases phosphorylate these amino acids not only in response to exogenous stress but also during G2 and M phases. Stabilization of Cdt1 in G2 sets the stage for Cdt1’s second cell cycle function in establishing robust microtubule-kinetochore attachments. 159
Degradation of Cdt1 can be prevented not only by MAPK-mediated phosphorylation but also by acetylation at lysines 24 and 29 by p300 and PCAF in asynchronously dividing 293T cells. 160 These acetylations were detected during early G1. During S phase, however, HDAC11 associated with and deacetylated Cdt1, promoting Cdt1 degradation. 160 It is unclear which E3 ubiquitin ligase Cdt1 is protected from by acetylation, but the proximity of Lys24 to Thr29 suggests that it could interfere with Skp2 binding, although this has not yet been tested, and the PIP degron that directs CRL4Cdt2 binding is also relatively close to Thr29 (amino acids 3-14). Further work is needed to fully understand the importance of Cdt1 acetylation/deacetylation during G1 and S phases.
Finally, Cdt1 inhibition is accomplished through the reaccumulation of geminin after APCCdh1 inactivation as cells transition from G1 to S phase and APC/C is inactivated. Geminin prevents the association of Cdt1 with the MCM complex and with Cdc6.9,10 Additionally, geminin accumulation disrupts the ORCA-Cdt1 interaction at origins. 161 Like Cdt1, ORCA is degraded during S phase, so its downregulation may also contribute to origin licensing inhibition. ORCA interacts with CRL4 and is polyubiquitinated during S phase; inhibiting CRL4 and DDB1 had no effect on ORCA stability, however, suggesting that multiple E3s could target ORCA for degradation. 161
Cdc6
Although many mechanisms are in place to prevent aberrant Cdt1 accumulation during S phase, there are also safeguards to inhibit other origin licensing factors. As cells exit G1, APCCdh1 is inactivated and no longer targets Cdc6 for proteasomal degradation (see above). As a consequence, Cdc6 protein levels are paradoxically much higher in S phase and G2 when licensing is blocked than during G1 when licensing should occur. Nevertheless, during S phase, Cdc6 is exported from the nucleus through a 2-step mechanism. First, the acetyltransferase Gcn5 acetylates Cdc6 at 3 lysine residues (92, 105, and 109). 162 These acetylation events are required for subsequent Cdc6 phosphorylation at Ser106 by Cyclin A-Cdk2, which is active because the p21 and p27 inhibitors are degraded in S phase (see below). Ser106 phosphorylation promotes CRM1-dependent nuclear export of Cdc6, and, of course, cytoplasmic Cdc6 cannot participate in origin licensing.163,164 Cdc6 persists in the cytoplasm and is eventually degraded by APCCdh1 in the next G1 phase.
PR-Set7
PR-Set7 is required for proper S phase progression and origin firing. Similar to Cdt1, PR-Set7 levels sharply decrease during S phase. 84 Treatment with proteasome inhibitors to block degradation revealed that PR-Set7 associates with PCNA at replication foci. 90 Moreover, PR-Set7 has recently been shown to be a target of the CRL4Cdt2 ubiquitin ligase that relies on PCNA as a cofactor for substrate recognition.82,165,166 PR-Set7 has 2 PIP boxes, but only one functions as the PIP degron.55,82,165 Just as with Cdt1, removal of polyubiquitinated PR-Set7 from chromatin is dependent upon p97/UFD1. 156 Inhibition of CRL4Cdt2 or deletion of the PR-Set7 PIP degron results in aberrant PR-Set7 accumulation during S phase and rereplication that is dependent on the catalytic activity of PR-Set7.79,165
E2F and the mixed-lineage leukemia methyltransferase
As cells transition from G1 to S phase, both E2F1 and MLL are targeted for degradation by SCFSkp2 through Skp2 binding to their N-termini.111,167 Degradation of E2F1 is presumed necessary since prolonged E2F1 activity leads to induction of apoptosis.168-170 Additionally, E2F1 degradation prevents aberrant expression of
Cyclin-CDK Regulation after the G1/S Transition
Cyclin D1
As cells reach the G1/S transition, Cyclin D1-Cdk4/6 is inactivated through proteasome-mediated degradation of Cyclin D1. This degradation is necessary for proper S phase progression in part because overexpressed Cyclin D1 can bind PCNA and prevent DNA replication.172,173 Phosphorylation of Cyclin D1 at Thr286 is required for its nuclear export and subsequent cytoplasmic ubiquitination and proteolysis; mutation of Thr286 to alanine therefore results in nuclear accumulation of Cyclin D1.174,175 ATM and ATR kinases are responsible for Cyclin D1 Thr286 phosphorylation during S phase.
176
Interestingly, nuclear accumulation of active Cyclin D1/Cdk4 indirectly promotes Cdt1 stabilization in S phase through transcriptional repression of
The SCF E3 ubiquitin ligase has been implicated in targeting phosphorylated Cyclin D1 for proteolysis at the G1/S transition. Studies conducted in various human cancer cell lines identified multiple F-box proteins that promote Cyclin D1 ubiquitination by the SCF complex including Fbx4,178,179 Fbxw8,
180
and Fbxo31.
181
Downregulation of Fbx4 promotes the accumulation of phosphorylated Cyclin D1 in the nucleus, and this accumulation facilitates oncogenic transformation.
178
Tumors isolated from
Cyclin E
As cells progress into late S phase, Cyclin E is targeted for degradation by 2 mechanisms. The first is through SCF-mediated ubiquitination and the second is through CRL4- mediated ubiquitination. Cyclin E- Cdk2 auto-phosphorylates at Thr62 and Thr384, and these phosphorylations promote interaction with the SCF adaptor protein Fbxw7, resulting in Cyclin E ubiquitination and degradation.186-188 Phosphorylation at these 2 residues is maximally detected during S phase and mitosis, respectively.
189
Mutation of the homologous residues to alanines in a knock-in mouse model resulted in aberrant proliferation and aneuploidy, presumably due to perturbations from inappropriate Cyclin E-Cdk2 activity during late S phase and G2.
190
The second mechanism for Cyclin E degradation is through ubiquitination by CRL42. Knockdown of
CKIs: p21 and p27
In addition to SCFSkp2-Cks1 activity, CRL4Cdt2 also promotes destruction of p21 during S phase.192-197 As for the Cdt1 and PR-Set7 targets of CRL4Cdt2, ubiquitination of p21 in S phase requires interaction with DNA-loaded PCNA. P21 contains a PIP degron near its C-terminus through which it interacts with both PCNA and Cdt2 (reviewed in Abbas & Dutta
198
). Interestingly, like Cdt1, p21 is also a stress MAP kinase substrate,
130
and MAP kinase activation blocks the replication-dependent degradation of human p21
158
by an as-yet unknown mechanism. In contrast to p21, p27 does not contain a PIP degron or even a minimal PIP-box and therefore cannot interact with CRL4Cdt2. SCFSkp2 constitutes the main p27 degradation mechanism during S phase. P27 is, however, still degraded in a CRL4-dependent manner as evidenced by its accumulation after knockdown of DDB1 or Cul4 and reduced p27 levels upon overexpression of
Summary of S-phase events
The regulatory mechanisms discussed above are all in place to prevent inappropriate origin licensing and subsequent rereplication during the S and G2 cell cycle phases. These regulatory mechanisms are disrupted in many human cancers, and in many cases those disruptions not only drive tumor initiation but also likely contribute to the progression to metastatic disease and/or the development of chemotherapy resistance. After the G1/S transition, cells inhibit origin licensing by exporting Cdc6 from the nucleus, inhibiting and degrading Cdt1, and degrading PR-Set7. Simultaneously, origins initiate DNA synthesis as a consequence of high Cyclin-Cdk2 activity. Amplification or overexpression of genes encoding D-type and E-type Cyclins are frequent events in human cancers, and the high CDK activity from these genetic perturbations not only inactivates Rb but also disrupts the normal fluctuations in CDK activity that maintain replication precision and genome stability. It is clear that protein degradation plays an integral role in regulating the G1 to S transition and preventing rereplication.
Transformed cells show a higher propensity to undergo rereplication than normal cells do. For example, HeLa cells undergo low and constitutive rereplication even during unperturbed cell cycles, 203 and many tumor-derived cell lines can be readily induced to rereplicate whereas normal cells are relatively resistant to such induction.141,151,204,205 Replication licensing proteins are overexpressed in a number of human cancers, in part due to deregulation of the Rb-E2F pathway and higher expression of genes encoding replication licensing proteins.206-208 Even very modest overproduction of the origin licensing factors Cdt1 or Cd6 promotes tumorigenesis in cultured human and mouse cells as well as in mouse xenograft models, presumably due at least in part to low levels of rereplication and consequent genome instability.151,209-214 Unregulated expression of replication licensing factors is one potential reason for higher rereplication in cancer cell lines,206-208 but other mechanisms such as oncogene signaling (e.g., Ras) may also contribute to this difference. 215 In addition to promoting oncogenesis, aberrant regulation of replication licensing also contributes to gene amplification through increased genome instability. Gene amplification promotes resistance of cancer cells to therapeutic agents, increasing the likelihood of developing second cancers.14,142
The Ubiquitin-Proteasome System as a Target for Anticancer Therapies
Given the previously discussed importance of ubiquitin-mediated proteolysis in cell cycle regulation, the ubiquitin-proteasome system (UPS) and the specific proteins involved are promising targets of anticancer therapies. These regulatory pathways are frequently mutated in diverse cancer types. Many drugs have been developed to target the UPS, and several are currently being used in the clinic to successfully treat many different cancers. Initial efforts to develop proteasome inhibitors for cancer therapies were met with skepticism as these drugs were predicted to be highly toxic to normal cells. Surprisingly, cancer cells are particularly sensitive to proteasomal inhibition.216,217 The mechanism for this sensitivity is not fully understood, but in the context of this discussion, cancer cells already suffering from perturbed cell cycle control may not tolerate additional deregulation and still maintain chromosome integrity, whereas normal cells are better able to tolerate such pressure.
The boronic-acid derivative bortezomib was the first proteasome inhibitor designed for cancer treatment. 218 It specifically and reversibly inhibits the active sites in the 26S proteasome. Proteasome inhibition promotes the upregulation of p21 and p27 and suppression of proinflammatory response genes resulting in apoptosis in tumor cells.219,220 Combination therapies with bortezomib have increased cancer cell sensitivity to other cancer drugs and led to the development of bortezomib derivatives such as carfilzomib.216,219 For example, a combination of melphalan, a DNA damaging agent, and bortezomib was successful in treating multiple myeloma patients. 221 Carfilzomib as well as bortezomib is now being used effectively to treat multiple myeloma and relapsed mantle cell lymphoma.222-225 These proteasome inhibitors sensitize cancer cells to other chemotherapeutic agents by abrogating the DNA damage response, upregulating the CKIs p21 and p27, and promoting apoptosis.219,226
The most recent effort to design a novel proteasome inhibitor specifically targets the CRL ubiquitin ligases. All members of the CRL ubiquitin ligase family require the addition of NEDD8, a 9-kD ubiquitin-like protein, for full activity. Neddylation of the cullin proteins produces a conformational change that enhances interaction with the activated E2 enzyme as well as ubiquitin transfer and may prevent association with the negative regulator CAND1.227,228 MLN4924 is the first drug that specifically inhibits the NEDD8-activating enzyme (NAE) to prevent activation of CRL E3 ubiquitin ligases. In the first proof-of-principle study, MLN4924 treatment of many different cancer cell types induced apoptosis and prevented tumor progression in mouse xenograft models. 229 This result was later attributed to high levels of rereplication that could be partially suppressed by Cdt1 knockdown, indicating that MLN4924 treatment promoted stabilization of Cdt1 through inhibition of CRL4Cdt2 activation. 230 Additionally, MLN4924 treatment was found to increase tumor sensitivity to ionizing radiation in both pancreatic and breast cancers.231,232 Several Phase I clinical trials are currently underway.
Footnotes
Acknowledgements
The authors thank members of the Cook laboratory for helpful comments on the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
