Abstract
In addition to genetic disorders, epigenetic alterations have been shown to be involved in cancer, through misregulation of histone modifications. Miswriting, misreading, and mis-erasing of histone acetylation as well as methylation marks can be actually associated with oncogenesis and tumor proliferation. Historically, methylation of Arg and Lys residues has been considered a stable, irreversible process due to the slow turnover of methyl groups in chromatin. The discovery in recent years of a large number of histone Lys demethylases (KDMs, belonging to either the amino oxidase or the JmjC family) totally changed this point of view and suggested a new role for dynamic histone methylation in biological processes. Since overexpression, alteration, or mutation of a number of KDMs has been found in many types of cancers, such enzymes could represent diagnostic tools as well as epigenetic targets to modulate for obtaining novel therapeutic weapons against cancer. The first little steps in this direction are described here.
In addition to genetic alterations, it is now well established that epigenetic dysfunctions can be involved in the pathogenesis and development of cancer. Epigenetic regulation of gene expression involves two major mechanisms, DNA methylation at the CpG islands of gene promoters and chromatin remodeling. Chromatin folding and its switch from heterochromatin, transcriptionally silent, to euchromatin, transcriptionally active, and vice versa, is finely tuned by three categories of proteins: (1) enzymes that add chemical marks to histone substrates, the so-called writers such as histone acetyltransferases (HATs) and histone methyltransferases (further distinct into protein arginine methyltransferases [PRMTs] and histone lysine methyltransferases [HKMTs]); (2) enzymes that remove such chemical covalent modification from histone substrates—namely, “erasers” such as histone deacetylases (HDACs) and histone lysine demethylases (KDMs, the lysine-specific demethylases [LSDs], and the Jumonji C [JmjC] families; Fig. 1); and (3) proteins that recognize and react to specific modified histone residues at epigenetic code level, the “readers.”1-4
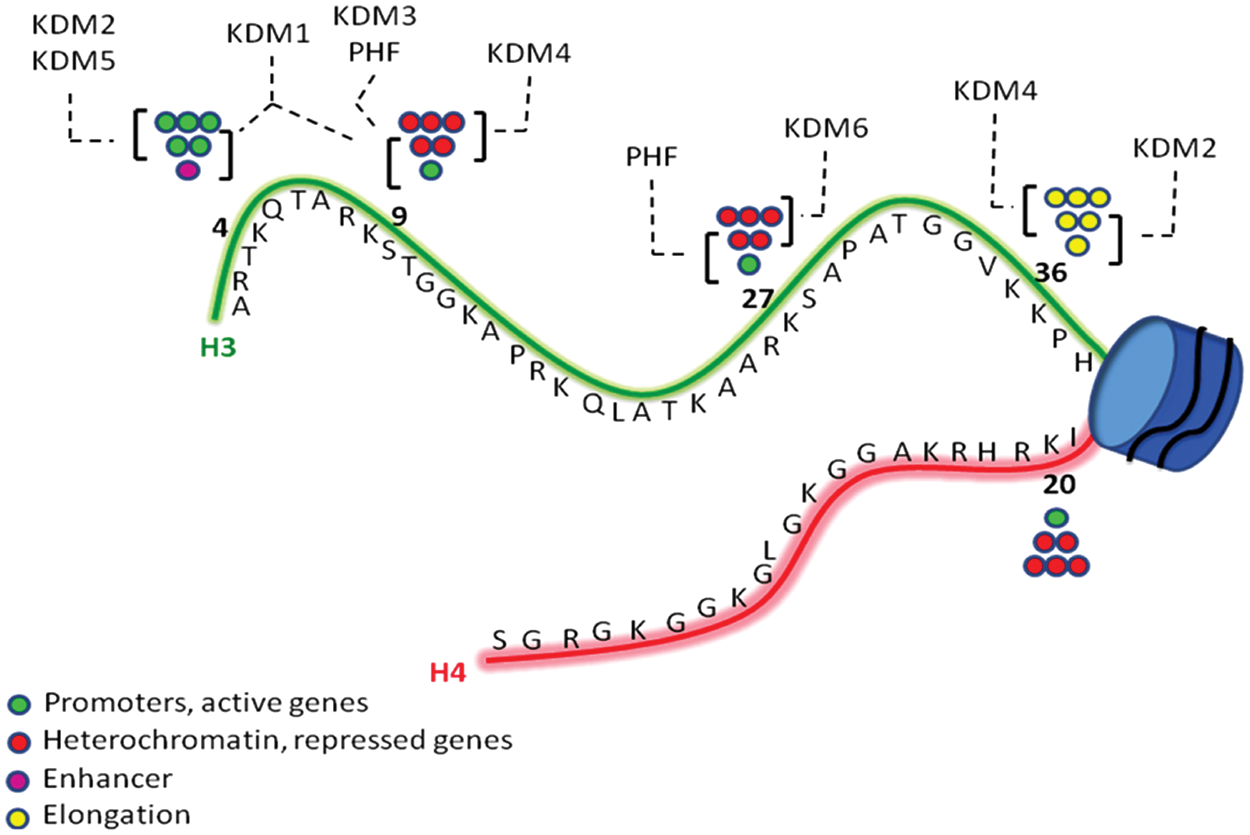
Schematic representation of H3 and H4 histone tails, with the most important Lys residues involved in mono-, di-, and trimethylation (circles). The most frequent KDM families involved in demethylation of specific methyl markers are shown.
Aberrant expression of writer or eraser enzymes has been implicated in the course of tumor initiation and progression, and indeed 2 HDAC inhibitors, vorinostat (suberoylanilide hydroxamic acid, SAHA) and romidepsin (depsipeptide, FK-228), have been approved by Food and Drug Administration (FDA) for the treatment of refractory cutaneous T cell lymphoma and actually are in clinical trials, alone or in combination, for a variety of both hematological and solid tumors. 5
KDMs are the most recently discovered families of erasers. Methylation of lysine residues at histone substrates was long retained an irreversible reaction. In 2004, the first protein showing mammalian KDM activity was reported to be a flavin-containing amino oxidase (AO) that specifically demethylated mono- or dimethylated lysine 4 at histone H3 (H3K4me1 and H3K4me2) and has been named LSD1 (lysine-specific demethylase 1)/KDM1A (Fig. 2A).6,7 More recently, a second flavin-dependent H3K4me1/2 demethylase called LSD2/AOF1/KDM1B has been identified in mammals. 8 In 2006, a larger and more versatile family of KDMs structurally different from LSD1 was identified, all containing as a signature motif the JmjC domain and all working through a Fe2+/2-oxoglurate (2-OG) mechanism for catalysis (Fig. 2B).9,10 The reaction goes on generating a superoxide radical by the complex Fe2+/2-OG, which hits the C2 atom of 2-OG, leading to its decarboxylation to succinate and formation of a Fe4+-oxo species. Afterwards, the Fe4+-oxo species abstracts a hydrogen from the N-methyl group of the amine substrate while it is reduced to Fe3+. The last two steps involve the generation of a carbinolamine intermediate on the substrate, which leaves formaldehyde and the amine with one methyl group less (Fig. 2B).
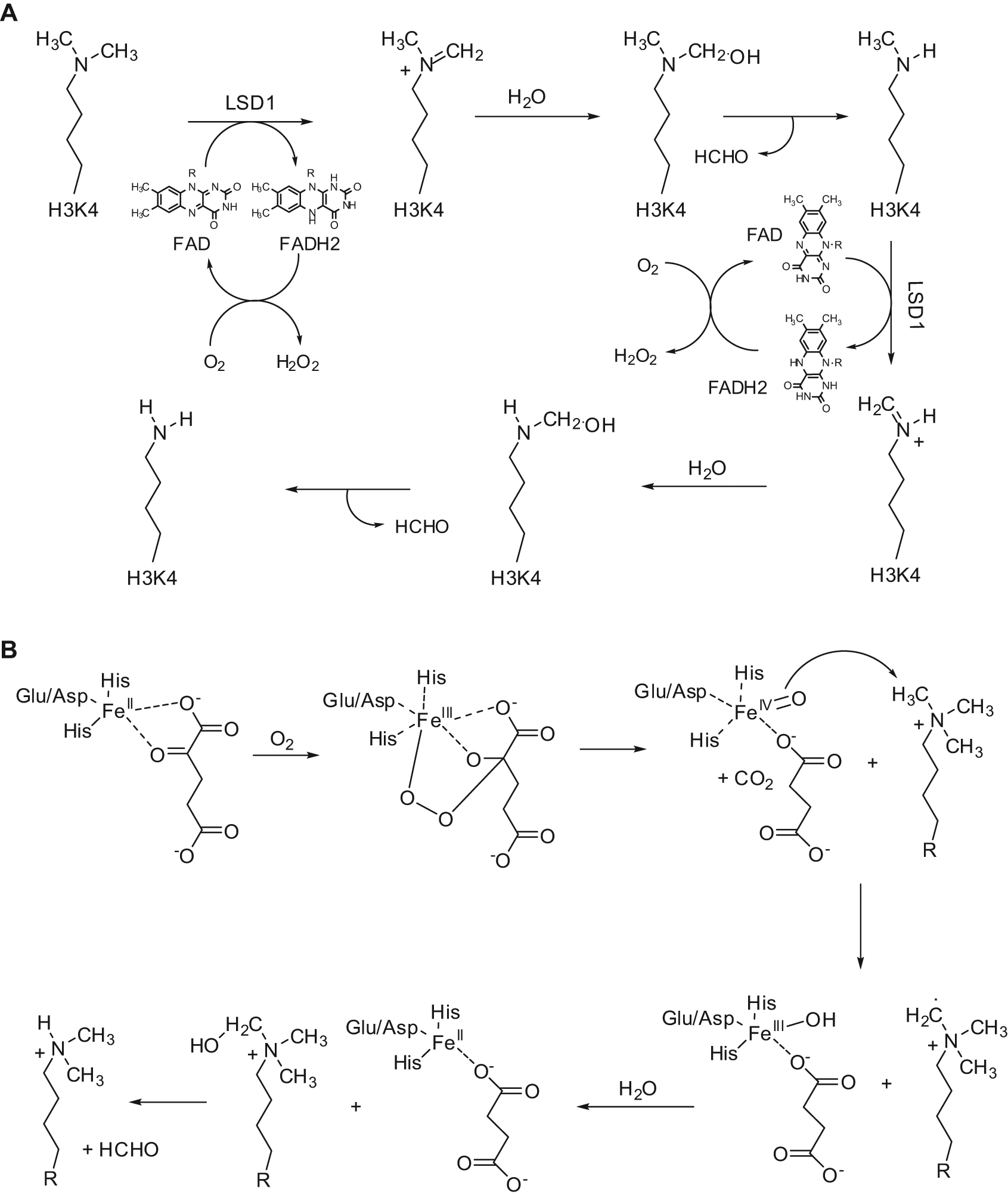
Catalytic mechanism of LSD1 (A) and JmjC (B) enzymes.
Recently, overexpression or mutation of KDMs has been linked to mis-erased histone methyl modifications and, from a clinical point of view, to many types of cancer (Table 1). Thus, such enzymes could represent new potential diagnostic tools as well as therapeutic targets in oncology, and compounds able to inhibit KDMs could be of considerable interest as novel anticancer agents.
Histone Lysine Demethylases (KDMs): Specificity, Transcriptional Effects, and Potential Links to Cancer

NR, not reported.
The Lys-Specific Demethylase LSD1
LSD1/BHC110/AOF2/KDM1A was identified for the first time as a stable component of the BRAF-HDAC transcriptional co-repressor complex containing also the REST co-repressor, CoREST, already known for a repressive role for neuronal genes in nonneuronal cells.11-14 LSD1 consists of an N-terminal SWIRM domain and an AO domain containing 2 sites to bind the methylated substrate and the flavin adenine dinucleotide (FAD) cofactor, respectively. Between these 2 sites in the AO domain, there is the active site of the enzyme, and from them 2 antiparallel α-helices project away as a unique feature forming the Tower domain, crucial for the interaction of LSD1 with CoREST and for the ability to demethylate nucleosomal substrates. 15 Indeed, LSD1 alone only demethylates H3K4 in histone substrates lacking associated DNA and requires CoREST to catalyze nucleosomal demethylation.14,16,17 The reaction starts with generation of an iminium cation intermediate from the methylated amino group of Lys, which is in turn hydrolyzed to yield a carbinolamine that spontaneously degrades to give formaldehyde and the demethylated amine (Fig. 2A). Since such mechanism of demethylation requires a protonated nitrogen to initiate the reaction, the demethylation is limited to dimethyl and monomethyl Lys. The FAD cofactor, reduced during the reaction to FADH2, is reoxidized by O2 in a following step that leads to production of H2O2 (Fig. 2A).7,18,19 The X-ray structure of the complex between LSD1 and H3K4me2 has been solved, and the observed tight constraints on the H3 N terminus account for no more than 3 amino acids to be present on the N-terminal side of the methyllysine in the active site of the enzyme, near to the FAD cofactor. This, combined with sequence-specific interactions between the active site and Arg2, Thr3, and Gln5, provides an explanation for the observed specificity of LSD1 toward the H3K4me1 and H3K4me2 substrates.20-22 When bound to androgen receptor (AR), LSD1 seems to change its substrate specificity: under AR signaling, protein kinase C β1 (PKCβ1) is recruited at the receptor and phosphorylates threonine 6 at histone H3 (H3T6). 23 Such histone H3 modification blocks the H3K4 demethylating activity of LSD1, which in this case catalyzes the H3K9me1 and H3K9me2 (2 repressive markers) demethylation, playing a role of co-activator.24,25 Such a role seems to be corroborated by the cooperation of LSD1 and JMJD2C, a H3K9me3 demethylase, into a specific demethylase complex on AR target genes, which was found to co-localize with AR in normal prostate as well as in prostate carcinomas. 26 Thus, depending on the its binding partners and its substrates, LSD1 seems to be able to have a role as a co-repressor (H3K4 demethylation) or a co-activator (H3K9 demethylation). In addition, LSD1 has been shown to act on nonhistone substrates. LSD1-mediated demethylation of K370me2 in the tumor suppressor p53 prevents its interaction with the co-activator 53BP1 (p53-binding protein 1) and represses its transcriptional activation and apoptosis induction. 27 Demethylation of DNMT1 by LSD1 stabilizes the enzyme from protein degrading and is essential for maintaining global DNA methylation in embryonic stem (ES) cells.28,29 Interestingly, many types of cancer show upregulation of both DNMT1 and LSD1, but a real link between the 2 epigenetic enzymes in such pathologies has to be established. The retinoblastoma protein 1 (RB1) regulator myosin phosphatase target subunit 1 (MYPT1) is also demethylated by LSD1 at Lys442, losing in such way its stability and leading to enhancement of RB1 phosphorylation. 30
In 2009, the second FAD-dependent demethylase, AOF1/LSD2/KDM1B, was identified in mammals. 8 Similar to LSD1, LSD2 contains a conserved SWIRM domain required for its catalytic activity and specifically demethylates H3K4me1 and H3K4me2; nevertheless, its repressing activity seems to be unrelated to the demethylase function. 31 Different from LSD1, LSD2 lacks the Tower domain and thus it is not able to bind CoREST; it forms active complexes with euchromatic histone methyltransferases such as G9a and NSD3 as well as cellular factors involved in transcription elongation rather than with HDACs, and it is localized at the gene body level rather than at the promoters. 32 With respect to LSD1, LSD2 has a more restricted expression pattern, it being abundant essentially in growing oocytes and being required for de novo DNA methylation of some imprinted genes. 33 Thus, considering the involvement of both the 2 flavin-containing amino oxidases LSD1 and LSD2 with DNMTs and de novo DNA methylation (see also above and below), it seems feasible that a real functional link between DNA methylation and histone demethylation would exist. Very recently, LSD2 has been reported to promote H3K9me2 in addition to H3K4me2 demethylation, leading to control of stimulus-induced recruitment of NF-κB to the MDC and IL12B promoters and activation of these inflammatory genes. 34
LSD1 in Cancer
Aberrant expression of LSD1 has been shown in many types of cancers. In particular, LSD1 expression is upregulated in bladder, small cell lung, and colorectal clinical cancer tissues when compared with the corresponding nonneoplastic tissues. 35 In bladder carcinogenesis, LSD1 is highly overexpressed even in tumors at an early grade, thus suggesting LSD1 to be one of the initiators of the whole process. Although estrogen receptor (ER)–positive breast tumors are efficiently treated with antihormonal therapy, ER-negative breast tumors are usually treated with nonselective cytotoxic drugs, and they generally have a less favorable prognosis. LSD1 expression has been found strongly upregulated in ER-negative breast cancers, so LSD1 could be suggested as a predictive biomarker for aggressive tumor biology and tumor recurrence during therapy. 36 Pharmacological (by tranylcypromine, clorgyline) downregulation of LSD1 in ER-negative breast cancer led to growth inhibition, and LSD1 silencing by siRNA resulted in increased expression of the tumor suppressor p21WAF1/CIP1 and in downregulation of the pro-proliferative genes CCNA2 and ERBB2. 36 LSD1 expression was also highly upregulated in poorly differentiated neuroblastoma, it being strongly associated with adverse clinical outcome and inversely correlated with differentiation. 37 Conversely, LSD1 was not expressed in benign ganglioneuroblastomas/ganglioneuromas or in nonmalignant cells such as stromal tissue or infiltrating leukocytes. Downregulation of LSD1 in vitro with both siRNA and monoamine oxidase (MAO) inhibitors (pargyline, clorgyline, tranylcypromine) in SH-SY5Y cells led to growth inhibition and differentiation with an increase of H3K4 methylation, and tranylcypromine treatment in vivo reduced the growth of neuroblastoma xenograft in nude mice, 37 indicating that LSD1 may provide not only a predictive marker for aggressive biology but also a novel therapeutic target for the treatment of aggressive cancer.
On the other side of the coin, LSD1 was shown to be recruited by the Mi-2/nucleosome remodeling and deacetylase (NuRD) complex that in such way possesses at the same time ATPase, deacetylase, and demethylase activities, influencing various pathways comprising transforming growth factor β (TGFβ) signaling, cell growth, and migration and invasion. 38 Through a negative correlation with the TGFβ1 level, highly relevant in epithelial-mesenchymal transitions and invasion, LSD1 suppressed breast cancer metastatic potential in vitro and in vivo. Thus, depending on a variety of factors such as the biological context, the type and the grade of the tumor, and the presence of different substrates, LSD1 seems to have either oncogenic or tumor-suppressive functions, and further studies are needed to better manage this point.
Putative Mechanisms for LSD1-Mediated Oncogenesis
Mechanisms by which overexpression of LSD1 leads or contributes to tumor formation could involve its capacity to silence tumor suppressor genes as a transcriptional co-repressor, mainly through H3K4 demethylation. The physical association of LSD1 with HDACs at the co-repressor complex contributes to repress transcription of common sets of genes. This interplaying activity has been confirmed by experiments made on LNCaP prostate cancer cells treated with pan- (vorinostat) and class I selective (HDAC42, entinostat) HDAC inhibitors. 39 Such compounds led to increased H3K4 methylation levels and suppression of expression of the JmjC demethylase RBP2 and LSD1 through downregulation of Sp1 expression. Moreover, shRNA-mediated silencing of the class I HDAC isozymes 1, 2, 3, and 8, but not that of the class IIb isozyme HDAC6, mimicked the drug effects on H3K4 methylation and H3K4 demethylases, which could be reversed by ectopic Sp1 expression. 39 Further evidence of the role of LSD1 as a transcriptional co-repressor involved in cancer development came from the recruitment, from the CoREST/HDACs/LSD1 complex, of the oncoprotein ZNF217, able to negatively regulate the tumor suppressor gene p15INK4B. 40 Similarly, the DNA binding factor SNAIL, which correlates with high tumor grade and nodal metastasis and is also predictive of a poor outcome in patients with breast cancer, interacts through its SNAG domain with the AO domain of LSD1/CoREST to repress E-cadherin in a variety of cancer cell lines and forms the ternary complex SNAIL/LSD1/CoREST in 116 primary breast cancer samples.41,42 Functionally, LSD1 is linked to DNA methylation and subsequent gene silencing. DNMT3L, a noncatalytic paralog of de novo DNMT3A and -3B, specifically interacts with the histone H3 tail only when H3K4 is unmethylated, and thus the formation of a complex LSD1/DNMT3L/DNMT3A is highly reliable. 43 Alternatively, LSD1 has been reported to demethylate the Lys residue of DNMT1, thus preserving it from degradation. 28 Interestingly, LSD1 and DNMT1 are both upregulated in many types of cancers, but whether the first is responsible for the increased expression of the latter is still unknown. However, further evidence of this DNMT1-LSD1 interplay came from the effect of combination treatment of LSD1 inhibitors and DNMT inhibitors against colon cancer xenograft mice, resulting in a great inhibition of the growth of the tumor. 44 Another possibility for explaining the oncogenic potential of LSD1 is its effect on the tumor suppressor p53. p53 is a nonhistone substrate that is demethylated by LSD1 at Lys370. Methylated Lys370 in p53 is important for its binding to the transcriptional co-activator p53-binding protein–1 (PBP-1), and its demethylation led to inhibition of the p53 apoptotic activity, whereas the knockdown of LSD1 increased p21WAF1/CIP1 and MDM2, well-known p53 target genes. 27 A further putative oncogenic mechanism of LSD1 resides into its capability to trigger Myc-induced transcription through transient H3K4 demethylation. 45 The local DNA oxidation given by the LSD1 catalytic activity and the following production of hydrogen peroxide gave the recruitment of 8-oxoguanine-DNA glycosylase (OGG1) and of the nuclease Ape1 on the E-box chromatin sequence, leading to the assembly of the transcription initiation complex of the oncogene Myc. 45
LSD1 Inhibitors
Substrate Analogues
From the knowledge that LSD1 is a FAD-dependent AO that catalyzes the oxidative demethylation of monomethyl- and dimethyl-K4 residue at histone H3, as well as H3-tail peptides requiring at least 15 amino acid residues for efficient reaction, the propargyl- Lys-derivatized peptide
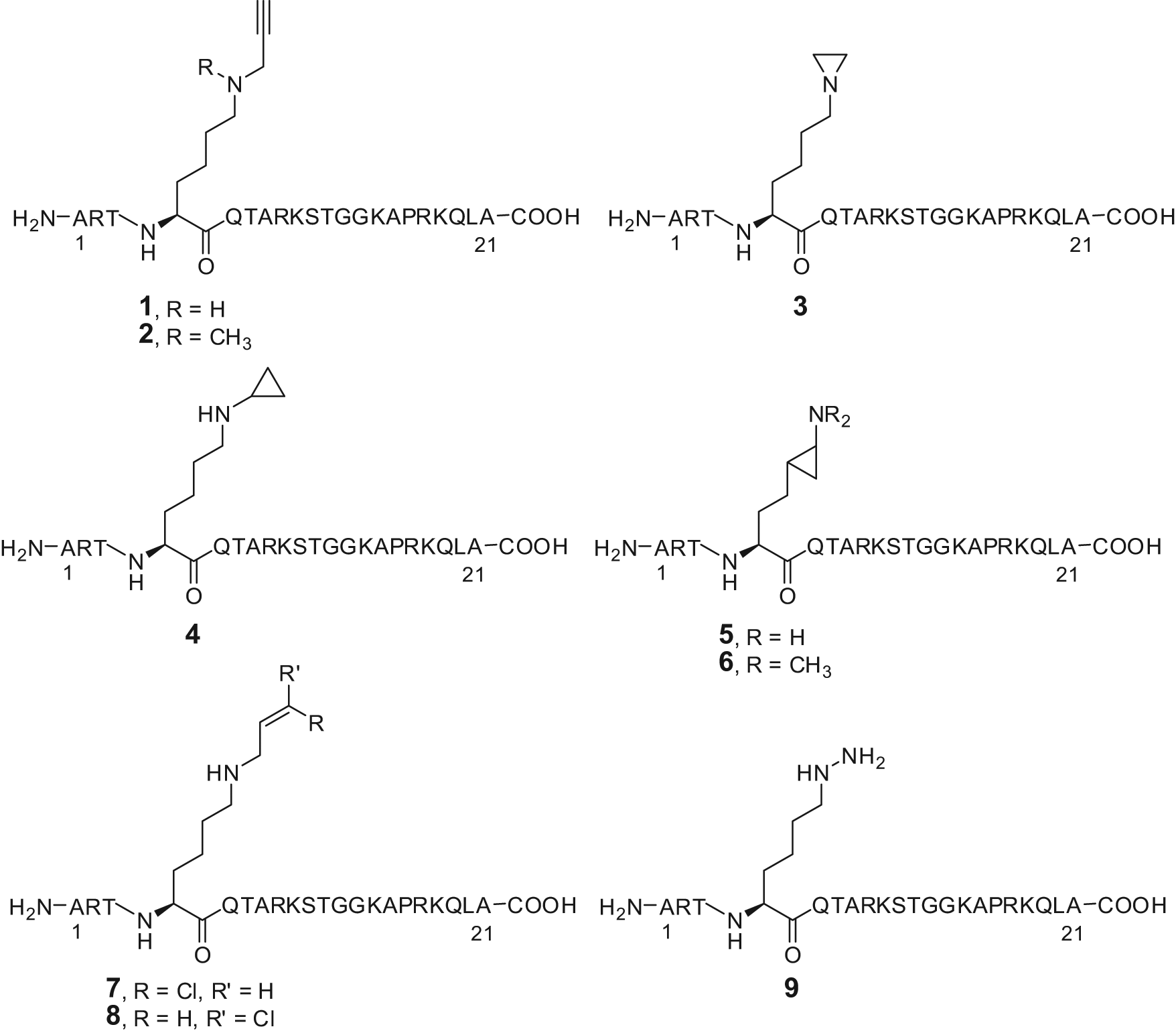
Substrate analogues as LSD1 inhibitors.
Mechanistic studies made on
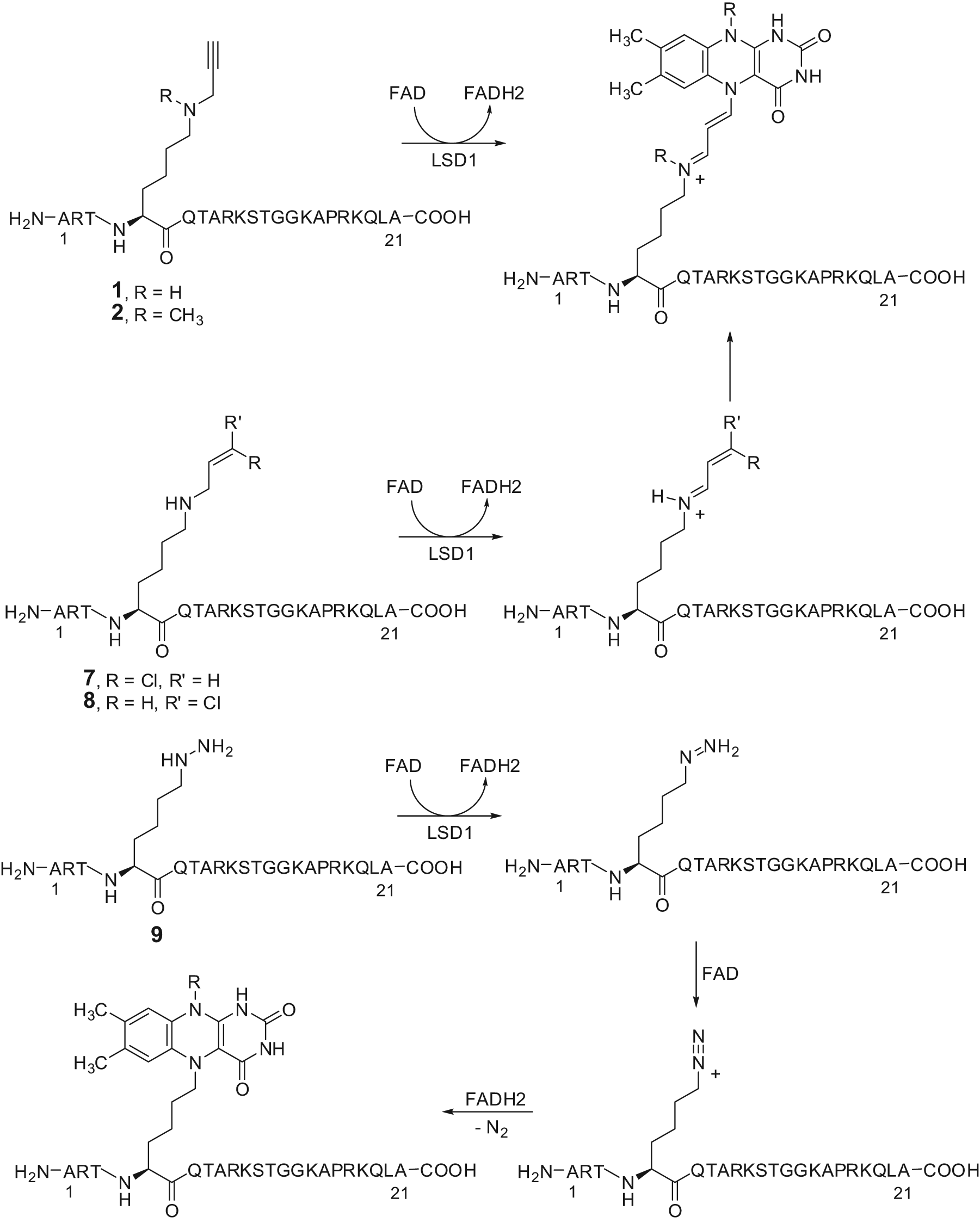
Proposed mechanism of inactivation of LSD1 by substrate analogue inhibitors.
Although aziridine and exo- and endo-cyclopropylamine units (compounds
Polyamine Analogues
The AO domain of LSD1 shares 20% sequence similarity with those of the FAD-dependent MAO and PAO (polyamine oxidases such as spermine oxidase [SMO] and N1-acetylpolyamine oxidase), with the catalytic domains of LSD1 and SMO showing 60% similarity in amino acid sequences. As compounds bearing guanidine groups were shown to inhibit both SMO and other polyamine oxidases, a pioneer series of (bis)guanidines and (bis)biguanides, previously synthesized as potential antitrypanosomal agents,
49
were tested as inhibitors of LSD1 (Fig. 5).
50
Among them, compounds
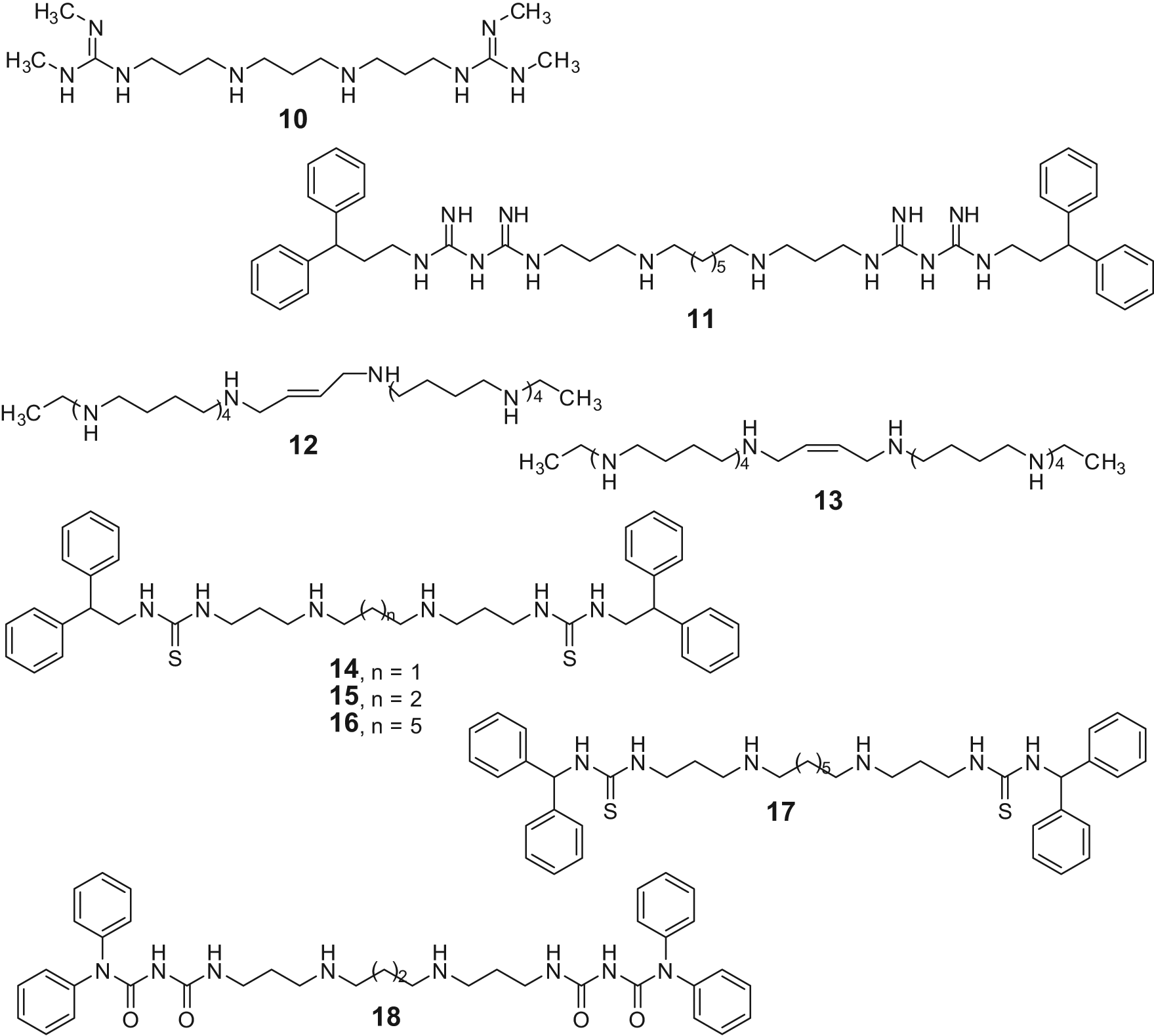
Polyamine analogues as LSD1 inhibitors.
Anti-MAO-Based Small Molecules
Due to the high sequence similarity between LSD1 and MAO, some known anti-MAO agents were tested against LSD1. Pargyline (Fig. 6), a propargylamine-based anti-MAO initially suggested as a valuable LSD1 inhibitor prototype, failed in further studies to inhibit the Lys demethylase.21,52 Phenelzine (Fig. 6), an anti-MAO agent bearing a hydrazine moiety, was initially reported as a weak LSD1 inhibitor, and when reinvestigated on its inhibiting activity, it resulted in having highly improved potency against LSD1 (Ki = 17.6 µM), similar to that displayed versus MAOs. 47
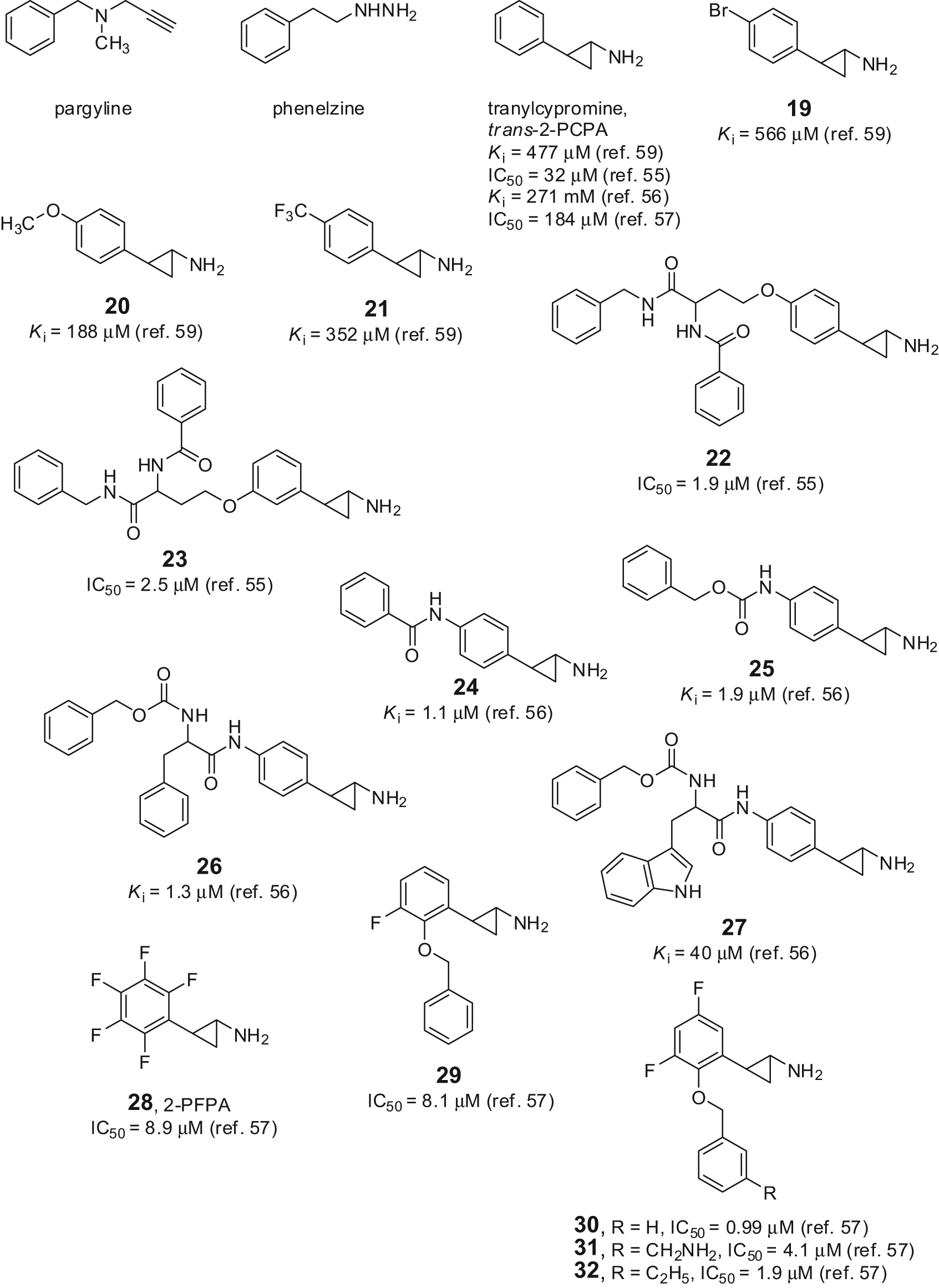
Anti-MAO-based LSD1 inhibitors.
Tranylcypromine (trans-2-phenylcyclopropyl-1-amine, trans-2-PCPA) (Fig. 6) was reported to inhibit LSD1 with a Ki value ranging from 477 to 22 µM, depending on the type of assay, the time of addition, the substrate, and the enzyme source used for testing.52-58 Nevertheless, trans-2-PCPA became a useful lead to develop highly potent small-molecule inhibitors of LSD1. Some trans-2-PCPA analogues carrying 4-bromo, 4-methoxy, and 4-trifluoromethoxy substitutions at the benzene ring led to compounds (
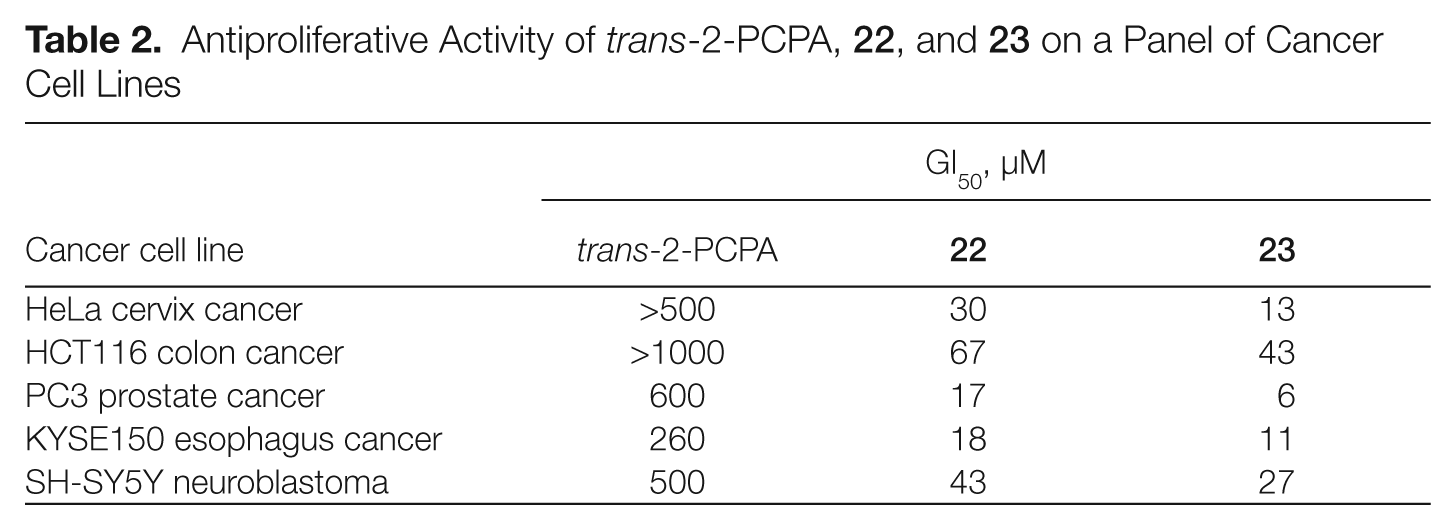
Antiproliferative Activity of trans-2-PCPA,
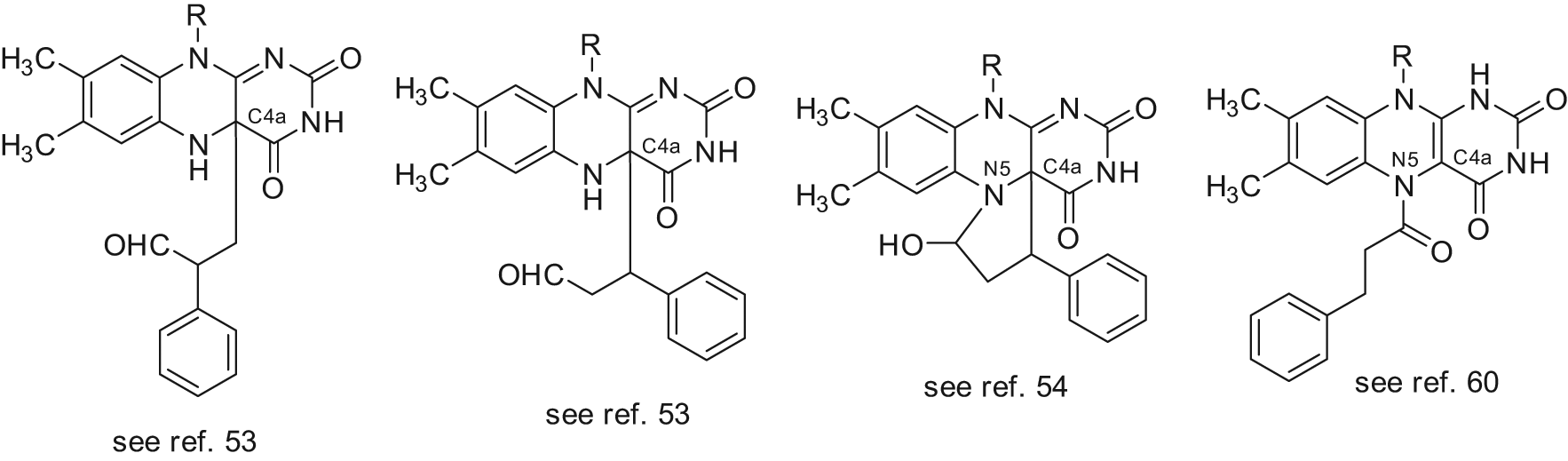
Proposed structures for the flavin adenine dinucleotide (FAD)–tranylcypromine adducts in the LSD1 complex.
As trans-2-PCPA is the racemic mixture of 2 enantiomers, (+)-trans-2-PCPA and (–)-trans-2-PCPA, we prepared large amounts of trans- and cis-2-PCPA, separated the 2 trans- and 2 cis-enantiomers, and tested them against LSD1 and MAOs, determining their binding to FAD through X-ray crystallography. 56 Thus, we found that the 2 trans-enantiomers of 2-PCPA were slightly more potent than the 2 cis congeners against LSD1, with (–)-trans-2-PCPA being 1.7-fold more efficient than (+)-trans-2-PCPA (Fig. 7). Conversely, against MAO-B, (+)-trans-2-PCPA was 20-fold more potent than the corresponding (–) enantiomer, highlighting the crucial role of stereochemistry in MAO-B inhibition by trans-2-PCPA. From X-ray crystal structures, it was clear that (–)-trans-2-PCPA connects its carbonyl carbon in a covalent bond with the flavin N5 atom, positioning its phenyl ring in the core of the substrate-binding pocket, above the flavin ring. Differently, (+)-trans-2-PCPA links the flavin N5 atom through the phenyl-substituted carbon of the inhibitor, pointing the phenyl ring away from the flavin ring (Fig. 8). Due to the higher potency shown by the (–) enantiomer, when the racemic trans-2-PCPA is crystallized with LSD1, the enzyme selects this enantiomer for the binding with FAD, and the N5-carbonyl adduct is obtained, according to the literature.56,60
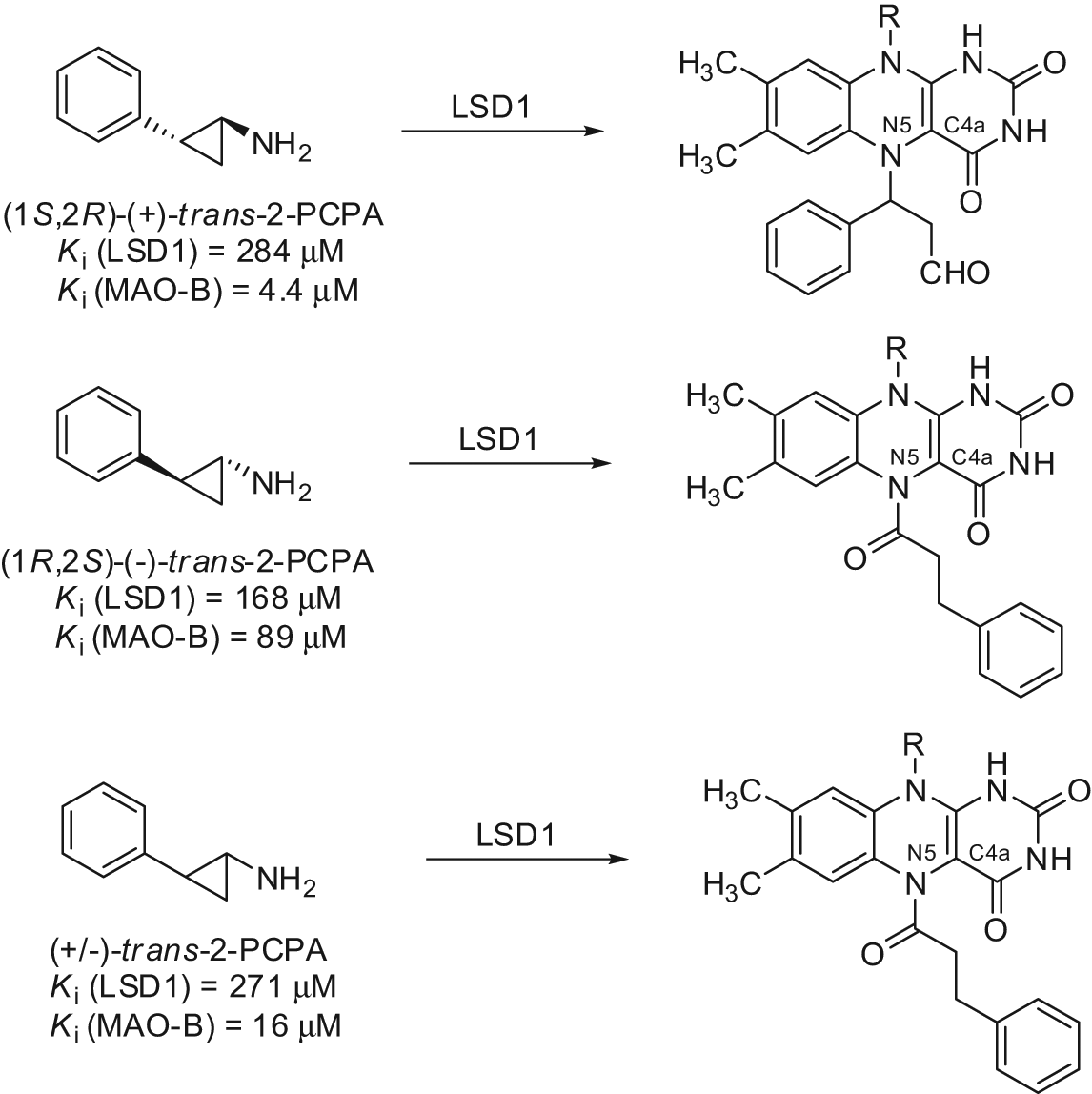
Structures of flavin adenine dinucleotide (FAD)–tranylcypromine adducts obtained by the trans-2-PCPA enantiomers.
LSD1 Activators?
Two immunomodulatory drugs, pomalidomide and lenalidomide (Fig. 9), have been reported to induce p21WAF1 expression in Burkitt lymphoma and multiple myeloma cell lines, leading to cell cycle arrest. This p21WAF1 induction is generated by modification of the chromatin structure at the p21 promoter involving a switch from H3K9 methylation (a repressive mark) to H3K9 acetylation (an activation mark of transcription). As LSD1 silencing reduced the observed pomalidomide and lenalidomide-induced p21 upregulation, LSD1 has been suggested to be involved in this effect of p21 induction through demethylation of H3K9, and pomalidomide and lenalidomide could act as direct LSD1 activators or indirectly through activation of a third partner required for LSD1 activation. 61
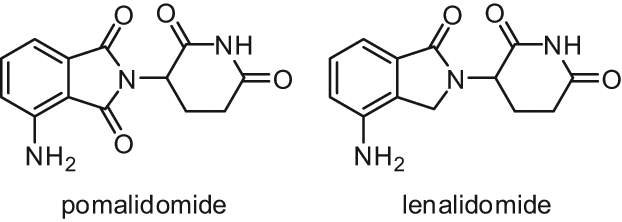
Structures of pomalidomide and lenalidomide.
The Jumonji C Family of Lysine Demethylases
The second and largest class of histone lysine demethylases (KDMs) to be discovered is the Fe+2/2-OG-dependent family of KDMs.9,10,62 These enzymes belong to a much larger enzyme superfamily, the 2-OG oxygenases, whose members (about 70-80 genes in the human genome) catalyse a diverse range of oxidation reactions, using 2-OG, molecular oxygen, and Fe2+ as co-substrates/co-factors (Fig. 2B). 63 In humans, the two main reactions catalyzed by 2-OG oxygenases identified so far are protein/nucleotide hydroxylation and N-methyl demethylation. 64 In particular, the 2-OG-dependent histone lysine demethylases are members of the Jumonji family of 2-OG oxygenases, which have common structural features (e.g., with respect to 2-OG binding residues) and comprise nonhistone modifying enzymes such as factor inhibiting HIF (FIH). 65 The Jumonji protein family is named this way because it contains the conserved Jumonji C (JmjC) catalytic domain that was first identified in the so-called Jumonji protein (JARID2). 66 In fact, jumonji means cruciform in Japanese, and the gene was so named because mice with a genetrap inserted in the Jumonji locus develop an abnormal cross-like neural tube. 66 Among the 30 JmjC domain-containing proteins identified so far within the human genome, about 20 have been published to demethylate specific lysines in the histone proteins.
The JmjC histone lysine demethylases catalyze the hydroxylation at the carbon of the Nε-methyl group to give an unstable hemiaminal intermediate that rapidly breaks down, leading to the release of formaldehyde and the demethylated lysine residue. 67 Unlike LSD1, the hydroxylation-based mechanism of the JmjC KDMs does not require a protonatable lysine ε-amine group, enabling these enzymes to demethylate all 3 lysine methylation states (tri-, di- and monomethylation) at H3K4, H3K9, H3K27, and H3K36, as well as H1K26. 67 Six different subfamilies (JMJD1s, JMJD2s, JARID1s, UTX/Y-JMJD3, PHFs, and FBXLs) of JmjC histone demethylases have been identified, which have different histone sequence and methylation state selectivities.67,68 For instance, KDMs of the JmjC domain-containing 2 (JMJD2) subfamily are selective for the demethylation of the tri- and di-Nε-methylation states of specific lysines on histone H3, whereas other subfamilies (e.g., the PHF and FBXL subfamilies) are selective for the di- and mono-Nε-methylated states and do not accept the trimethylated state. Recent crystallographic analyses have shown that variations in the size of the active site region binding the Nε-methyl group are in part responsible for the methylation state selectivity. 69 Selectivity for a particular histone sequence seems to be conferred by binding interactions directly with the catalytic domain and, at least for some enzymes, by the presence of other noncatalytic domains that target the catalytic one to particular residues, as recently demonstrated for PHF8.69-71 In fact, this histone demethylase has a plant homeobox domain (PHD), in which binding the Nε-trimethylated form of the K4 of the histone H3 increases the activity of the catalytic domain with regard to demethylation of the Nε-dimethylated H3K9. Mounting evidence (especially works with peptides) suggests that some JmjC demethylases have also nonhistone substrates, a conceivable finding given the rising data on nonhistone proteins that are Nε-methylated. 72
Despite the functional characterization of many JmjC histone demethylases being still at a relatively early stage, recent evidence suggests for these enzymes many important biological roles ranging from the regulation of cellular differentiation and development to the control of neuronal function, from senescence process modulation to the control of genome stability and cancer cell proliferation.64,67,68 Jumonji histone demethylases are being targeted for inhibition by small molecules, both with a view to developing “chemical probes” potentially useful in functional assignments and for the eventual development of pharmaceuticals, with the most likely application being cancer.18,73 In fact, the misregulation of JmjC KDMs has significantly being implicated in cancer initiation and progression, and this will be reviewed here by analyzing the involvement of different subfamilies of the JmjC histone lysine demethylases.
The KDM2 Cluster (FBXL Subfamily)
KDM2A/JHDM1A (JmjC domain–containing histone demethylation protein 1A)/FBXL11 (F-box and leucine-rich repeat protein 11) was the first JmjC domain–containing protein shown to be a histone lysine demethylase. Human KDM2A and KDM2B/JHDM1B/FBXL10 have been shown to catalyze H3K36me2/me1 demethylation.74,75 In addition, mammalian FBXL10 has been suggested to act as an H3K4me3 demethylase,76,77 but reports on its activity toward H3K4me3 are conflicting.74,76 Both proteins contain an F-box domain in addition to 2 leucine-rich repeat (LRR) domains. 76 Retroviral insertional mutagenesis within the FBXL10/JHDM1B gene has been shown to cause lymphoma in BLM (Bloom syndrome RecQ protein-like-3 DNA helicase)–deficient mice, indicating KDM2B as a candidate tumor suppressor. 78 Other studies have proposed FBXL10, along with FBXL11, as a putative proto-oncogene. 79 Expression data from human cancers show overexpression of KDM2B in lymphomas and adenocarcinomas 79 but reduced expression of KDM2A and KDM2B in prostate cancer and in the most aggressive of the primary brain tumors, the glioblastoma multiform, respectively.76,80 Cell culture studies support the view that KDM2 proteins might have context-dependent pro- and anti-oncogenic functions.
KDM2A can suppress NF-κB-dependent growth of colon cancer cells, 81 whereas KDM2B has been shown to inhibit proliferation of tumor cells correlating with repression of rRNA genes. 76 By contrast, KDM2B prevents premature senescence in MEFs (mouse embryonic fibroblasts) through a mechanism that likely involves repression of the Ink4b gene.74,77,82 MEFs ectopically expressing KDM2A or KDM2B bypass stress-induced senescence, depending on an intact JmjC domain, and KDM2B cooperates with Ras in oncogenic transformation.74,79 In addition to the proliferative control, KDM2 proteins are important for genomic stability. KDM2B has been demonstrated to protect cells against ultraviolet (UV)–induced apoptosis, oxidative stress, and spontaneous mutagenesis,78,83,84 whereas depletion of FBXL11 was reported to influence chromosomal segregation through effects exerted in the heterochromatic areas of the genome. 80 Taken together, mammalian KDM2 demethylases have been suggested to have context-dependent tumor-promoting and tumor-inhibiting functions.
The KDM3 Cluster (JMJD1 Subfamily)
KDM3A/JHDM2A/JMJD1A (Jumonji domain–containing 1a)/TSGA (testis-specific gene A) is a histone demethylase specific for H3K9me2/me1. 85 Two JMJD1A homologues exist in human cells: KDM3B/JHDM2B/JMJD1B and JMJD1C/JHDM2C/TRIP8 (thyroid receptor interacting protein 8). JMJD1A features an LXXLL motif that is a signature involved in nuclear receptor interactions. 86 The expression of JMJD1A is very significant in testes and has been implicated in demethylation of H3K9me2 of AR target genes. 85 Thus, JMJD1A was found to interact with the AR in a ligand-dependent manner. Inhibition of JMJD1A expression in the prostate cancer cell line LnCaP led to an increase in H3K9me2 in a subset of AR target genes, including PSA, NKX3.1, and TMPRSS22, and a decrease in their expression. 85 These results show that JMJD1A acts as a co-activator of AR-mediated transcription.
Studies of genetrap and knockout mice have demonstrated important roles for KDM3A in germ cell development and metabolism.87-90 JMJD1A is highly and dynamically expressed during spermatogenesis, and male Kdm3a genetrap mice are infertile, with small testes and a severe reduction in sperm count. This defect has been linked to KDM3A positively regulating expression of genes involved in sperm chromatin condensation and maturation through demethylation of H3K9me2/me1.88,89 Kdm3a knockout mice also exhibit an adult-onset obesity phenotype, and in this context, KDM3A has been implicated in the transcriptional control of metabolic genes in muscle and adipose tissues.87,90 Little is known about the implication of JMJD1s in tumorigenesis. Frequent deletion of 5q31 locating the JMJD1B gene or reduced expression of JMJD1C in various malignancies suggests possible roles of KDM3 proteins in tumor suppression.91,92 Recent studies have shown that JMJD1A is the target of hypoxia-inducible transcription factors, and under hypoxia, JMJD1A increases a subset of hypoxia-inducible genes enhancing tumor growth. 93
The KDM4 Cluster (JMJD2 Subfamily)
Although the potential role of JMJD1s in cancer remains controversial, the JMJD2 subfamily members are mainly regarded as oncogenes. Their oncogenic potential could result from demethylating heterochromatic H3K9me3/2, an important mark for formation and maintenance of heterochromatin and somehow of genomic stability. In fact, KDM4 (JMJD2) proteins were the first published demethylases that showed activity toward trimethylated lysines. There are 4 members of the KDM4 cluster in mammalian cells: KDM4A/JHDM3A/JMJD2A, KDM4B/JHDM3B/JMJD2B, KDM4C/JHDM3C/JMJD2C/GASC1 (gene amplified in squamous cell carcinoma 1), and KDM4D/JHDM3D/JMJD2D. The JMJD2 proteins catalyze the demethylation of H3K9me3/me2 and/or H3K36me3/me2, with the substrate specificity differing between subfamily members.94-98 Moreover, the KDM4 proteins have recently been demonstrated to demethylate H1.4K26me3/me2. 99
Mounting evidence correlates human KDM4 family members—especially KDM4C—to malignant transformation. Significantly, amplification of the KDM4C locus has been reported for esophageal squamous carcinomas, medulloblastomas, and breast cancers, and amplification of KDM4B is detected in medulloblastomas.100-102 Gene expression analysis further shows that JMJD2A-C is overexpressed in prostate cancer. 94 Depletion of JMJD2C from cancer cells, expressing high levels of it, resulted in inhibition of cell growth of several tumor cell lines,26,94,101 whereas overexpression of GASC1 in human nontransformed mammary epithelial cells resulted in phenotypic alterations that are hallmarks of neoplastic transformation, including growth factor–independent proliferation and anchorage-independent growth in soft agar. 101 Introduction of the JMJD2C gene in normal breast MCF10A cells could increase higher capacity to generate mammospheres, a phenotype of cancer stem cells, suggesting that JMJD2C acts as a transforming gene. 101 However, the genomic targets and cellular functions of the mammalian KDM4 subfamily members are still for the most part uncharacterized. To date, different studies have shown that KDM4C can function as a transcriptional activator by removing the heterochromatic H3K9me3 at promoter regions, which is generally associated with transcriptional repression. 26 Analysis of genes altered by overexpression of GASC1 in MCF10A- GASC1 cells revealed NOTCH1 as a target gene of GASC1. 101 Induction of NOTCH signaling promotes self-renewal of normal human mammary stem cells. This may support the finding that the phenotype of cancer stem cells was increased in MCF10A-GASC1 cells. The involvement of GASC1 in tumorigenesis has been supported further by a recent report demonstrating the functional interaction between GASC1 and AR in prostate carcinoma. 26 It was known that LSD1 and JMJD1A are also required for transcriptional activation of AR responsive genes and for proliferation of prostate cancer cells. Although LSD1 and JMJD1A only demethylate mono- and dimethylated H3K9, GASC1 is especially capable of efficiently demethylating trimethylated H3K9, inducing a robust cooperative stimulation of AR transcriptional activity. Thus, specific modulation of GASC1 activity alone, or in combination with LSD1, may be a promising therapeutic strategy to control AR activity in tissues where AR has a pivotal physiological role. 26 Clearly, these observations suggest also critical functions not only in cancer but also during development. In summary, growing evidence suggests important roles for human JMJD2B and JMJD2C in tumorigenesis, indicating these proteins as potential targets for the development of new-generation anticancer agents.
The KDM5 Cluster (JARID1 Subfamily)
The JARID1 subfamily, specific for the demethylation of H3K4me3/2, encompasses 4 enzymes: KDM5A/JARID1A (Jumonji, AT-rich interactive domain 1A)/RBP2 (retinoblastoma-binding protein 2), KDM5B/JARID1B/PLU-1, KDM5C/JARID1C/SMCX (selected mouse cDNA on the X), and KDM5D/JARID1D/SMCY. 67 Despite the fact that all members of KDM5 cluster catalyze the demethylation of the same histone mark, they appear to have exclusive functional properties probably because of their different expression profiles and presence in distinct protein complexes. RBP2 is broadly expressed in diverse tissues and has been linked by target gene mapping and functional studies to regulation of differentiation, cell cycle progression, and mitochondrial function.103-107 The KDM5A interaction partners identified so far include RBP-J (recombination signal-binding protein–Jk), Sin3, c-Myc, the tumor suppressor pRB (retinoblastoma protein), and PcG proteins of the PRC2 complex, which harbor H3K27 methyltransferase activity.105,106,108-110 Little is known at present about the role of JARID1A/RBP2 in human cancers, and the few available reports are contradictory. RBP2 has been shown to be a key effector of pRB-mediated cell cycle withdrawal and differentiation by interacting with the tumor suppressor pRB. 103 Recently, RBP2 protein was shown to be an integral part of the core Notch-RBP-J repressor complex, contributing to switch off the Notch signaling. 109 In contrast to its tumor suppressor function, a recent report showed that it is overexpressed in gastric cancer and that its inhibition leads to cellular senescence of gastric and cervical cancer cells by de-repressing cyclin-dependent kinase inhibitors such as p27, p21, and p16. 107 Recently, enhanced Jarid1A/Rbp2 expression was found to contribute to drug tolerance with respect to a receptor tyrosine kinase inhibitor in a non–small cell lung carcinoma (NSCLC) cell system, whereas RBP2 knockdown significantly reduced the number of the drug-tolerant population. 111 These drug-tolerant NSCLC cells displayed reduced levels of H3K4me2 and H3K4me3, which is reminiscent of the prognostic value of reduced H3K4me3 in NSCLC. This also indicates that acquired drug resistances are accompanied by epigenetic changes, and combining a chromatin-modifying agent with a single, rationally targeted agent would prevent the development of drug resistance.
Although significant expression of PLU-1 (JARID1B) is limited to testis and only low levels of PLU-1 are seen in other normal adult tissues, its expression increases markedly in pathological conditions such as malignant breast cancer, metastatic prostate cancer, and testis cancer.112,113 Based on its overexpression in cancer, PLU-1 was suggested as a testis cancer antigen. Supporting an oncogenic potential at a cellular level, PLU-1 knockdown slowed down the proliferation of MCF-7 breast cancer cells and tumor growth of mammary carcinoma cells in a syngenic mouse mammary tumor model. 114 Like LSD1, PLU-1 was shown to be involved in silencing a series of tumor suppressor genes previously linked to breast cancer, including SFN (14-3-3-σ), BRCA1, CAV1, and HOXA5. Although this is consistent with a transcription repressive role for this protein, JARID1B can also act as a transcriptional co-activator for the androgen receptor.112,113 Involvement in hormone signaling is consistent with high expression levels in testis, ovary, and mammary glands of pregnant females. 112
JARID1C/SMCX (KDM5C) has been implicated in X-linked mental retardation and epilepsy. 115 However, its role in the pathogenesis of human papillomavirus (HPV)–associated cancers was recently identified by a genome-wide siRNA screen that found, among other genes, JARID1C/SMCX as one of the E2-dependent regulators of HPV oncogene expression. 116
The KDM6 Cluster (UTX/Y-JMJD3 Subfamily)
The KDM6 cluster in human cells consists of KDM6A/UTX (ubiquitously transcribed X chromosome tetraticopeptide repeat protein), UTY, and KDM6B/JMJD3. KDM6A and KDM6B are histone demethylases specific for H3K27me3/me2, whereas no activity has been reported for UTY so far.117-120
The tri- and dimethylation of H3K27, which are catalyzed by the polycomb group (PcG) proteins, are important repressive histone marks. The PcG genes have been characterized as oncogenes and are frequently overexpressed or amplified in cancer. Their oncogenic potential is mainly mediated through PcG-mediated H3K27 methylation and epigenetic inactivation of the INK4A-ARF locus.121,122 The INK4A-ARF locus encodes tumor suppressor genes p16INK4A and p14ARF, which are key regulators of cellular senescence.
Both the H3K27 demethylases KDM6A and KDM6B are suggested as tumor suppressor genes by functioning antagonistically to the oncogenic PcG proteins. Although JMJD3 and UTX have the same catalytic activity, they most likely have, as the JARID1 proteins, different biological functions. Overexpression of either leads to cell cycle arrest in fibroblasts, dependent on intact catalytic domains.123-125 Although knockdown of KDM6A confers an immediate growth advantage to fibroblasts, 123 depletion of KDM6B enables MEFs to overcome the senescence barrier.124,125 The growth arrest induced by KDM6A has been linked to transcriptional activation of components of the pRB pathway. In fact, a genome-wide chromatin occupancy analysis has shown that many pRB-binding proteins, including HMG-box protein 1 (HBP1), are UTX target genes. 123 UTX, by means of preventing H3K27 methylation and silencing HBP1, is able to enforce cell cycle arrest by pRB. Consistently, UTX depletion results in elevated levels of proliferation. UTX, the first reported mutated histone demethylase gene associated with cancer, is a frequent target of somatic mutations in human cancers with high prevalence in multiple myeloma, esophageal squamous cell carcinomas, and renal cell carcinomas.126,127
It has been shown that JMJD3, but not UTX, contributes to the activation of INK4A-ARF by removing H3K27me3 during induction of oncogene- or stress-induced senescence.124,125 KDM6B expression in fibroblasts is induced by oncogenes of the RAS-RAF pathway, suggesting that it could provide an important barrier to tumorigenesis. Accordingly, KDM6B levels are high in melanocytic nevi, which constitute pre-neoplastic lesions, whereas data mining indicates that KDM6B expression is decreased in subsets of human cancers.124,125 Moreover, it appears that a large part of the genetic lesions leading to p53 loss likewise causes the deletion of JMJD3. The JMJD3 gene is located on chromosome 17, in close proximity to the p53 tumor suppressor gene. The allelic loss at 17p13.1, including both p53 and JMJD3, was also significantly correlated with more aggressive tumor behaviour. 128 Taken together, genetic lesions and/or decreased expression of JMJD3 might contribute to the development of some human cancers, most likely by epigenetic silencing of the INK4A-ARF tumor suppressor locus.
JmjC Histone Demethylase Inhibitors
Potent and specific inhibitors of the Jumonji C domain–containing histone lysine demethylases have not been identified yet, and the available structure-activity relationship (SAR) data are limited.129-131 However, the efforts to target the cofactors essential for the activity of this class of enzymes have provided first promising results.129-131 The 2-oxoglutarate analogue, N-oxalylglycine (NOG), can weakly inhibit JMJD2C and the catalytic core of JMJD2A (Fig. 10). 132 JMJD2A was also found to be specifically inhibited by disruption of its Zn2+ binding site by Zn-ejecting compounds such as disulfiram and ebselen (Fig. 10). 133 Interestingly, Zn-ion ejection seemed to be selective for the JMJD2 subfamily as other KDMs do not contain a Zn-binding site near their catalytic domain. 133 In a recent study employing binding analyses by non-denaturing mass spectrometry (MS), dynamic combinatorial chemistry coupled to MS, turnover assays, and crystallography, a set of N-oxalyl-D-tyrosine derivatives was investigated for the inhibition of the JMJD2 (KDM4) subfamily of histone demethylases. Some of the inhibitors were shown to be selective for JMJD2 over the hypoxia-inducible factor prolyl hydroxylase PHD2. 130 Other 2-OG mimetics endowed with a promising JMJD2 inhibitory potency are the 2,4-pyridindicarboxylic acid (PDCA), a fragment-size inhibitor with (sub)micromolar activity against JMJD2s and some of its derivatives,129,131 and a 2,2′-bipyridyl derivative containing a 4-carboxylate group (Fig. 10). 129 A recent screen of a small library of compounds provided as JmjC KDM inhibitors some flavonoid- and catechol-type molecules, which have the limitation of being promiscuous inhibitors affecting a wide range of enzymatic targets (Fig. 10). 134 Quantitative high-throughput screening of a ~236,000-member collection of diverse molecules led to the identification of 8-hydroxy-5-carboxyquinoline as a potent inhibitor of JMJD2 enzymes via chelation of the active-site iron and endowed with activity against JMJD2A in cell-based studies. 135 The well-known HDAC inhibitor suberoylanilide hydroxamic acid (SAHA) was shown to be also a micromolar inhibitor of the recombinant enzyme JMJD2E (annotated as a pseudogene) that, for its close analogy in terms of sequence identity to the catalytic domain of the other JMJD2 enzymes, is easily amenable to large-scale inhibition studies (Fig. 10). 129 Recently, some hydroxamic acids bearing as a unique feature an N-propionic acid unit have been reported by Hamada and co-workers 136 as selective inhibitors of JMJD2A and JMJD2C over PHD1 and PHD2 enzymes (Fig. 10). Two cell-permeable pro-drugs of the best inhibitor of this series displayed a synergistic cell growth inhibition in combination with a tranylcypromine-based LSD1 inhibitor in different cancer cell lines (LNCaP, PC3, and HCT116), suggesting clinical potential for this type of combination in anticancer chemotherapy. 136
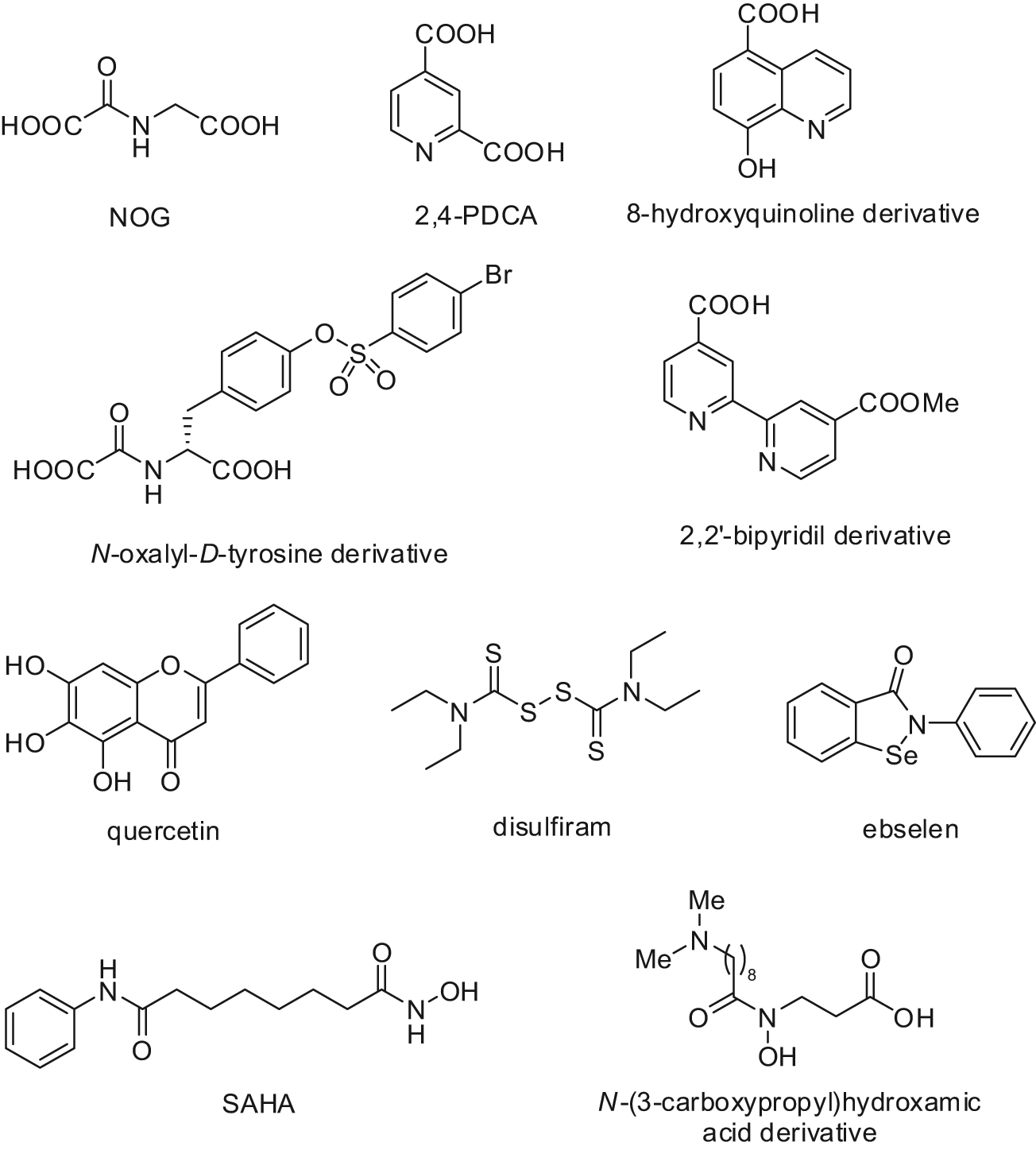
Structures of JmjC KDM inhibitors.
Footnotes
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) received no financial support for the research, authorship, and/or publication of this article.
