Abstract
Drug resistance remains a clinical challenge in cancer treatment due to poor understanding of underlying mechanisms. We have established several drug-resistant prostate cancer cell lines by long-term culture in medium containing chemotherapeutic drugs. These resistant lines displayed a significant increase in side population cells due to overexpression of drug efflux pumps including ABCG2/BCRP and MDR1/Pgp. To uncover potential mechanisms underlying drug resistance, we performed microarray analysis to identify differentially expressed genes in 2 drug-resistant lines. We observed that POU5F1/OCT4, a transcription factor key to regulating pluripotency in embryonic stem cells, was upregulated in drug-resistant lines and accompanied by transcriptional activation of a set of its known target genes. Upregulation of OCT4 in drug-resistant cells was validated by RT-PCR and sequencing of PCR products as well as confirmation by Western blot and specific shRNA knockdown. Analysis of the regulatory region of
Introduction
Although screening and early detection have dramatically increased survival rates, prostate cancer remains the second leading cause of cancer deaths among men in the Western world. Hormone ablation is often achieved through surgical or pharmacological means as a standard treatment approach for most advanced prostate cancer patients. This therapy aims to reduce androgen levels, which are believed to support prostate cancer cell growth and proliferation. A majority of patients initially respond well to this treatment but inevitably relapse and develop an incurable castration-resistant disease within a few years. 1
Few treatment options are currently available for castration-resistant prostate cancer. Interchalating agents, alkylating compounds, and microtubule stabilizers are the main classes of drugs used in the clinic at this stage. 2 While most offer only palliative effects, the latter group that includes taxanes has shown promising clinical response in combination with other drugs like prednisone. 3,4 However, intrinsic or acquired resistance can often lead to failure of this secondary treatment. Taxane resistance has been attributed to a number of factors including increased efflux pump activity, selection of cells with increased survival advantages, or enhanced DNA-damage repair. Overexpression of ATP-binding cassette (ABC) transporter efflux pumps can offer a protective effect from chemotherapeutic drugs as a result in enhanced efflux of xenobiotics out of the cell. Taxanes such as docetaxel are direct substrates for P-glycoprotein (Pgp), so they can easily be exuded by cancer cells expressing a high level of this protein. 5,6 Because docetaxel exerts its mode of action by preventing microtubule depolymerization, alterations in β-tubulin expression or signaling can also lead to decreased patient sensitivity to the drug. 7 Understanding such pathways of chemoresistance is critical not only for resensitizing tumor cells to therapeutic drugs but also for prevention of this process.
OCT4/POU5F1 is a well-established transcription factor critical for maintaining pluripotency in embryonic stem cells. It remains unclear what roles if any OCT4 serves in somatic cells or during carcinogenesis. A novel function of OCT4 in tumorigenesis was proposed when its ectopic expression induced dysplastic growth of epithelial tissue.
8
It has also been described as a marker for germ cell tumors.
9
OCT4 was reported to be reactivated in cancer cell lines,
10,11
yet other groups have not been able to detect such upregulation.
12,13
One possible explanation for these discrepancies is that the
We previously established drug-resistant derivatives from the CWR-R1 prostate cancer cell line. 20 In an effort to characterize these lines, we examined the differentially expressed genes that may underlie their drug resistance and enhanced tumorigenicity. Side population and microarray analysis indicated that several stem-like genes were preferentially expressed in drug-resistant cells. Of interest, we noted that embryonic stem cell marker OCT4 was overexpressed compared to the drug-sensitive parental line. We confirmed OCT4 expression using multiple analyses. We further showed that OCT4 in drug-resistant lines is functional in regulating transcription of its known target genes. Knockdown of OCT4 in drug-resistant lines demonstrated its biological significance as it drastically attenuated tumor growth. This study provides an interesting insight into how prostate cancer cells may function during advanced disease progression and how they may be targeted to increase survival rates during secondary chemotherapy.
Results
Drug-resistant prostate cancer cells overexpress drug efflux pumps and exhibit an increased side population
In our previous publication, we developed drug-resistant prostate cancer cell lines to examine the role of Pim-1 kinases in chemoresistance. 20 These cells developed resistance to docetaxel (DTX) or mitoxantrone (MX) after maintaining clinically relevant doses in culture media. Both of these chemotherapeutic drugs are commonly used to treat castration-resistant prostate cancer. Side population (SP) was first analyzed to characterize these lines and determine what percentage of cells possesses a drug-resistant phenotype (Fig. 1). The SP is defined as the cells that exhibit enhanced efflux of DNA-binding dye Hoechst 33342 dye and is often considered to have similar characteristics to stem cells. 21 As expected, both docetaxel-resistant (R1/DTX) and mitoxantrone-resistant (R1/MX) lines exhibited a significant increase in SP percentage as a result of their constant exposure to drugs. The SPs of R1/DTX and R1/MX after adding drug efflux pump blockers verapamil or Ko143, respectively, were effectively eliminated. Conversely, there was little change in SP when treating R1/DTX with Ko143 or R1/MX with verapamil. Treating the parental CWR-R1 cells with either verapamil or Ko143 had no significant effect on the already small SP. This suggested that docetaxel-resistant cells preferentially utilize MDR1 as their dominant efflux pump, whereas mitoxantrone cells predominantly use BCRP. This observation was confirmed by flow cytometry and immunofluorescence (data not shown), which agreed with current literature regarding known substrates of these efflux pumps. 22,23
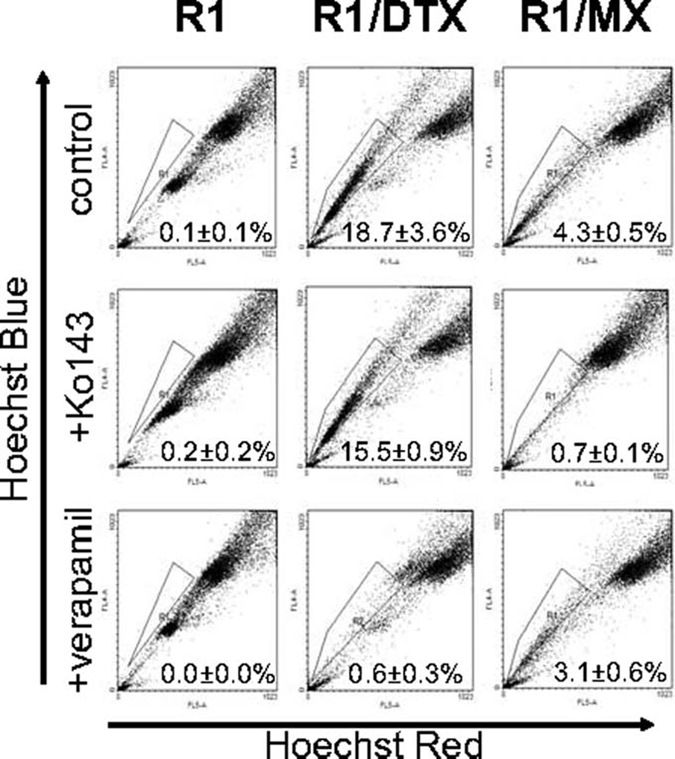
Drug-resistant prostate cancer cells have an expanded side population. Flow cytometric side population analysis was performed as described in Materials and Methods. Verapamil and Ko143 were used as blockers of MDR- and BCRP-expressing populations, respectively. A significant change of SP population in drug-resistant lines (R1/DTX and R1/MX) compared to the parental line CWR-R1 (R1) was detected.
Drug-resistant prostate cancer cells overexpress embryonic stem cell marker POU5F1/OCT4
Microarray analysis was performed to identify unique genes commonly altered in both drug-resistant cell lines compared to their drug-sensitive parental line. Only genes that were altered in both lines at least 2-fold with
Differentially Expressed Genes in Drug-Resistant Cell Lines
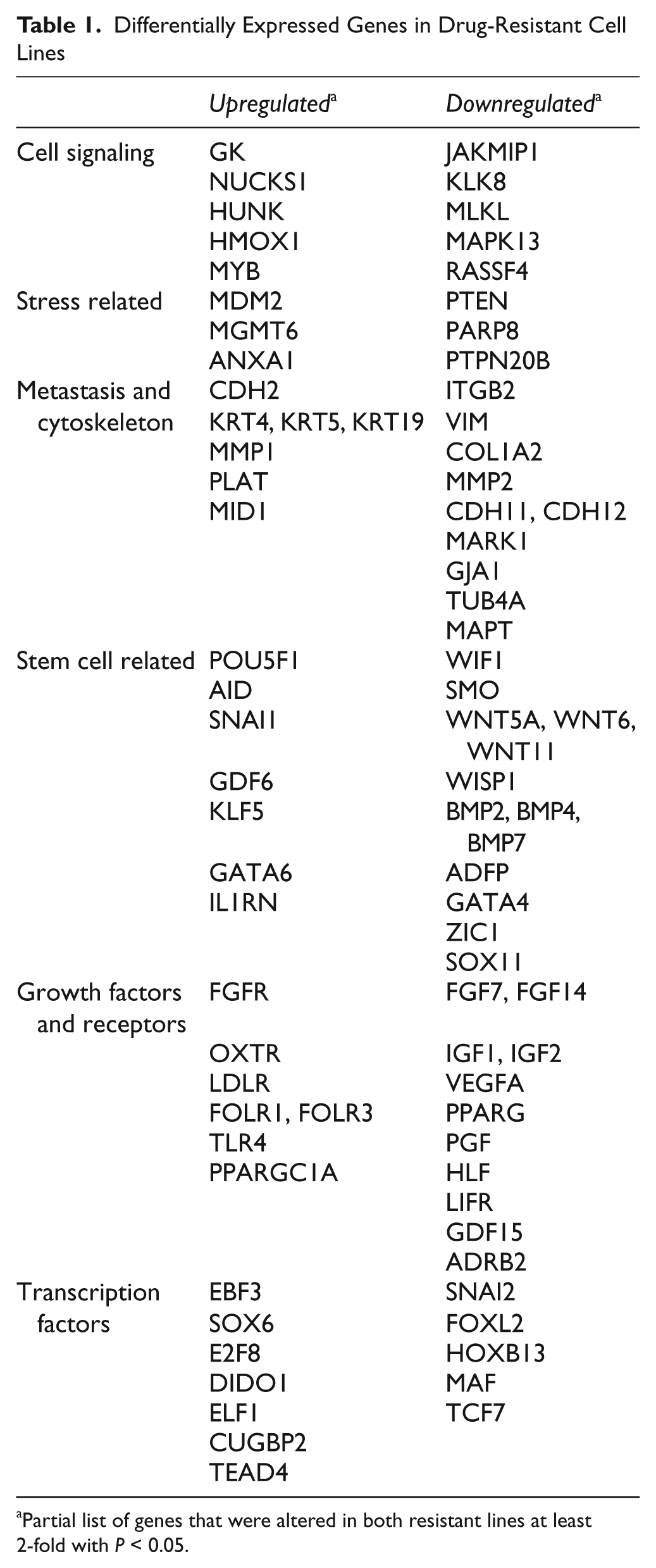
Partial list of genes that were altered in both resistant lines at least 2-fold with
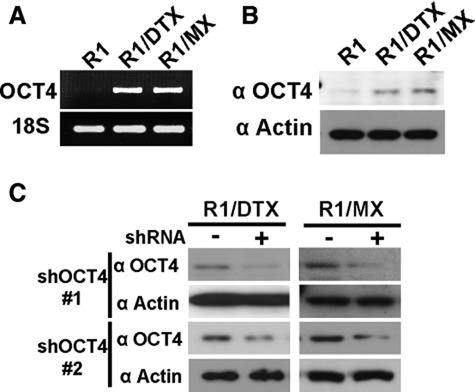
OCT4 expression is upregulated in drug-resistant cells. (
OCT4 is transcriptionally active in drug-resistant lines
Because OCT4 is known to be silenced through genomic methylation within its regulatory regions during differentiation of embryonic stem cells,
24
we proceeded to test whether methylation status of regulatory regions of the
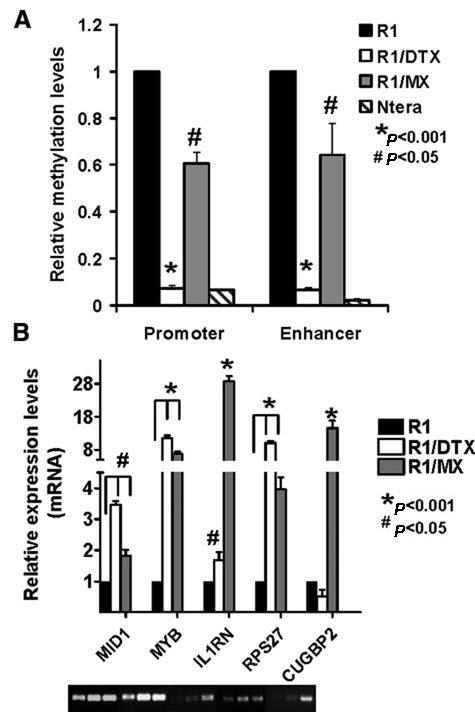
OCT4 is transcriptionally active in drug-resistant cells. (
R1/DTX and R1/MX exhibit enhanced tumorigenicity in vitro and in vivo
To test whether the drug-resistant cells with re-expressed OCT4 protein have tumorigenic growth advantages compared to the parental line, we first examined their ability to self-expand in soft agar assays. As shown in Figure 4A, R1/DTX and R1/MX were more clonogenic than their drug-sensitive parental line. Furthermore, inoculation of as few as 10 R1/DTX or R1/MX cells into castrated male SCID mice was sufficient to form tumors after 8 weeks (Fig. 4B), while the parental R1 cells could not induce detectable tumor growth under similar conditions. These data suggested that the resistant lines are enriched for tumor-initiating cells. We hypothesized some of the genes differentially regulated in these drug-resistant lines, such as those identified in our microarray analysis, may be responsible for their enhanced tumorigenicity.
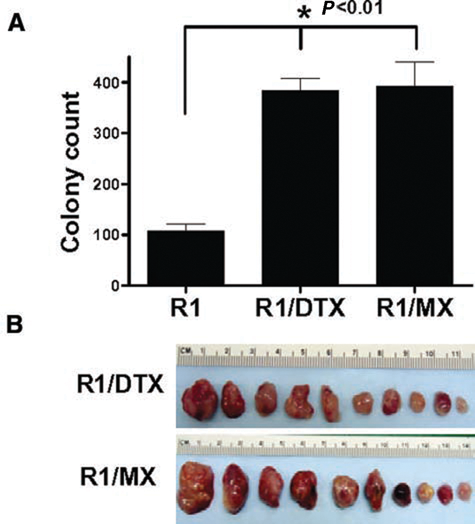
Drug-resistant cells are more tumorigenic. (
Expression of OCT4 plays a critical role in growth and survival of drug-resistant prostate cancer cells
To test whether OCT4 re-expression plays a role in drug-resistant activity in R1/DTX and R1/MX cells, we treated the resistant cells with either control shRNA or OCT4 shRNA and examined the effect of OCT4 knockdown on their growth
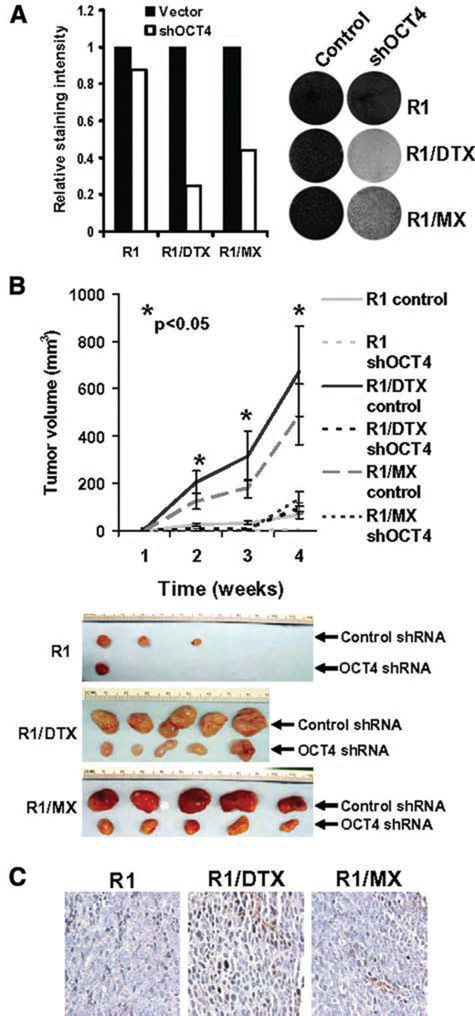
Expression of OCT4 plays a critical role in growth and survival of drug-resistant cells. (
Discussion
Treatment of prostate cancer remains a clinical challenge since there are limited options for patients with castration-resistant tumors. In addition, these patients will inevitably develop resistance to cytotoxic agents used during advanced stage. Our data suggest that this process may enrich more aggressive tumor-initiating cells in recurrent tumors that are increasingly difficult to target. Therefore, a better understanding of mechanisms of drug resistance is needed to improve treatment efficacy for secondary prostate cancer therapy. Identifying pathways underlying chemoresistance to use as targets offers an opportunity for patients to maintain or regain sensitivity to drugs used during combination therapy.
In this study, we have characterized the CWR-R1 cell line and its derivatives, which have developed resistance to docetaxel and mitoxantrone, two different agents commonly utilized to treat advanced recurrent prostate cancer. These drug-resistant cell lines were established by maintaining clinically relevant doses of the chemotherapeutic drugs in culture media over a period of months. As expected by virtue of their acquired drug resistance, these cell lines exhibited an enhanced SP compared to their parental line. Analysis of other tumor types by flow cytometry suggests potential cancer stem cells or tumor-initiating cells may reside within the SP.
27
Soft agar assays and
A number of genes differentially regulated in the drug-resistant lines were identified by microarray analysis. Although DTX and MX have quite different mechanisms of action, both resistant lines shared numerous changes over their derivative CWR-R1. We focused on several stem cell–related genes following SP analysis and found that POU5F1/OCT4 was among those upregulated in R1/DTX and R1/DTX compared to their parental line. Although POU5F1/OCT4 expression in cancer remains controversial, we demonstrated that OCT4 expression was enhanced in both drug-resistant cell lines using multiple analyses including RT-PCR, sequencing, Western blot, and shRNA knockdown. We also showed that the “re-expression” of OCT4 was associated with reduced methylation in the regulatory region of
Consistent with our observations, a recent genome-wide study on prostate cancer showed that
The implications of this study are of great significance to prostate cancer therapy. Unfortunately, the mortality rate of advanced prostate cancer remains high since patients often relapse and become resistant to androgen deprivation therapy. Chemotherapeutic drugs such as taxanes remain a viable option for treatment, but resistance to cytotoxic drugs can become another hurdle to overcome. Therefore, combination treatment targeting both the bulk tumor as well as chemoresistant tumor-initiating cells could provide a greater increase in survival rates. There is a critical need to better understand mechanisms of drug resistance in order to identify novel targets to improve treatment efficacy. Our data suggest that OCT4 may be one such target, and its use may be especially relevant to aggressive drug-resistant cancers.
Materials and Methods
Cell culture and constructs
CWR-R1 lines were cultured in RPMI 1640 medium (Mediatech Inc., Manassas, VA) supplemented with 10% heat-inactivated fetal bovine serum, 1% penicillin/streptomycin, and maintained in a 37°C incubator at 5% CO2. Drug-resistant cell lines were developed as previously described 20 and maintained in medium containing 100 nM docetaxel (Fluka, Sigma-Aldrich, St. Louis, MO) for R1/DTX or 20 nM mitoxantrone (Sigma, Sigma-Aldrich) for R1/MX. Lentiviral constructs expressing OCT4 shRNAs were purchased from Sigma, which contains target sequence CCGGTCATTCACTAAGGAAGGAATTCTCGAGAATTCCTT CCTTAGTGAATGATTTTT or Addgene plasmid 12197 (Cambridge, MA) with target sequence GGATGTGGTCCGAGTGTGGTT.
Side population analysis
The method previously described by us for analyzing side population was utilized.
32
One million cells were suspended in 1 mL DMEM containing 2% fetal calf serum (FCS) and 10 mM HEPES. The cell suspension was incubated with 1 mg/mL Hoechst 33342 (Sigma) at 37°C in the presence or absence of Pgp inhibitor verapamil (50 µM) or BCRP inhibitor Ko143 (1 µM). After 90 minutes of incubation, cells were centrifuged at 4°C and resuspended in cold HBSS with 2% FCS and 10 mM HEPES. Prior to analysis, propidium iodide (2 µg/mL) (Sigma) was added to discriminate dead cells. Samples were then analyzed by BD LSR II 4-laser flow cytometer (BD Biosciences, San Jose, CA). The Hoechst dye was excited at 355 nm, and the fluorescence profile was measured in dual wavelength analysis (405/30 nm and 670/40 nm). The side population was analyzed as mentioned in references.
33,34
Two independent measurements were performed, and a significant change in SP population was determined by the Student
Microarray analysis
Total RNA was collected from cells and analyzed using the RNA 6000 Nano kit on an Agilent Bioanalyzer (Agilent Technologies, Palo Alto, CA). RNA was fluorescently labeled according to the standard protocol designed for the Agilent Human Whole Genome Expression System. Fluorescent dyes (Cye5 and Cye3) were incorporated into the amplified cRNA during a linear amplification step. Approximately 2 µg total RNA from drug-resistant lines was labeled with the Cye5 dye. RNA from the R1 parental cells was labeled with the Cye3 dye and served as a common reference, while the other labeled samples were cohybridized in a classic 2-color hybridization scheme. Hybridization was carried out using the conditions specified by the manufacturer. The Agilent microarrays contained 44k 60mers in sense orientation. Scanned images were processed for quality assessment using the Agilent Feature Extraction software. Preprocessed, normalized expression values were imported into the BRB-ArrayTools (National Cancer Institute, Bethesda, MD) software for visualization and analysis. A list of genes was compiled that composed of statistically significant changes that were least 2-fold increases or decreases in expression.
Real-time PCR
Total RNA was extracted from confluent 100-mm plates of CWR-R1 and drug-resistant cells using TRIzol reagent (Invitrogen, Carlsbad, CA). Approximately 5 µg total RNA was treated with DNase I (Ambion, Austin, TX) and then used for reverse transcription using the iScript kit (Bio-Rad Laboratories, Hercules, CA). PCR amplification of the resultant was carried out using FastStart Taq Polymerase (Roche, Basel, Switzerland), following the program: 10-minute denaturation at 94°C, 35 cycles at 94°C for 30 seconds, 58°C for 30 seconds, and 72°C for 45 seconds. PCR products were run on 1% agarose (Invitrogen) gel with a 100-bp DNA ladder. OCT4 primers used were the following: ACACCTGGCTTCGGATTTCG (forward) and GGCGATGTGGCTGATCTGCT (reverse). Bio-Rad GelDoc imagining system was used to capture images of PCR products separated by 1% agarose gels.
Western blot and antibodies
Immunoblotting was carried out as described previously. 36 Mouse monoclonal anti-OCT4 antibody was from Santa Cruz Biotechnology (sc5279) (Santa Cruz, CA). Monoclonal anti-actin antibody (sc8432) were both from Santa Cruz Biotechnology. All experiments shown are representative of at least 3 experiments.
Soft agar assay
Soft agar assays were set up as described previously. 37 Briefly, approximately 1 × 104 cells from CWR-R1 or its drug-resistant derivatives were plated in a 24-well plate and allowed to incubate for 8 days. Plates were stained with iodonitro tetrazolium blue (2 mg/mL) and quantified using an automated reader from Microbiology International (Frederick, MD) under an inverted microscope. Only colonies measuring between 50 to 200 µm were counted by the reader.
In vivo xenograft models
The tumor growth of CWR-R1 and its derivates in xenograft models was carried out as described previously,
38
using the cell number as indicated in the figures. For tumor initiation experiments, 10 cells suspended in 25 µL medium were mixed with 25 µL of Matrigel (BD Biosciences, Franklin Lakes, NJ) and then subcutaneously (s.c.) injected in flanks of castrated NOD/SCID mice. For knockdown experiments, 104 cells were mixed with 100 µL of Matrigel (BD Biosciences) at 48 hours postinfection and then s.c. injected as above. Tumor volumes were measured weekly and calculated using the formula 0.5236 × r1
2 × r2, wherein r1 < r2. Differences in tumor sizes formed on both flanks were compared by the paired
Immunohistochemical analysis
The Vectastain Elite ABC Kit (Vector Laboratories, Burlingame, CA) was used for immunohistochemical staining according to the protocol supplied by the manufacturer using a monoclonal OCT4 antibody as described previously. 39
Footnotes
Acknowledgements
The authors thank Dr. C.W. Gregory for kindly providing CWR-R1 cells and Dr. Douglas Ross for supplying Ko143 for SP analysis.
The author(s) declared no potential conflicts of interest with respect to the authorship and/or publication of this article.
This work was supported in part by grants from DOD (W81XWH-08-1-0174) and Maryland Stem Cell Research Fund to Y.Q. and DOD Pre-doctoral Fellowship (W81XWH-08-1-0068) to D.E.L.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
