Abstract
Numbers of wild Asian elephants (
Introduction
The Asian elephant (
For genetic variation studies, a good marker is one with high genetic variability and the ability to generate multilocus data from the genome under study (Anne, 2006). Intersimple sequence repeat (ISSR) are segments of DNA that are flanked at both ends by microsatellite sequences. Dominant markers usually produce multiple DNA fragments in a single reaction, yielding a large number of loci across the genome with unknown DNA sequences of the target regions. ISSR markers have a high reproducibility capability compared with random-amplified polymorphic DNA (RAPD), where DNA fragments are amplified by polymerase chain reaction (PCR) using short synthetic primers. It is possible that DNA amplification from ISSR with longer primers and at higher annealing temperatures may be better than those of RAPD, which can result in mismatches between RAPD primer and template, and a total absence of PCR product. Thus, the RAPD results can be more difficult to interpret than ISSR (Askari, Abadi, & Baghizadeh, 2011; Zietkiewicz, Rafalski, & Labuda, 1994). Moreover, ISSR markers are more economical and less time-consuming compared with Amplified Fragment Length Polymorphism, a technique based on selective PCR amplification of restriction fragments from a total digestion of genomic DNA which makes developing locus-specific markers from individual fragments difficult and requires the use of different kits adapted to the size of the genome being analyzed. To date, ISSR marker analyses have been applied primarily to plant genetics (Bentley, Barker, & Dold, 2015; Li, Zhong, Sui, Ji, & Ying, 2013; Ng & Tan, 2015; Tiwari et al., 2015) but are beginning to be used to estimate the genetic diversity and for individual identification in animals, particularly in domestic species like fish, cattle, buffalo, goats, and sheep (Askari et al., 2011; Aytekin, Ozdil, Zulkadir, Boztepe, & Sariyel, 2011; Maltagliati, Lai, Casu, Valdesalici, & Castelli, 2006; Moradi, Rostamzadeh, Rashidi, Vahabi, & Ferahmand, 2014; Nesteruk, Makarova, Svishcheva, & Stolpovsky, 2015; Srinulgray & Piyapattanakorn, 2009; Stolpovsky et al., 2011). Genetic diversity or genetic structure in Asian elephants from different regions of India, Bhutan, Sri Lanka, Laos, Vietnam, and Malaysia has been evaluated based on mitochondrial DNA or nuclear microsatellites (Ahlering et al., 2011; Elliza, Shukor, Othman, & Md-Zain, 2015; Fernando, Pfrender, Encalada, & Lande, 2000; Vidya, Fernando, Melnick, & Sukumar, 2005a, 2005b; Vidya & Sukumar, 2005; Vidya, Varma, Dang, Van Thanh, & Sukumar, 2007). Likewise, genetic diversity in both captive and wild Asian elephants in Thailand has been evaluated on the basis of mitochondrial DNA or nuclear microsatellites (Fickel, Lieckfeldt, Ratanakorn, & Pitra, 2007; Thitaram et al., 2010; Thongchai et al., 2011). Genetic diversity was found to be high for captive elephants reintroduced into protected areas in northern Thailand (Thitaram et al., 2015), and among tourist elephants where there can be significant movement of animals between camps (Thitaram et al., 2010). Moreover, a historical report of captive elephant transportation found considerable movement of elephants among South East Asian countries (Lair, 1997; Thitaram et al., 2010) and selection of tame bulls and cows for tourist purposes.
Today, many captive bulls are subfertile; thus, only one or a few bulls are used for breeding in each camp, with occasional transfer of elephants among camps for breeding (Thitaram, 2012; Thongtip et al., 2004). This type of breeding selection bias might eventually lead to inbreeding in some camps. Although genetic diversity based on microsatellite analysis indicated good diversity among camp elephants in northern Thailand (Thitaram et al., 2010), it has been over 8 years since the study was conducted. It has been further suggested that alternative sources of nuclear genes could be used to enhance the characterization of population structure in Asian elephants. To date, no genetic studies utilizing ISSR analyses have been conducted in this species. Thus, the goal was to conduct a follow-up study to reassess genetic diversity of elephants residing in several camps in Chiang Mai province using ISSR markers to provide more information on elephant genetic health and prevent future inbreeding problems that might occur within individual elephant camps in the region.
Materials and Methods
Sample Collection
Blood samples (2 ml) from 97 captive Asian elephants (78 females and 19 males) maintained at seven elephant camps in Chiang Mai province (Table 1; Figure 1) were collected from an ear vein into 5-ml ethylenediaminetetraacetate tubes. This study was approved by the Animal Use Committee of the Faculty of Veterinary Medicine, Chiang Mai University, Thailand in 2018 (S32/2561).

Distribution of seven elephant camps in the study area by province.
Number of Samples in Each Elephant Camps Which Separated Male and Female.

DNA Extraction
Whole blood samples were used for extraction of genomic DNA using the RBC Bioscience™ Real genomics DNA extraction kit for Blood/Bacteria/Cultured cells (RBC Bioscience Corp., New Taipei, Taiwan) following manufacturer instructions. First, fresh blood in ethylenediaminetetraacetate was mixed with RBC lysis buffer to lyse the cells, and a cell pellet was obtained. Next, proteinase K (10 mg/ml) was added and mixed with the cells by vortexing and then incubation at 65°C overnight until the sample lysate was clear. After incubation, absolute ethanol was added to the sample lysate. Finally, the standard elution volume was reduced from 100 µl to 50 µl to increase the DNA concentration. The concentration, yield, and purity of the DNA sample were determined using a Beckman Coulter DU® 730 spectrophotometer (Beckman Coulter, CA, USA). Once extracted and checked for purity, genomic DNA from the blood cells was kept at −20°C for further analysis.
ISSR Amplification
Twenty one ISSR primers from the University of British Columbia (Microsatellite UBC primer set 9, University of British Columbia, Vancouver, Canada) were screened, resulting in the selection of nine that produced reproducible bands for ISSR amplification in this study (Table 1). All samples were amplified individually by PCR using 25 µl of deionized water, 19.3 µl of 10 mM dNTP mix solution, which contained dATP, dCTP, dGTP, and dTTP, 0.5 µl (Vivantis, Selangor Darul Ehsan, Malaysia), 2.5 µl of 10× ViBuffer S (160 mM (NH4)2SO4, 500 mM Tris-HCl [pH 9.2 at 22°C], 17.5 mM MgCl2, and 0.1% Triton™ X-100), 0.2 µl of 5u Taq DNA polymerase (Vivantis, Selangor Darul Ehsan, Malaysia), 0.5 µl of ISSR primer, and 2 µl of 10 ng template DNA. For every PCR reaction, deionized water was used instead of DNA templates as a negative control. PCR amplifications were performed in PTC-200 The DNA Engine® Thermal Cycler (Bio-Rad Laboratories, Inc., CA, USA) under the following conditions: predenaturation at 95°C for 5 min, followed by 38 cycles consisting of a denaturation step at 94°C for 45s, annealing step at 57°C for 45 s, and extension step at 72°C for 2 min. The last step involved a final extension at 72°C for 10 min. The PCR products were stained by RedSafe™ Nucleic acid staining solution (iNtRON Biotechnology, Gyeonggi-do, South Korea) and then separated electrophoretically on 1.5% agarose gel (PanReac AppliChem ITW companies, Darmstadt, Germany) by MINI GEL II tank electrophoresis (Select BioProducts, NJ, USA) containing 1× Tris-acetate-ethylenediaminetetraacetate (TAE) buffer at 130 V for 40 min and visualized by UV light in a UV Transilluminator gel dock (Major Science, CA, USA) to detect the amplification and record results by digital photography. A 100 bp DNA Ladder RTU (GenedireX, Inc., Miaoli, Taiwan) was used as a DNA marker for measuring DNA size or molecular weight of the PCR products. Bands that were observable after gel electrophoresis were scored independently by two persons at least 2 times for greater precision and confirmed the results from interrater repeatability.
Statistical Analysis
We analyzed results from ISSR amplification products from male and female elephants separately and also from the overall population. Only bands or loci that were clearly observed and unambiguous from the ISSR-amplified fragments were scored in a binary symbol for band 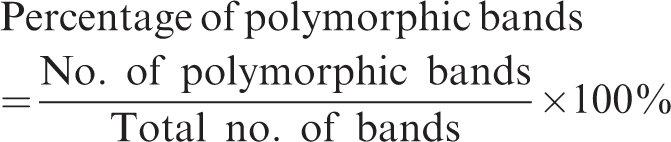
Interpopulation genetic variation in this study was determined by the observed number of alleles (
Results
ISSR Polymorphism
Preliminarily, all 21 primers were screened at three different annealing temperatures (55°C, 57°C, and 59°C), twice in each condition, which found that at an annealing temperature of 57°C, 9 primers in the microsatellite UBC primer set 9 of the University of British Columbia produced informative results that were reproducible. This made them suitable ISSR primers to use as genetic markers to assess genetic variation and differentiation, or fingerprinting, of the 97 captive Asian elephants in this study. Another 12 ISSR markers did not provide informative results due to some optimization problems.
As shown in Table 2, using nine ISSR markers, 88 DNA fragments were produced in male captive Asian elephants, of which 37 (42.05%) were polymorphic; the UBC808 primer produced the most polymorphism (63.64%). In female elephants, 115 DNA fragments were obtained, of which 83 (72.17%) were polymorphic. Again, the UBC 809 primer produced the most polymorphism (91.67%), indicating substantial genetic diversity at the population level.
The Nucleotide Sequences of Intersimple Sequence Repeat Primers, Number of Total Bands and Polymorphic Bands, and Percentage of Polymorphic Bands in Both Male and Female Captive Asian Elephant.

Genetic Variation in Captive Asian Elephants
Genetic variation of male and female captive Asian elephants from each camp is shown in Table 3. The mean observed number of alleles (
Diversity Indices for Intersimple Sequence Repeat Markers Evaluated on Captive Asian Elephant.

Genetic Differentiation Among Elephant Camps
Pairwise Population Matrix of Nei’s Genetic Distance Are Below the Diagonal and Pairwise Population

Pairwise Population Matrix of Nei’s Genetic Distance Are Below the Diagonal and Pairwise Population

Pairwise Population Matrix of Nei’s Genetic Distance Are Below the Diagonal and Pairwise Population

Phylogenetic Dendrograms
Phylogenetic dendrograms of individual Asian elephants identified 27 clusters based on allelic frequency (Figure 2(a)) and showed that each elephant camp differed, with 4, 2, 6, 8, 7, 11, and 3 main clusters observed in Camps A, B, C, D, E, F, and G, respectively (Figure 3). Moreover, the camps divided into two main clusters: Camps D and E and Camps F, C, B, A, and G (Figure 2(b)).

The phylogenetic dendrogram of individual captive Asian elephants (a) and the phylogenetic dendrogram of elephant camps (b).
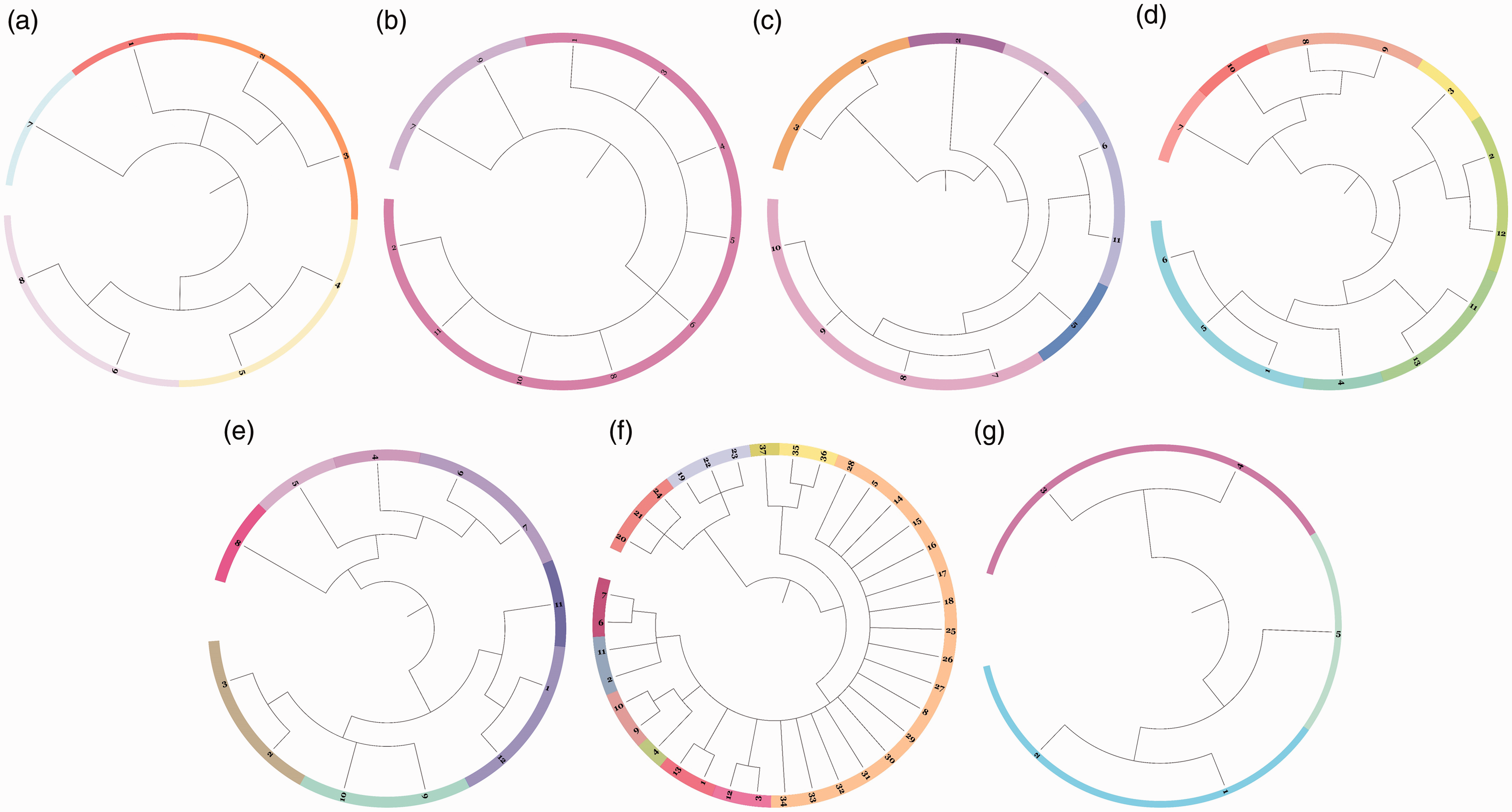
The phylogenetic dendrogram of individual captive Asian elephants in Camp A, B, C, D, E, F, and G (a–g, respectively).
Discussion
This is the first study to apply ISSR marker analysis to assess genetic variation of captive Asian elephants based on the mean allelic frequency and found good genetic variability within and between tourist camps in Chiang Mai, Thailand. The results showed that the expected heterozygosity (He) was high in every elephant camp. This high heterozygosity likely reflects the varied origins of elephants brought into the camps and possibly also the fairly routine transfer of elephants between camps (Luan, Yuh Chiang, & Gong, 2006; Wang, Zheng, Toure, Dandekar, & Kafatos, 2001; Wisely, Buskirk, Fleming, McDonald, & Ostrander, 2002). The Shannon’s information index values, which were 2.126 in females, 1.366 in males, and 2.145 for the entire population, were higher than other studies in domestic animals, such as cattle, sheep, buffaloes, and goat, which range from 0.18 to 0.75 (Askari et al., 2011; Aytekin et al., 2011; Moradi et al., 2014). The percentage of polymorphic bands also showed a high level of variability in female elephants (72.17%), with a lower value (42.05%) for males, but both indicated high genetic diversity across elephant camps in Chiang Mai, similar to a previous genetic study using microsatellite analyses (Thitaram et al., 2010). The genetic variance within captive Asian elephants further showed high levels of genetic variation (percentage of polymorphic bands = 57.08%,
Genetic diversity reflects the total number of genetic characteristics and is related to the number of genes and their alleles within individuals and can influence adaptability and distribution of a species in diverse habitats. Genetic differentiation is the accumulation of differences in allelic frequencies between completely or partially isolated populations due to evolutionary forces and is important for understanding selection or genetic drift in species like the Asian elephant. Fixation index is a classical statistic for summarizing population differentiation, which measures the difference in heterozygosity among populations (Yamamichi & Innan, 2012). A fixation index value above 0.25 indicates high genetic differentiation among a population. The fixation index values in our population were negative, so no genetic subdivision was apparent between the populations. Pairwise
An individual dendrogram was generated using the clustering method of Unweight Pair-Group Method with Arithmetic Mean. This dendrogram indicated that all captive Asian elephants fell into 27 clusters using the nine ISSR markers. In some clusters, only one captive Asian elephant was assigned, which could be due to captive Thai elephants being transported between provinces and also between the countries of Thailand and Myanmar. The finding of many different clusters within camps suggests good breeding management. The elephant camps separated into two clusters: Camps F, C, B, A, and G and Camps D and E, which may also be useful for breeding management and suggests that camps in different clusters should cooperate and transfer genetically valuable bulls or cows for breeding.
The value of genetic diversity in this study was higher compared with studies using other dominant markers, such as RAPD in the Moxoto goat breed population (0.22–0.33) and Amplified Fragment Length Polymorphism for Italian goats (0.21–0.24; Ajmone-Marsan et al., 2001; Olivera et al., 2005). Further studies of genetic variation and differentiation of Asian elephants should be performed with a greater number of individuals and in other areas of the country. We conclude that ISSR markers are an efficient method for discriminating genotype at the molecular level and can successfully be used to estimate the population level polymorphism and genetic diversity. The markers also provided a good comprehension of genetic variability but showed no genetic differentiation among these captive elephants. In this study, of the 21 primers initially tested, only 9 produced clear reproducible fragments or bands. Thus, the suggestion is that a greater number of biologically relevant ISSR primers that produce reproducible unambiguous bands should be applied in future studies to obtain more accurate results. In conclusion, this study showed the potential of ISSR markers to evaluate genetic diversity in captive Asian elephants and proved an attractive alternative to other molecular markers.
Implications for Conservation
Inbreeding and its effects on lowering genetic diversity could negatively impact successive generations of captive Asian elephants. Therefore, information about genetic diversity and variation has become more important in the past decade. This is especially true for elephants in Thailand because there is no documentation of pedigrees similar to the studbooks used in western zoos to guide breeding recommendations. Thus, although this study revealed a high diversity among captive Asian elephants within and between the study camps today, attention will need to be paid in the future to ensure this variability is maintained. This is especially important given the limited availability of breeding bulls. The results of this and future studies can be useful for better management of breeding across camps to further conserve the genetics of these valuable populations.
Footnotes
Acknowledgments
The authors would like to thank all of the owners, managers, and mahouts at the Baan Chang elephant camp, Mae-Taeng elephant camp, Mae-Sa elephant camp, Maetaman elephant camp, Baan Chang Thai elephant camp, Chokchai elephant camp, and Elephant Nature Park for their cooperation. The authors also thank the anonymous reviewers for their suggestion for improving this manuscript and many who colleagues participated in field research and data collection.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed the receipt of the following financial support for research, authorship and/or publication of this article: This work was supported by The Royal Jubilee PhD program, Thailand Research Fund, The Excellence Center in Veterinary Bioscience, and the Center of Excellence in Elephant and Wildlife Research, Chiang Mai University, Thailand.
