Abstract
The use of pesticides, fertilizers, copper compounds, and antibiotics to control microbial diseases in crops has caused adverse effects on human health and the environment, as well as the emergence of multidrug-resistant phytopathogenic bacteria and fungi. In this sense, to minimize reliance on agricultural chemicals and protect the environment, current agricultural practices are integrating life sciences technologies, such as genomics, big data, and artificial intelligence. For instance, the discovery and design of antimicrobial peptides integrate these areas. These peptides have become ideal candidates for promoting sustainable agricultural models. They are small, amphipathic, and polycationic molecules with a complex and diverse mode of action, which makes it difficult for microbes to develop resistance. In this review, we present advances in artificial intelligence and computational approaches for antimicrobial peptide design, along with a pipeline for designing antimicrobial peptides that utilize computational and artificial intelligence strategies extensively employed in our research group. Besides, we summarize advances in peptide research as a promising strategy for sustainable agricultural practices.
Keywords
Introduction
Humans’ survival is always dependent on agriculture. Unfortunately, numerous problems threaten the proper functioning of agriculture, including climate change, land degradation, biodiversity loss, soil erosion, and pollution. All these problems have led to the introduction of the term “sustainable agriculture,” which aims to raise awareness in society about good agricultural practices that ensure the survival of humans and other living beings without adversely affecting ecosystems and the environment. 1 Additionally, the problem of antimicrobial resistance, a common issue recognized in the medical field, also affects the agricultural sector. The use of pesticides, copper compounds, and antibiotics to control microbial diseases in crops has been causing adverse effects on human health and the environment, leading to the emergence of multidrug-resistant phytopathogenic bacteria and fungi.2,3
As an alternative, antimicrobial peptides (AMPs) have been proposed. AMPs are small, amphipathic, and polycationic molecules; most contain between 10 and 50 amino acids in length and are part of the immune systems of most living beings, acting as the first line of defense against bacteria, parasites, viruses, and fungi.4,5 The primary mechanism of action of AMPs is to disrupt the integrity of the plasma membrane of microorganisms, generating an immediate lethal effect. Furthermore, some AMPs exhibit additional mechanisms, such as inhibition of protein synthesis and interference with essential enzymatic functions, providing a multifaceted strategy that hinders microbial adaptation and the emergence of resistance. 6 Despite their small size, AMPs have demonstrated remarkable potential as antimicrobial agents for various applications, including the management of plant diseases. 7 In addition to their ability to combat pathogens, these AMPs possess a wide range of therapeutic properties, including antioxidants and immunomodulatory effects. AMPs also possess low toxicity and short half–lives, which prevent their accumulation in the soil and plant tissues over long periods of time, making them ideal candidates for promoting sustainable agricultural models. 8
Sustainable agriculture practices must incorporate knowledge from diverse areas, including social, biological, economic, and environmental sciences. In this sense, current agricultural practices have been transforming, integrating life sciences technologies like genomics, big data, and artificial intelligence (AI), giving rise to the term “agriculture 4.0”, 9 making clear the need for minimizing the reliance on agricultural chemicals, thereby protecting the environment from pollution and diminishing secondary effects associated with the indiscriminate use of these substances. 10 Specifically, AI is the study of processes and practical aspects that enable the development of computational systems capable of performing intelligent tasks and solving problems. 11 In this sense, AI and its subfields, such as machine learning (ML) and deep learning (DL), are essential tools that enable the design of synthetic molecules, including AMPs.
Currently, many AMPs have been identified, and thousands more have been designed; the latter known as synthetic AMPs, can be designed using computational and AI strategies, leading to the advantage of directing the design toward a more specific antimicrobial activity, like the design for a specific Gram-positive or Gram-negative bacteria, virus, or fungi, as well as improving their physicochemical characteristics to obtain better stability in some fluids, facilitating the future product formulation.12–14 Besides the extensive study of AMPs in the medical field, these studies serve as a basis for understanding their mechanisms of action, design, and applications in other fields, such as agriculture and animal husbandry. In this sense, this review presents recent advances in AI approaches for AMPs design, along with a pipeline for design using computational and AI strategies extensively employed in our research group. Additionally, we summarize advances in peptide research as a promising strategy for sustainable agricultural practices.
Antimicrobial Peptides (AMPs)
AMPs are small amphipathic and polycationic molecules, and most are between 10 and 50 amino acids in length. The majority of AMPs exhibit isoelectric points above 7, are enriched in positively charged amino acids, display pronounced cationic properties (+1 to +9), and have a relative hydrophobic moment higher than 50%.15,16 Naturally, AMPs are part of the immune systems of most living beings, acting as the first line of defense against bacteria, parasites, viruses, and fungi, representing a reservoir of potentially bioactive compounds known as peptides chemical space. 17 AMPs were discovered in the early twentieth century. In 1922, Alexander Fleming discovered lysozyme, considered the first AMP recognized. Then, in 1939, René Dubos isolated gramicidin from Bacillus brevis, which exhibited high efficacy against Gram-positive bacteria. Later, several small peptides with broad–spectrum antimicrobial activity were identified. However, the big protagonism started in the 1980s with the discovery of defensins in mammals, magainins in amphibians, and cecropins in insects. This interest continued, and in the 1990s, AMPs from insects, plants, and microorganisms were isolated.18,19
According to their origin, AMPs can be synthetic or natural. Natural AMPs are classified according to their site of synthesis within cells (ribosomal or non-ribosomal), their net charge (cationic or anionic), their structure (α-helix, β-sheet, a mixed αβ structure, and non-αβ structure), and their organism of origin. Naturally, most AMPs are encoded by genes, produced through secondary metabolism, or synthesized by non-ribosomal synthases, while others are active after post-translational modifications.13,20 Ribosomal peptides originate from genes and are typically produced through the enzymatic cleavage of precursor proteins. In contrast, nonribosomal peptides are synthesized by complex, multi-domain enzymes known as NonRibosomal Peptide Synthases (NRPSs). In bacterial systems, these NRPSs are generally encoded within a single operon, whereas in eukaryotic organisms, they are found within gene clusters. Each operon or gene cluster typically directs the biosynthesis of a single peptide. 21
Structurally, most natural AMPs fall into two categories: linear forms or non-linear forms stabilized by disulfide bonds. The non-linear peptides typically adopt β-sheet conformations, with one or more disulfide bridges providing structural stability. Breaking the disulfide bonds in these peptides drastically diminishes their antimicrobial effectiveness. Meanwhile, linear AMPs vary considerably in chain length, hydrophobicity, and overall charge distribution. Most studies on this group were conducted with amphipathic α-helical peptides. For instance, incorporating proline or substituting D-amino acids reduces their antibacterial activity. In contrast, when several D-amino acids are incorporated, the resulting diastereomers exhibit high antibacterial activity while being less cytotoxic to mammalian cells. 22
Regarding the organisms’ producers, AMPs are known as defensins (produced by animals, including humans), cecropins (produced by insects), cathelicidins (produced specifically by mammals), magainins (amphibians), clavanins (tunicates), caenopores (nematodes), penaeidins and crustins (crustaceans), and bacteriocins (produced by bacteria). Furthermore, plants produced several AMPs against different pathogens (ie, bacteria, fungi, and herbivorous pests) known as thionins, cyclotides, and defensins. Various fungi can also produce AMPs, such as several peptaboils like plectasin in the case of Pseudoplectania nigrella.4–6,18,23,24
Mechanisms of Action
In comparison to the cell membranes of multicellular plants and animals, microbial membranes are negatively charged due to the display of negatively charged phospholipid headgroups to the cell's external environment, like phosphatidylglycerol. Meanwhile, the outer leaflet of the cell membranes of multicellular microorganisms presents zwitterionic phospholipids such as phosphatidylcholine and segregates phospholipids with anionic head groups like serine on the inner face. 25 This difference makes AMPs’ mechanism of action unique and selective, involving their attraction to membranes through opposite electrical charges, followed by hydrophobic interactions. 13 This interaction destabilizes the membranes, leading to rupture and subsequent cell death. Specifically, in Gram-negative bacteria, which are exceptionally proficient at developing resistance and highlighted as priority pathogens by the World Health Organization (WHO), 26 AMPs initially interact with the anionic lipopolysaccharide outer membrane, disrupting its structure and gaining access to the inner membrane. Meanwhile, in Gram-positive bacteria, AMPs interact with the anionic teichoic acids that integrate the proteoglycan envelope accumulating on the surface and then gain access to the inner cytoplasmic membrane. In the case of fungi, AMPs can inhibit or kill by enhancing fungal cytolysis, interacting with the fungal cell membrane and components of the fungal cell wall to damage its structure and function. For instance, phosphatidylcholine is an abundant component of the fungal cell wall that exhibits anionic characteristics. These mechanisms are described in Figure 1A. Besides, AMPs can interfere with the synthesis of the fungal cell wall, interacting with mitochondria and influencing metabolic processes.15,25,27,28 On the other hand, antiviral peptides primarily interact with viral receptors in host cells, thereby affecting viral attachment and entry; others target glycoproteins on the viral envelope, leading to the inactivation of the virus. 29

The interaction mechanism between AMPs and cell walls and membranes. A. The specific interaction with Gram-positive and Gram-negative bacterial cell walls and fungal cell walls. B. The four interaction models between AMPs and membranes. The toroidal-pore model, the barrel-stave model, the carpet-like model, and the aggregate model. Created in BioRender. Blandon, L. (2026) https://BioRender.com/bp2ul3e
Four models describe the interaction between AMPs and membranes: the toroidal pore, barrel–stave pore, carpet, and aggregation model. These are described in Figure 1B. In the toroidal model, the hydrophilic and hydrophobic regions of the cell membrane bind to the polar and nonpolar regions of the AMPs, respectively, thereby opening a pore. In the barrel stave model, the AMPs aggregate and insert into the microbial membrane like a barrel stave, in the carpet model, electrostatic interactions between the membrane and the anionic phospholipid head group cause the AMP to envelop the membrane surface in a carpet-like shape, releasing nanometric-size vesicles, and in the aggregate model unstable channel structures are generated that permit the direct transit of ions and larger molecules, without inducing substantial depolarization or compromising membrane integrity.13,15 However, the type of interaction model depends on the peptide structure; in this sense, AMPs with helical structures tend to adopt the carpet model, while β-sheet AMPs tend to adopt either a barrel-stave or toroidal pore model. While extended/random coil AMPs don’t present a defined secondary structure, they don’t have a defined mechanism of action. In this sense, they can disrupt cell membranes, interact with lipid bilayers, or penetrate microbial cells without damaging the membrane and thereby affect cellular processes, such as protein, DNA, or RNA synthesis.8,30 An example case is proline-rich AMPs (PrAMPs), which can pass through the membrane without causing damage, but can be lethal for bacteria by inhibiting protein synthesis. 31 According to the latest, some AMPs can enter cells directly or via endocytosis. Once inside the cells, they can block DNA replication, RNA synthesis, protein synthesis, hinder cell wall synthesis, and hinder cell division. Besides, AMPs can interact with mitochondria, affecting ATP efflux, oxidation of phospholipids and macromolecules, and mitochondrial respiration, thereby damaging mitochondria and plasma membranes and ultimately leading to cell death. 15 The main AMPs mechanisms of action inside the cells are described in Figure 2.

Intracellular targets of AMPs independent of membrane disruption. Created in BioRender. Blandon, L. (2026) https://BioRender.com/e50qh96
Another activity is the antibiofilm activity. Naturally, bacteria can be found as free cells (planktonic) or forming multicellular communities called biofilms. The biofilms enable bacteria to remain sessile in specific environments, such as hydrothermal hot springs, freshwater rivers, or deep-sea vents. In biofilms, bacteria are encapsulated in a matrix composed of polysaccharides, extracellular DNA, lipids, proteins, and water. This matrix makes them resistant to physical and chemical factors, including UV damage, metal toxicity, and antibiotics, among others. In general, AMPs can affect biofilms by several methods, such as altering the homeostasis of the intracellular environment, inhibiting microbial cell adhesion, and down-regulating the extracellular matrix.32,33
Due to this complex and diverse mode of action, the development of resistance to AMPs is slower compared to conventional antibiotics. Documented cases exist, primarily in clinical settings, highlighting the need for prudent design and use. Resistance development can occur due to a mutation that changes the composition or lipid organization of the cell membrane or the cell wall composition.34,35 Besides, some bacteria have acquired resistance by producing specific proteases such as the “omptins” of enterobacteria. 36
Artificial Intelligence (AI) and Antimicrobial Peptides (AMPs)
AI is the study of processes and practical aspects that enable the development of computational systems capable of performing intelligent tasks. 11 It unites data sets with computational sciences to solve problems and make decisions. 37 To achieve this purpose, AI employs algorithms and mathematical models and has been used in peptide discovery and design. Specifically, AI and computational methods have emerged as a new approach to antibiotic discovery, bringing the search for new antibiotics into the virtual realm. 38 In AMP discovery and design, AI is utilized on two fronts: for the de novo design and for predicting biological activity. For these purposes, the main methods of AI include ML, DL, genetic algorithms (GA), linguistic models (LM), and pattern insertion algorithms (PI). 39
Rational Design of Antimicrobial Peptides (AMPs)
Rational design involves creating peptides with high antimicrobial activity and a range of properties that enhance functionality in various application fields, including low toxicity and high stability. Structure-based design has historically been the first method used to computationally design bioactive peptides. This method is based on the knowledge of the 3D structure of the targeted site of action, which leads to the design of new peptides by modeling their interactions with this site using docking and molecular dynamics. Structure design has been primarily based on helical AMPs, as their properties can be mapped out and studied using helical wheels, as well as the polar and nonpolar phases, allowing for the easy visualization of the distribution of charged amino acids. Additionally, chemical modifications, motifs, and features can be introduced into the parent sequences of helical peptides, thereby improving their antimicrobial activity. Conversely, this type of design is limited mainly because of its applicability to only one specific structure type. Another approach in peptide design is molecular dynamics, which can be used to obtain key peptide features. Its main principle of simulation is based on the classical Newtonian equations of motion for all the atoms in a system.40,41
Currently, computational and AI methods have been developed, along with various pipelines tailored to the specific aims of each study or purpose. In general, these methods can be classified into two categories: those that start from a known base or template, such as a proteome, a known peptide, or genomic data, and the de novo methods, which aim to create AMP using only sequences and patterns. In the first case, peptides can be designed using physicochemical approaches or a known sequence template. Physicochemical approaches are widely employed in the design of α-helical peptides; the most common are hydrophobicity, charge, and the hydrophobic moment. The use of a known sequence template in the design aims to generate novel AMPs from a known sequence, which may or may not be active, with the goal of reducing size, improving selectivity, and increasing antimicrobial activity. Whereas de novo methods can generate several sequences with no apparent relation using amino acid patterns or frequencies.42,43 In this type of design, LM and PI algorithms are widely employed. LM uses regular grammar rules to generate a new peptide sequence by considering each amino acid as a word and forming a phrase. 43
Another approach is the use of GA, which is a type of evolutionary algorithm that can be used for peptide optimization. It involves modifying the amino acid sequence to enhance its activity and stability. GAs are inspired by natural selection and genetics. Their main components are representation, initial population, and fitness function. Briefly, the representation denotes the problem, and the GA creates the initial population as a random candidate solution, and the fitness function evaluates the best solution. In this sense, for a determined peptide, the fitness function can be based on the calculation of specific properties like isoelectric point, amphiphilicity, and secondary structure, among others. 17
Machine Learning (ML)
ML is a subfield of AI that involves algorithms that are trained with datasets. Algorithms used in ML are classified as supervised, unsupervised, and semi-supervised. Supervised ML algorithms are trained with data containing the outcome of interest; some of the most used for designing and development are random forest models (RF), support vector machine (SVM), and fuzzy K-nearest neighbor (FKNN) regression. Unsupervised ML algorithms detect hidden outputs in unlabeled data. These algorithms use dimensionality reduction tests, hierarchical and k-means clustering, such as non-negative matrix factorization (NMF), principal component analysis (PCA), and t-distributed stochastic neighbor embedding (t-SNE). Alternatively, a semi-supervised algorithm is a hybrid of supervised and unsupervised. 17
In AMP design and properties prediction, supervised algorithms have been extensively used. 17 In contrast to physical models, ML models are data-driven, meaning they infer a relationship between two factors or properties (X and Y) through statistical learning using characterized experimental databases. In this sense, a supervised learning algorithm must be trained to map an input to a specific output. The input can be molecular fingerprints, peptide sequences, descriptors, structure-based features, or a combination of these. The output typically consists of a label class, such as active or non-active, a sequence representing a synthetic AMP, and a value indicating the peptide's biological activity. Specifically, ML models for activity prediction are framed within the quantitative structure–activity relationship (QSAR) models. Some of the ML algorithms implemented in AMPs classification and prediction models are: k-nearest neighbor, naïve Bayes, hidden Markov models (HMM), SVM, neural networks (NN), zero-shot learning (ZSL), and RF.44–46
For the de novo design, several DL approaches have been developed, mainly based on structural aspects. DL is a subset of ML that uses different types of neural networks. Some of the neural networks used in DL are convolutional neural networks (CNNs), artificial neural networks (ANNs), recurrent neural networks (RNNs), graph convolutional networks (GCNs), and the most used for the design of AMPs variational autoencoders (VAEs), and generative adversarial networks (GANs),39,46 for instance, Deep – AmPEP30 developed by Yan et al in 2020, 47 involves the prediction of short peptides (≤30 amino acids) using DL combining CNN and Reduced Amino Acid Composition strategy (RAAC) which clusters the amino acids based on evolutionary information. On the other hand, an alternative DL strategy for the de novo design of AMPs involves the application of generative models. A recent example is AMPGen, introduced by Jin et al in 2025, 48 which integrates evolutionary information and a diffusion-driven generative model to generate novel AMP sequences. AMPGen is composed of three main components: a generator, a discriminator, and a scoring module. The generator utilizes a pre-trained autoregressive diffusion model that captures protein evolutionary information from multiple sequence alignments. Then, the sequences generated are filtered with an XGBoost-based discriminator using physicochemical properties, and a final score is obtained using Long Short-Term Memory (LSTM), a type of RNN. Another recent approach using a DL-based RF diffusion method for the de novo design was reported by Vázquez Torres et al in 2025, 49 who designed proteins with thermal stability and high binding affinity that neutralized a lethal neurotoxin from the 3FTx snake venom family. A summary of the main AI fields used for AMPs design is shown in Figure 3.

Summary of the main artificial intelligence (AI) fields used for the antimicrobial peptides (AMPs) design. Created in BioRender. Blandon, L. (2026) https://BioRender.com/jlp65ig
Databases, Prediction, Design, and Classification Tools
According to the antimicrobial peptide database (APD) in its sixth version (https://aps.unmc.edu/, accessed January 2026), there are more than 5000 AMPs, including natural, synthetic, and predicted. 50 This has led to the creation of databases with all this information. These databases also offer search, analysis, and biological activity prediction tools, assisting in the identification, prediction, and design of new AMPs.13,14 These databases can be divided into three categories: i) databases containing peptide sequences with proven biological activity, ii) databases that provide not only the peptide sequence but also prediction and design tools, and iii) databases with tools for identifying possible AMPs encoded in proteomes or genomes. The most commonly available databases are described in Table 1.
Accessible Databases, Prediction, Design, and Classification Tools Dedicated to Antimicrobial Peptides (AMPs) (Accessed January 2026).

The first and most crucial peptide database is APD, which started in 2003. The first APD version started with 525 peptide entries. 62 Since 2011, APD has primarily cataloged natural peptides that have been proven to have antimicrobial activity. In 2025, APD evolved to its sixth version, which includes the current consolidated database, a broadened design, property prediction, functional options, and an innovative pipeline for peptide information. Besides, the APD contains information on peptide activity, toxicity, structure, post-translational modifications, mechanism of action, recombinant production, and clinical trials. To be included in the APD6 database, the AMPs can be natural, synthetic, or predicted and must fulfill the following conditions: first, their amino acid sequence must be known; second, they must exhibit verified antimicrobial properties; third, the peptides should comprise a maximum of 200 amino acids. 50
In the following years after the APD's appearance, other manually curated databases have been created. One is the Data Repository of Antimicrobial Peptides (DRAMP), which contains a rich collection of AMPs sequences, a total of 30,260 structures in its last version, including activities, physicochemical attributes, patent status, clinical relevance, and reference information. 59 In 2016, Singh et al created an integrated database containing peptides collected from 20 databases and two peptide datasets available in the public domain, known as “A database of Structurally Annotated Therapeutic Peptides (SATPdb). 61 Another database is The Database of Antimicrobial Activity and Structure of Peptides (DBAASP) that provides the sequences of AMPs, along with information on chemical modifications, 3D structures, biological activity, and toxicities. In its latest version (3.0), it features over 15,700 entries. On the other hand, the Collection of Antimicrobial Peptides (CAMPR4) provides curated information on natural and synthetic AMPs. In its last version, it includes separate prediction algorithms for natural and synthetic AMPs. It contains approximately 24243 sequences, 933 structures, 2143 patents, and 263 AMP family signatures. Besides, it provides tools for rational design, AMP prediction, sequence alignment, and pattern creation. 56 On the other hand, the database dbAMP contains a total of 33,065 AMP sequences and has been curated from over 5200 publications. It also includes predicted three-dimensional structures of AMPs and contains an efficient tool for predicting the half-life of AMPs and their hemolytic toxicity, along with an AMP optimization platform and an AMP finder, a tool that searches for potential AMPs in proteomes and genomes. 57 Others, like PepMultiFinder, 60 serve to search for possible AMPs in proteomes, and the AntiBP2 server helps identify antibacterial peptides in a protein sequence. 52
Some databases are more specific, like InverPep, a database of invertebrate antimicrobial peptides that contains 702 experimentally validated AMPs from invertebrate species with their source, secondary structure, physicochemical properties, and biological activity. 23 In addition to databases, some servers have been developed with different types of algorithms to predict the specific biological activity of a determined peptide sequence, like ClassAMP, AmpClass, and AMP Scanner. The last version of the AMP Scanner has been developed with a deep neural network model.29,45,51 Other web servers seek to evaluate a more specific activity, like Antifp and AVPdb. Antifp defined as a prediction server for antifungal peptides, was developed to predict and design antifungal peptides, with a dataset consisting of 1459 antifungal peptides. 53 Meanwhile, AVPdb was developed to target about 60 medically important viruses. It is a manually curated database that contains experimentally verified 2683 AVPs, including 624 modified AVPs. 54 Besides, dbAMP provides tools for the prediction of antibacterial and antiviral activity. 57 On the other hand, BAGEL is a mining server to identify gene clusters involved in the biosynthesis of bacteriocins and Ribosomally synthesized and Post-translational modified Peptides (RiPPs) from bacterial (meta-) genomic DNA. 55
A proposed pathway for the rational design of AMPs starting from proteomes is described in the following section, along with a step-by-step description, and proposes the use of certain databases and their tools.
A Proposed Pathway for Rational Design
Nowadays, high-throughput sequencing technology has opened the door to exploring new AMPs. Biological data, such as proteomes, genomes, and metagenomes, can serve as the basis for identifying new AMPs. Specifically, the encrypted bioactive peptides in precursor or mature proteins have been documented.
63
For instance, milk fermentation with proteolysis is an efficient approach to obtain bioactive peptides.
64
Besides, this digestion can also be made in silico, and extinct proteins can be mined as a promising source of AMPs.65–67 In this section, a pathway for the rational design of AMPs is proposed, beginning with proteome scanning. This procedure is summarized in Figure 4.
Data Mining:
One of the most common approaches is to start with a proteome. Proteomes from all the natural kingdoms can be obtained from the Universal Protein Resource (Uniprot; https://www.uniprot.org/), a leading global data resource for protein sequence and functional information. It unites the information contained in the three central protein databases (TrEMBL, SwissProt, and Protein Information Resource (PIR)). In general, Uniprot is composed of the Uniprot Archive (UniPArc), a historical collection of protein sequences, the Uniprot Reference Clusters database (UniRef), and the Uniprot knowledge base (Uniprot-KB). Protein sequences are stored in the Uniprot knowledgebase, which is divided into UniprotKB / Tr EMBL, containing the automatically annotated sequences, and Uniprot KN / SwissProt, containing manually curated and annotated sequences. In 2022, Uniprot was awarded the Global Core Biodata Resource status (https://globalbiodata.org).
68
The National Center for Biotechnology Information (NCBI) maintains another important protein sequence database. This is a compilation of entries from other databases like Swissprot, the Protein Data Bank (PDB), the PIR database, protein translations of the GenBank database, among others.
69
Scan the proteomes for antimicrobial peptides candidates’ sequences:
Once the proteomes of interest had been selected, they could be analyzed to obtain AMPs sequence candidates with tools like PepMultiFinder.
60
This tool enables the inference of possible AMPs sequences based on their physicochemical properties. The most common physicochemical properties to be considered to obtain AMP candidates with PepMultiFinder are the overall charge, isoelectric point, percent hydrophobicity, Boman index, and the polar angle. Values within the following intervals are recommended
70
:
Charge from +2 to +5 Hydrophobicity from 40% to 65% Boman index −1 to +1 Polar angle from 100 to 160 ° Isoelectric point from 9 to 14 Besides, if the aim is to obtain α-helical peptides, it is recommended to exclude cysteine to prevent disulfide bond formation. Aspartic acid and glutamic acid could be avoided because their presence can reduce the positive charge of the resulting peptides.
Antimicrobial activity prediction:
The prediction systems used to evaluate the antimicrobial activity, as mentioned before, are based on ML models. One of the most used databases is CAMPR4 that contains a prediction tool that enables the users to do three tasks: i) prediction of antimicrobial activity by four different models, ii) prediction of an antimicrobial region within a peptide sequence, and iii) generation of a combinatorial list of sequences from a defined sequence and the prediction of the effect of a single residue substitution on the antimicrobial activity.46,71 Other databases were described in the previous section and in Table 1.
Structural optimization of peptide sequences:
Structural optimization is a crucial step to address the challenges of limited stability and reduced activity in natural peptides. Various structural optimization approaches have been established, such as amino acid substitution, cyclization techniques, and the design of peptide mimics. First, it is essential to consider that the initial interaction between AMPs and microbial membranes occurs through electrostatic and hydrophobic interactions. In this sense, a positive charge is crucial for initial attraction to the negatively charged membranes. Then, the hydrophobic bulk guides the insertion of the AMPs into the membrane, facilitating subsequent disruption or damage. So, the increase in hydrophobicity and cationic nature may enhance the antimicrobial activity.
12
Regarding substitution, replacing amino acids is one strategy; for instance, increasing the presence of lysine and arginine makes AMPs more cationic. The presence of tryptophan leads to the formation of amphipathic helical structures.
72
Once the peptide sequence is optimized, the researcher can return to step 3 to evaluate if the antimicrobial activity was improved. Then, other biological activities, such as cytotoxicity, can also be evaluated to confirm the low toxicity of the designed AMP. Additionally, modifications can be visualized using tools such as Heliquest,
73
Type–Peptide,
74
and Alphafold.
75

Procedure proposed for the rational design of antimicrobial peptides (AMPs) using proteomes as templates. Created in BioRender. Blandon, L. (2026) https://BioRender.com/vewgenk
Antimicrobial Peptides (AMPs) in Agriculture
According to the causal agent, plant diseases can be infectious or non-infectious. Infectious plant diseases can be caused by microbial agents. 76 Pesticides, fertilizers, copper compounds, and antibiotics commonly threaten these diseases. Unfortunately, the biomagnification of these compounds affects the health of both animals and humans,77,78 and in most cases, no effective sustainable control treatments and measures have yet been disclosed. Specifically, the use of antibiotics has led to the emergence of multidrug-resistant microorganisms.2,3,78
The central plant pathogenic bacteria genera are Clavibacter, Agrobacterium, Pseudomonas, Erwinina, Streptomyces, Xyllela, Ralstonia, Dickeya, Pectobacterium, and Xanthomonas.76,79 Several species of these bacteria are the causative agents of diseases that affect economically significant plants. For instance, Pseudomonas syringae pathovars are the causal agents of the bacterial speck of tomato, the bleeding canker of horse-chestnut, and bean halo blight. Xanthomonas species, like Xanthomonas campestris pathovars, cause diseases of agronomic importance, such as the black rot of crucifers (Xanthomonas campestris pv. campestris), and the cassava bacterial blight, a disease endemic in tropical and subtropical areas caused by Xanthomonas axonopodis pv. manihotis. Erwinia amylovora causes fire blight disease of quince, raspberry, apple, pear, blackberry, and many wild and cultivated rosaceous ornamentals. Xylella fastidiosa is the causative agent of almond leaf scorch disease, citrus variegated chlorosis, and Pierce's disease of grapevine.79–81 On the other hand, some fungi like those belonging to the genera Geotrichum, Fusarium, Alternaria, Botrytis, Sclerotina, Collectotrichum, Magnaporthe, Penicillium, Puccinia, Aspergillus, Blumeria, Cochliobolus, Micosphaerella, and Rizhopus, as well as oomycetes, are among the most devastating.82–84 For instance, the Fusarium oxysporum species infects more than 100 different hosts, such as banana, melon, tomato, and cotton, 84 and Magnaporthe oryzae is responsible for rice blast, the most devastating disease affecting rice crops.84,85
Plants express various AMPs, either constitutively or in response to microbial attack, which are active mainly against phytopathogens. These AMPs are cysteine-enriched compounds positioned in plant tissues most vulnerable to infection, including roots, leaves, and seeds. Their structures encompass α-helical and β-sheet conformations as well as exceptionally stable cyclic arrangements. 86 Plants’ AMPs can be classified into different families based on their tertiary structure, including thionins, snakins, hevein-like AMPs, defensins, lipid transfer proteins (LTPs), knottins, and cyclotides. 13 Nevertheless, the situation previously described leads to the search for other molecules and alternatives for plant protection. In this sense, AMPs from other sources are considered a promising choice due to their low toxicity, high efficiency, and short half-lives, which prevent their accumulation in the soil over long periods of time. The following sections aim to provide a brief overview of the research conducted since 1980, starting with natural AMPs, which have served as a known template for the design of synthetic analogs or derivatives. The use of AI for design is also discussed. The most well-known peptides with a significant role in plant protection are summarized in Table 2.
Antimicrobial Peptides (AMPs), Derived, Analogs, Hybrids, and Synthetics, with Proven Significant Biological Activity (Minimum Inhibitory Concentration (MIC) < 20 µM) Against Phytopathogenic Bacteria and Fungi.

acetylation (Ac), tosylation (Ts), benzoylation (Bz), and benzylation (Bn).
Cecropin and Melittin Derivatives, Analogs, and Hybrids
The interest in the application of AMPs against phytopathogenic bacteria and fungi started with the discovery of the natural antibacterial peptides cecropin and melittin. These peptides, as well as their derivatives, analogs, and hybrids, were the first tested against these microorganisms. Cecropins play a crucial role in the immune response of insects 92 ; meanwhile, melittin is a hemolytic peptide found in bee venom. 101 Cecropins comprise three forms A, B, and D, and were initially isolated from the hemolymph of the cecropia moth Hyalophora cecropia. Cecropin A and B were first reported to have antibacterial activity against pathogens of clinical interest. 102 Then, studies were conducted against bacteria and fungi, not only with natural peptides but also with derivatives and analogs designed to enhance antimicrobial activity, reduce size, and prevent degradation by proteases. For instance, in 1992, Andreu et al 103 synthesized short cecropin A-melittin hybrids, finding that these hybrids exhibited more antibacterial activity against clinical pathogens compared to the natural ones. Then, in 1998, Cavallarin et al 92 synthesized cecropin-A-derived (Pep2 and Pep3) and cecropin A-melittin hybrid peptides (Pep1 and Pep4), which are smaller than the natural cecropin A. Those were tested against Phytophthora infestans, Fusarium moliniforme, and F. orxysporum, finding that P. infestans was the most sensitive to the shortened cecropin A peptides. The same authors also discovered that the bioactivity of the peptides was influenced by protein extracts from the leaves of tobacco (Nicotiana tabacum) and tomato plants (Solanum lycopersicum), with the highest tolerance observed in extracts from tomato plants, suggesting that tobacco should not be used for the transgenic expression of cecropin A genes. Then, in 2006, Ferre et al 95 tested Pep3 and 22 analogs against E. amylovora, X. vesicatoria, and P. syringae, finding growth inhibition and bactericidal effects at low micromolar concentrations. Specifically, one of the analogs (BP76) showed the highest bactericidal activity, the lowest cytotoxicity, and the lowest susceptibility to protease degradation compared to Pep3. In 2009, the same authors continued their research with Pep 3 and its analogs, testing their antifungal activity against F. oxysporum, P. expansum, A. niger, and R. stolonifer, and found the most active compounds to be BP21 and BP34. Also, these peptides showed sporicidal activity against P. expansum conidia. 82 On the other hand, the first report on the use of a specific peptide against a plant virus was in 1995, made by Marcos et al 104 who synthesized analogs of melittin inhibitors of the tobacco mosaic virus.
The antimicrobial activity of cecropin B and its derivatives against agriculturally relevant microorganisms has also been extensively studied. For instance, in 1992, Nordeen et al 87 tested cecropin SB-37 derived from cecropin B against bacterial phytopathogens that affect tomato, potato, tobacco, sugar beet, and soybean plants. Also, they tested the peptide against protoplast cell lines from some of these plants, finding a big difference between the lethal concentration against bacteria and the protoplasts, being the first minor, suggesting that the introduction of a modified cecropin B gene into the plants will be secure. Then, in 1993, Jaynes et al 88 used the binary vector pB1121 to express cecropin SB-37 and Shiva - I, both derived from cecropin B, in transgenic tobacco plants, finding that the disease severity caused by a virulent strain of P. solanacearum over the first transgenic progeny plants expressing Shiva - I was reduced. In the same year, Jia et al 88 obtained transgenic potato plants expressing cecropin B, and some plants showed resistance against P. solanacearum. In 1995, Florack et al 105 . transformed tobacco plants with three different cecropin B encoding genes, finding that plants harboring a full-length cDNA–derived construct with the insect signal peptide had the highest cecropin B mRNA levels. Nevertheless, the cecropin B peptide wasn’t detected in any of the transgenic plants, probably because of the plant's endogenous proteases. In 1997, Owens and Heutte 90 synthesized a cecropine B structural analogue (MB39) and compared the susceptibility of cecropine B and MB39 to protease degradation in vitro using leaf intercellular fluid from various crops. They found that the half-life of the analogue was higher, and its antibacterial activity was conserved. In the same year, Huang et al 106 transformed tobacco plants with an MB39 coding sequence, resulting in the absence of disease symptoms. Figure 5 describes the first antimicrobial peptides derived from cecropin A, B, and melittin that served as a starting point for the generation of more derivatives and analogs.
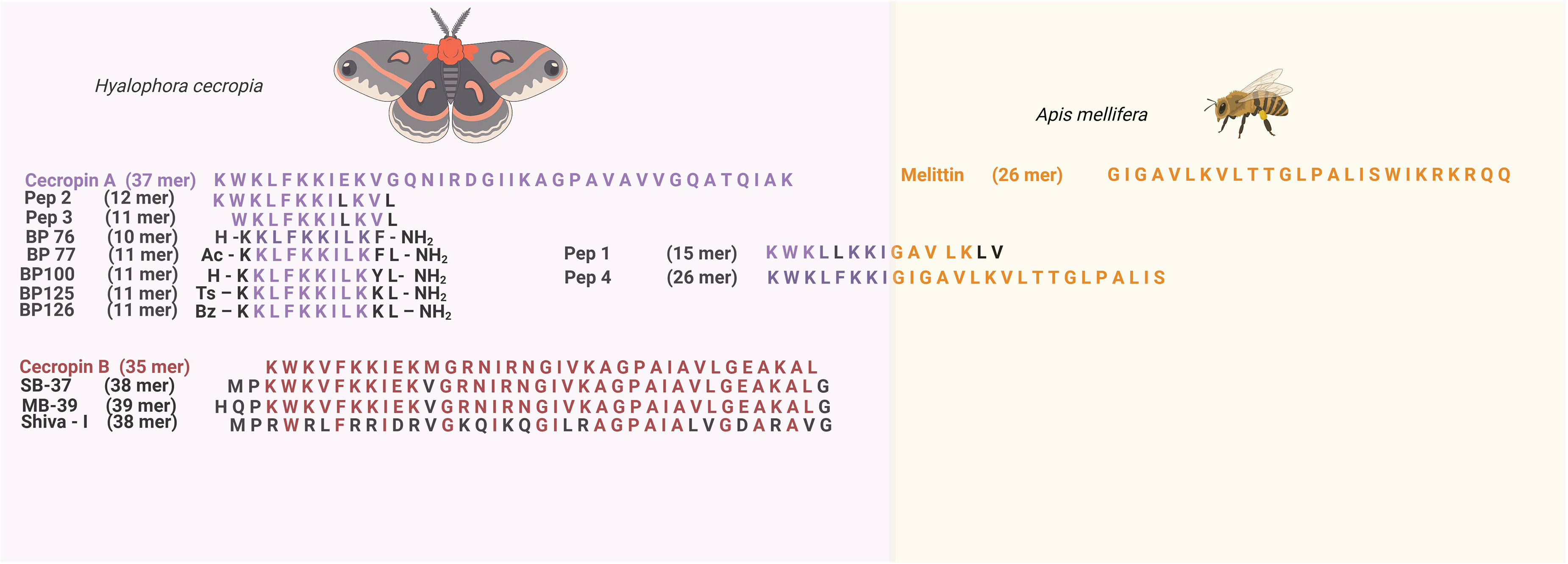
Description of the first antimicrobial peptides derived from cecropin A, B, and melittin that served as a starting point for the generation of more derivatives and analogs. Created in BioRender. Blandon, L. (2026) https://BioRender.com/rozofyc.
Synthetic Chemistry and Synthetic Combinatorial Libraries (SCL)
Synthetic chemistry and synthetic combinatorial libraries (SCL) techniques enable the generation of large numbers of peptide analogs and new peptide sequences with antimicrobial activity, starting from a known template. 107 For instance, in 1995, Powell et al 89 synthesized AMPs that mimicked the amphipathic α-helix found in magainins. The design started with ESF1, which mimicked the magainin PGLa, a peptide from the granular gland of the frog Xenopus laevis, 108 then analogs of ESF1 were designed (ESF4, ESF5, ESF6, ESF12, ESF13, ESF17). These peptides were tested in vitro against conidia of Cryphonectria parasitica, F. oxysporum f. sp. lycopersici, and Seploria musiva, as well as against the bacteria A. tumefaciens, E. amylovora, and P. syringae. The three fungi were affected by the peptides ESF1, ESF5, ESF6, and ESF12. ESF12 also affected A. tumefaciens, E. amylovora, and P. syringae. On the other hand, ESF1, ESF5, ESF6, and ESF12 didn’t inhibit the germination of pollen from Salix lucida and Castanea mollisima, and ESF12 also didn’t affect the germination of pollen from Malus sylvestris and Lycopersicon esculentum
A library can be defined as a collection of chemical compounds that can be obtained by parallel synthesis methods or by combinatorial chemistry. By comparison, combinatorial libraries consist of all possible combinations of a class of compounds and represent extensive collections (thousands to millions of compounds) prepared simultaneously in a highly systematic manner.
109
In 1997, Reed et al
91
constructed a synthetic peptide combinatorial library to obtain peptides active against the phytopathogenic fungi F. oxysporum f. sp. lycopersici, Rhizoctonia solani, Ceratocystis fagacearum, and Pythium ultimum. Their strategy consisted of an iterative process that started with a 26-series from a 400 D-amino acid peptide mixture, identifying a promising D-hexapeptide (FRLKFH) and a D-pentapeptide (FRLHF). Additionally, microscopy studies indicated that the fungal cytoplasmic and nuclear membranes were rapidly compromised. Similarly, in 2000 López – García et al
110
identified the hexapeptide PAF19 from a peptide library and synthesized either D or L amino acid. This peptide demonstrated antimicrobial activity against the fungal postharvest pathogens Botrytis cinerea, Penicillium italicum, and P. digitatum
Regarding the BP76 peptide derived from cecropin A that exhibited antimicrobial activity against Erwinia amylovora and Xanthomonas axonopodis pv. vesicatoria, and Pseudomonas syringae pv. syringae. 95 In 2007, Badosa et al 96 used a combinatorial chemistry approach to prepare a 125-member library of synthetic peptide analogs of BP76. The strategy consisted of incorporating amino acids with varying hydrophobicity and hydrophilicity at positions 1 and 10 and varying the N-terminus. These analogs were tested in vitro against E. amylovora, X. axonopodis pv. vesicatoria, and P. syringae pv. syringae finding four (BP77, BP100, BP125, and BL126) with the highest antibacterial activity. Then, these peptides were tested in vivo for their preventive effect against E. amylovora in apple and pear flowers, and peptide BP100 was found to be the most promising, more potent than BP76 and only slightly less effective than streptomycin. Continuing with BP100 Badosa et al in 2013 112 employed this peptide to generate a library of 40 analogues with the necessary characteristics to be expressed in plants. The strategy consisted of adding an AGPA hinge with a glycine residue at the N-terminus as a protease recognition site, and introducing a KDEL unit or tag54-2 at the C-terminus to enable retention in the endoplasmic reticulum and subsequent recognition by a complementary epitope. 112 In the same year, Torcato et al 113 aimed to improve the BP100 activity against Gram–positive bacteria and designed two analogs, R-BP100 and RW-BP100, which also showed activity against Gram-negative bacteria. These peptides were tested against clinical pathogens with good results. Then, in 2021, Mariz Ponte et al 80 took BP100 and RW-BP100 together with CA-M previously designed by Andreu et al in 1992, 103 and tested against eight strains of P. syringae pv. actinidiae, and the three peptides showed high antimicrobial activity. Then, Mendes et al, also in 2021 114 tested the five AMPs RW-BP100, CA-M, 3.1, D4E1, and Dhvar-5 against eight strains of E. amylovora, the AMPs with the highest antibacterial activity were RW-BP100, CA-M, and BP100. In the same year, the same authors tested mixtures of these three AMPs against three strains of E. amylovora with different genotypes and virulence, finding minimum inhibitory concentration (MIC) values lower than those reported in the previous study.
As discussed in the previous section, most studies on AMPs have used α-helical peptides. Nevertheless, non-linear peptides also showed high antimicrobial activity. In 2000, Oren and Shai 22 studied the effects of cyclization on the selectivity between bacterial and mammalian cells, finding that cyclization increased this selectivity and the activity toward Gram-negative bacteria. For instance, in 2006, Monroc et al93,94 designed de novo cyclic cationic peptides, aiming to increase bacterial selectivity over mammalian cells. They found that the peptide BPC10L was the most active against X. vesicatoria and P. syringae. Then, the same authors used BP10L as a lead to discover new cyclic decapeptides with antibacterial activity and low toxicity to eukaryotic cells, employing a combinatorial chemistry methodology involving two libraries. They finally proposed the structure c(X1X2X3X4KKKLQ) with X being K or L, yielding peptides with optimized activity profiles against E. amylovora, X. vesicatoria, and P. syringae. In the same year, these researchers continued to improve the antimicrobial activity of the cyclopeptides and prepared a library of 56 cyclic decapeptides, finding that cyclopeptides BPC194 and BPC198 had better antimicrobial activity. 94 These cyclopeptides served as templates for the posterior synthesis of lipopeptides, which are regarded as promising agents for the design of novel peptide-derived pesticides. They are composed of a short (six to seven D- and L-amino acids) linear or cyclic peptide sequence, and a lipid tail. 115 On the other hand, it has been reported that incorporating an acyl chain at the N-terminus can enhance antimicrobial activity, serving as a membrane anchor. Additionally, the presence of the acyl chain could protect the lipopeptides against proteolytic degradation.97,100 In this sense, Vila et al in 2016 97 used the cyclopeptide BPC194 as a template for designing new cyclolipopeptides that incorporate an acyl chain. Finding BPC702 as the best cyclopeptide, which incorporates D-Lys 5 acylated with a butanoyl group, active against X. axonopodis pv. vesicatoria, P. syringae pv. syringae and F. oxysporum. Besides, BPC702 showed no hemolysis and the lowest necrosis levels in tobacco leaves. Then, in 2017, Güell et al 98 studied the effects of adding triazolyl amino acids to the template BPC194 on the antibacterial activity. The addition of triazolyl amino acids can contribute to resistance to metabolic degradation and enhance antimicrobial activity. In this case, the peptides BPC548 and BPC550 were the best with antibacterial activity against X. axonopodis pv. vesicatoria and P syringae pv. syringae as well as low hemolysis and low phytotoxicity 98 . Continuing with the improvement of the antibacterial activity against the same bacteria, Camó et al 99 in the same year studied the effect of incorporating a tryptophan residue using combinatorial chemistry, taking into account that the indole side chain of the tryptophan facilitates the membrane partition through the formation of hydrogen bonds and cation interactions. The best peptides were BPC086 W and BPC108 W. Regarding BP100, in 2018, Oliveras et al 100 developed a collection of 36 lipopeptides derived from BP100, identifying five (BP370, BP378, BP381, BP387, and BP389; see Table 2) that were highly active against diverse phytopathogenic bacteria and fungi.
Discovery and Design of Antimicrobial Peptides (AMPs), from Biological Data and AI for Sustainable Agricultural Purposes
Since the release of the Brundtland Report in 1987, the notion of sustainable agriculture has gained importance, emerging alongside the broader framework of sustainable development. This concept highlights the necessity to develop an agriculture able to continually provide food and other resources to a growing world population while preserving the environment, supporting economic viability, and ensuring social equity for future generations. 1 Together, these changes are paving the way for Agriculture 4.0—a future greener, more innovative, highly efficient, and free from conventional industrial boundaries model. Entering the twenty-first century, agriculture has increasingly taken on a multidisciplinary character, as cutting-edge fields in the life sciences—such as genomics, proteomics, structural biology, and molecular imaging merge with emerging areas like nanotechnology, information science, big data, and AI. 9 In this context, several studies have been conducted to refine and enhance AMPs, alongside the development of biopesticides to strengthen crop resistance against a wide range of diseases. Advances in chemical synthesis, targeted amino acid substitutions, and fusion protein creation are being explored to address current limitations. By merging synthetic biology with AI, AMPs are evolving into programmable ‘smart agrochemicals,’ paving the way for sustainable strategies that promote plant health and boost agricultural productivity. 116
Besides the discovery and rational design of AMPs starting from proteome data mining and high-throughput sequencing technology, the large amount of biological data obtained with these technologies has led to the exploration of new AMP sources. A remarkable example is the discovery of non-conventional peptides derived from unannotated coding sequences, such as introns, intergenic regions, and untranslated regions. These types of peptides were discovered more than twenty years ago, and some studies have shown that they play a role in plant development and environmental responses. In animals, they are involved in various cellular processes, including embryogenesis, muscle function, calcium transport, and stress resistance. 117
On the other hand, recent advances in genomic technology have led to a more thorough exploration. For instance, a method that integrates mass spectrometry with genome mining has been developed, known as peptidogenomic. Specifically, peptidogenomic combines peptidomic analysis—leveraging computational mass spectrometry techniques—with genomic data to identify contiguous amino acid sequences derived from tandem mass spectrometry (MS/MS) spectra.118,119 In 2020, Wang et al 114 developed a peptidogenomic pipeline for the large-scale identification of non-conventional peptides in plants. Then, in 2021, Tian et al tested these peptides against phytopathogenic fungi, finding promising results. 120 Another approach using genomes is in the study of Kishi et al in 2018, 121 which explored citrus genomes to search for alternatives to treat citrus canker caused by X. citri subsp. citri and huanglongbing (HLB or citrus greening disease) caused by Candidatus Liberibacter asiaticus (CaLas), finding the peptide Citrus-amp1 encrypted in the genome of Citrus sinensis with high antibacterial activity, making it a possible candidate for the transformation of citrus plants resistant to these diseases.
Antimicrobial Peptides (AMPs) Synthesis and Production for Agricultural Purposes
AMPs are commonly synthesized chemically, mainly for research purposes. Nevertheless, chemical techniques such as solid-phase synthesis (SPPS) have evolved to enable large-scale production. Besides, AMPs can be used for agricultural purposes in two ways: by transforming plants to support the breeding of resistant cultivars or for heterologous expression and by developing a topical product, for example, to extend the postharvest shelf life. In this context, four main strategies have been reported for AMPs production: chemical synthesis, plant transformation, and the heterologous expression in plants, microorganisms, and insect cells.
Chemical Synthesis
Chemical synthesis has been one of the most common strategies during the research stages, enabling in vitro or in vivo experiments once a new AMP is discovered or designed. Peptides can be designed using solution-phase or SPPS; the latter is the most common method because it enables the synthesis of longer and more complex peptides. SPPS has progressed from a laboratory technique to an industrial-scale process, leading to enhanced peptide production yields. In SPPS, solid supports such as polystyrene resins are employed to anchor and stabilize the growing peptide chain. 122 The synthesis is made in the C—N direction. During this process, peptides are protected by adding groups to their side chains or N-termini; the most common protecting groups are tert-butoxycarbonyl (Boc) and 9-fluorenylmethyloxycarbonyl (Fmoc). In addition, peptide libraries can be obtained using a SPOT array, which immobilizes peptides on membranes acting as a solid support, thereby allowing the simultaneous synthesis of several peptides. Finally, the synthesized peptide is released from the solid support, and the protecting groups are removed. Then, mass spectrometry can be used to confirm the correctness of the synthesis. Furthermore, the synthesis time can be reduced using microwave irradiation during the coupling and N(α)-deprotection process.18,27 Three types of peptides can be obtained by chemical synthesis: i) the retro-inverse peptides, the reverse sequence of the parent L-amino peptide; ii) peptides where all amino acids are D-amino acids, and iii) peptides with partial D-amino acids. However, AMPs are commonly synthesized using D-amino acids, which makes them less susceptible to degradation by naturally occurring proteases and avoids the activation of the host immune response. Despite their potential applications, D-amino acids are scarcely present in natural systems and cannot be incorporated through ribosomal translation. As a result, chemical synthesis remains the predominant strategy for their generation.27,123 Nevertheless, chemical synthesis is expensive and limited in large-scale production, which is why other methods, such as plant-based production and heterologous expression in microorganisms and plants, have been explored. Meanwhile, the chemical production of peptides with fewer than 10 amino acids has been studied, leading to the discovery of some effective short cationic AMPs. This strategy also allows the introduction of nonproteogenic amino acids. 12
Use of Plants
Plants serve both as a source for developing resistant cultivars and as biofactories capable of producing high quantities of AMPs. The most common strategy for plant transformation is through Agrobacterium tumefaciens, with tobacco (Nicotiana tabacum) as the most extensively studied host.88,105,106,124,125 Besides tobacco, rice, and potato plants are also frequently used. Recently, in 2024, Khalifeh-Kandy et al 125 reported the expression of roseltide rT1 128, a plant peptide with antimicrobial and therapeutic properties, in tobacco plants. The protein extracts of the transform plants showed efficient antimicrobial properties against Pseudomonas syringae, Pectobacterium carotovorum, and Ralstonia solanacearum. Nevertheless, there are some different reports. In 2001, Liu et al 126 successfully generated transgenic Royal Gala apple (Malus × domestica Borkh.) shoots carrying a modified cecropin SB37 gene fused to the secretory signal sequence of barley α-amylase, regulated by a wound-inducible osmotin promoter derived from tobacco, resulting in transgenic plants resistant to Erwinia amylovora. Information on some of the AMPs expressed in plants, the vectors used, and the target pathogens is shown in Table 3.
Some Antimicrobial Peptides Expressed in Plants, the Vectors Used, and the Target Pathogens.

It is important to note that for the heterologous expression in plants, the peptide accumulation in particular tissues is needed, and for this, it is necessary to have the incorporation of retention signals as well as the presence of target sequences and protease cleavage amino acids to facilitate the detection of peptides, so structural modifications are needed without affecting the peptide bioactivity. 132 For instance, in 2009, Barrell and Conner 127 expressed a chimeric magainin gene in potato (Solanum tuberosum). The gene encoded an analogue of magainin II called magainin D. Magainin D incorporated a histidine-to-arginine substitution, which increases the peptide's stability against protease cleavage, and three alanine substitutions to increase antimicrobial activity. “The magainin D gene sequence was ligated into a chimeric expression construct designed to transport the resulting peptide into the intercellular space. After the transformation, certain potato lines possessed significantly greater resistance to the soft rot disease caused by Erwinia carotovora. Gene fusion is another approach. In 2018, Badrhadad et al 130 transformed tobacco plants with an AMP encoding gene from alfalfa seeds (alfAFP) 133 fused to the C-terminal part of the chitin-binding domain of the rice chitinase-encoding gene. Extracts containing the recombinant peptide showed activity against Fusarium solani. Also, in 2020, Khademi et al 131 constructed a recombinant peptide using dermaseptin B1 (obtained from Phyllomedusa bicolor) 134 fused to Avr4 effector protein chitin-binding domain from Cladosporium fulvum and transformed tobacco plants. Production of the recombinant peptides conferred strong protection against Alternaria alternata, Alternaria solani, Fusarium oxysporum, and Fusarium solani in transgenic plants. Besides, Transgenic plants were more resistant to Pythium and Fusarium infections.
Regarding the peptide BP 100 and its derivatives, in 2017, Montesinos et al 128 continued their research and designed a derivative of BP100, the peptide BP178, and produced it using rice (Oryza sativa) seeds, recovering it from the seed endosperm, obtaining a yield of 21 μg/g. The produced peptide retains its biological activity, and the transgenic seedlings exhibited resistance to pathogen infection (Dickeya sp.), suggesting that BP178 was also biologically active in the rice seed. Then, Baró et al in 2022 129 used a Potato virus X (PVX) based vector system to express genes encoding peptides, BP134 and BP178, both BP100 derivatives, in Nicotiana benthamiana to treat Xylella fastidiosa infections. X. fastidiosa is a Gram-negative bacterium that inhabits the xylem vessels of nearly 600 plants from 85 taxonomic families. To date, there is no efficient cure for diseases associated with this bacterium, primarily because of the pathogen's remarkable genetic adaptability, its confinement to xylem tissues, and the asymptomatic nature of infected host plants during the initial phases of infection. Finally, the researchers found that the plant protection was more efficient with BP178; however, this protection was mediated by the induction of the plant defense.
Use of Microorganisms
In the case of microbial-derived AMPs, most NRPs produced by NRPSs can be advantageous for large-scale production using microorganisms. In this case, enzymatic synthesis and recombinant expression are the most common methods. 8 For instance, some microbial AMPs have potential as plant protectants, such as fengycin, mycosubtilin, and tailocins. 8 In 2007, Montesinos et al 24 described AMPs produced by bacteria and fungi as crucial for plant disease control, such as bacteriocins from plant-associated bacteria, peptaibols from several fungi, and cyclopeptides produced by bacteria, fungi, and cyanobacteria.
Bacteria are widely used as hosts for heterologous expression because they grow quickly, are inexpensive to cultivate, and enable high-yield production. Nonetheless, specific bacterial systems struggle to secrete antimicrobial peptides, form disulfide bonds, and carry out complex post-translational modifications, such as glycosylation. Recombinant AMP production using bacteria has mainly been reported in Escherichia coli and Bacillus subtilis. 122 E. coli-based expression provides high AMP production yields. Nevertheless, the extracellular nature of these peptides and the absence of an expression reporter are challenges to the ideal expression scenario in which intracellular accumulation is desired. In addition, AMP production in E. coli is frequently accompanied by endotoxin contamination, particularly lipopolysaccharides, which can exert harmful effects in vitro and in cell culture systems. Because of its low endotoxin content, B. subtilis is considered much safer than E. coli for AMP production. In this case, the presence of secretory pathways enables the secretion of AMP or the AMP fusion protein. 27
For library construction, yeast two-hybrid is one of the techniques highly employed. The target protein is designated as the ‘bait’ and is fused to a heterologous DNA-binding domain, whereas the candidate peptide, referred to as the ‘prey’, is linked to a transcriptional activation domain. Upon interaction between the peptide and the target protein, a functional transcription factor is reconstituted through the physical association of the DNA-binding and activation domains. This chimeric transcription factor subsequently initiates transcription by activating the promoter of a reporter gene. The resulting gene expression can be quantitatively assessed using colorimetric enzymatic assays or qualitatively evaluated through selective growth-based screening. 135
Another approach is the production of peptide aptamers, which are short peptides (8-20 amino acid residues) with specific, high-affinity binding to targets, consisting of a biologically inactive scaffold protein and diverse peptides immobilized on the scaffold at the ends. Their mode of action involves the suppression of intracellular protein activity and the disruption of protein–protein interaction. Commonly, peptide aptamers are selected from a library that consists of randomly generated amino acids. For instance, in 2020, Colombo et al 136 used the yeast two-hybrid approach to construct a combinatorial 8-amino acid peptide aptamer library with the aim of obtaining an aptamer peptide with antimicrobial activity against the oomycete Plasmopara viticola (Berl. & de Toni), which attacks grapevine green tissues, causing Downy mildew. For this purpose, the NoPv1 peptide aptamer was designed using the enzyme P. viticola cellulose synthase 2 (PvCesA2) as a target, as cellulose biosynthesis plays a crucial role during pathogen infection. Then, the cytoplasmic soluble portion of PvCesA2 was fused to the GAL4 DNA-binding domain (BD) and used as bait. It was demonstrated that the NoPv1 peptide aptamer prevents P. viticola grapevine leaf infection and germ tube formation. Additionally, this peptide can counteract the growth of Phytophthora infestans, the causal agent of late blight in tomato and potato, possibly due to the sequence similarity between P. viticola and P. infestans cellulose synthase enzymes. But bacteria can also be used for peptide aptamer production. In 2019, Xu et al 85 designed the peptide aptamer SNP-D4 to inhibit the spore development of Magnaporthe oryzae. In this case, the researchers constructed peptide aptamer libraries using the bacterial two-hybrid system using calmodulin (CaM) of M. oryzae as the bait, a critical regulator in the Ca2+ signaling pathway implicated in fungal infection. The library consisted of the scaffold protein Staphylococcus aureus nuclease and an exposed surface loop of 16 random amino acids. Finally, SNP-D4 was expressed and produced using an E. coli strain and purified. Extracts containing the molecule and the pure SNP-D4 were tested, and ultimately, SNP-D4 was identified as a promising antagonist of CaM and can be used for controlling fungal infections.
Insect Cells
The baculovirus-based expression platform, which employs insect cells, has been widely used to generate diverse heterologous proteins, including AMPs. Insect cells provide essential benefits for recombinant protein production, including efficient post-translational modifications and proper folding. However, their ability to perform glycosylation is restricted. Different strategies have been explored to address this issue, including inserting mammalian genes into insect cells to enable the synthesis of more complex glycans, knocking out the gene encoding the trimming enzyme, and applying CRISPR-based engineering to modify insect cells. 122
Formulation and Delivery System Strategies
The development of diverse formulations—including suspension agents, microemulsions, and encapsulated suspensions—serves to shield peptide molecules from degradation caused by environmental factors such as moisture, ultraviolet radiation, temperature fluctuations, and metabolic enzymes. Specifically, drug delivery systems enable precise delivery of therapeutic agents to target sites through controlled-release platforms such as hydrogels, cubosomes, and nanocarriers, thereby enhancing drug utilization efficiency. When combined with innovative drug delivery systems approaches, readily degradable peptides can be applied in precision agriculture. 137
Moreover, the challenges associated with AMP bioavailability can be addressed through innovative drug delivery approaches, particularly nano-delivery systems. Within these platforms, AMPs may be incorporated via two main strategies: passive loading, in which they are encapsulated within nanocarriers, and active loading, in which they self-assemble on the carrier's surface. Employing diverse nanocarriers—including liposomes, micelles, metallic and polymeric nanoparticles, nanofibers, mesoporous particles, carbon nanotubes, hydrogels, and dendrimers—has proven effective in improving the antimicrobial activity, stability, and target specificity of AMPs. 39
Economic Feasibility
The use of AI for the discovery or rational design of AMPs can reduce costs in the initial phase of studies by analyzing large numbers of peptide sequences using classification tools, filtering, and finally selecting the best candidates for synthesis and continuing with future in vitro and in vivo studies. Nevertheless, there are some limitations related to the reliance on data, which can lead to the amplification of biases, since AI models depend strongly on the scope and quality of the information used for training. For instance, the availability of quantitative measurements of AMP activity, like MIC values, in public datasets is limited. However, there are some public datasets like GRAMPA with MIC values.116,138 Another important limitation is the activity cliff phenomenon, observed when a pair of molecules differing by a small structural alternation exhibit a high difference in their biochemical activities. This phenomenon is extrapolated to AMP studies, and can affect the optimization and predictive modeling. 138
Numerous limitations regarding antimicrobial activity, biocompatibility, non-target toxicity, bioavailability, and production cost hinder the implementation of AMPs in agriculture. Aiming for cost reduction, heterologous expression has been one of the most promising strategies. One of the most common limitations is protease degradation. Also, several difficulties arise during downstream processes. For instance, the presence of disulfide bonds and significant hydrophobicity often induces protein misfolding and reduces solubility, necessitating the use of hazardous organic solvents and costly chromatographic techniques for purification. Furthermore, the relatively low molecular weight of AMPs complicates the detection and characterization of impurities arising during production. 123
Regulation and Current Products
In considering the regulatory landscape for AMP applications, a critical distinction arises between stable peptide expression within plants and topical delivery. Plants genetically engineered to synthesize peptides are automatically categorized as GMOs and are subject to the specific regulatory regimes of each jurisdiction. Conversely, innovative topical formulations and their routes to commercialization are encompassed within the established framework governing new plant protection products. 78
Currently, 18 peptides have been commercialized as green agents for plant protection. Notably, the bioinsecticide Spear®, developed from a spider-venom neuropeptide, received the U.S. Presidential Green Chemistry Challenge Award (Small Business category) in 2020 and was honored with the Best New Biological Agent Award in 2021. 137 Besides, BLAD, a peptide generated during the germination of Lupinus albus, has been commercialized in two formulations: ProBlad® Verde, developed by the U.S. company Sym-Agro, and ProBlad Plus™, produced by the Portuguese firm Consumo em Verde. Problad Plus™ (https://www.cev.com.pt/) is a biofungicide containing 20% BLAD that targets susceptible fungal pathogens by disrupting cell walls and internal membranes. It demonstrates efficacy against diseases such as powdery mildew and gray mold and is recommended for crops including strawberries, grapes, tomatoes, and stone fruits (eg, peaches and cherries). ProBlad® Verde (https://sym-agro.com/problad/) employs a unique multisite mode of action to control both fungal and bacterial diseases. It is registered for use on crops such as vine berries, herbs, coffee, leafy greens, and mustard plants, providing protection against pathogens including coffee leaf rust, white mold, gray mold, powdery mildew, anthracnose, bluegrass leaf rust, leaf blight, and Rhizopus. Furthermore, ProBlad® Verde has been granted exemption from the European Union's Maximum Residue Limit (MRL), enabling producers of high-value export crops to comply with increasingly strict regulatory standards. 137
Ecological Consequences
In addition to the well-recognized issue of antimicrobial resistance, the extensive use of antibiotics has resulted in substantial volumes of contaminated wastewater containing antibiotic residues. In this sense, AMPs have emerged as a promising solution not only to the problem of antimicrobial resistance but also to the environmental problem. One of the main aspects that makes the topical application of AMPs promising is their short half-lives, which could indicate no ecological impact compared to conventional antibiotics, or at least minimal impact. Besides, the effect on beneficial microorganisms is a questionable concern. Both aspects need to be considered during AMP-based product development.
Recently, Gimranov and colleagues 2 evaluated the synthetic peptides CA-M, BP100, RW-BP100, and 3.1 against plant growth-promoting bacteria and biological control agents. Their findings showed that the MICs required to affect beneficial Bacillus and Pseudomonas strains were higher than those needed to exert bacteriostatic or bactericidal activity on the phytopathogens tested, suggesting that these AMPs could be environmentally safe. Nevertheless, beneficial microorganisms are not the only ecological aspect to consider. For instance, in 2022, Wang et al 139 highlighted that the AMP Polymyxin B (PMB) has been detected in wastewater and tested the influence of PMB on aerobic denitrification. Their study revealed that PMB reduced nitrate removal efficiency and shifted denitrifiers toward converting nitrate into biomass rather than releasing it as gaseous nitrogen.
Conclusions
Human survival is dependent on agriculture. Since humanity has domesticated animals and plants, everything has changed, leading to the formation of settlements that later became cities. Plant diseases are not a current topic; they have been occurring since ancient times, leading to significant economic and farming problems. Naturally, plants are part of an ecological equilibrium where plant diseases help to keep this balance. Nevertheless, because of many years of domestication, many valuable crops and ornamental plants are currently susceptible to diseases and unable to survive without human intervention. With all this in mind, humanity needs to consider solutions that ensure its survival, but these solutions must be accompanied by proper control and equilibrium. For instance, compounds like antibiotics, which help improve our life expectancy by fighting infections that could once be fatal, have also become solutions to treat infectious diseases in plants. Unfortunately, the ignorance of proper management and the urgency for survival have led to environmental and antibiotic resistance problems.
To manage these problems, new agricultural practices have emerged, such as agriculture 4.0, which integrates sustainable agricultural practices and the life sciences with areas like big data. In this context, AMPs have become a promising solution for plant infectious diseases; these molecules can be discovered, designed, and optimized with computational and AI tools. Research with AMPs started in the clinical field; all these studies have been ideal for the comprehension of their mechanisms of action, as well as the understanding of properties like bioavailability and stability. While it is true that the action over the pathogens, like the membrane interaction, is the same in all the scenarios, there are considerable differences in the development of a clinical and an agricultural product. Starting with the production method, which can involve the use of plants, the proper formulation for a topical product is required. Besides, while no or minimal ecological impact is expected, it is important to take this into mind and always lead studies that control or elucidate what is happening when these kinds of products are employed. Additionally, researchers should not overlook conventional control methods, such as crop rotation, prior to adopting new technologies, as this may be the way to achieve proper management.
Footnotes
Author Contributions
LB and AM writing—original draft preparation, SO —review and editing. All authors have read and agreed to the published version of the manuscript.
Funding
Thanks to the Ministerio de Ciencia, Tecnología e Innovación (Minciencias, contract 212-2024, Orchids Program for Women in Science) and the Universidad Nacional de Colombia sede Medellín, Hermes research grant 61927.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
