Abstract
Objective
This study aimed to isolate and identify bioactive secondary metabolites, including a new phenazine derivative, from a marine-derived Streptomyces sp. CNQ-031 belonging to the MAR4 clade, and to evaluate their antibacterial activity.
Methods
The marine-derived Streptomyces sp. was cultivated and extracted using ethyl acetate. The crude extracts were subjected to chromatographic separation to afford a new natural product, pontophenazine (
Results
Four compounds were isolated, including a new phenazine derivative, pontophenazine (
Conclusion
The discovery of pontophenazine (
Introduction
Natural products are a fundamental source of novel chemical diversity and an essential component of the pharmaceutical industry. Oceans, which cover 71% of the Earth's surface, contain 97% of the planet's water, and shelter nearly 87% of all animals and plants, are abundant sources of undiscovered organisms, including microorganisms. Among these, the exploration of rare actinomycetes within unexplored marine environments presents a promising avenue for discovering unprecedented chemical frameworks exhibiting a range of biological activities. These encompass anti-inflammatory, antibacterial, neuroprotective, antiparasitic, and anticancer properties. 1
Marine actinomycetes, particularly the Streptomyces lineage, have mainly originated from marine environments
2
and represent abundant sources for the discovery of interesting chemical scaffolds with bioactivity. The streptomycete clade MAR4, a distinct marine actinobacterial lineage, exhibits significant taxonomic diversity and excellent proficiency in producing diverse hybrid terpenoid skeletons such as polyketide-terpenoid molecules (particularly naphterpins D and E).
3
Furthermore, MAR4 strains possess average five 2-amino-4-boronobutanoic acid (ABBA) prenyl-transferases per strain compared to other streptomycetes. Based on their substrate specificity, ABBA prenyl-transferases may be separated into two distinct group and are only present in secondary metabolism.4,5 One family of ABBA, phenol/phenazine prenyl-transferases catalyses the prenylation of phenazines,
6
naphtoquinones,
7
benzodiazepines
8
and hydroxybenzoates.
9
The hybrid isoprenoids produced by MAR4 strains, including prenylated polyketides, pyrroles, and phenazines, are of considerable interest due to their potent biological activities.
10
One of the strains belonging to this group, CNQ-031, isolated from marine sediment collected from La Jolla, California, has been reported to produce a phenazine derivative (
Further investigation of the chemical components of this strain has resulted in the discovery of a new phenazine derivative, pontophenazine (
Experimental and Methods
General Experimental Procedure
The UV spectra were measured with a V-730 UV visible spectrophotometer (Jasco, Easton, MD, USA) using a path length of 1 cm. The infrared (IR) spectra were recorded on a Varian Scimitar Series (Palo Alto, CA, USA) using KBr disks. The NMR spectra were acquired with a Bruker NMR spectrometer (Bruker, Middlesex, MA, USA, 300 MHz for 1H NMR) in MeOD-d4 using a solvent signal as an internal reference (δH 3.31 ppm) and a JEOL NMR spectrometer (JEOL Ltd, Tokyo, Japan, 500 and 125 MHz for 1H and 13C NMR, respectively) in DMSO-d6 using a solvent signal as an internal reference (δH 2.50 ppm and δC 39.5 ppm). Low-resolution liquid chromatography/mass spectrometry (LC/MS) data were obtained on the Agilent Technologies (Santa Clara, CA, USA) 6120 quadrupole and Waters (Milford, MA, USA) Micromass ZQ LC/MS systems using a reversed-phase column (Phenomenex Luna C18(2), 100 Å, 50 mm × 4.6 mm, 5 μm) at a flow rate of 1.0 mL/min at the National Research Facilities and Equipment Center (NanoBio Energy Materials Center) at Ewha Womans University. High-resolution mass analysis was conducted by a JMS-700 mass spectrometer (JEOL Ltd, Tokyo, Japan) at Seoul National University. The fractions were purified by a Waters 616 quaternary HPLC pump and a Waters 996 photodiode array detector using a Phenomenex Luna C18(2) (100 Å, 250 mm × 10 mm, 5 μm) reversed-phase HPLC column.
Collection and Phylogenetic Analysis of Strain CNQ-031
The marine-derived actinomycetes strain, CNQ-031, was isolated from a sediment sampled off the coast of California. It was identified as the Streptomyces sp. MAR4 clade based on a 16S rRNA gene sequence analysis. The gene sequence data are available from GenBank (deposit number KC261620).
Fermentation and Extraction
The CNQ-031 strain was cultured in 40 L of a 2.5 L Ultra Yield Flask, each flask containing 1 L of medium SYP SW (10 g/L of soluble starch, 2 g/L of yeast, and 4 g/L of peptone dissolved in 1 L seawater), at 27 °C with shaking at 120 rpm for 7 days. The culture medium was extracted with EtOAc (40 L overall), and the EtOAc-soluble fraction was then evaporated in vacuo to yield 5.84 g of crude extract.
Isolation and Purification of Compounds
The organic extract of CNQ-031 (5.84 g) was fractionated with C-18 open column chromatography with a step gradient from 20% to 100% MeOH in DW to obtain nine fractions. The third fraction was further subjected to reversed-phase HPLC with 39% aqueous acetonitrile (Phenomenex Luna C18(2), 250 × 100 mm, 2.0 mL/min, 5 μm, 100 Å) to obtain 5.0 mg of pontophenazine ( Pontophenazine (
Antibacterial Activity Testing (MIC Assay)
Antibacterial activity was evaluated against a panel of Gram-positive and Gram-negative bacteria. The Gram-positive bacteria used were Bacillus subtilis KCTC1021, Staphylococcus aureus KCTC1927, Kocuria rhizophila KCTC1915, and Micrococcus luteus SC0560. The Gram-negative bacteria included Escherichia coli KCTC2441, Klebsiella pneumoniae KCTC2690, Salmonella typhimurium KCTC2515, Pseudomonas fluorescens SND204, Pseudomonas aeruginosa SNC165, and Agrobacterium tumefaciens SND195.
All bacterial strains were grown overnight in the appropriate media (Mueller-Hinton Broth for the KCTC strains, tryptic soy broth for S. aureus and M. luteus, Kings broth for P. aeruginosa and P. fluorescens, and marine broth for A. tumefaciens). Cultures were adjusted to a 0.5 McFarland standard (1.5 × 108 CFU/mL).
The test compounds and positive controls (vancomycin, ampicillin, kanamycin, and rifampin) were dissolved in DMSO at a concentration of 256 μg/mL, and DMSO was used as the negative control. Each compound (100 μL) was added to the first well of a sterile 96-well plate containing 50 μL of the respective growth medium, followed by two-fold serial dilutions to obtain final concentrations of 128, 64, 32, 16, 8, 4, 2, 1, 0.5, and 0.25 μg/mL.
Subsequently, 50 μL of the adjusted bacterial inoculum was added to each well, resulting in a final inoculum density of 5.0 × 105 CFU/mL. Plates were incubated at 37 °C for 18–24 h. MIC values were determined by visual examination of bacterial growth inhibition; clear wells indicated growth inhibition, whereas turbid wells indicated bacterial growth. All experiments were performed in triplicate, and the results were reproducible within experimental error; therefore, no statistical analysis was applied.12,13
Biofilm Inhibition Activity
The biofilm inhibition activity was tested using P. fluorescens SND204, P. aeruginosa SNC165, A. tumefaciens SND195, S. aureus KCTC1927, and M. luteus SC0560 as the biosensors. After cultivating the bacteria in the same way as in the quorum sensing test, compounds were inoculated (256 µg/mL, 200 μL) in each culture and cultivated for 72 h at 37 °C. Briefly, 2 mL of media and 50 μL of the tested compound, positive controls (rifampin and kanamycin) and negative control (DMSO) were added into individual 10 mL test tubes. Each test tube was then inoculated with 4 μL of appropriately adjusted bacterial cultures and incubated at 37 °C for 72 h without shaking. After completing the incubation, the cultures were gently discarded using a 1 mL pipette, ensuring caution to avoid damaging the biofilm formed on the tube walls. Subsequently, the tubes underwent three washes with sterile distilled water. The bacteria constituting the biofilm were stained by adding 2.5 mL of 0.1% crystal violet for 1 h without shaking. The crystal violet dye was discarded, and the tubes were washed with sterile distilled water to eliminate any unbound dye. The tubes were then dried, and the biofilm was eluted using 2 mL of 100% ethanol. Thorough dissolution of the dye was ensured by shaking the tubes. Finally, the optical density (OD590nm) of the eluted samples was measured using a spectrophotometer. 14 All experiments were performed in triplicate, and the results were reproducible within experimental error; therefore, no statistical analysis was applied.
Result
Pontophenazine (
Analysis of 2D NMR spectroscopic data allowed the structural assignment of
Based on the molecular formula and the number of carbon signals observed in the 13C NMR spectrum, the chemical structure of
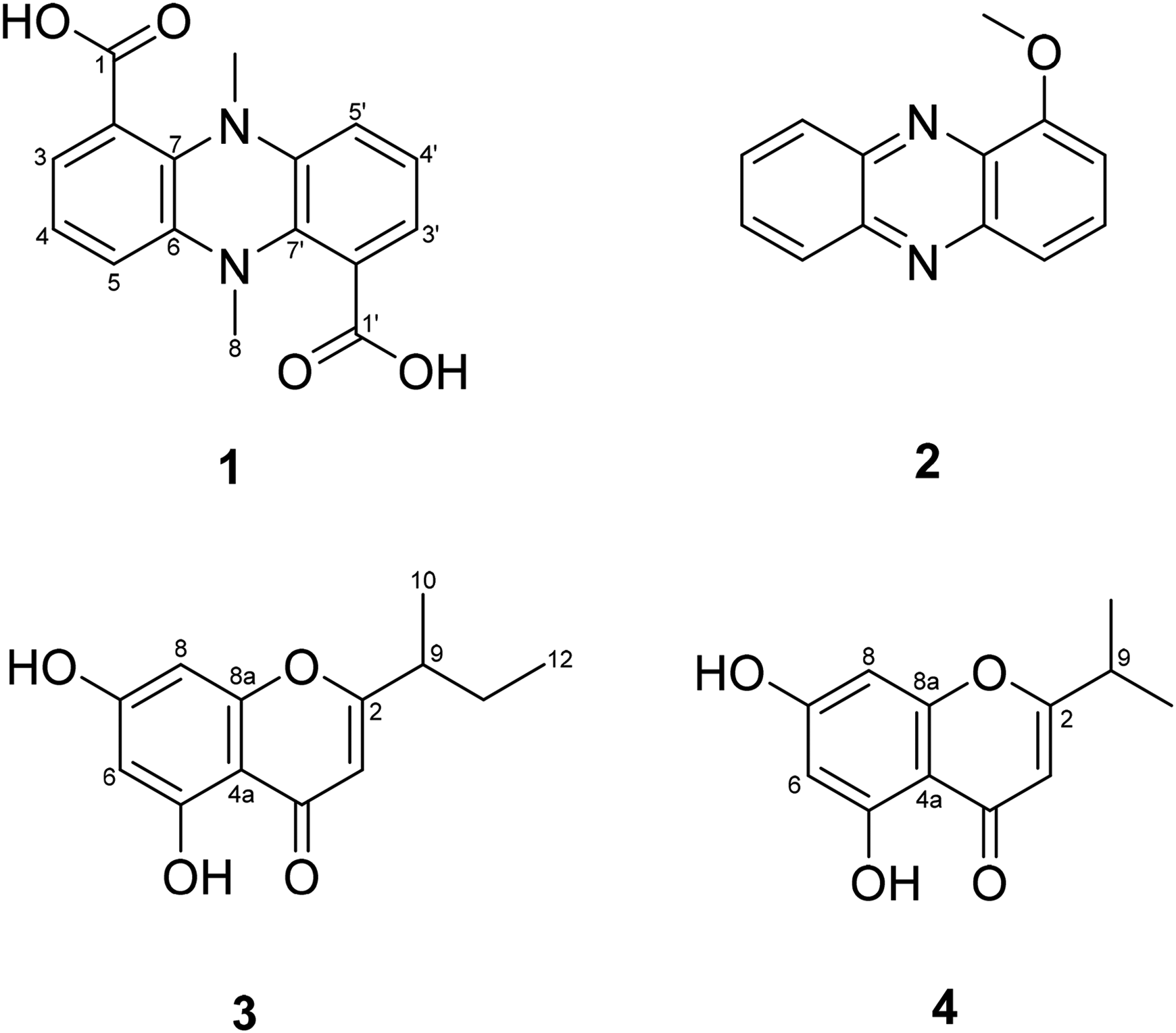
The chemical structures of
Compound
Pontophenazine (
Compounds
To explore the antibacterial properties of the isolated compounds, samples
Antibacterial Activities of Compounds

Each sample was tested in triplicate, and the experiment was repeated three times.
In the antibiofilm assay, compound

Biofilm inhibition activities of 1-4. Percentage of biofilm inhibition refers to the percentage of biofilm that was inhibited in the presence of the compounds, when compared to biofilm formation with only DMSO. Shown are the average results of three replicates and error bars represent the standard error of the mean (SEM).
Discussion
The biological properties of phenazines are determined by the positions and types of functional groups present in their molecules.
25
Phenazines and their derivatives function as electron shuttles, regulating the redox state of cells and influencing downstream gene expression associated with biofilm formation and bacterial survival.
26
Based on previous study,
18
the position of the substituents in phenazine derivatives greatly influences their antibacterial activity. Moreover, according to the QSAR analysis of the antibacterial mechanism of phenazines conducted by Udumula et al
27
2017, it is anticipated that compounds with higher water solubility and lower EHOMO energies may have more potent antibacterial action. Compound
Biofilm formation is a crucial virulence and persistence strategy for many pathogenic bacteria, as cells embedded in an extracellular polymeric substance (EPS) matrix show increased tolerance to antibiotics and host immune responses compared to their planktonic counterparts.29,30 This enhanced tolerance contributes to chronic and relapsing infections and is recognized as an important factor in antimicrobial resistance.31,32 In this study, compound
Although, pontophenazine (
Conclusions
A new natural product, pontophenazine (
Supplemental Material
sj-docx-1-npx-10.1177_1934578X251405973 - Supplemental material for Pontophenazine, a new Phenazine Analog from Marine-Derived Streptomyces sp. CNQ-031
Supplemental material, sj-docx-1-npx-10.1177_1934578X251405973 for Pontophenazine, a new Phenazine Analog from Marine-Derived Streptomyces sp. CNQ-031 by Chaeyoung Lee, Prima F Hillman, Ji Young Lee, Dong-Chan Oh and Songyi Lee, William Fenical, Sang-Jip Nam in Natural Product Communications
Footnotes
Ethical Approval Statement
This article does not involve studies with human participants or animals. Ethical approval was therefore not required for this research.
Statement of Informed Consent
There are no human subjects in this article, and informed consent is not applicable.
Author Contributions
Compound isolation, structure elucidation, activity test and manuscript writing, C.L., P.F.H.; activity test J.Y.L.; writing—review and editing, C.L., P.F.H., D.-C.O.; supervision and conceptualization, S.L., W.F., S.-J.N. All authors have read and agreed to the published version of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by Korea Basic Science Institute (National Research Facilities and Equipment Center) grant funded by the Ministry of Education (2020R 1A 6C 101B194 to S.-J.N.); the National Research Foundation of Korea (NRF) grant funded by the Korean government (MSIT; No. NRF-2022R 1A 2C 1011848 to S.-J.N.). This research was supported by the Global Learning & Academic Research Institution for Master's·PhD students, and Postdocs (LAMP) Program of the National Research Foundation of Korea (NRF) grant funded by the Ministry of Education (No. RS-2023-00301702). Isolation of the bacterium was a result of the financial support from the US National Cancer Institute (grant CA R37044848 to W.F.).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Interests
Data available on request from the corresponding author.
Reporting Guidelines Statement
Reporting guidelines such as CONSORT or ARRIVE do not directly apply to this in vitro study; however, we have followed best practices for transparent and reproducible reporting.
Supplemental Material
Supplemental material for this article is available online.
Note
This article does not contain any studies on human or animal subjects, and the statement of human and animal rights is not applicable.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
