Abstract
Background
The growing threat of drug-resistant bacterial infections demands the discovery of bioactive compounds from natural sources. This study explored the chemical diversity generated through the cocultivation of two fungal strains, Aspergillus pseudocaelatus and Trichoderma gamsii, isolated from the rhizospheres of medicinal plants.
Methods
Co-culture fermentation on rice medium was employed to induce unique hybrid metabolites alongside other bioactive metabolites. LC-QTOF/MS metabolomic profiling combined with molecular network-based dereplication identified terpene–amino acid conjugates, while genome mining revealed biosynthetic potential.
Results
Time-course experiments demonstrated that prolonged incubation enhanced metabolite yields, with coculture significantly increasing secondary metabolite production versus monocultures. Chromatographic isolation and spectroscopic characterization yielded two terpene–amino acid hybrids (
Conclusion
These findings underscore the value of microbial interactions in activating cryptic biosynthetic pathways and expanding the chemical space accessible from microbial sources. This work contributes to the growing field of microbial natural product discovery by demonstrating that fungal‒fungal interactions can be strategically leveraged to access active molecules with promising bioactivities, supporting ongoing efforts to address the global challenge of antibiotic resistance.
Introduction
Bacterial infections caused by multidrug-resistant (MDR) pathogens are a growing global health threat, driving the search for novel antimicrobials from natural sources. 1 Bacterial infections contribute to an altered immune state characterized by chronic low-grade inflammation, immune dysregulation, and disrupt the balance of the gut microbiome (dysbiosis), potentially contributing to inflammation and metabolic dysfunction.2,3 This compromised immune environment increases susceptibility to infections, including those caused by MDR pathogens.
Fungal secondary metabolites have long been a rich source of antibiotics, and cocultivation strategies have emerged as powerful tools to activate otherwise silent biosynthetic pathways. 4 Microbial natural compounds play essential roles in cell signaling, crosstalk, defense, and microbial evolution. They facilitate interactions among microorganisms within communities and with their abiotic environment elements.5,6 Owing to their remarkable structural and functional diversity, these compounds serve as ideal sources for developing therapeutic drugs, antimicrobial agents, and other bioactive agents. 7 While the structures and biosynthetic pathways of these compounds are often well characterized, microbes typically exist in complex microbiomes, where chemical signaling can be influenced by community-specific dynamics and modifications. These powerful chemical and biological factors help elucidate this poorly understood aspect of microbial chemical development. 6
Hybrid metabolites are compounds that originate from diverse biosynthetic pathways and are subsequently joined through a conjugation process, increasing their chemical diversity and biological activity. Among these compounds, terpene-amino acids and meroterpenoids have attracted attention in recent years, driven by the discovery of notable examples such as the anthelmintic CJ-12662, the insecticidal paeciloxazine, and aculene A. In the biosynthesis of these hybrid metabolites, single-module nonribosomal peptide synthetases (NRPSs) play a key role in the esterification step, facilitating the fusion of modified terpene backbones and amino acid-derived moieties. 8 The clear chemical benefit of combining terpene and amino acid biosynthesis is the enrichment of compounds with nitrogen atoms, which are rare among hydrocarbon-derived terpenoids. 9
The aim of this study stems from the need to discover natural resources capable of producing compounds effective against drug-resistant bacteria. Here, we developed a coculturing process for two fungal strains to screen their potential for the production of diverse compounds. This was achieved through metabolomic screening of fungal extracts via an in-house established library of microbial hybrid metabolites. Two terpene-conjugated amino acids were isolated and evaluated for their antibacterial activity.
Experimental
General Experimental Procedures
LC/MS grade reagents and solvents were used. Acetonitrile, methanol, and formic acid were purchased from Wako Pure Chemicals Ltd (Osaka, Japan). Ultrapure water was produced via a Milli-Q integral system equipped with a 0.22-μm point-of-use membrane filter cartridge (EMD Millipore, Billerica, MA, USA). HPLC analysis was carried out via the Agilent 1260-LC system equipped by DAD detector (Agilent Technologies). HR-ESI-MS data for compounds were recorded on a quadrupole time-of-flight mass spectrometer (Agilent QTOF-LC-MS, Agilent Technologies, USA). A JNM-ECS400 spectrometer (JEOL, Tokyo, Japan) or Bruker Avance 600 MHz (Bruker, Germany) was used to record 1D (1H and 13C NMR) and 2D NMR (HSQC, HMBC, COSY) experiments using CD3OD or DMSO-d6 as solvents. Chemical shifts (δ) were expressed as parts per million (ppm).
Microbial Materials and Fermentation
The fungal strains Aspergillus pseudocaelatus (MG772677) and Trichoderma gamsii (KX685665) were recovered from the rhizosphere areas of the medicinal plants Aloe vera and Ocimum basilicum, respectively, their antimicrobial activity and genomic richness made these strains particularly promising candidates for cocultivation-based natural product discovery, according to.10,11 The culture and morphological of fungal characterizations were done according to their macro and microscopic characteristics and compared with species characteristics of the standard taxonomic guidelines to identify the fungal genera and species as well as genetic analysis based on 18S rDNA.10,11 Rhizopus sp. was recovered from the rhizosphere of Japanese cypress (Chamaecyparis obtusa). The strains were maintained on the surface of potato dextrose medium (PDA) slants at 4 °C. The bacterial strain Bacillus subtilis ATCC 6633 was obtained from the American Type Culture Collection and maintained on the surface of nutrient agar medium (NA) slants at 4 °C.
Genome Mining of Microbial Genera
Available genomes of the fungal strains Aspergillus pseudocaelatus (A), Trichoderma gamsii (T), and Rhizopus sp. (R). In addition, the bacterial strain Bacillus subtili (B) was obtained from the database of the National Center for Biotechnology Information (NCBI) (https://www.ncbi.nlm.nih.gov/). Genome mining was conducted via antiSMASH software version 8.0. (https://antismash.secondaryspecializedmetabolites.org/) to identify the gene putatively encoding the biosynthesis of specialized metabolites. 12
Cocultivation and Extraction Process
Seed culture of each strain was prepared on PDA medium incubated for 7 days at 28 °C in static conditions. For co-culture fermentation, two 6 mm diameter discs of each strain was applied to an autoclaved flask containing 2.5 g of rice in 50 mL of water with ratio 1:1 of inoculum. For mono- and mixed cultures, three biological replicates of each condition were incubated until the desired period. The yield was then harvested after 0, 7, 14, 21 and 28 days of incubation at 28 °C by adding 100 mL of ethyl acetate, sonicated for 30 min, extracted three times, and then concentrated under vacuum. 10
Extraction and Isolation of Compounds
The large-scale fermentation of (A) and (T) for 28 days yielded 8 g of the EtOAc extract, which was subjected to chromatographic fractionation via Biotage equipped with flash column chromatography (Biotage Selekt, Sfär HC Duo column, 30 μm, 50 g) and eluted with n-hexane containing increasing proportions of EtOAc and MeOH to obtain 14 fractions (Fr1-14). Further purification steps were applied, as shown in Figure 1, to obtain four pure compounds (1- 4).

Isolation of compounds 1-4 from the ethyl acetate extract of AT.
LC-QTOF/MS Profiling
The microbial extracts were analyzed via an Agilent 1290 UHPLC system coupled with an Agilent G6545 hybrid Q-TOF/MS system. Chromatographic separation was performed via an Agilent 2.1 × 100 mm C18 Poroshell column at 40 °C with a flow rate of 0.2 mL/min. The gradient of the mobile phase was A: 0.1% formic acid in H2O and B: 0.1% formic acid in ACN starting from 5%-9% B from 1-4 min. The gradient increased to 50% B in 2 min. from 50%-70% in 9 min. This was followed by increasing from 70%-95% in 1 min and holding for 4 min. Finally, the gradient was returned to 5% B in 4 min. For each sample, 2 μL of a 1 mg/mL mixture was injected. The dual ESI source was adjusted under the following conditions: nebulizer gas pressure: 50 psi; drying gas flow rate: 12 L/min; drying gas temperature: 300 °C; and sheath gas flow rate: 11 L/min with a sheath gas temperature of 350 °C. For the parameters of the scan sources, Vcap was set at 3500 V in positive mode, the skimmer voltage was adjusted to 65 V, the octopole voltage was 750 V, and the fragmentor voltage was 130 V. The data were collected in both centroid and profile modes in the extended dynamic range (2 GHz). Calibration and tuning of the instrument were performed following procedures recommended by the manufacturer. Accurate mass spectra were obtained in the m/z range of 100-1700, and all ion MS/MS acquisition was applied with 5 scans/sec. The desired mass accuracy of the recorded ions was confirmed via continuous internal calibration during analysis with reference solution signals at m/z 121.0509 (protonated purine) and 922.0098 (protonated hexakis [1H, 1H, 3H-tetrafluoropropoxy]) in positive mode. 13 For structure elucidation, MS/MS spectra were predicted via CFM-ID 4.0 and compared with the experimental MS/MS spectra.
Data Analysis of the Mass Profile
MassHunter Profinder software B.10.00 with recursive molecular feature extraction (rMFE) was used for nontarget screening. In rMFE, all ions with similar elution profiles and m/z values were extracted as molecular features (MFs). For peak picking, the MS1 and MS2 tolerances were set to 0.03 and 0.1 Da, respectively, in centroid mode for each dataset. The data between 0.5 and 24 min were collected within the mass range of 100-1700 m/z. The extracted data were exported in Files in compound exchange (.cef format) for further analysis via the Agilent Mass Profiler Professional (MPP) software package (version 15.1) (Agilent Technologies, Santa Clara, CA, USA) for further processing of the differential analysis, and the CSV file format was exported to MetaboAnalyst 5.0 for visualization and statistical analysis. 10
Molecular Network-Based Analysis
Tandem mass spectrometry molecular networks were created via the GNPS platform (http://gnps.ucsd.edu). The data (.d) were converted to the .mzML format with MS-Convert. The converted files were used to generate an MS/MS molecular network via the GNPS data analysis workflow version release 30. The precursor and fragment ion mass tolerances were set to 0.02 Da, and the product ion tolerance was 0.02 Da. Networks were generated using six minimum matched ion fragments, a minimum cluster size of 2 and a cosine score of 0.7. The remaining parameters were kept at default values. The library spectra were filtered in the same manner as the input data. All matches kept between network spectra and library spectra were required to have a score above 0.7 and at least 6 matched peaks. 14
Antimicrobial Evaluation
The isolated compounds were prepared at a concentration of 100 µg/mL via nutrient broth dilution methods
15
for their antimicrobial efficacy against Gram-positive bacteria (Bacillus subtilis ATCC 6633 and Staphylococcus aureus ATCC25923) and Gram-negative bacteria (Escherichia coli JCM 1649). Bacterial strains were obtained from ATCC or Japan collection of microorganisms (JCM) and maintained on the surface of nutrient agar medium (NA) slants at 4 °C. The bacterial inoculum was standardized to 1 × 105 CFU·mL−1. Each well of a 96-well plate was filled with 5 µL and then 100 µL of nutrient broth media inoculated with 1 × 105 CFU·mL−1 bacterial strains in triplicate and added to each well. The plates were incubated at 37 °C for 18 h under static conditions. Sorbic acid was used as a positive control at a concentration of 100 μg/mL.
16
DMSO was used as a negative control. After the incubation period, the optical density was measured via a plate reader (BioTek Synergy HTX Multimode Reader) at OD600. The percent inhibition was calculated as follows:
Results
Microbial Coculture Combinations
Different combinations of coculturing experiments were designed to investigate the power of coculturing strategy for quantitative and qualitative enhancement of metabolite yield as shown in Figure 2 and Figures S1-7. The dual interaction after 4 weeks of incubation increased the yield of the extract and the pairwise fungal-fungal interactions, especially those of Trichoderma (T) and Aspergillus (A), followed by interactions with Rhizopus (R) and then interactions with Bacillus bacteria. The yields produced via pairwise interactions were greater than those produced via monoculture.

The metabolite yields of pairwise interactions between Trichoderma (T) and Aspergillus (A), Rhizopus (R) and Bacillus (B) bacteria after 28 days of interaction.
Metabolite Yield of Time-Dependent Fungal Cocultures
The total yield of metabolites increased gradually with increasing fermentation time; the fungal dual interaction improved the productivity of secondary metabolites, as shown in Figure 3. The metabolites yield of AT interaction elevated from 11.2 mg to 81.7 mg for 7 and 28 days of incubation, respectively. The same pattern for monoculture of Trichoderma (T) and Aspergillus (A) was observed. The yield of coculture of AT was higher than the yield of Trichoderma (T) and Aspergillus (A) in monoculture conditions.
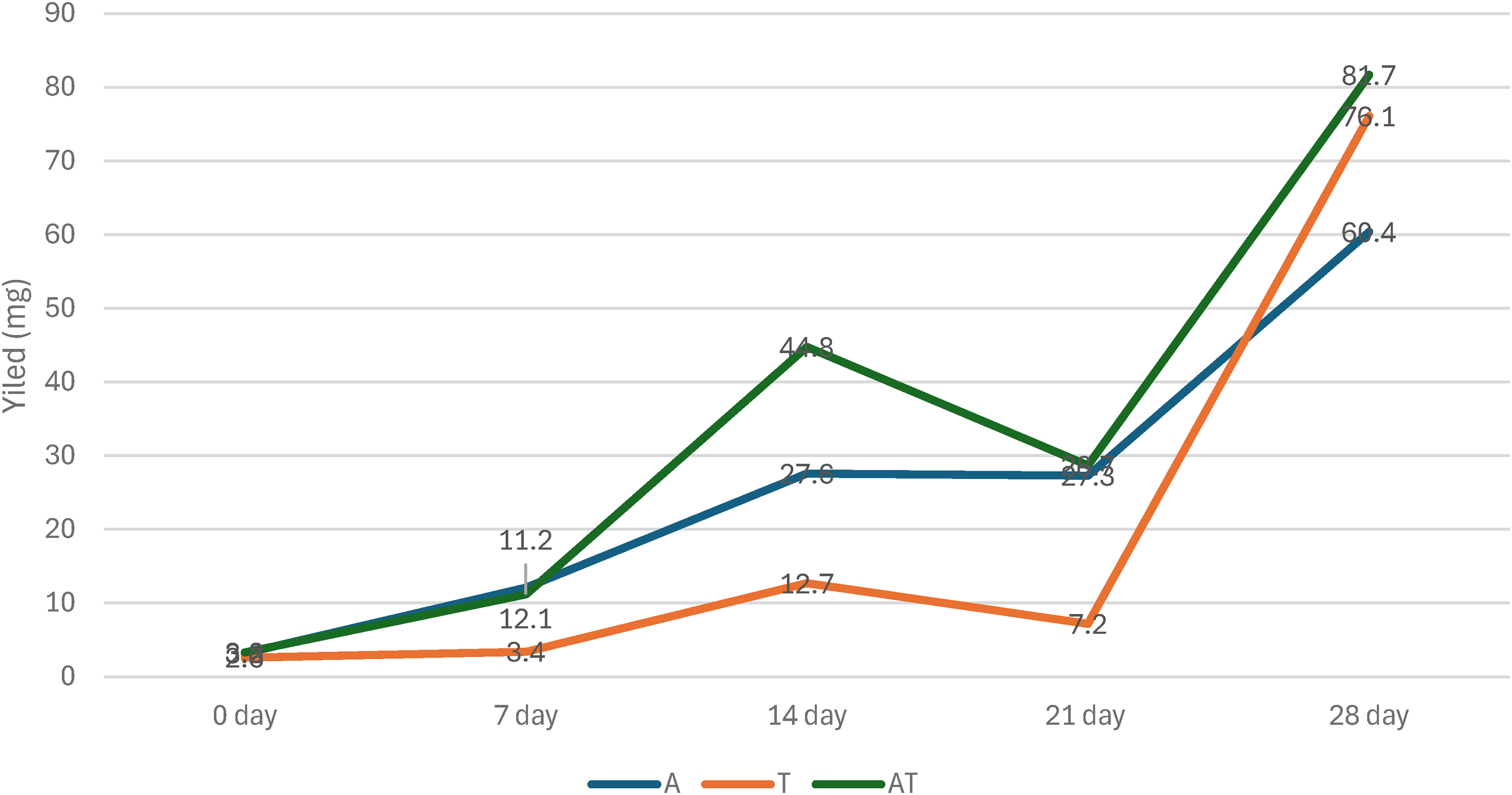
Metabolite yield of the interaction between Trichoderma (T) and Aspergillus (A) during different incubation periods (0-28 days).
Isolation and Identification
The cocultivation of (A) and (T) for 28 days on rice media extracted three times with ethyl acetate which produced 8 g. as shown in Figure 4 after vacuum drying as shown in Figure 4. The chemical investigation guided by molecular network analysis of the (AT) extract led to the isolation of two hybrid metabolites, namely, terpene-amino acid residue compounds (

Microbial culture and extracts of Aspergillus (A), Trichoderma (T), and pairwise (AT) fungi at 0 and 28 days of interaction.
Determination of Isolated Compounds
Talaroconvolutin A (
Berkedrimane A (
Compound
Phomaligol A (

Chemical structures of the isolated compounds (1-4).

Total ion chromatograms of A, T, and AT interactions.
Molecular Network Analysis
The molecular network of AT interactions constructed via GNPS included 1229 nodes connected with 1318 edges forming 35 clusters, as shown in supplementary data Figure S31. The second large cluster consisted of 21 nodes, including the node with a molecular weight of 231.138, which represents a building block unit of C15H19O2 for the hybrid molecule of drimane sesquiterpenoids conjugated with amino acid residues as shown in Figure 7. The propagation identification of this cluster resulted in the identification of five hybrid molecules, berkedrimanes A and B, and purpurides B and F in addition to minioluteumides A. The MS/MS fragmentation patterns of these molecules are presented in Figures 8–14 and Table 1.

Metabolic network cluster of hybrid metabolites produced by AT interactions.

MS/MS fragmentation spectrum of talaroconvolutin A.
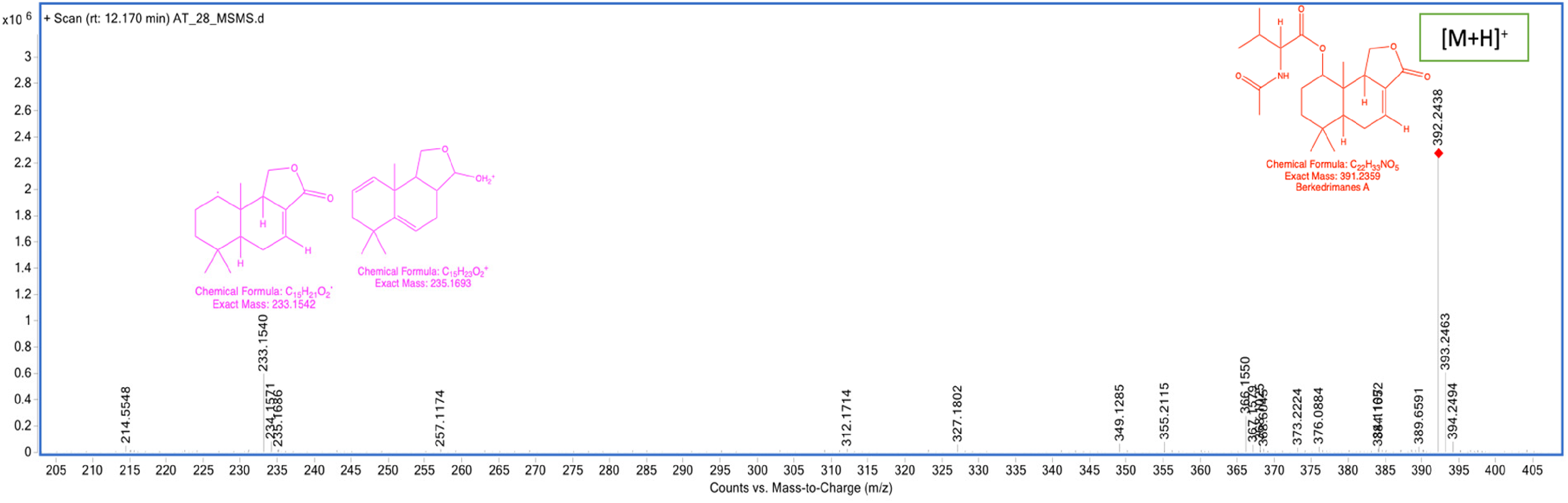
MS/MS fragmentation spectrum of berkedrimanes A.
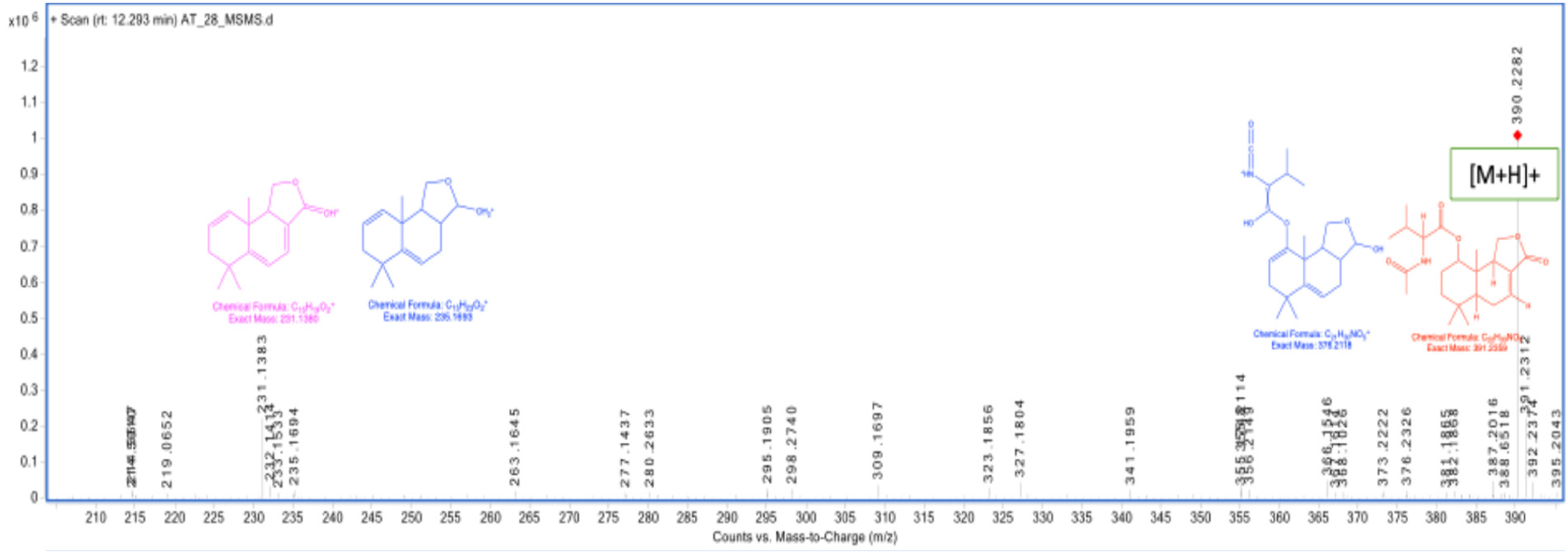
MS/MS fragmentation spectrum of purpuride B.

MS/MS fragmentation spectrum of talaminoid B.

MS/MS fragmentation spectrum of minioluteumides A.

MS/MS fragmentation spectrum of berkedrimanes B.

MS/MS fragmentation spectrum of purpuride F.
Tentative Identification of Other Hybrid Molecules in AT Interaction.

The relative quantification of detected hybrid metabolites was calculated based on the relative peak area of 1 mg of EtOAc extract. The berkedrimanes B represented the highest abundance in the profile of AT interaction followed by berkedrimanes B, then purpuride F then talaroconvolutin A, as shown in Figure 15.

Relative abundance of compounds in clusters associated with AT interactions. Relative peak area/1 mg extract weight-averaged abundance calculated from three biological replicates (n=3 ± standard error).
Biological Activity
The antimicrobial potential of the isolated compounds 1, 2, 4 was assessed. While compound 3 (spiculisporic acid) a well-characterized biosurfactant with extensively documented antimicrobial properties against pathogenic bacteria, was not tested as its bioactivity profile has been comprehensively reported in previous studies. The antibacterial activities of compound

Antibacterial efficacy of isolated compounds (1, 2 and 4) at a concentration of 100 μg/mL. Sorbic acid at a concentration of 100 μg/mL was used as a positive control (PC), which was calculated from three biological replicates (n=3 ± standard deviation).
Discussion
Metabolite Yield Enhancement During Microbial Co-Culturing
Microbial co-culturing is a powerful strategy to enhance metabolite yield and diversity. Nonaka et al 25 reported that cocultivation of Penicillium pinophilum FKI-5653 with Trichoderma harzianum FKI-5655 led to a remarkable increase in the production of Secopenicillide C. The yield increased from 23.2 mg·L-1 in monoculture to 87.4 mg·L-1 under coculture conditions, representing a 3.7-fold improvement. This increase is due to the significant impact of fungal–fungal interactions on the activation and upregulation of secondary metabolite biosynthesis-related gene clusters. Wu et al 26 demonstrated that, compared with monoculture conditions with a 1:1 inoculation ratio, coculturing Bacillus amyloliquefaciens and Trichoderma asperellum significantly increased the production of antibacterial compounds, further increasing specific amino acid yields. Zohair et al 10 found that cross interaction between bacteria and fungi, coculturing between Trichoderma and Bacillus produced 18 unique chemical features relative to monoculture and coculture of Aspergillus and /or Bacillus. Coculturing is a powerful strategy for activating silent biosynthetic gene clusters (BGCs) that remain unexpressed under standard laboratory conditions. In natural environments, microbes constantly interact, and these interspecies interactions can serve as cues to trigger the expression of cryptic or silent gene clusters, leading to the production of novel secondary metabolites 27 or improve the yield of sepcific products. Metabolomic profiling revealed several additional hybrid metabolites, including Berkedrimanes B, Talaminoid B, Purpuride F, and Minioluteumides A. While these compounds represent promising targets for future investigation, their isolation was hindered by challenges including structural similarity leading to chromatographic separation difficulty, and the prioritization of resources toward obtaining comprehensive structural and biological data for representative compound classes. Nevertheless, the detection of this diverse array of hybrid metabolites underscores the chemical richness activated through fungal cocultivation and validates the approach as a strategy for accessing rare natural product scaffolds.
Effect of Genetic Material and Species Selection
Zohair et al 10 revealed that genome analysis of Aspergillus sequence contained 95 regions of biosynthesis-related gene clusters (BCGs) expressed for a diverse range of chemical classes, including T1PKs, T3PKs, 28 regions of terpenes, indoles, NRPs, RiPP-like metabolites, and fungal RiPPs. Genome analysis of Trichoderma revealed 41 regions for the BCGs of T1PKs, terpenes, and NRPs, whereas the Bacillus bacterial genome included 14 BCGs for glycocin, epipeptide, T3PKs, and NRPs. The genome of Rhizopus includes 16 genomic regions, including four terpene-precursor regions, seven terpene regions, three NRPS-like regions, betalactone regions, and fungal-RiPP and NI-siderophore regions as shown in supplementary data (Figure S32). The unique BGCs present in the genomes of each microorganism govern their potential to produce secondary metabolites. In coculture, genetic factors not only control a microbe's inherent metabolite profile but also determine responsiveness to cocultivation signals. Some gene clusters remain silent in monocultures but are activated through interspecies interactions either via competitive, cooperative, or signaling pathways. 28 Different microbial combinations can elicit the production of distinct classes of metabolites, trigger mutually induced biosynthesis, cross-biotransformation, and, in rare cases, horizontal gene transfer of metabolic capabilities. 29 In our study we found that, the combination with different microbial species affected the production of hybrid metabolites. The detected hybrid molecule production was specific to such microbial species. As shown in Figure S33.
Chemical Importance of Hybrid Metabolites
Terpene–amino acid conjugates represent an exceptionally rare class of natural products, with only limited examples reported in the literature. This study reports the first isolation of two hybrid terpene through a cocultivation approach and their first detection from the A. pseudocaelatus–T. gamsii interaction. Notably, these metabolites were absent in monoculture fermentations, indicating that interspecies communication specifically activates their biosynthesis. Drimane sesquiterpenoids conjugated with an N-acetyl-L-valine group are rare compounds in nature that are produced by Penicillium spp,13,30 and few such hybrid natural compounds have been detected thus far. The first examples of these hybrid compounds were purpuride, reported from Penicillium purpurogenum in 1973, 31 and the others included berkedrimanes A and B from P. solitum, 31 purpurides B–D from P. purpurogenum JS03-21 32 and P. sp. ZZ1283, 33 and minioluteumides A–D from P. minioluteus PILE 14-5. 13 Berkedrimanes A and B inhibit the signal transduction enzymes caspase-1 and caspase-3 and reduce the production of interleukin-1β. 31 Berkedrimane B, purpuride, and purpurides B–D were found to have antifungal activities against Candida albicans. Purpurides B–D also have antibacterial activities.32,33 The clear chemical importance of integration of terpene and amino acid biosynthesis is enrichment of molecules with nitrogen atoms, which are rare among hydrocarbon-derived terpenoids. 34
The Diverse Chemical Structures of Hybrid Metabolites and Their Biological Efficacy
Hybrid metabolites exhibit remarkable chemical diversity, arising from the merging or modification of biosynthetic pathways of distinct organisms or molecular scaffolds. This diversity is reflected in their structural motifs, which may combine features from multiple parent compounds.
35
Compound

Structure–activity relationships of compounds (1-3).
The cocultivation strategy represents a promising approach for the discovery of hybrid metabolites and bioactive compounds. However, several limitations require further investigation. Chromatographic challenges arising from structural similarities prevented the complete isolation of all tentatively identified hybrid metabolites. Moreover, the molecular mechanisms underlying metabolite induction—including specific chemical signals, transcriptional networks, and the responsible biosynthetic gene clusters—remain uncharacterized. Finally, the biosynthetic origin of the hybrid compounds, whether from a single organism or through metabolic cooperation, has not been definitively established. Despite these constraints, this work validates cocultivation as a viable strategy for accessing rare bioactive natural products and provides valuable direction for future mechanistic and translational studies.
Conclusion
This study establishes fungal cocultivation as an effective strategy for discovery secondary metabolites through activation of silent biosynthetic gene clusters. The interspecies interaction between Aspergillus pseudocaelatus and Trichoderma gamsii induced structurally unique terpene-amino acid conjugates, demonstrating that microbial competition triggers chemical diversity. The moderate-to-strong antibacterial activities observed against Staphylococcus aureus and Bacillus subtilis highlight the therapeutic potential of these hybrid scaffolds, particularly given their selectivity toward Gram-positive pathogens a feature valuable for targeted antimicrobial development. By systematically leveraging fungal–fungal interactions guided by metabolomic analysis, and genome mining techniques, this strategy offers a promising avenue for discovering antimicrobial scaffolds needed to address the global challenge of drug-resistant infections.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X251405381 - Supplemental material for Fungal–Fungal Cocultivation Unveils Bioactive Metabolites, Including Rare Terpene–Amino Acid Hybrids, with Antibacterial Activity
Supplemental material, sj-docx-1-npx-10.1177_1934578X251405381 for Fungal–Fungal Cocultivation Unveils Bioactive Metabolites, Including Rare Terpene–Amino Acid Hybrids, with Antibacterial Activity by Moustafa M Zohair, Ahmed H El-Desoky, Fahd M Abdelkarem, Yhiya Amen, Ahmed A El-Beih and Kuniyoshi Shimizu in Natural Product Communications
Footnotes
Acknowledgments
The first author (M.Z.) is funded by a full scholarship from the Ministry of Higher Education of the Arab Republic of Egypt, and he would like to express his deepest gratitude for the financial support.
Ethical Approval
Ethical Approval is not applicable for this article.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Authors’ Contribution
MZ and KS designed the study and wrote the manuscript; MZ conducted the chemical and microbiological parts as well as the metabolomic data interpretation. AHE, FMA, YA and AAE elucidated the compounds structure and revised the manuscript. All authors participated in the critical reading of the final manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Statements and Declarations
Figures 5 and ![]() included in this manuscript were created by the authors. No copyrighted material was reproduced from external sources.
included in this manuscript were created by the authors. No copyrighted material was reproduced from external sources.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

