Abstract
Background
Fungal infection predominantly damages agricultural practices, and conventional chemical fungicides and insecticides are applied to control it, which extensively harms human health and the environment. Some bacterial species can control fungus by lysing its outer chitin layer.
Objectives
The present research aimed to isolate microorganisms capable of producing chitinase, thus acting as a highly effective biocontrol agent in combating fungal phytopathogens.
Methods
Two chitinase-producing bacterial strains were successfully isolated and screened from soil samples from a fish market environment. The process involved the aseptic collection of soil samples, followed by serial dilution to facilitate microorganism isolation. The bacterium exhibited optimal extracellular chitinase enzyme production following a 72-h incubation period at a temperature of 30 °C in a chitinase detection medium containing 0.5% chitin. Validation of chitinase production was confirmed through a clear zone assay, thus verifying its chitinase-producing capacity.
Results
Among the various isolated strains, isolates S3C1 and S3C3 demonstrated the highest chitinase activity, leading to their selection for further investigation. Comprehensive morphological and biochemical tests were conducted on these two isolates to assess their characteristics and capabilities. These tests established that both isolates were gram-negative, rod-shaped bacteria. Through genetic sequencing of the 16S rRNA gene, both organisms were identified as Klebsiella variicola exhibiting a remarkable similarity of 99% with S3C1 and S3C3 respectively. The bacteria exhibited maximum chitinase synthesis at optimal circumstances, which were determined to be a temperature of 30 °C and a pH of 7, after a 48-h incubation period. The bacteria exhibited robust antifungal activity during bioassays, demonstrating their capability to suppress the growth of fungal pathogens (specifically, Fusarium oxysporum) in vitro.
Conclusion
This research suggests a promising alternative to synthetic fungicides in agricultural practices, fostering a sustainable approach to disease management.
Introduction
Biocontrol agents’ antagonistic activity often involves the breakdown of host structures through the secretion of extracellular lytic enzymes. 1 Chitinase (EC 3.2.1.14) holds a significant role among these lytic enzymes, contributing to the biological control of various plant diseases through the degradation of chitin polymers present in the cell walls of fungus. 2 This enzymatic activity impacts various aspects of fungal growth, including germ tubes and hyphal tips. 3 Several chitinases have been successfully isolated and various bacteria have been genetically cloned to express these enzymes.1,4 In the last ten years, chitinases have attracted growing interest because of their wide array of potential uses.5,6 These applications encompass direct microbial regulation, indirect use of purified proteins, and gene modification strategies.7,8
Plant diseases inflicted by pathogenic fungi lead to substantial agricultural losses, affecting 10-20% of crop productivity. The broad spectrum of fungi—over 10,000 types—can harm numerous plant species.9-11 Macrophomina phaseolina is responsible for root diseases in over 500 plant species. These diseases encompass dry root rot, stalk rot, stem canker, and charcoal rot. 12 Fusarium species lead to damping-off and wilt diseases. Likewise, Rhizoctonia solani causes damping-off and root rot in more than 2000 plant species. 13 Managing these diseases, especially in high-temperature regions, proves challenging and leads to significant crop losses. Considering the environmental concerns associated with chemical pesticides, there is a growing interest in identifying alternatives for suppressing soil-borne plant diseases. 14 Chemical pesticides have triggered environmental and health-related issues, contributing to up to 14% of occupational injuries.15,16 Although numerous microorganisms have demonstrated antagonistic properties against soil-borne plant diseases, only a limited number have been commercially adopted as biocontrol agents. 17
Extracellular lytic enzymes lyse host structures in biocontrol agents via an antagonistic effect. 18 Fungal cell walls contain chitin polymer, which is degraded by chitinase (EC 3.2.1.14) and contribute to biological control of plant diseases. This cell wall breakdown affects fungal growth, particularly hyphal tips, and germ tubes. 5 The identification and cloning of genes encoding chitinases in bacteria have further advanced our understanding. 19 The versatile applications of chitinases have propelled them into the spotlight in recent years, whether utilized directly for microbial regulation, via purified proteins, or gene modification strategies.20,21
This study aims to isolate and characterize endochitinase-producing microbes with potential as biocontrol agents against fungal phytopathogens. We also investigated the optimal culture conditions for fungicide production by these isolates, alongside assessing their thermal and pH stability. Furthermore, the antifungal properties of this enzyme were evaluated through antifungal assays.
Materials and Methods
Isolation of Chitinase-Producing Microorganisms
A total of forty soil samples were gathered from diverse areas of fish markets and lakes in Jhenaidah, Kushtia, and Jashore regions of Bangladesh. Then 5 gm of soil samples were diluted in 50 ml sterilized distilled water, and gently stirred at 150 rpm for 30 min. Serial dilution was performed up to 10 times, and the samples were plated onto chitinase-detection agar (CHDA) plates. CHDA medium was composed of K2HPO4 (0.3 gm/L), MgSO4.7H2O (0.5 gm/L), KH2PO4 (0.7 gm/L) FeSO4. .7H2O (0.1 gm/L), MnCl2 (0.001 gm/L) peptone (5 gm/L), colloidal chitin (5 gm/L) prepared from chitin powder 22 and bacteriological agar (16 gm/L) at pH 7. Following an incubation period of three days at 30 °C, clear halos were observed around colonies. The colonies were isolated and preserved as glycerol stocks at −80 °C for subsequent investigations.
Morphological and Biochemical Characterization
The identification of the isolated chitinase-producing bacteria was conducted by morphological, biochemical and growth attributes following the guidelines outlined in Bergey's Manual of Systematic Bacteriology. 23 The morphological characteristics were assessed through techniques like viable cell count, and Gram staining. The biochemical profiling encompassed a series of tests, such as the potassium hydroxide (KOH) test, oxidase test, catalase test, indole test, urea hydrolysis test, and cultivation agar media was designed for selection and differentiation, including MacConkey agar (Lactose fermentation test), Eosin Methylene Blue (EMB) agar (a selective medium that allows isolation and identification of gram-negative bacteria), Simmons Citrate agar (Citrate utilization test), mannitol salt agar (Salt tolerant test), Bismuth Sulfite Agar (BSA) (highly selective for Salmonella species and actively inhibits most other bacteria) and starch hydrolysis test.24-29 The isolates S3C1 and S3C3 were identified based on size, shape, color, texture, etc. To optimize physical growth parameters such as temperature and pH, the development of secluded chitinase-producing bacteria was inspected under various conditions, including growth at 45 °C, alongside different temperature and pH settings on colloidal chitin (CC) medium.
Growth Optimization of Isolated Endochitinase-Producing Bacteria: Effect of Temperature and PH on Growth
The pH of the adsorption medium is the key parameter influencing absorption capacity. The medium was adjusted to pH 5-9, and bacterial strains were inoculated under contamination-free conditions in a laminar air cabinet. After inoculation, flasks were incubated on an orbital shaker at 150 rpm for 72 h, and growth was monitored by measuring optical density with a spectrophotometer at 660 nm. Nutrient broth (NB) medium with pH 7 was used for isolated bacteria, and incubation occurred in a biosafety level 2 cabinet. Their growth efficiency was measured at various temperatures ranging from 28 °C to 40 °C over 72 h.
Molecular Identification
Genomic DNA was isolated by boiling method as described by Rawool et al 30 and used a template for PCR using 27R and 1492 R primer.31,32 The 16S rRNA gene was then amplified through PCR using an automated thermal cycler (PTC-200, M J Research Inc.). 33 The PCR process comprised 35 cycles, involving denaturation at 92 °C for 1 min and 10 s, followed by annealing at 48 °C for 30 s, and the extension for 2 min and 10 s at 72 °C. Following amplification, the PCR products were purified before being sent to INVENT Technologies in Malaysia for cycle sequencing. The 16S sequences were aligned by CodonCode Aligner software. The aligned sequences of the 16S rRNA gene were matched against the existing sequences within the GenBank database (https://www.ncbi.nlm.nih.gov/genbank/) using BLAST, and multiple sequence alignment tools to identify the isolated chitinase-producing bacteria. The sequence was submitted to the GenBank database.
Phylogenetic Analysis
The 16S rRNA sequence of the isolated chitinase-producing bacteria and other chitinase-producing bacteria were collected to evaluate the phylogenetic relationship. Finally, a neighbor-joining (NJ) phylogenetic tree was constructed for the bacterial isolates using MEGA 11 software. 34
In Vitro Antifungal Activity
The evaluation of antifungal efficacy against Fusarium oxysporum was conducted using a dual culture method. 35 Circular fungal mycelium discs were positioned at the center of potato dextrose agar (PDA) plates, adjacent to bacterial strains. After an incubation of 48 h at 28 °C, the inhibition of fungal growth was quantified as a percentage in comparison to the control.
Statistical Analysis
All studies were performed in triplicates and data were analyzed and plotted using Microsoft Excel (Office 2013). Mean values and standard deviation were calculated by Microsoft Excel.
Results
Screening of Chitinase-Producing Bacteria from Soil Samples of Fish Markets and Lakes
Chitinase-producing microbes were successfully isolated and the colonies were cultivated on colloidal chitin-containing chitinase detection media plates, resulting in diverse clear zones after incubation for three days at 30 °C temperature. The microbial cultures underwent further cultivation for the identification of chitinase-producing bacteria, confirmed by the presence of clear zones indicating chitinase activity (Figure 1). Following an incubation period of three days at 30 °C, clear halos were observed around six colonies. Among these, primarily four colonies were selected depending on their clear zones in the chitin hydrolysis assay.

Cultivation of Microbes on Colloidal Chitin-Containing Chitinase Detection media with Clear Zones of Chitin Hydrolysis.
Identification of Isolated Chitinase-Producing Bacteria
Among these four colonies, chitinase-producing bacteria S3C1 and S3C3 were selected based on the extent of clear zones produced during hydrolysis assays (Table 1). The highest zone of chitinolysis is shown around the colony of S3C1 (18 mm) and S3C3 (12.5 mm) (Figure 2). A clear zone around the bacterial colony indicates the chitinase-producing ability of the bacteria. Two isolates, designated as S3C1 and S3C3, were preserved as glycerol stocks at −80 °C for subsequent investigations.
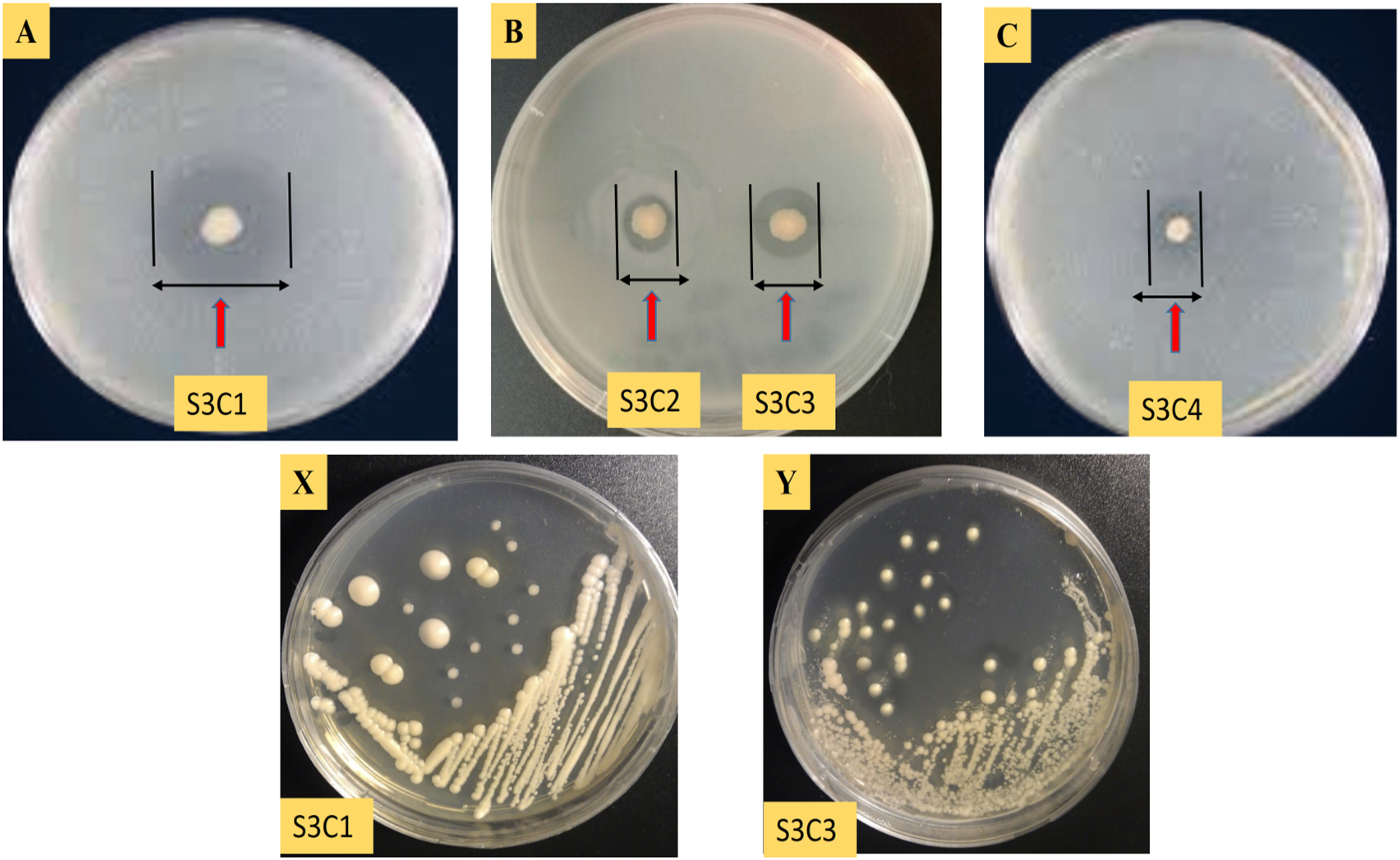
Zone of Chitinolysis Assay for (A) S3C1, (B) S3C2 and S3C3, (C) S3C4. Pure Culture of Isolated Endochitinase Bacteria on Colloidal Chitin (CC) Containing Medium, (X) S3C1 and (Y) S3C3.
The Result of Chitin Hydrolysis Assay of Isolated Chitinase-Producing Bacteria with Zone of Chitinolysis.

Morphological and Biochemical Characterization
The pure isolated colonies were grown on different selective media and they were identified based on character and growth appearance. Maintaining a suitable plating method, the result was observed after 48 h, both isolates S3C1and S3C3 showed creamy white, round, large colonies with observed 89 × 1015 / 1 ml and 44× 1020 / 1 ml cell density. These isolated chitinase-producing bacteria demonstrated attributes of gram-negative, small rod-shaped bacteria. Positive results were observed in the potassium hydroxide test, catalase, urease, citrate utilization, and lactose fermentation. Conversely, these microbes yielded negative outcomes in tests like oxidase, indole, BSA, salt tolerant test, and starch hydrolysis test (Figure 3). A comprehensive overview of their morphological, and biochemical properties is presented in Table 2.

Morphological and Biochemical Test Results of Chitinase Enzyme-Producing Bacterial Isolates. Here, left one represents S3C1 results and the right one represents S3C3 results. Both (a) and (i) showed gram-staining results of S3C1 and S3C3 respectively. Both isolates are gram-negative and rod- shaped. Figure (b) and (j) denotes the positive results of KOH test. KOH test is done to differentitate between gram-positive and gram-negative bacteria based on their cell wall structure, with gram-negative bacteria lysing in KOH solution. The catalase test results of both isolates are presented in Figures (c) and (k). The presence of bubbles for bacterial strains indicates positive results in the catalase test. (d) and (l) show the negative results of both isolates S3C1 and S3C3 for oxidase test. Organisms lacking cytochrome c as part of their respiratory chain do not oxidize the reagent, leaving it colorless within the limits of the test, and are oxidase-negative. Both isoaltes showed negative results in the indole test. Results are shown in Figures (e) and (m). A positive result is shown by the presence of a red or red- violet color in the surface alcohol layer of the broth. A negative result appears yellow. In Figures (f) and (n), both isolates produced color around the colony on MacConkey agar media so that was lactose fermenting. Both isoates S3C1 (g) and S3C3 (o) showed growth on EMB agar indicating, this is gram-negative bacteria. Both the isoaltes showed bright pink color in Figures (h) and (p). Thus, it showed a positive result for the urea hydrolysis test. The citrate test result is presented in the Figures (i) and (q). After bacterial inoculation, the blue color indicates that the isolates are citrate-positive. Figure numbers (i), (j), (k) and (r), (s), (t) showed negative results for both isolates on BSA, Mannitol Salt Agar, and Starch Hydrolysis test respectively. After studying all the morphological and biochemical assay, isolated endochitinase bacteria S3C1 and S3C3 may exhibit the characteristics of Klebsiella Genus.
Morphological and Biochemical Characteristics of Chitinase-Producing Bacterial Isolates, S3C1 and S3C3.

Growth Optimization of Isolated Endochitinase-Producing Bacteria
Effect of Temperature and pH Variations on the Growth of Isolated Endochitinase-Producing Bacteria
The growth temperature preferences of isolates S3C1 and S3C3 were investigated across a temperature range from 28 °C to 40 °C and maximum growth occurred at 30 °C after 48 h of incubation for both S3C1 and S3C3. The growth was monitored by measuring optical density with a spectrophotometer at 660 nm. Recorded data are presented in Figure 4.

Effects of Temperature Variations on the Growth of the Isolates (a) S3C1, and (b) S3C3. The Highest Growth was Recorded at 30 °C After 48 h and the Lowest Growth at 28 °C After 12 h of Incubation.
Additionally, the optimal growth conditions were determined for pH 5–9 at an optical density of 660 nm. The isolated endochitinase-producing bacteria exhibited a limiting growth temperature of 60 °C, while their maximum growth was noted at 30 °C. The effects of pH on the growth of bacterial isolates are illustrated in Figure 5.

Effects of pH Variations on the Growth of the Isolates (a) S3C1, and (b) S3C3. Maximum Growth Occurred at pH 7 After 48 h of Incubation for Both S3C1 and S3C3 Isolates and Minimum Growth was Recorded at pH 5 After 12 h of Incubation.
Molecular Identification of Isolates
The 16S rRNA gene amplification by PCR yielded a distinct band at approximately 1.5 kb. Verification of the isolate's classification was achieved through 16S rRNA gene sequencing. Figure 6 illustrates the gel electrogram band of the amplified PCR products and chromatogram of sequenced bacterial isolates. All sequenced data were deposited in NCBI GenBank (https://www.ncbi.nlm.nih.gov/), revealing 99% similarity of both S3C1 and S3C3 with Klebsiella variicola (Table 3).

(a) Gel Electropherogram Showing a Band of Approximately 1.5 kb Length of the Amplified PCR Product of 16S rRNA Gene of the Isolates, [M Denotes DNA Ladder (Marker). A Denotes Isolate S3C1 and B Denotes Isolate S3C3]. (b) Chromatogram of 16S rRNA Gene Sequencing of the Bacterial Isolate S3C1, and (c) Chromatogram of 16S rRNA Gene Sequencing of the Bacterial Isolate S3C3.
Klebsiella variicola Produced a Significant 99% Identity with the 16S rDNA Gene Sequence of the Isolate S3C1 and S3C3 Respectively.

Phylogenetic analysis by fast minimum evolution method (illustrated in Figure 7), further affirmed that both isolates S3C1 and S3C3 were most closely aligned with Klebsiella variicola. The isolated chitinase-producing Klebsiella variicola showed a close relationship with other Klebsiella species.

Phylogenetic Tree Showing Relationships Between Isolate S3C1, Isolate S3C3, and Several Other Strains of the Bacterium Based on Their 16S rRNA Gene Sequences.
Antifungal Activity
Both isolated chitinase-producing Klebsiella variicola, S3C1 and S3C3 displayed antifungal activity against the phytopathogenic fungus F. oxysporum after 48 h of incubation. Such species can be used to control pests and are known as biological control agents. Both the isolates, S3C1and S3C3 showed antifungal activity because the bacteria suppressed the growth of fungal phytopathogen. Table 4 represents the observed growth inhibition which was 51.78% and 49.26% for F. oxysporum strains by S3C1 and S3C3 respectively indicating the potential of these isolates as effective antifungal agents.
In-Vitro Antagonistic Activity of Isolates S3C1 and S3C3 Against Fusarium oxysporum by Dual Culture Method.

The results of antifungal activity of bacterial isolates, S3C1 and S3C3 via chitinase enzyme production are illustrated in Figure 8.

The Results of the in-vitro Antifungal Activity Test of Isolates: (a) Control, (b) S3C1, and (c) S3C3. Both Endochitinase Bacteria can Effectively Control Fusarium oxysporum Fungal Phytopathogen.
Discussion
The isolation and screening of endochitinase-producing bacteria played a pivotal role in this study. Chitin, a fundamental component of various natural systems including aquatic environments, undergoes degradation and hydrolysis by bacteria, making them significant contributors to this process. 36 The enzymatic hydrolysis of chitin generates products of varying sizes, endowed with multifaceted applications such as fungicides, pesticides, and medical treatments.37-39
The regulatory mechanism governing endochitinase enzyme expression in bacteria involves an inducer as well as a repressor system. Chitin acts as an inducer, while glucose often operates as a repressor. 40 In a queue, the present study utilized colloidal chitin as a substrate to stimulate chitinase production from diverse bacteria. Detection of chitinase production was achieved through the appearance of clear zones surrounding bacterial colonies. Among the isolates, S3C1 and S3C3 exhibited substantial chitinase activity when cultivated in colloidal chitin-containing media, warranting their selection for further investigation. Subsequent identification revealed that both isolates S3C1 and S3C3 were classified as Klebsiella variicola. Notably, the culture broth of Klebsiella variicola was grown aerobically in a chitinase production medium, and exhibited anti-fungal activity, effectively inhibiting the growth of pathogenic fungi.
Morphological and biochemical assessments reaffirmed the gram-negative, rod-shaped nature of the isolates. Positive results were obtained for the KOH test, EMB agar, urea hydrolysis, citrate utilization, catalase, and lactose fermentation, while negative outcomes emerged for, oxidase, indole, starch hydrolysis, and mannitol salt agar tests, aligning with the Bergey's Manual of Systematic Bacteriology (taxonomy). 30 °C temperature was set as the optimal temperature, pH 7 was maintained as the desired pH, and the incubation period was 48 h.
The chitinase activity plays a pivotal role as a biocontrol agent against deadlier fungi, like – Aspergilllus species, Fusariuim species, Trichoderma species, Alterneria species. In this study, fungal phytopathogen is effectively controlled at about 51.78% and 49.26% in in-vitro bioassay for F. oxysporum strains by S3C1 and S3C3 respectively indicating the potential of these two isolates as effective antifungal agents. The production of microbial chitinases attracted great attention in the last few decades and microorganisms which produce a complex of mycolytic enzymes like chitinase, are considered to be possible biological control agents.
Serratia marcescens is a gram-negative, rod-shaped bacterium that was also isolated from the soil sample. Serratia marcescens is capable of producing chitinase enzymes and degrading chitin at the substrate level. Showed positive results for the indole test, citrate utilization test, and the hydrolysis of chitin while negative for the catalase test, methyl red test, and others. 41
The production of chitinases is not exclusive to bacteria; Aspergillus species, particularly A. terreus, have garnered attention for their chitinase activity. 42 Similarly, bacteria such as Serratia marcescens, Acinetobacter ASK18, and Aeromonas hydrophila have been identified for their chitinolytic capabilities.43,44
Furthermore, Cellulosimicrobium funkei and Streptomyces laurentii were found to excel in the degradation of chitin, displaying chitinase activity when grown under optimal conditions. 45 The 30 °C temperature and pH 7 observed in this investigation resonates with the optimal conditions for various other bacterial species such as Enterobacter sp., Zymomonas sp., Bacillus sp., B. cereus, and Streptomyces sp..46,47 Each bacterial species exhibits distinct optimization conditions to achieve peak enzyme production. Physiochemical characterization of the produced chitinase enzyme and determination of specific enzyme activity are essential parts of this study. However, these were not done which indicates the limitations of the study to understanding the structure, characteristics, activity, and molecular weight of the enzyme. 48 Klebsiella variicola is a diazotrophic soil bacterium and a member of Klebsiella pneumoniae species complex, worldwide used as a bio-fertilizer. Rice, bananas, and sugarcane plants are some prominent sources of this bacterium. It can be utilized in wastewater treatment, bioremediation, and biodegradation. Furthermore, it fixes nitrogen, produces ammonia, acetic acid, and acetoin, and has alkalotolerance and halo-tolerance activities.49,50 However, certain strains of K. variicola can act as opportunistic pathogens and have been associated with battlefield-related trauma, infection in patients with comorbidities or immunosuppressive patients, endodontic infections, etc.51-53 This raises concerns regarding the safety of plant pathologists. A great extent of biosafety measures should be taken by researchers, and pathologists while handling it and they have to ensure that the final endochitinase enzyme doesn’t pose any harm to human.
Harnessing chitinase-producing bacteria as biological control agents holds promise as a sustainable alternative to harmful insecticides for safeguarding plants against chitin-containing pathogens like fungi.
While our current study lays a crucial foundation for understanding the biocontrol potential of the two identified Klebsiella variicola strains as biocontrol agents, additional research is essential to uncover their full potential to solidify our research. Although our main aim for this investigation was to identify and isolate endochitinase-producing bactetia and assess its potential antifungal activity against a specific fungus (Fusarium oxysporum), we are planning to expand this project. In the future, we are aiming to include a wide range of fungal species and evaluate the antifungal activities in potted plants and greenhouse settings to explore isolated bacterial strains’ antifungal mechanisms broadly.
Conclusion
Infection of pathogenic fungi not only hampers human health but also severely damages crops and thus results in the decline of the economy. Two strains of gram-negative bacteria Klebsiella variicola named S3C1 and S3C3 were isolated from fourteen local area soil samples. Klebsiella variicola would further be subjected to purification of the enzyme produced, and hence the active principle could be evaluated as an effective biocontrol agent of fungal phytopathogens. Morphological and biochemical tests confirmed that both isolates were gram-negative, rod-shaped bacteria which showed positive results for EMB agar test, urea hydrolysis test, catalase test, lactose fermentation, citrate utilization, MacConkey agar test, while it showed a negative result for oxidase test, indole test, starch hydrolysis test, mannitol salt agar test, and bismuth sulfite test. Then the final identification was done using molecular techniques via 16S rRNA gene sequencing. These isolated bacterial strains effectively controlled Fusarium oxysporum fungi by excreting endochitinase enzyme. The selection of antagonistic microorganisms to perform biological control is considered an alternative practice of synthetic fungicides. In the era of biotechnology, microbes are most useful for mankind. They are exploited for antibiotic production, enzyme preparation, biomass and single-cell protein synthesis, production of medicinal components (amino acids, vitamins, steroids), and waste management. Microbial enzymes have multidimensional uses starting from the food industry to pollution control. Among the microbial enzymes, chitinase has a promising market value. The demand for microbial chitinase is increasing day by day. Several agro-industry used chitinase as a fungicide to reduce the production cost of their crops. This step is an eco-friendly process and a great progress towards a safe world. An approach to protein engineering makes the enzyme more thermostable, oxidant resistant, and chelator insensitive along with a better pH profile. All these factors increase the potential uses of chitinases in different sectors. Chitinases are efficiently used in photographic products, cement, pharmaceuticals, chelating agents of heavy metals, cosmetics, and single-cell protein production. In most cases, chitinases from bacterial sources act as a fungicidal compound. Thus, chitinases are fairly excellent industrial enzymes and the demand will always be high in the near future. So, this study offers a sustainable solution for protecting plants from chitin-containing pathogens such as fungi.
Footnotes
ORCID iDs
Ethical Considerations
The Animal Welfare and Experimentation Ethics Committee of the Biological Sciences Faculty of Islamic University, Kushtia approved this study.
The study needed Ethical approval due to the following reasons and for the following areas:
We collected the soil samples from regular fish markets in the local area. We isolated the chitinase-producing bacteria from the fish market soil samples, maintaining the recommended biosafety levels. We evaluated the efficiency of the isolated chitinase-producing bacteria against fungal phytopathogens. Fungi are the main culprits for many plant diseases and cause huge damage to plants. We tried to control the fungal pathogen by using chitinase-producing bacteria, which are usually non-pathogenic to humans and other animals.
For these, we needed to follow some ethics and thus needed ethical approval.
Author Contributions
Md. Rezuanul Islam, Mohammad Abu Hena Mostofa Jamal, and Md. Khasrul Alam: Conceived and designed the experiments; Wrote the paper.
Munzura Khatun, Mst. Lutfa Khatun: Performed the experiments; Wrote the paper.
Md. Al-Amin Milon, Masuma Anzuman: Analyzed and interpreted the data; Wrote the paper.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Ministry of Science and Technology (MoST), Government of the People's Republic of Bangladesh (Grant No: 39.00.0000.009.99.024.22-193/BS-239).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
Data associated with this study has been deposited at “NCBI DNA sequence database” under the accession number (OR596960; OR594155).
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
