Abstract
Objective
This study aimed to screen and identify central genes associated with the progression of non-alcoholic fatty liver disease (NAFLD) to hepatocellular carcinoma (HCC), to establish prognostic and clinical prediction models based on the expression of these genes, and to evaluate the intervention of a traditional Chinese medicine (TCM) formula—SPXJF—on the development of NAFLD into HCC through network pharmacology analysis and in vitro and in vivo validation.
Methods
Datasets for analysis were retrieved from multiple public databases using bioinformatics techniques. Through network pharmacology analysis, SPXJF's active components and their related targets were assessed, while in vitro experiments validated the inhibitory effects of SPXJF on the growth and migration of HepG2 cells.
Results
The results revealed that the expression levels of the two central genes, regulator of cell cycle gene (RGCC) and atypical chemokine receptor 3 (ACKR3), were closely tied to the survival prognosis of HCC patients. Furthermore, SPXJF significantly inhibited the proliferation and migration of HepG2 cells.
Conclusion
RGCC and ACKR3, as hub genes, significantly influence HCC patient survival during “NAFLD-HCC” progression. The risk scoring system and prediction model perform well. In vitro experiments showed that SPXJF could inhibit the proliferation and invasion of hepG2 cells. These findings support the discovery of molecular targets for early diagnosis and drug intervention in HCC. They also underscore TCM's potential in anti-HCC strategies and clinical application.
Introduction
Hepatocellular carcinoma (HCC) is recognized as one of the most prevalent malignancies worldwide and ranks as the third leading cause of cancer-related mortality globally. According to the GLOBOCAN 2020 report, HCC registers over 900 000 newly diagnosed cases annually, with approximately half of these cases reported within the Chinese population. 1 Unfortunately, the overall prognosis for HCC remains dismal, necessitating effective prevention and intervention strategies. 2 HCC and intrahepatic cholangiocarcinoma (ICC) represent the primary classifications of primary liver cancer. Additionally, angiosarcoma, hemangiosarcoma, and hepatoblastoma, although less common, are also encompassed within this category. It is pertinent to mention that HCC originates from hepatocytes, the predominant parenchymal cells of the liver, while ICC originates from the bile duct. HCC constitutes more than 80% of primary liver cancer cases globally. 3 This study identifies the main risk factors for HCC, including hepatitis B or C virus infection, chronic alcohol addiction, obesity-related non-alcoholic fatty liver disease (NAFLD), and diabetes mellitus. 4 NAFLD constitutes a variety of liver conditions, extending from the comparatively benign steatosis to the more harmful non-alcoholic steatohepatitis (NASH). Notably, NAFLD represents the prevailing liver ailment, exhibiting a worldwide prevalence of approximately 25%. 5 NAFLD is recognized as a major contributor to HCC, closely linked to metabolic disorders such as insulin resistance, hyperlipidemia, and type 2 diabetes. 6 Numerous treatment modalities are accessible for HCC in its early stages, including surgical resection, liver transplantation, and local ablation. Transarterial chemoembolization therapy is also viable in the intermediate stages. Nevertheless, a considerable portion of HCC patients are diagnosed during the middle to late stages, severely curtailing treatment options and leading to less favorable prognoses. 7 Given the criticality of early detection, this research aims to construct an HCC risk scoring system and clinical prediction model through the use of bioinformatics, specifically addressing NAFLD-HCC progression. The primary objective is to evaluate the survival prognosis of HCC patients and provide theoretical support for early diagnosis, emphasizing the significance of early intervention in improving patient outcomes. By addressing the key clinical challenges associated with HCC, including screening for hub genes of NAFLD progression to HCC and developing predictive models for HCC survival prognosis based on these hub genes, this study endeavors to contribute to the advancement of knowledge and enhance the management of HCC in clinical settings. Traditional Chinese medicine (TCM), with its holistic approach and principles of syndrome differentiation and treatment, offers unique advantages in early intervention and throughout the treatment of HCC. A formulation called “Shipi-Xiaoji recipe” was tailored to strengthen the spleen and disperse accumulation. This prescription, comprising Astragalus, Ginseng, Poria, Ligustrum lucidum, Epimedium, Pyrite, Peach Kernel, Ground Beetle, Curcuma, Pinellia, Prunella, Gecko, and Spiny Jujube Kernel, notably improved the clinical survival benefits for patients with HCC.
Cancer represents a prominent etiology of morbidity and mortality on a global scale. Consequently, an imperative arises to ascertain novel biomarkers or distinguishing characteristics conducive to the timely identification and prognostication of this disease. 8 Building upon prior clinical and experimental studies conducted by the team, this study endeavors to further explore molecular targets related to early diagnosis, staging, and intervention of HCC. To achieve this, the study employs the “NAFLD-HCC” progression as a starting point and utilizes bioinformatics to construct an HCC risk scoring system and clinical prediction model. These tools aim to assess the survival prognosis of HCC patients, while concurrently investigating the molecular targets and mechanisms of action associated with SPXJF intervention in NAFLD-HCC progression. The ultimate goal is to provide essential theoretical support for early HCC diagnosis, identify molecular targets for drug intervention, and offer insights into the direction for anti-HCC measures and the clinical integration of Chinese medicine.
Materials
Cells
HepG2 cells were purchased from Guangzhou Gineo Biotechnology Co.
Animals
15 SPF-grade male SD rats, body mass 220∼240 g, purchased from Hunan Slake Jinda Laboratory Animal Co. Ltd, animal qualification certificate No. SCXK (Xiang) 2019-0004. All experimental animals were handled in accordance with the standard of “Quality Management Measures for Experimental Animals” issued by the State Science and Technology Commission, approved by the Animal Ethics Committee of Hunan University of Traditional Chinese Medicine, and approved by the Animal Ethics Committee of Hunan University of Traditional Chinese Medicine. The approval number is LL2022070501.
Drugs and Reagents
SPXJF consisted of Astragalus, Ginseng, Poria, Chasteberry, Epimedium, Epimedium, Shihmian, Peach kernel, Tupelo, Prevotella, Ulmus, Fagopyrum, Sempervirens, Lobelia, and Gecko, all of which were purchased from the Affiliated Hospital of Hunan Academy of Traditional Chinese Medicine. Penicillin-streptomycin solution (No. PB180120), Dulbecco's modified Eagle's medium (DMEM) medium (No. PM150210), 0.25% trypsin (No. PB180226), phosphate-buffered saline (PBS, No. PB180327), and fetal bovine serum (FBS, No. 164210) were purchased from Wuhan Pnosay Life Science Co. kit (Item No. C0038) and EdU Cell Proliferation Kit (Item No. C0071S); anti-regulator of cell cycle gene (RGCC) (Item No. 19556-1-AP), anti-ACKR3 (Item No. 20423-1-AP), anti-β-actin (Item No. 81115-1-RR) primary antibodies, and HRP-coupled secondary antibodies (Item No. SA00001-2) were purchased from Wuhan Sanying Biotechnology Co.
Instruments
The 3427 Cell Culture Box (ThermoFisher, USA), BHC-1300IIA/B2 Bio-Clean Safety Cabinet (Suzhou Purification Equipment Co., Ltd), Cytation3 Enzyme Labeling Instrument (Molecular Devices, USA), Eclipse Ts2-FL Inverted Fluorescence Microscope (Nikon, Japan), CP2251 Electronic Balance (Sartorious Stedim Biotech, Germany), T100TM Thermal Cycler Fluorescence Quantitative PCR Instrument (Bio-Rad, USA), Protein Electrolyzer (Nikon) CP2251 Electronic Balance (Sartorious Stedim Biotech, Germany), T100TM Thermal Cycler Fluorescence PCR (Bio-Rad, USA), Protein Electrophoresis and Transfer Instrument (Bio-Rad).
Methods
Screening and Identification of “NAFLD-HCC” Hub Genes
The GSE63067, GSE99807, and GSE76427 datasets were downloaded from the Gene Expression Omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo/). In addition, gene expression profiles and clinical data from 374 HCC and 50 normal control samples were obtained from The Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov/) (TCGA-LIHC). Immunohistochemical picture collected from the Human Protein Atlas (HPA) database (https://www.proteinatlas.org/). To explore differentially expressed genes (DEGs) within the GSE63067 and GSE99807 datasets, we employed the limma R package. 9 The threshold for the DEGs was set as P-value <0.05 and |log 2 fold change (FC)| ≥1. By using the R package (ggplot2), we take the intersection of two gene expression profiles, which can be identified as “NAFLD-HCC” hub genes. The intersecting DEGs of NAFLD and HCC were used for further analysis.
Prognostic and Clinical Prognostic Modeling of “NAFLD-HCC” hub Genes
The survival analysis based on median hub gene expression was conducted on 373 HCC patients from TCGA-LIHC using the R packages survival and survminer. A log-rank P-value <0.05 was considered the threshold for statistical significance. Additionally, to evaluate the predictive efficacy of hub genes for 1-, 3-, and 5-year overall survival (OS) in HCC patients, the R package timeROC was utilized to generate time-receiver operating characteristic curves (time-ROC). Prediction accuracy was assessed by the area under the curve (AUC) values of the Time-ROC curve.
To calculate the risk score, multivariate Cox regression analysis was employed to analyze the combined effect of “NAFLD-HCC” hub gene expression on survival, utilizing survival outcome and survival time as dependent variables. For survival analysis, we utilized the R package survminer. We conducted the survival analysis using the GSE76427 dataset as external validation data. The comparison of survival between the two groups was performed using the Kaplan-Meier curve and log-rank test with the R package survminer.
Subsequently, univariate and multivariate Cox regression analyses were conducted to ascertain the independence of the predictive effect of the prognostic model from clinical factors. The predictive effect encompassed the risk score, age, gender, BCLC stage, and TNM stage. Following this, a nomogram integrating risk scores and other clinical factors was constructed to forecast the likelihood of 1-, 3-, and 5-year OS using the R packages RMS and survival. A P-value <0.05 was considered the threshold for statistical significance. Calibration analysis was then performed to validate the nomogram prognostic model, assessing its predictive power.
Network Pharmacologic Analysis of the SPXJF Intervention in the Progression of “NAFLD-HCC”
We used network pharmacology to further investigate the relationship between the formula for consolidating the spleen and eliminating accumulation and the genes driving the progression of “NAFLD-HCC.” The active ingredients of the SPXJF were retrieved from TCMSP 10 (https://tcmsp-e.com/), TCMID 11 (http://47.100.169.139/tcmid/search/), and TCMIP 12 (http://www.tcmip.cn/TCMIP/index.php/Home/Index/index.html) databases. The SMILES codes representing the chemical structures of the active fractions in Spleen Dissipating Drink were obtained from the PubChem database 13 (https://pubchem.ncbi.nlm.nih.gov/) and subsequently inputted into the Swiss Target Prediction database 14 (http://www.swisstargetprediction.ch/) to retrieve the drug targets associated with the spleen-eliminating drink.
Spearman correlation analysis was conducted using the R language limma package 9 to assess the relationship between the “NAFLD-HCC” hub genes RGCC and ACKR3. The screening criteria for correlation coefficients were set as |cor| > 0.3 and P < 0.05. The intersection of RGCC, ACKR3, and the drug targets of SPXJF allowed for the identification of the targets of SPXJF that intervene in the progression of NAFLD-HCC (ie, the targets affected by SPXJF intervention). These intervening targets were then plotted in conjunction with the targets of SPXJF. Finally, a co-expression heatmap was generated to illustrate the co-expression patterns of RGCC and ACKR3.
Protein interaction analysis of the intervention targets of SPXJF was conducted using the STRING database. 15 Subsequently, a protein-protein interaction (PPI) was constructed using Cytoscape 3.9.1. To gain insight into the functional implications of the gene sets, functional annotation was performed through GO (Gene Ontology) and KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway enrichment analysis.
In Vitro Validation of SPXJF Intervention in the Progression of “NAFLD-HCC”
Drug-Containing serum Preparation
Each herbal slice of SPXJF was placed in a decoction pot, to which distilled water was added in a 1:1 ratio followed by stirring. The mixture was soaked for 30 min and then brought to a boil. After achieving a boil, the concoction was decocted for an additional hour. The resulting decoction was then poured out and filtered. The herbal residue was subjected to two more cycles of decoction, following the aforementioned procedure. The decoctions were combined and concentrated to obtain a final concentration of 2.0 g/mL, which was then stored in a refrigerator at 4 °C.
Sprague-Dawley rats were randomly assigned into two groups: the control group and the SPXJF group. The concentrated decoction of SPXJF was administered orally to the rats in the SPXJF group, with the dosage calculated to be 12.42 g·kg−1·d−1, over a continuous 7-day period. The control group was gavaged with an equivalent volume of distilled water. One hour after the last gavage, blood was sampled from the abdominal aorta, and serum was isolated. Serum samples from rats within the same group were pooled, heat-inactivated at 56 °C in a metal bath for 30 min, and then filtered through a 0.22 μm filter to prepare SPXJF medicated serum, which was made free of bacteria through this process.
Cell Culture
Cells were in DMEM (Gibco BRL, Grand Island, NY, USA) with 10% fetal bovine serum (Hyclone, Logan,UT, USA), 1% penicillin, 1% streptomycin, and 1% MEM non-essential amino acid. The cells were incubated in a humidified atmosphere of 95% air and 5% CO2 at 37 °C.
Cell Inhibition Rate by Cell Counting Kit-8
Approximately 2 × 105 cells/mL were seeded into 96-well culture plates and treated with or without SPXJF at 48 h. After treatment, 100 μL of the new media was replaced and continuously 10 μL of Cell Counting Kit-8 (CCK-8) solution was added into the wells and the well plate was incubated at 37 °C for 1 h. The absorbency at 450 nm was measured using a Bio-Rad micro-plate reader. Cell inhibition rate was calculated according to the following equation. Cell inhibition rate (%) = (ODcontrol − ODsample)/(ODsample − ODblank) × 100%
EdU
HepG2 cells, during their logarithmic growth phase, were seeded within a 24-well plate at a density of 3 × 104 cells per well. The cells were cultured in media containing various concentrations (0%, 1%, 5%, 10%) of SPXJF, creating distinct groups: control, SPXJF-L, SPXJF-M, SPXJF-H, and were incubated at 37 °C for 24 h. For the assessment of cellular proliferation, an 5-ethynyl-2′-deoxyuridine (EdU) cell proliferation assay kit was utilized according to the manufacturer's instructions. Each well received 500 μL of the diluted EdU working solution and incubation continued for an additional 3 h. Subsequently, the culture medium was discarded, and the cells were fixed at room temperature with 4% paraformaldehyde for 15 min. The fixative was then removed, and the cells were washed three times with PBS, with each wash lasting 3 min. Permeabilization of the cells was achieved with a solution containing 0.3% Triton X-100 in PBS for 10 min at room temperature. Following the removal of the permeabilization solution, the cells were washed twice with PBS, with each wash lasting 3 min, and then stained with Hoechst 33342 for nuclear visualization. Fluorescent micrographs were captured where EdU-positive proliferative cells appear in red, and Hoechst 33342 stained viable cells are visible in blue. The relative quantity of EdU was determined by calculating the ratio of EdU-positive cells to the total number of viable cells.
Transwell
Cells were seeded onto a 6-well plate and subjected to a 24-h intervention. Following trypsinization and centrifugation, cells were resuspended in serum-free DMEM to prepare a 200 µL suspension containing 1 × 105 cells. This cellular suspension was then introduced into the upper chamber of a Transwell apparatus previously coated with Matrigel. To the lower chamber, 600 µL of media supplemented with 10% fetal bovine serum was added. After incubating for 24 h, the cells were fixed and stained with crystal violet. Transmembrane cell migration was observed and quantified under a microscope, and the invasive cell count was determined by ImageJ.
Western Blotting
Total proteins were isolated from HepG2 cells treated with SPXJF ranging from 0% to 10% for 48 h. After quantitation using the BCA method, the samples were subjected to SDS-PAGE and subsequently transferred onto a PVDF membrane. The membrane was then blocked, followed by an overnight incubation with the primary antibody at a dilution of 1 : 5000. This was succeeded by a 1-hour incubation with the secondary antibody at a dilution of 1 : 3000. The chemiluminescent detection method was employed to visualize the target proteins. For relative quantification analysis, ImageJ was utilized.
Statistical Analysis
All statistical analysis in this study was presented as values with a standard deviation. (As the mean ± SD) The statistical mean differences were achieved with one-way ANOVA to assess the significance and all statistical methods were performed using GraphPad Prism 10.0 software (CA, USA).
Result
Identification of Differentially Expressed Genes in NAFLD and HCC
The visualization of normalization results for GSE63067 and GSE99807 datasets is in Figure 1, along with the comparison of DEGs in NAFLD compared to normal tissues, and NAFLD-HCC compared to NAFLD tissues:

The common differentially expressed genes (DEGs) between NAFLD and normal tissues/ NAFLD-HCC. (a, d) Box plot results depict good tissue sample homogeneity and comparable baselines for the GSE63067 and GSE99807 datasets. (b, e) Principal component analysis (PCA) plots demonstrate that samples compared by GSE63067 and GSE99807 are well-differentiated and exhibit significant differences. (c, f) Uniform manifold approximation and projection (UMAP) plots illustrate better differentiation and significant differences among the compared samples from GSE63067 and GSE99807 datasets. (g) The volcano plot showing 102 DEGs from GSE63067. (i) The volcano plot showing 828 DEGs from GSE99807. (h, j) The heatmap illustrating the expression of DEGs based on GSE63067 (h) and GSE99807 (j). (k) Visualization of hub genes (eg, RGCC, ACKR3) implicated in NAFLD-HCC progression. Abbreviations: NAFLD: non-alcoholic fatty liver disease; HCC: hepatocellular carcinoma.
This figure outlines the visualization components and their significance in analyzing normalization results and DEGs in the GSE63067 and GSE99807 datasets, shedding light on the molecular alterations associated with NAFLD and NAFLD-HCC progression.
The Validation of two hub Genes in the TCGA-LIHC Dataset
We conducted further investigations into the relationships between the expression of hub genes and the survival of 374 patients diagnosed with HCC, utilizing clinical information from the TCGA-LIHC dataset. Our findings reveal a significant elevation in the expression levels of RGCC and ACKR3 among HCC patients (Figure 2a). Moreover, analysis of data from the HPA demonstrates a high expression of RGCC and ACKR3 at the tissue protein level in HCC patients (Figure 2c). Importantly, a high expression of RGCC is significantly correlated with longer OS in the aforementioned cohort of 374 HCC patients (Figure 2d). Conversely, an elevated expression of ACKR3 is significantly associated with shorter OS in the same group of patients (Figure 2d). The results of time-ROC survival prediction analysis (Figure 2e) indicate that RGCC and ACKR3 exhibit predictive value in estimating 1-year, 3-year, and 5-year survival in patients with HCC. Specifically, RGCC demonstrates better efficiency in predicting 3-year survival (AUC = 0.361), while ACKR3 shows better efficiency in predicting 1-year survival (AUC = 0.663).

The validation of two hub genes expression in HCC patients from the TCGA-LIHC dataset. (a) Unpaired sample test for hepatocellular carcinoma tissue versus adjacent normal tissue from the TCGA-LIHC dataset (Figure 2a, P < 0.001 or P < 0.05). (b) Paired sample test for paracancerous liver tissue with corresponding hepatoma tissue from the TCGA-LIHC dataset (P < 0.001 or P < 0.05). (c) Immunohistochemical results of two genes in the HPA database for patients with hepatocellular carcinoma and normal tissues. (d) Kaplan-Meier curves for overall survival of RGCC and ACKR3, respectively. (e)Time-ROC curves showing survival at 1, 3, and 5 years of RGCC and ACKR3, respectively. Abbreviations: TCGA: The Cancer Genome Atlas; HPA: Human Protein Atlas; time-ROC: time-receiver operating characteristic.
Validation of the Prognostic Signature and Construction of a Nomogram
To investigate the relationship between hub genes and patient outcomes, RGCC and ACKR3 were subjected to univariate and multivariate Cox proportional hazards regression analyses. The dependent variables in these analyses were survival outcome and survival time. Based on these analyses, a prognostic signature composed of the two genes was established. The risk score was calculated using the following formula: Risk Score = −0.376 × RGCC(exp) + 0.182 × ACKR3(exp) + 0.772. The time-ROC curve was used to assess the predictive efficacy of the risk score for 1-, 3-, and 5-year OS in patients with HCC (Figure 3a). The AUC values for 1-, 3-, and 5-year survival prediction were 0.655, 0.676, and 0.616, respectively. Based on the 3-year survival prediction, the AUC value of the prognostic model outperformed others. Therefore, a cut-off value of 0.028 at 3 years was used as the grouping condition. HCC patients from the TCGA-LIHC dataset were divided into low-risk and high-risk groups, with 174 cases in the high-risk group and 199 cases in the low-risk group. The survival probability of the high-risk and low-risk groups was compared (Figure 4b). The picture illustrates that patients with higher risk scores tended to have a poorer survival prognosis, with a hazard ratio (HR) of 1.94 (95% confidence interval [CI] = 1.37-2.77). In the GSE76427 dataset, 155 patients with HCC were divided into high-risk and low-risk groups using a cut-off value of 0.028. Time-ROC (Figure 3c) and Kaplan-Meier curves (Figure 3d) both validated the results of the TCGA-LIHC datasets. As depicted in the picture, patients with higher risk scores exhibited a worse survival prognosis, with HR = 3.97 (95% CI = 1.73-9.10).
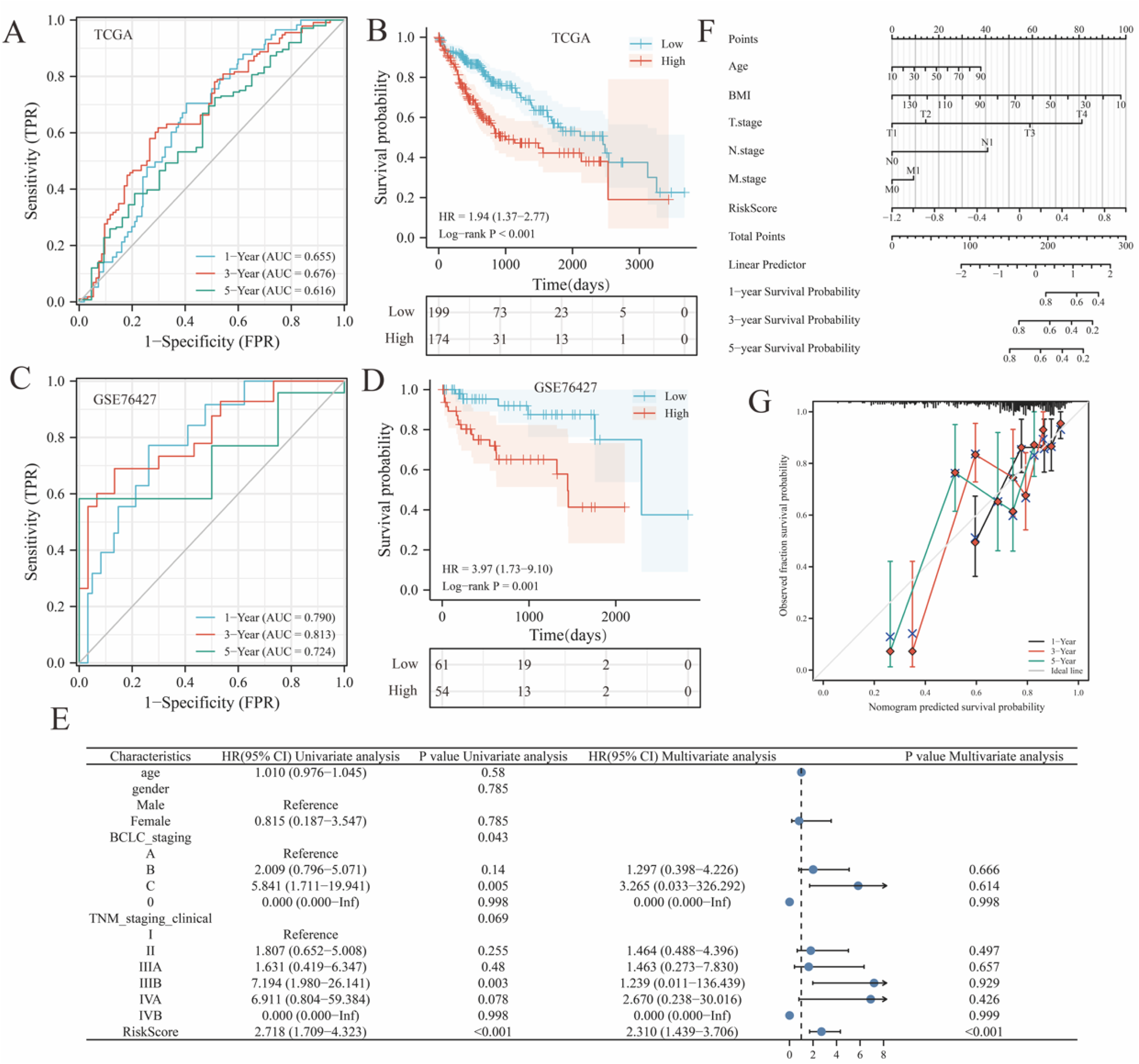
Validation of the prognostic signature and construction of nomograms. (a, b) Time-ROC curves showing survival at 1, 3, and 5 years for 373 patients in the TCGA-LIHC dataset (a) and GSE76427 dataset (b), respectively. (c, d) Kaplan-Meier curves for survival prognosis in the TCGA-LIHC dataset (c) and GSE76427 dataset (d), respectively. (e) Determining the independent factors of a prediction system. (f) The comprehensive nomogram for 1-, 3-, and 5-year survival probability in HCC patients. (g) Calibration analysis results for Nomogram plots. Abbreviations: HCC: hepatocellular carcinoma; TCGA: The Cancer Genome Atlas; time-ROC: time-receiver operating characteristic.
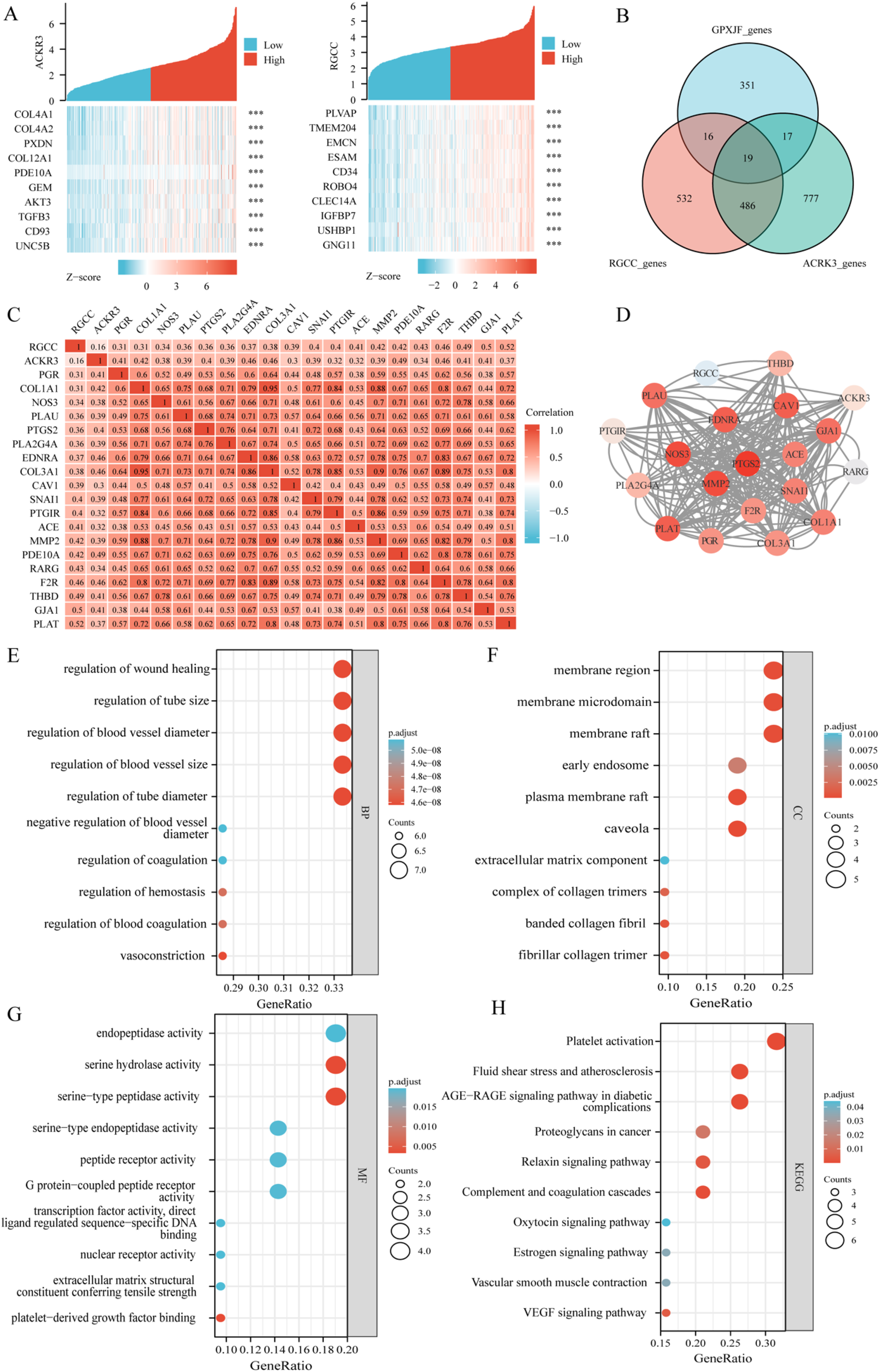
Network pharmacology-based investigation of SPJXF intervention in the progression of NAFLD-HCC. (a) The top 10 genes exhibiting the highest correlation with RGCC and ACKR3 co-expressed genes. (b) Venn diagram depicting the intersection of RGCC, ACKR3, and the drug targets of Shipi-Xiaoji Recipe. (c) Heat map illustrating the expression correlation between RGCC, ACKR3, and the 19 intervention targets of Shipi-Xiaoji Recipe. (d) Protein-protein interaction (PPI) results of the 21 targets. (e-h) Results of the Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis. Abbreviations: NAFLD: non-alcoholic fatty liver disease; HCC: hepatocellular carcinoma.
Then, we performed univariate Cox proportional hazards regression analysis to identify significant clinical features associated with prognosis, such as age, gender, BCLC staging, TNM staging (clinical), and risk score. Among these variables, only BCLC staging, TNM staging (clinical), and risk scores in the prognostic model were found to significantly impact the prognosis of patients with HCC. To determine the independent prognostic factors for HCC patients, we conducted multivariate Cox proportional hazards regression analysis. The results revealed that only the risk score remained a significant and independent prognostic factor for predicting outcomes (HR = 2.310, 95% CI = 1.439-3.706, P < 0.001) (Figure 3e). A comprehensive nomogram was constructed to provide individualized predictions of 1-, 3-, and 5-year OS by incorporating prognostic features such as age, gender, BCLC staging, TNM staging (clinical), and risk score (Figure 3f), which has a C-index = 0.668 (0.632-0.704). This suggests a good predictive capability. Calibration analysis showed (Figure 3g): the predicted 1-, 3-, and 5-year survival probabilities from the Nomogram aligned closely with the actual observations.
Network Pharmacology-Based Investigation of SPXJF Intervention in the Progression of NAFLD-HCC
Integration of data from TCMSP, TCMID, TCMIP, Pubchem, and Swiss Target Prediction databases resulted in the identification of 403 unique drug targets associated with SPXJF. Notable targets included PTGS2, PLAT, and MMP2. Utilizing Spearman correlation analysis on the TCGA-LIHC cohort, a total of 1053 RGCC co-expressed genes and 1299 ACKR3 co-expressed genes were identified. The top 10 genes with the strongest correlations were PLVAP, TMEM204, EMCN, ESAM, CD34, ROBO4, CLEC14A, among others (Figure 4a). By intersecting RGCC, ACKR3, and three sets of NAFLD drug targets (Figure 4b), a total of 19 NAFLD intervention targets were obtained. The expression levels of “NAFLD-HCC” progression genes RGCC and ACKR3 were positively correlated with the 19 targets of NAFLD-HCC (P < 0.001), and a correlation expression heat map was generated (Figure 4c). To investigate the potential mechanism of action of SPXJF targets, the PPI analysis revealed 201 interactions among the 21 shared targets (Figure 4d). Notably, PTGS2, MMP2, and NOS3 exhibited the strongest interactions. These targets exhibited molecular functions such as enzymatic activity, G protein peptide receptor-coupling activity, and nuclear receptor activity (Figure 4g), and were involved in mediating pathways such as VEGF signaling, platelet activation, and AGE-RAGE signaling (Figure 4h). Furthermore, they regulated important biological processes including scar healing, vascular neogenesis, and vascular diameter regulation (Figure 4e).
In Vitro Validation of SPXJF Intervention in the Progression of “NAFLD-HCC”
To investigate the toxic effects of SPXJF on HepG2 cells and to determine the intervention concentrations of SPXJF-containing serum, the study designed experimental groups with blank serum, fetal bovine serum, and SPXJF-containing serum at concentrations of 1%, 5%, 10%, 15%, and 20%. Interventions were carried out on the cells for 24 and 48 h. The results indicated that there was no significant difference in cell viability between the blank serum and fetal bovine serum. However, SPXJF exhibited a notable concentration-dependent inhibitory effect on the viability of HepG2 cells (P < 0.05), as depicted in Figure 5a. Based on these findings and referencing prior research from our research group, subsequent cell experiments employed 1% SPXJF-containing serum as the low-dose group (SPXJF-L), 5% SPXJF-containing serum as the medium-dose group (SPXJF-M), and 10% SPXJF-containing serum as the high-dose group (SPXJF-H).

In vitro validation of SPXJF intervention in the progression of “NAFLD-HCC.” (a) effect of SPXJF on HepG2 cell viability. (b, c) Assessment of cell proliferation through EdU. (d, e) Assessment of cell invasion through Transwell Assay Experimentation. (f) The protein expression of RGCC and ACKR3 by Western blot experiments. (ps. * indicates a statistically significant difference with P < 0.05. ** indicates a statistically significant difference with P < 0.01.). Abbreviations: NAFLD: non-alcoholic fatty liver disease; HCC; hepatocellular carcinoma; EdU: 5-ethynyl-2′-deoxyuridine.
To further explore the proliferative and invasive actions of SPXJF on HepG2 cells, EdU proliferation assays and transwell chamber experiments were conducted. The experimental setup included blank serum, SPXJF-L, SPXJF-M, and SPXJF-H groups. The EdU assay employs a red fluorescent marker to identify actively proliferating cells, while Hoechst 33342 serves to stain all viable cells, emitting a blue fluorescence. The results of the EdU assay indicated that the quantity of viable cells in the SPXJF group was significantly reduced in comparison to the control group, with the number of proliferating cells also diminishing concomitant with increases in SPXJF concentration. Notably, the group receiving the highest dosage of SPXJF exhibited the smallest number of viable and proliferating cells, with these findings achieving statistical significance (P < 0.05), as depicted in Figure 5b and c. In evaluating the invasive capabilities of the cells, a transwell assay was employed. The results demonstrated that the control group had a higher number of cells at the bottom of the chambers, whereas the SPXJF group exhibited fewer cells. When compared to the control group, the difference was statistically significant (P < 0.05), with the high-dose SPXJF group showing the most pronounced difference (P < 0.01), as depicted in Figure 5d and e. These findings suggest that SPXJF is capable of inhibiting the proliferation and invasion of HepG2 cells.
Additionally, to delve further into the influence of SPXJF intervention on the transition from non-alcoholic fatty liver to hepatocarcinogenesis, Western blot experiments were performed to detect the protein expression of RGCC and ACKR3 following SPXJF treatment (Figure 5f). Findings revealed that compared to the control group, SPXJF suppressed RGCC and ACKR3 protein expression in a dose-dependent manner, suggesting that SPXJF may modulate the progression from non-alcoholic fatty liver to liver cancer by regulating RGCC and ACKR3.
Discussion
Unhealthy dietary habits and lifestyle choices have been attributed to the increased incidence of NAFLD, as dietary patterns undergo a structural transformation with the continuous improvement in living standards. 16 NAFLD refers to a collection of clinical syndromes characterized by the gradual buildup of fats in the liver, resulting in lipid degeneration, NASH, and liver fibrosis. 8 These conditions result from long-term excessive lipid deposition in the liver, often influenced by various factors, including lifestyle and dietary habits. Importantly, NAFLD serves as a significant risk factor for the development of HCC. Epidemiological evidence highlights that uncontrolled NAFLD can contribute to the progression of HCC, posing a grave threat to patients’ life and well-being. 17 Therefore, there is an urgent requirement to develop innovative therapeutic agents that can hinder the progression of NAFLD to HCC, with the aim of lowering the incidence and mortality rates associated with HCC. A gene screening-based approach for the prevention and treatment of HCC development offers a promising avenue to identify key genes involved in this process. Additionally, developing new drugs within the framework of the “window shifting” treatment mode for HCC can contribute to enhancing the survival benefits of patients and mitigating the burden of HCC.
To address the issue of controlling the progression from NAFLD to HCC and to prevent the development of HCC, identifying driver genes as entry points holds promise. By pinpointing key genetic factors responsible for the transition from NAFLD to HCC, targeted interventions and therapies can be developed to halt or slow down disease progression, ultimately improving patient outcomes and reducing the burden of HCC worldwide. Advancing the therapeutic “window” for intervention through the development of novel medications aimed at the comprehensive management of HCC progression could significantly reduce both the incidence and mortality rates associated with HCC, thus enhancing patient survival outcomes. TCM, with its holistic approach and principles of syndrome differentiation and treatment, offers unique advantages in early intervention and throughout the treatment of HCC. A formulation called “Shipi-Xiaoji recipe” was tailored to strengthen the spleen and disperse accumulation. This prescription, comprising Astragalus, Ginseng, Poria, Ligustrum lucidum, Epimedium, Pyrite, Peach Kernel, Ground Beetle, Curcuma, Pinellia, Prunella, Gecko, and Spiny Jujube Kernel, notably improved the clinical survival benefits for patients with HCC.
Building upon this premise, the present study employs bioinformatics approaches to identify the driver genes in the progression from NAFLD to HCC, examines their correlation with the survival prognosis of HCC patients, and establishes a prognostic risk scoring system and a clinical predictive model. In conjunction with network pharmacology methods, potential targets and mechanisms of action are sought for the early prevention and comprehensive intervention of the “NAFLD-HCC” continuum utilizing the traditional formulation known for invigorating the spleen and dissolving accumulation. Through differential analysis and intersection, it has been discovered that RGCC and ACKR3 regulate throughout the entire “NAFLD-HCC” progression. Gene and protein expression results in liver cancer tissues and adjacent non-tumor tissues of HCC patients indicate a significant upregulation of RGCC and ACKR3 in liver cancer tissues. To delineate the relationship between driver genes in “NAFLD-HCC” progression and the survival prognosis of HCC patients, Kaplan-Meier and time-ROC curve analyses suggest a significant correlation between the expression of the driver genes RGCC and ACKR3 and survival prognosis. In HCC patients, low expression of RGCC and high expression of ACKR3 portend poorer survival prognosis, potentially indicating shorter survival duration.
Given the significant correlation between the “NAFLD-HCC” progression driver genes and the survival prognosis of HCC patients, we have developed a dual-gene risk scoring system comprising RGCC and ACKR3, which can effectively evaluate the survival prognosis of HCC patients. Moreover, this system has been validated with superior efficacy in the external dataset GSE76427. Independent prognostic analysis, incorporating clinical information such as the patient's age, TNM clinical staging, and BCLC staging, has identified the risk score as an independent risk factor for the prognosis of HCC patients.
To delve deeper into the clinical predictive value of the risk scoring system developed from “NAFLD-HCC” progression genes, we have created a Nomogram clinical predictive model by integrating the HCC patient's age, BMI, and TNM clinical staging. This model provides a robust theoretical foundation for the prediction of 1, 3, and 5-year survival rates for HCC patients, offering prospects for early intervention in HCC.
This study employs the concept of “anticipatory treatment window” in the context of HCC management and utilizes network pharmacology to identify 403 potential pharmacological targets of the SPXJF. This prescription is found to effectively intervene in the entire progression of “NAFLD-HCC” by influencing 19 key molecular targets, among which the interactions between PTGS2, MMP2, and NOS3 proteins are particularly robust. The underlying mechanisms may relate to the functions exerted by the SPXJF at various cellular components such as cell membranes, mitochondrial membranes, and cytoplasm. These functions include enzymatic activity, G protein-coupled receptor activity, and nuclear receptor activity, which mediate pathways like VEGF signaling, platelet activation, and the AGE-RAGE signaling pathway. These actions regulate biological processes such as wound healing, angiogenesis, and vascular tone adjustment. The findings shed light on the direction of TCM in the comprehensive and staged prevention and treatment of HCC.
RGCC is a pivotal gene involved in the complement response. It is also known as C13orf15 or RGC-23. The RGCC gene is ubiquitously expressed across a multitude of tissues and plays a critical role in regulating the cell cycle, augmenting cellular proliferation and differentiation, as well as modulating immune responses. RGCC is expressed by tumor cells and plays a dual role in cancer, functioning as either a tumor promoter by endorsing malignancy initiation, progression, invasion, metastasis, and angiogenesis, or as a tumor suppressor. 18 ACKR3 is an atypical chemokine receptor which is overexpressed in numerous cancer phenotypes, playing a pivotal role in the modulation of tumor cell proliferation, migration, angiogenesis, and the acquisition of drug resistance. Consequently, this receptor engenders an environment conducive to cancer progression and metastasis. 19 Meanwhile, ACKR3 has been found to regulate lipid levels within adipose tissues. Existing research indicates that expression of ACKR3 escalates in adipose tissue amidst obesity, and its pervasive ablation precipitates hyperlipidemia in murine models. 20
To ascertain the impact of the SPXJF on the progression from NAFLD to HCC, cellular experiments were conducted. Observations indicated that SPXJF effectively inhibited the proliferation and invasion of HepG2 cells. To evaluate the influence of icariin on the viability of HepG2 cells, experiments employing the CCK-8 and EdU incorporation assay revealed that SPXJF exerted a dose-dependent inhibitory effect on the proliferation of these cells. Moreover, Transwell chamber assays demonstrated the capacity of SPXJF to suppress the invasion of HepG2 cells. Through application of Western blot analysis, the expression of key NAFLD-HCC-related genes RGCC and ACKR3, as previously identified via bioinformatics, was validated. Results suggested that SPXJF mitigated the expression of RGCC and ACKR3 proteins in a dose-dependent manner. This intimates that SPXJF may regulate the progression from NAFLD to HCC by modulating the expression of genes RGCC and ACKR3. At present, there is also a lot of evidence that TCM can effectively treat NAFLD and liver cancer. For example, Zhang Xiaobo et al 21 demonstrated that TCM improved NASH in mice by inhibiting NLRP3 inflammasome, reducing inflammation and hepatic steatosis. The treatment of liver cancer with TCM has been included in the guidelines of The Chinese Society of Clinical Oncology (CSCO) and can be used for patients with advanced HCC and poor liver function. It is recommended by top-level experts (based on Class 2A evidence).
In summary, the present study employs bioinformatics and network pharmacology approaches to focus on the progression process of “NAFLD to HCC” and the “early intervention” treatment paradigm, identifying RGCC and ACKR3 as correlated with the progression of “NAFLD to HCC” and with HCC prognosis. Additionally, we have successfully constructed two joint-gene risk scoring systems and clinical prediction models which, when combined with the clinical characteristics of HCC patients, can effectively predict survival at 1 year, 3 years, and 5 years. Consequently, these models serve as a constructive complement to the existing clinical TNM staging system for forecasting the survival prognosis of patients with HCC. Our analysis suggests that the entire intervention mechanism of the SPXJF formula may be associated with its regulation of co-expressed genes, mediation of AGE-RAGE and VEGF signaling pathways, molecular functions pertaining to enzymatic activity, and its influence on angiogenesis in HCC. Furthermore, through in vitro experiments, it has been demonstrated that SPXJF can inhibit the proliferation and invasion of HepG2 liver cancer cells and can suppress the expression of RGCC and ACKR3 proteins in a dose-dependent manner. This offers theoretical support for identifying molecular targets for early diagnosis and pharmacological intervention in HCC, as well as guiding the direction for the anti-HCC potential of Chinese medicine and its clinical translation.
However, there are still several limitations to this research. Firstly, the data used is derived from TCGA and GEO databases, which, while representative of a subset of liver cancer patients, cannot be generalized to all populations. Secondly, due to experimental constraints, only one liver cancer cell line was selected, which may not fully represent the various subtypes of liver cancer, and thus, the experimental results may not be broadly applicable to all forms of HCC. Lastly, the conditions of in vitro experiments, such as culture conditions, oxygen, and nutrient supply, differ significantly from the complex microenvironment within which tumors grow in patients’ bodies, so the results may not fully predict the therapeutic efficacy and safety when applied clinically. We will continue to refine our research in these areas in subsequent experiments.
Footnotes
Acknowledgements
The completion of this article would not have been possible without the support and assistance of numerous individuals, to whom I extend my sincerest gratitude. Special thanks are due to Mr Zhibin Wang for his invaluable suggestions and thorough mentoring concerning the experimental section of this work. His professional advice and meticulous attention significantly enhanced the accuracy and efficacy of the experiments conducted. Although academic protocol prevents his inclusion in the list of authors, his substantial contributions merit my profound appreciation.
Authors Contributions
Jingting Zhang wrote the manuscript; Hongyao Chen and Wei Peng collected the data. Renyi Yang designed the in vitro experiments; Kexiong Li and Puhua Zeng involved in the formal analysis and designated the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical Approval is not applicable for this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China (82074425, 82074425), Natural Science Foundation of Hunan Province, China (2023JJ40400, 2023JJ30364, 2023JJ30448, 2021JJ40310), Key Project of Hunan Provincial Administration of Traditional Chinese Medicine (A2023042), Hunan Provincial Traditional Chinese Medicine Research Program (D2022010), Innovation Project for Postgraduate Students at Hunan University of Chinese Medicine (2022CX43), Hunan Province Science and Technology Top Leading Talent Project (2020173, D2022064), Hunan Provincial Graduate Research and Innovation Project (CX20230835), Health Research Projects in Hunan Province (20232605) and Graduate Human University of Traditional Chinese Innovation (2022CX43, 2022CX195, 2023CX26).
Statement of Human and Animal Rights
All handling of experimental animals adhered to the standards set forth in the “Regulations on Quality Management of Experimental Animals” issued by the National Science and Technology Committee. The procedures were approved by the Animal Ethics Committee of Hunan University of Traditional Chinese Medicine, with approval number LL2022070501.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
