Abstract
Objectives
The current study was conducted to evaluate the antibacterial potential of leaf and fruit extracts of Zanthoxylum armatum against two pathogenic bacterial isolates, Staphylococcus aureus and Staphylococcus epidermidis.
Methods
Twelve commercially available antibiotics were tested S. aureus and S. epidermidis by antimicrobial susceptibility test (AST). Qualitative analysis of phytochemicals was performed to evaluate the presence of certain secondary metabolites. The activity of Z. armatum extracts against S. aureus and S. epidermidis was measured as a maximum zone of inhibition exhibited by each leaf and fruit extract. An in-silico study was conducted on flavonoids and alkaloids to show their binding affinity with the PBP2a receptor protein of S. aureus and TcaR of S. epidermidis.
Results
The AST revealed that S. aureus was resistant to Penicillin, Ampicillin, Clindamycin, Vancomycin, Rifampicin, Novobiocin, and Oxacillin, whereas S. epidermidis was resistant to Streptomycin, Oxacillin, Tetracycline, and Novobiocin. Qualitative analysis of phytochemicals resulted in the presence of Saponins, fixed oils, flavonoids, alkaloids, starch, and fatty acids in both leaf and fruit extracts. The maximum zone of inhibition against S. aureus was produced by methanolic leaf extracts of Z. armatum and chloroform fruit extracts. For S. epidermidis, the best activity was exhibited by benzene leaf extracts and methanolic fruit extracts. An in-silico study showed that flavonoids Nitidine and Nevadensin exhibited binding affinity with the PBP2a receptor protein higher than selected antibiotics, ie, Penicillin, Chloramphenicol, and Oxacillin. TcaR of S. epidermidis interacted with Tambuletin, followed by Nitidine and Kaempferol.
Conclusion
After in vitro testing, in silico analysis advised extracting and purifying the bioactive components from Z. armatum extracts that showed significant interaction with bacterial virulence proteins for use as natural antibiotics against antibiotic-resistant bacteria.
Keywords
Introduction
Overuse and misuse of antimicrobial drugs have resulted in selective antibiotic resistance, encouraging the development of new, better alternatives. The coagulase-positive, gram-positive coccus, Staphylococcus aureus (S. aureus) is a frequent colonizer of the human population and is responsible for significant morbidity and mortality worldwide.1,2 About 20% to 30% of humans carry S. aureus in their nose predominantly, as well as commonly in the skin, throat, axillae, groin, and intestine. More intricate and in-depth research is being done on the interaction between the host microbiota and the prospective pathogen, emphasizing the part other commensals play in influencing S. aureus colonization.2–4 The notorious, S. aureus can cause a variety of ailments, from minor skin and wound infections to sepsis or multiorgan failure. 5 S. aureus holds a unique position because of its combo of strong plasticity and relatively high virulence, which allow it to adapt to a variety of environmental situations. S. aureus strains have developed defenses against practically all antimicrobial medications used in therapy. The most significant is resistance to the beta-lactams, glycopeptides, and oxazolidinones that are most frequently used to treat Gram-positive infections. 6 Staphylococcus epidermidis (S. epidermidis) is a coagulase-negative bacteria that is a member of the skin microbiome. S. epidermidis is not just a passive skin resident; it also actively primes the cutaneous immune response, maintains skin homeostasis, and inhibits the colonization of disease-causing opportunistic pathogens.7,8 Most infections linked to indwelling medical devices are caused by S. epidermidis. Due to its prevalence on human skin and ability to stick to catheter surfaces and create biofilms, S. epidermidis is frequently seen in this kind of infection. S. epidermidis serves as a repository of genes that may be passed on to S. aureus, boosting its pathogenic potential and drug resistance.9,10 Methicillin-resistant S. epidermidis (MRSE) is a term used to describe the 75% to 90% of S. epidermidis strains found in hospitals that are resistant to the antibiotic. Of note, some MRSE strains have been classified as multidrug-resistant (MDR) because they are resistant to multiple drugs.10–12
Natural compounds are widely accepted for having medicinal qualities, such as antibacterial and antifungal activities. 13 Medicinal plants play a critical role in meeting the need for innovative medications originating from remote markets. It is indeed possible to look back in time and discover proof that people formerly used medicinal plants to treat ailments. Over 10% of the various plant species are used in medicinal and cosmetic processes.14,15 They include a wide range of secondary metabolites, including simple phenols, flavonoids, alkaloids, tannins, and terpenes, among others, as several investigations of their chemical constituents have demonstrated. These bioactive phytochemicals are usually extracted using organic reagents. Many studies focused on the screening of plant antibacterial properties.16,17
The Zanthoxylum armatum plant (Figure 1), native to Kashmir, Rawalpindi, and the Northern areas of Pakistan, is a small tree or sizeable spiny shrub which sprouts in March and April. It belongs to the family Rutaceae and is known by local names such as Timur, Green Sichuan pepper, and winged prickly ash (English).18,19 The dried fruits of Z. armatum are consumed as a condiment and are an excellent source of spices. Additionally, fruits, roots, stems, and leaves are utilized as starting materials for medications that have antioxidant, analgesic, antibacterial, anticancer, and antiinflammatory properties.20,21 Different plant parts, including fruits, seeds, leaves, and bark, have been used to extract several phytocomponents, including alkaloids, flavonoids, terpenoids, phenols, and steroids. 22 The plant possesses antibacterial, antioxidant, antifungal, hepatoprotective, and allopathic activities, according to earlier investigations. It is also reported to function as a tonic, condiment, carminative, stomachic, and anthelmintic.23,24 Leaves, bark, and fruits are used as mouth fresheners and in tooth care and are also used in the cosmetic industry. Essential oils of plants are used in fragrances and flavoring agents for foods and beverages. Numerous bioactive phytochemicals isolated from Z. armatum are used for insect control.25,26 Timur is also used to treat everyday ailments such as toothache, fever, cough, and cold, giving warmth to the body.27,28
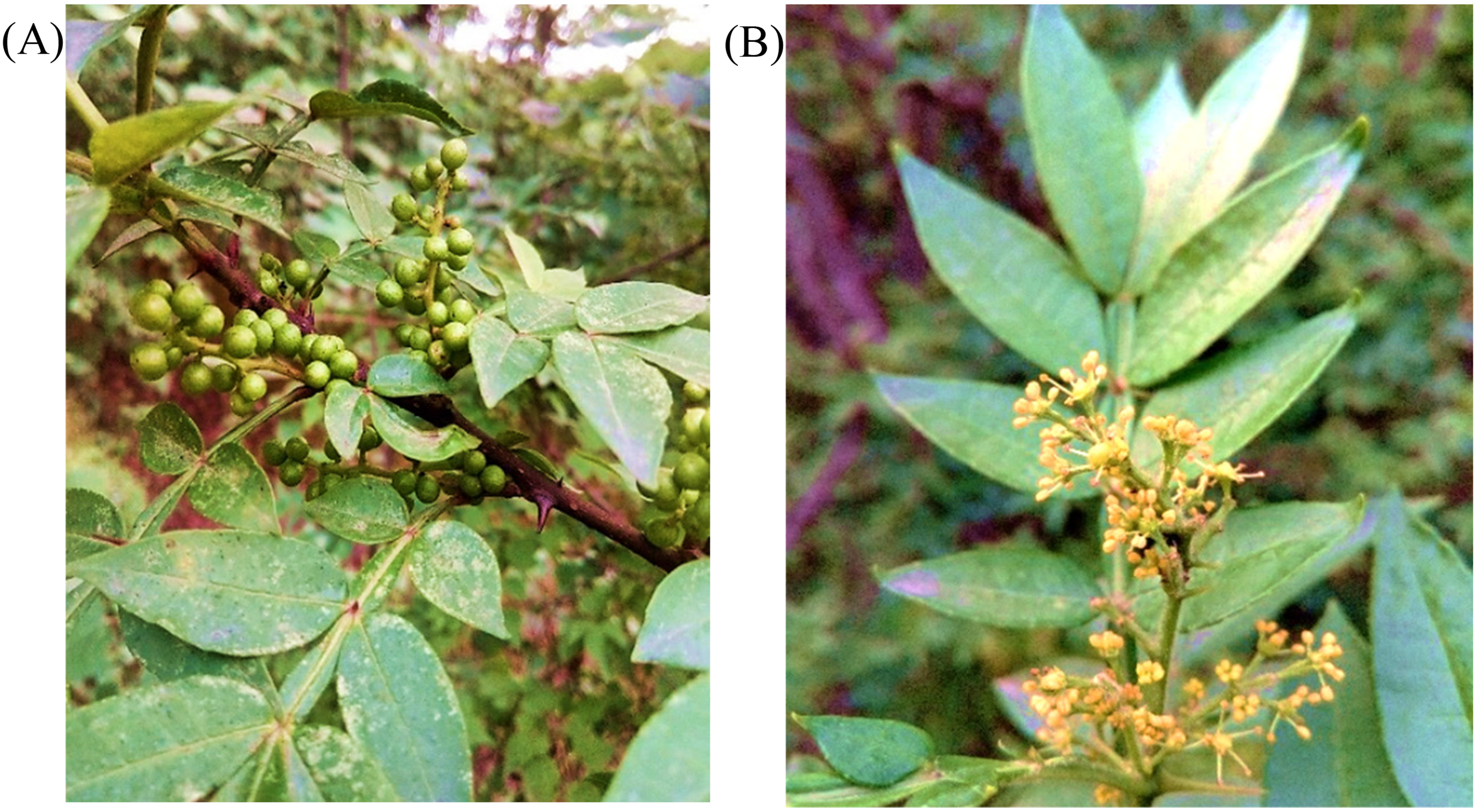

Ariel parts of Zanthoxylum armatum plant; (A) fruit and leaves (B) flowers.
To validate the age-old traditional ethnomedical practices, a firm connection must be made between traditional knowledge and modern research. Among the many natural substances found in plants, molecular docking has also been utilized to find novel inhibitors.29,30 Molecular docking has become a crucial tool in the drug discovery process. Additionally, these methods aid in the prediction of a compound's toxic and adverse effects. Many common medications, like remediffer, hydroxychloroquine, and chloroquine, have been studied in laboratories and found to have particular therapeutic effects. But the clinical pharmacological response is not particularly promising, and toxicity is still an issue that will inevitably result in detrimental side effects.31,32 The current study aims to explore the antibacterial potential of leaf and fruit extracts of Z. armatum against two pathogenic bacterial isolates, S. aureus and S. epidermidis and to evaluate the antibiotic resistivity pattern of S. aureus and S. epidermidis. Preliminary phytochemical analyses of methanolic and ethanolic leaves and fruit extracts were also performed to confirm the presence of certain phytocompounds. Computational analyses of phytochemicals from flavonoid and alkaloid classes were performed to evaluate their druglikeness, bioavailability, and toxicity potentials. Molecular docking of phytochemicals from alkaloid and flavonoid classes of compounds with potential drug targets of S. aureus and S. epidermidis was performed.
Material and Methods
Preparation of Plant Extracts
Leaves and fruits of Z. armatum used for this study were collected from Kotli, Azad Jammu and Kashmir, Pakistan. Plant samples were picked randomly, and rotten leaves and unripe or squeezed berries were removed. Later, plant parts were washed with distilled water 33 and blot-dried at room temperature in the shade. Twenty grams of each part (ie, leaves and fruits) was weighed and subjected to grinding in mortar to form a powder drug and stored in a tight container before extraction.
For solvent extraction, the crushed material was soaked in 200 mL of each solvent, ie, ethanol, 34 methanol, 35 chloroform, 36 benzene,36,37 and water 37 separately and shaken (120 rpm) at room temperature for 24 h. 38 In general, polar solvents such as water, methanol, and ethanol are used in the extraction of polar compounds, whereas nonpolar solvents such as hexane and dichloromethane are used in the extraction of nonpolar compounds.39–42 Then filtrates of each plant extract were collected using Whatman No. 1 filter paper. Extracts were dried under the heating drying oven at a temperature less than the boiling points of the solvents, ie, 40 °C for volatile solvents (ethanol, methanol, chloroform, benzene from Sigma-Aldrich USA) and 60 °C for aqueous extracts. 43 The dry weight of extracts was determined and dissolved each plant extract in dimethyl sulfoxide (DMSO Sigma-Aldrich USA). Two hundred fifty milligrams of each extract was dissolved in 10 mL of DMSO to obtain 25 mg/mL. 44 Reconstituted extracts were then passed through the bacterial filter of size 0.45 μm and stored in labelled Eppendorf tubes at 4 °C for further use. 45
Phytochemical Analysis
Major bioactive components of plants ie, steroids, alkaloids, carotenoids, flavonoids, tannins, terpenoids, and glycosides are the necessary starting materials for drug development. 46 Phytochemical analysis determines the presence of these active compounds in the plants. Z. armatum phytoconstituents determination in methanolic extracts was performed using standard procedures.46–51
Test for Saponins
Two-milliliter plant extract was mixed with 2 mL of distilled water and shaken vigorously in a test tube. The formation of stable foam indicates the presence of saponins.
Test for Fixed Oils and Fats
Approximately one drop of the extract was pressed between two filter papers to confirm the presence of fixed oils and fats. The oil stain on the paper indicated the presence of fixed oil.
Test for Proteins
One milliliter of plant extracts were treated with a few drops of concentrated nitric acid. The presence of proteins was confirmed by forming the yellow color.
Test for Phenols
Extracts were mixed with 3 to 4 drops of ferric chloride solution. The change of color to bluish black indicated the presence of phenol.
Test for Flavonoids
Lead acetate solution was added to 1 mL of extract. The formation of yellow precipitates was taken as a positive test for the presence of flavonoids.
Test for Betacyanin
One milliliter extract was added with 1 mL 2N NaOH and observed. Extracts that changed their color to yellow confirmed the presence of betacyanin.
Test for Starch
One milliliter extract was added with 1 mL iodine solution, and the formation of black color was taken as an indication of the presence of starch.
Test for Resins
One milliliter extract was treated with 1 mL acetone and 1 mL distilled water. Turbidity upon shaking indicates the presence of resins.
Test for Alkaloids
Two milliliters concentrated hydrochloric acid was added to 2 mL of each plant extract and heated. Then few drops of Mayer's reagent were added. The presence of yellow color or off-white precipitate indicated the presence of alkaloids.
Test for Gums and Mucilage
One milliliter of extract was taken, and 2 mL of absolute alcohol was added with constant stirring. White or cloudy precipitates indicated the presence of gums and mucilage.
Antimicrobial Susceptibility Tests
The present study used S. aureus and S epidermidis provided by the Department of Biosciences, University of Wah. Preliminary bacterial identification tests (Gram staining, catalase test, coagulase test, oxidase test and substrate utilization test) were performed for isolates and their glycerol stocks were stored at −40 °C till further use.
The findings of the antimicrobial susceptibility test (AST) are also used to evaluate the efficacy of preventative and infection control methods, such as surveillance of resistant microbes like methicillin-resistant S. aureus (MRSA). 52 A total of 12 commercially available antibiotics purchased from Liofilchem Italy and Oxoid were used to determine the susceptibility of bacteria (Table 1). Mueller–Hinton agar plates were inoculated with 5 μL of the bacterial culture, and the agar medium followed by coating with commercially made filter paper disks impregnated with specified quantities of an antibiotic. After that, the plates were incubated for 24 h at 35 °C to 37 °C in room temperature. The antibiotics spread into the agar during this incubation, and as the distance from the disk increases, the concentration of the antibiotics decreases. By measuring the width of the bacterial inhibition zones surrounding the antibiotic disks and comparing it to disk diffusion interpretative criteria updated yearly by CLSI, antibiotic susceptibility was ascertained. The terms “susceptible” (S), “intermediate” (I), and “resistant” (R) are used by both EUCAST and CLSI, and their meanings were formerly shared. While categories S and R continue to have a consistent interpretation in both standards and are simple to understand, Category I was difficult to understand because it combined several definitions into one: appropriate use when an antibiotic is physiologically concentrated at the site of the infection, appropriate use of a high dosage of the medication, etc.52,53
Description of Commercially Available Antibiotics Used in AST.

AST, antimicrobial susceptibility test.
Antibacterial Assay
The agar well diffusion method 54 was performed to check the antibacterial potential of Z. armatum extracts. Freshly grown cultures (at 0.7-0.8 O.D) were used for the antibacterial assay. The nutrient broth was inoculated with test microbes grown at 37 °C in a shaking incubator at 120 r/min. Culture density was recorded at 600 nm using a spectrophotometer.
Mueller-Hinton each agar plate was sectioned and labeled appropriately. After 10 min of solidification, the plates were swabbed with the 20 μL test microbiological cultures. The centers of the separated sections were punched with a 5 mm sterile borer. Melted soft agar was then used to seal the bottom of the well. Eighty micoliter of the test substances (plant extracts or DMSO) were then pipetted into the perforations.55,56
Selection of Receptor Proteins
MRSA is one of the leading causes of hospital- and community-acquired infections.57,58 The penicillin-binding protein 2a (PBP2a) found in S. aureus (MRSA) is responsible for accelerating the formation of peptidoglycan in the bacterial cell wall. 59 The primary strategy that MRSA has used to build clinical resistance to a wide range of β-lactam antibiotics is the production of penicillin-binding protein 2a. 60 According to the National Institutes of Health, biofilms are to blame for around 80% of microbial illnesses.61,62 The biofilm extracellular matrix of S. epidermidis has a critical component known as polysaccharide intercellular adhesion (PIA). 63 According to recent studies, the expression of the icaADBC operon, which is controlled by the transcription regulator TcaR, determines how much PIA is produced.64,65 TcaR is a multifunctional regulator and a member of the MarR gene family that contributes to teicoplanin and methicillin resistance.66,67
Molecular Docking
For docking analysis, 13 phytochemicals from Z. armatum 03 standard antibiotics were chosen, and their structures and physicochemical properties were noted from PubChem (Table 2). Crystal structures of two receptor proteins, the penicillin-binding protein 2a (PBP2a) of S. aureus (PDB ID: 1MWT) (Figure 2A) and S. epidermidis TcaR-methicillin (PDB ID: 3KP4) (Figure 2B), were downloaded from RCSB protein databank. AutoDock Tools 1.5.6 was used to perform molecular docking between receptor proteins and ligands. AutoDock Vina calculates the dimensions of grid boxes automatically, and conveniently estimates binding sites. In the present study, a grid box size of 40 × 40 × 40 (x, y, and z) and a grid spacing of 0.375 were used for both receptor proteins. The center of the grid for S. aureus PBP2a was set at positions 5.092, 37.973, and 47.653 for x, y, and z, respectively, while for S. epidermidis the centers (x, y, and z) were set at −22.315, −29.216, and −0.294, respectively. The docking results showed binding energy values in kcal mol−1, and the ligand with the most negative value was considered to have the highest binding affinity for the target protein. The top model (Model 1) was chosen from the nine distinct conformations and the binding interactions were shown in 2D by using Discovery Studio v2021.
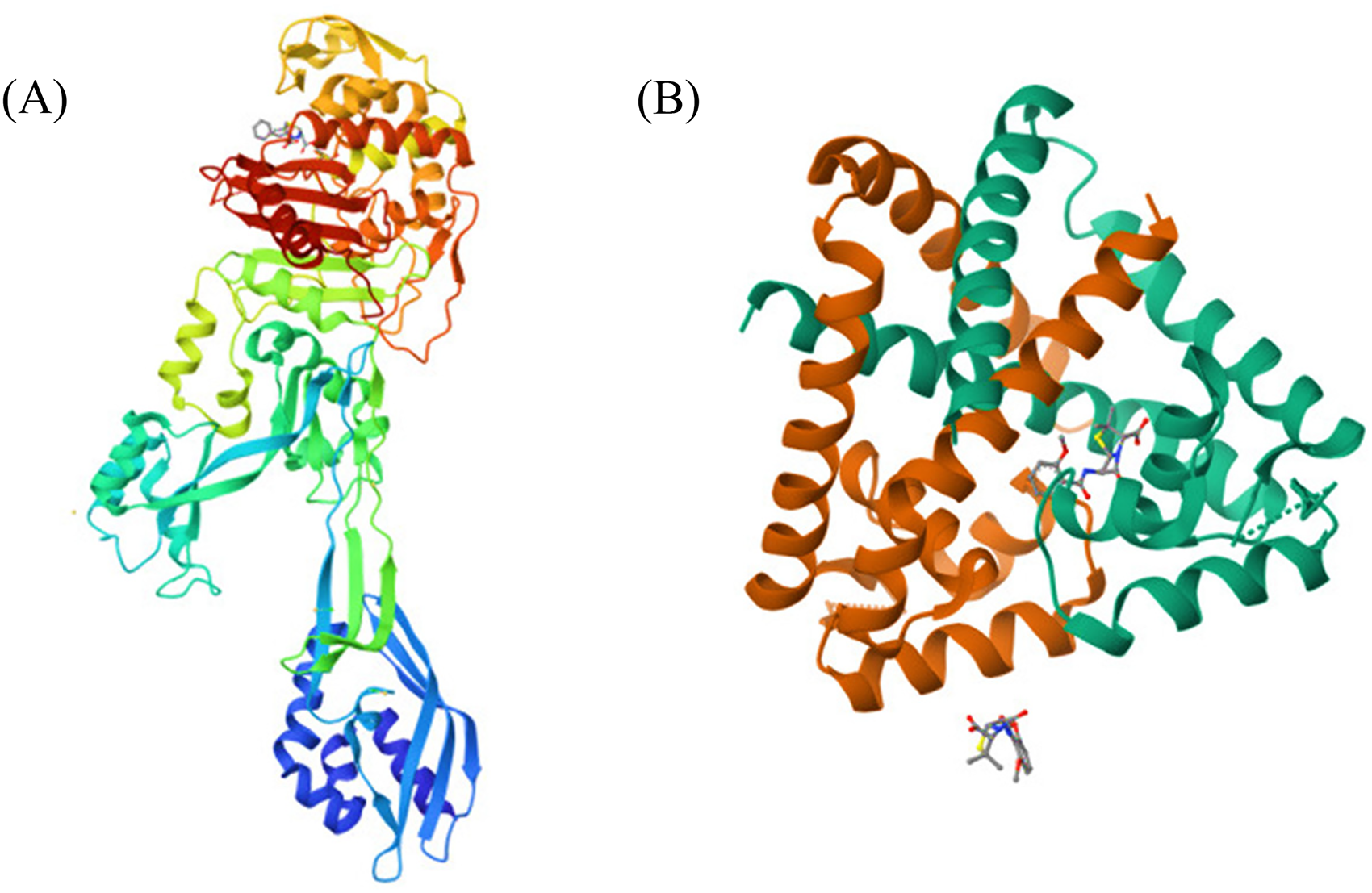
Crystal structures of two receptor proteins; (A) Penicillin-binding protein 2a (PBP2a) of Staphylococcus aureus (PDB ID: 1MWT), (B) Staphylococcus epidermidis TcaR-methicillin (PDB ID: 3KP4).
Physicochemical Properties and Structures of Phytochemicals from Zanthoxylum armatum and Standard Antibiotics.

Swiss ADME Predictions and Toxicity Analysis of Phytocompounds
Virtual screening of Z. armatum resulted in the selection of 13 phytochemicals belonging to alkaloid and flavonoid classes of compounds. The chemistry of phytochemicals and their physicochemical characteristics, including lipophilicity, solubility, pharmacokinetics, and drug similarity, were evaluated by SwissADME.74,75 Toxicity of each compound was estimated computationally by the Protox-II server.75,76
Statistical Analysis
Every statistical analysis was conducted using GraphPad Prism software Version 9 (https://www.graphpad.com/features) on data from three biological replicates, and the results were presented as the mean plus or minus the standard deviation and standard error mean (SEM).
Results
Phytochemical Analysis
Phytochemicals like alkaloids, flavonoids, phenols, fatty acids, amino acids, and many other compounds have been identified in Z. armatum ethanolic and methanolic extracts and summarized in Table 3. Phytochemicals like saponins, fixed oils, flavonoids, alkaloids, starch, and fatty acids were present in both leaves and fruit extracts of Z. armatum. Leaf extracts contained phenols, while they were absent in fruit extracts. Gums and mucilage were absent in both ethanolic and methanolic leaf extracts while being present in methanolic fruit extracts only. Resins, betacyanins, terpenoids and coumarins were present in fruits and absent in leaf extracts. Quinones, Steroids, Glycosides, Xanthoprotein Test, Carbohydrates, Phlobatannins, Emodins, Anthraquinones, Leucoanthocyanin, and Cordial Glycosides were completely absent in all extracts.
Phytochemical Analysis of Leaf and Fruit Extracts.

Key: Present (+), Absent (−).
Antimicrobial Susceptibility Testing Against Selected Antibiotics
Staphylococcal species tested for antimicrobial susceptibility (Supplementary Figure 1) revealed that S. aureus showed a maximum mean zone of inhibition recorded for ciprofloxacin ie, 29 mm followed by Chloramphenicol (ie, 28 mm), hence regarded as susceptible according to CLSI 2017/20. The CLSI standards also suggest that S. aureus is susceptible to kanamycin (23 mm), streptomycin (24 mm), and tetracycline (18 mm). The minimum zones (ie, 0 mm) were recorded for Penicillin, Ampicillin, Clindamycin, Vancomycin, Rifampicin, Novobiocin, and Oxacillin, therefore S. aureus was regarded as resistant against tested antibiotics (Figure 3A). For S. epidermidis, the maximum inhibition zone was recorded against Penicillin (ie, 35 mm) and minimum ie, 0 mm against Oxacillin and Streptomycin. According to standard guidelines available, S. epidermidis was identified as resistant against Tetracyclin (14 mm) and novobiocin (12 mm) while being sensitive against Ampicillin (27 mm), Clindamycin (23 mm), Kanamycin (16 mm), Ciprofloxacin (20 mm), Vancomycin (20 mm), Rifampicin (20 mm), and Chloramphenicol (25 mm) (Figure 3B).

Antimicrobial susceptibility testing against selected antibiotics; (A) zones of inhibition (mm) formed by Staphylococcus aureus against selected antibiotics, (B) zones of inhibition (mm) formed by Staphylococcus epidermidis against selected antibiotics.
Antibacterial Assay
Present findings demonstrated that leaf and fruit extracts of Z. armatum are worthy of antimicrobial activity against S. aureus (Figure 4A). The maximum zone of inhibition produced by leaf extracts of Z. armatum (Supplementary Figure S1) was by methanolic extracts ie, 13.33 ± 1.33 mm (M ± SEM). Ethanolic leaf extract also produced a significant zone ie, 12.67 ± 0.89 mm whereas aqueous extracts of leaves exhibited minimum activity (ie, 3.66 ± 3.667). Chloroform and benzene produced equal zones of inhibition ie, 11.33 mm and DMSO used as negative control did not produce any zone of inhibition. The major zone of inhibition produced by fruit extracts of Z. armatum against S. aureus was by chloroform extracts ie, 10 ± 5.12 mm (M ± SEM).

Antibacterial activity of leaf and fruit extracts of Zanthoxylum armatum; (A) zones of inhibition (mm) formed leaf and fruit extracts of Z. armatum against Staphylococcus aureus, (B) zones of inhibition (mm) formed leaf and fruit extracts of Z. armatum against Staphylococcus epidermidis.
For S. epidermidis (Figure 4B) the best activity was exhibited by benzene leaf extracts of Z. armatum ie, 12.33 ± 1.33 mm (M ± SEM) followed by ethanol (9.667 ± 4.842 mm), methanol (9.667 ± 4.910 mm), and aqueous (9.667 ± 4.910 mm) extracts. Chloroform extracts of leaves produced a minimal zone of inhibition (8.000 ± 4.041 mm) whereas no zone was seen for DMSO. The methanolic fruit extracts were found to exhibit maximum activity against S. epidermidis by producing a zone of inhibition of 13.67 ± 1.764 mm (M ± SEM) followed by ethanolic (13.00 ± 0.57 mm) and chloroform (13.00 ± 1.52 mm) fruits extracts. Benzene fruit extract produced a zone of 12.67 ± 0.33 mm whereas water extracts exhibited a minimum zone of inhibition ie, 4.33 ± 4.33. The details of zones of inhibition produced by leaf and fruit extracts of Z. armatum against S. aureus and S. epidermidis are given in Supplementary Table 1.
In Silico Studies
Docking
Docking interaction of selected phytoconstituents from Z. armatum and antibiotics with Penicillin-binding protein 2a (PBP2a) of S. aureus showed that Nitidine attached with the receptor protein and released the lowest energy ie, −8.2 kcal/mol indicating the highest binding affinity. The binding interactions suggest that amino acid residues Ala276, Tyr272, Lys319, Lys322, Lys290, and Lys273 of PBP2a are involved by conventional hydrogen bond, carbon hydrogen bond, pi-alkyl bond, alkyl bond and pi-pi T-shaped bond. Followed by nitidine are Nevadensin (−8.1 kcal/mol) and Tambuletin (−8.0 kcal/mol). These three compounds have shown binding affinity higher than selected antibiotics ie, Penicillin (−7.9 kcal/mol), Chloramphenicol (−6.5 kcal/mol), and Oxacillin (−7.9 kcal/mol). Details of docking analysis of all the phytocompounds and antibiotics with PBP2a of S. aureus are given in Table 4. The docking poses and the interactions of PBP2a with Nitidine, Nevadensin, and Oxacillin (antibiotic) are given in Figure 5 and the remaining compounds are given in Supplementary Figure 1.

The docking poses and the interactions of PBP2a with (A) Nitidine, (B) Nevadensin, and (C) Oxacillin.
Docking Interactions of Selected Phytoconstituents From Zanthoxylum armatum and Commercially Available Antibiotics With Penicillin-Binding Protein 2a (PBP2a) of Staphylococcus aureus.

Docking analysis of selected phytoconstituents from Z. armatum with the transcription regulator TcaR of S. epidermidis revealed that Tambuletin bound with greater binding affinity releasing binding energy of −9.1 kcal/mol followed by Nitidine (−9.0 kcal/mol) (Figure 6A, B). Tambuletin (−9.1 kcal/mol) interacted with TcaR amino acid residues Val19, Ile16, Asn17, His42, Asn20, Gln61, Val63, Ala24, Gln31, and Ala38 through conventional hydrogen bonds, pi-alkyl, and pi-anion bonds. Of selected antibiotics, oxacillin interacted with TcaR releasing −9.6 kcal/mol showing the highest binding affinity (Figure 6C) with amino acid residues Val63, Ala38, Ile16, Glu13, Arg110, Asn45, and His42. The lowest binding affinity was recorded for chloramphenicol with binding energy −7.1 kcal/mol. Details of docking analysis of all the phytocompounds and antibiotics with TcaR of S. epidermidis are given in Table 5.
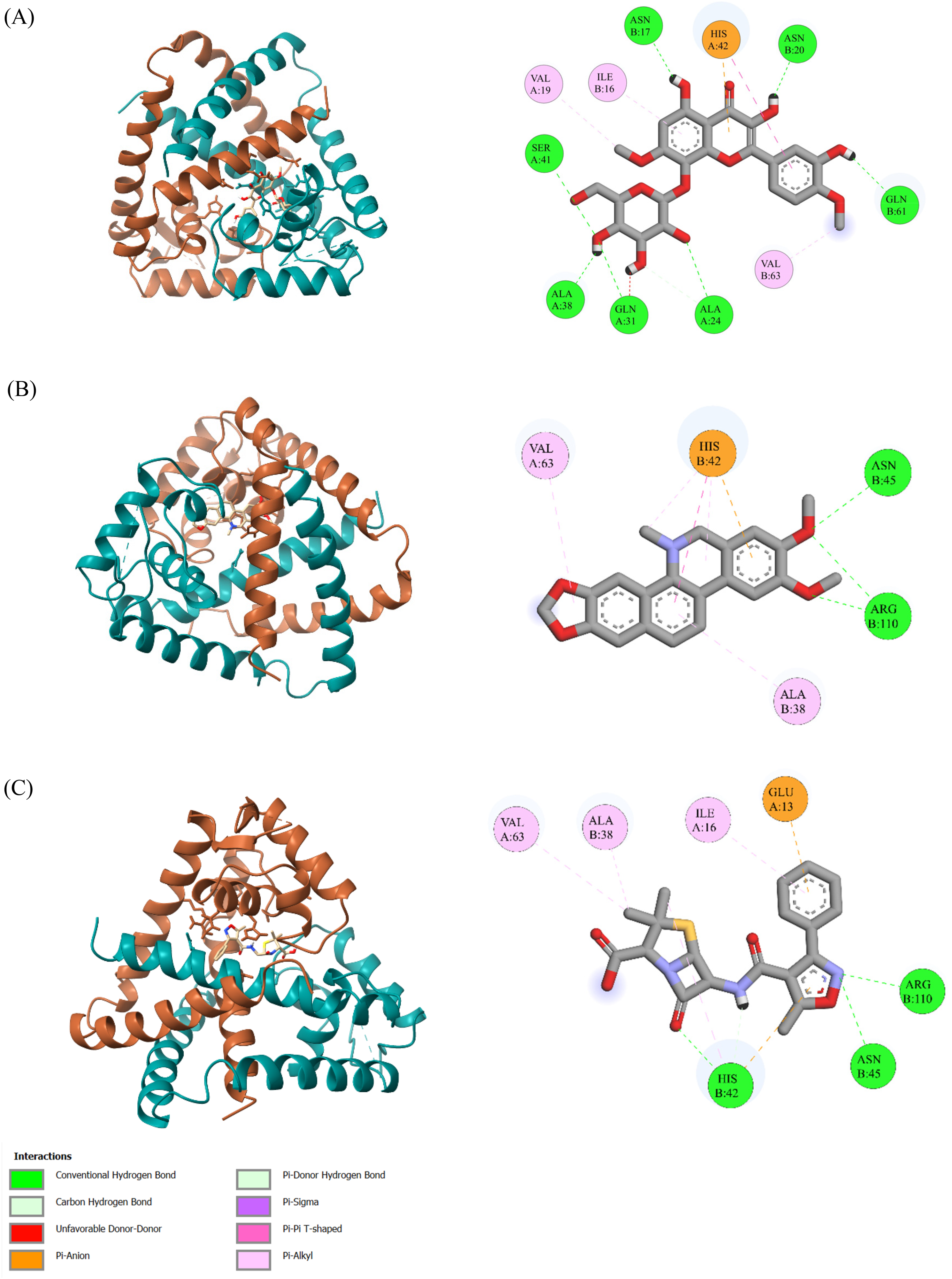
The docking poses and the interactions of TcaR with (A) Tambuletin, (B) Nitidine, and (C) Oxacillin.
Docking Interactions of Selected Phytoconstituents From Zanthoxylum armatum and Commercially Available With TcaR of Staphylococcus epidermidis.

Swiss-ADME and Toxicity Analysis of Phytocompounds and Antibiotics
Pharmacokinetics analysis of Phytocompounds of Z. armatum and antibiotics computationally estimated the GI absorption, blood–brain barrier (BBB) permeation, and cytochrome inhibition (Table 6). All selected phytochemicals except Tambuletin, and antibiotics showed high gastrointestinal (GI) absorption. None of the selected antibiotics and phytochemicals Nevadensin, Tambulin, Kaempferol, and Tambuletin showed BBB permeation while remaining phytocompounds were BBB permeant. Phytochemicals Magnoflorine, Nitidine, Sanguinarine, Tambuletin, and antibiotic oxacillin were P-gp substrates suggesting controlled drug delivery and lesser toxicity while the remaining phytocompounds and antibiotics were non-P-gp substrates which is ideal for improved bioavailability, and lowered risk of drug–drug interactions and multidrug resistance. Cytochromes are enzymes that are essential for the biotransformation and the metabolism of drugs. Skimmiamine ideally inhibited all classes of cytochrome while Haplopine, Nevadensin, Robustine, and Tambulin inhibited four out of five cytochrome classes. Penicillin, Chloramphenicol, and Oxacillin inhibited none of the Cytochrome classes.
Pharmacokinetics Analysis of Selected Phytoconstituents of Zanthoxylum armatum and Commercially Available Antibiotics.

Estimation of Lipinski's rule of five by SwissADME ensured that all of the selected phytocompounds except Tambuletin and antibiotics followed Lipinski's rule of five. Tambuletin has showed Lipinski's violations in molecular weight (508.43 kcal/mol), hydrogen bond acceptors (13), hydrogen bond donors (7), and TPSA (208.74 Å2) values. Moreover, Tambuletin was the only compound that exhibited a negative lipophilicity (Log Po/w) value and less bioavailability score. The details of Lipinski's rule of five, lipophilicity value and bioavailability score of all selected compounds and antibiotics are given in Table 7.
Evaluation of Lipinski's Rule of Five.

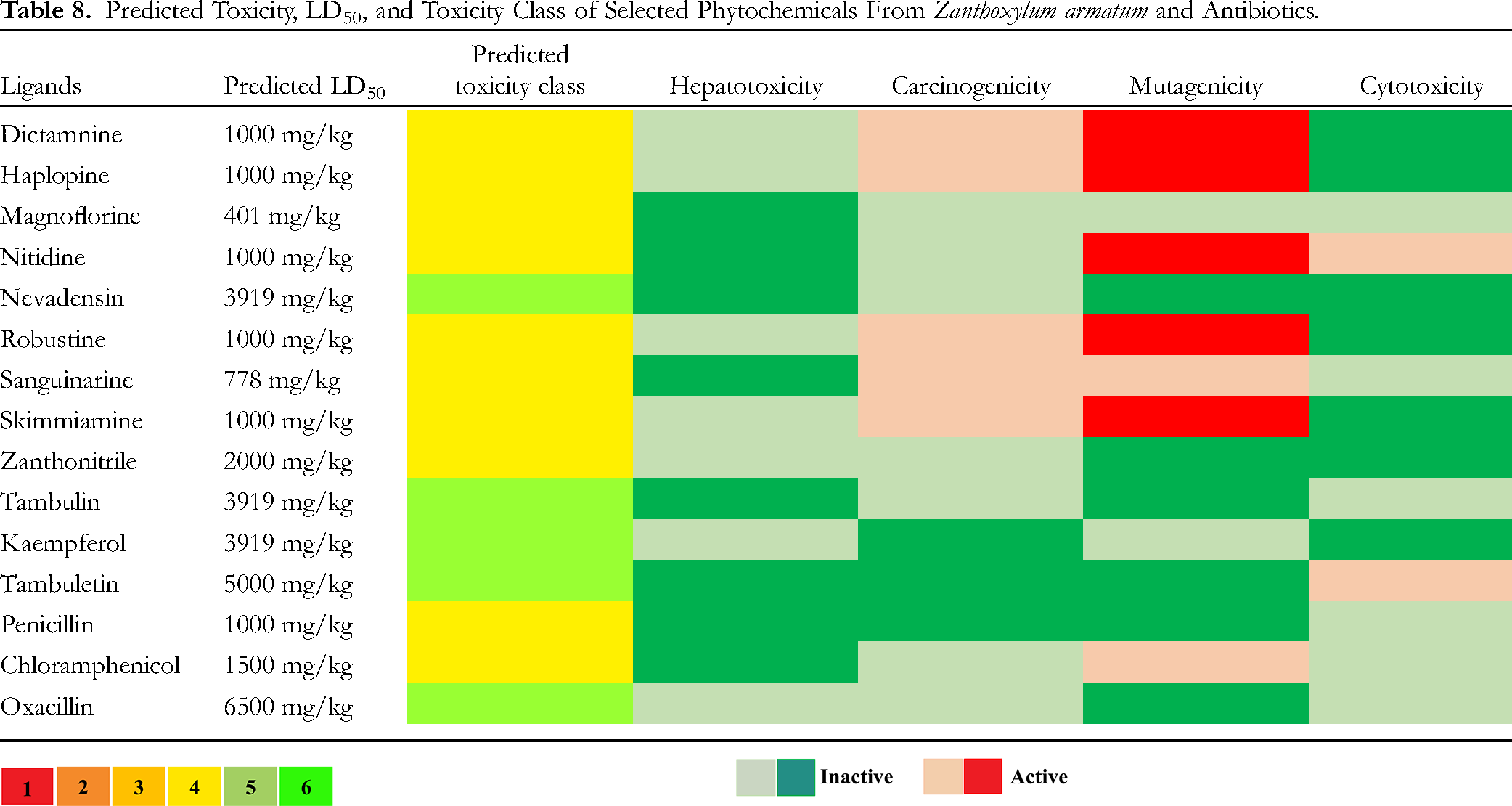
Evaluation of predicted toxicity, toxicity class, and dosage of selected phytochemicals and antibiotics (Table 8) suggest that all antibiotics and phytochemicals Magnoflorine, Nevadensin, Zanthonitrile, and Kaempferol were nontoxic in terms of hepatotoxicity, carcinogenicity, mutagenicity, and cytotoxicity. Dictamnine, Haplopine, Robustine, Sanguinarine, and Skimmiamine were found mildly active for carcinogenicity and mutagenic effects. Nitidine was found to be active for mutagenicity and cytotoxicity, and Tambuletin was also found to be somewhat carcinogenic. Among the understudy antibiotics, chloramphenicol was found to be mutagenic. Nevadensin, Tambulin, Kaempferol, and Tambuletin belonged to toxicity class 5 while the remaining phytochemicals belonged to Class 4. Antibiotics penicillin and chloramphenicol belonged to Toxicity Class 4 whereas oxacillin was found in Class 6.
Predicted Toxicity, LD50, and Toxicity Class of Selected Phytochemicals From Zanthoxylum armatum and Antibiotics.

Discussion
The use and misuse of antibiotics globally is a hot and contentious issue that has resulted in developing antibiotic resistance in microbes. 77 In the 1950s and early 1960s Penicillin-resistant S. aureus was declared a pandemic 78 and the emergence of methicillin-resistance was reported in 1961 79 with the mortality rate associated with MRSA being approximately 20%. 80 A key factor in the fast emergence of antibiotic-resistant S. aureus is the acquisition of resistance genes from other S. aureus strains or even from other genera. 81 Multiple antibiotic resistance was conferred in MDR S. epidermidis strains by the abundance of plasmid-borne genes and sequence variants that they had accumulated through time. 82
Approximately 61% of the novel drugs developed in the late 20th and early 21st centuries to treat infectious diseases, such as cancer, were derived from natural or phytochemical compounds. In many underdeveloped nations, traditional plant-based pharmaceutical formulations are used to treat and prevent human illnesses.83,84 The study of microbiology and the treatment of many diseases caused by infectious agents such as bacteria have advanced significantly, but new strains of pathogenic bacteria and drug-resistant bacteria continue to spread and cause epidemics, making the development of novel antibiotics necessary. Currently, scientists are reevaluating whether it makes sense to use cutting-edge technology to study medicinal plants in an effort to identify new bioactive molecules that can address significant public health concerns.84–86
The goal of the study that is being presented is to determine if commercially available antibiotics can effectively combat the resistance of clinically relevant bacterial species, such as S. aureus and S. epidermidis, while also potentially leveraging the antibacterial properties of a local medicinal plant identified Z. armatum. Previous studies suggest that MRSA consistently demonstrates resistance to a variety of antibiotics, such as penicillin, methicillin, oxacillin, cefoxitin, amoxicillin-clavulanic acid, amoxicillin-sulbactam, quinolones, macrolides, cephalosporins, tetracycline, and chloramphenicol.87–89 AST in this study revealed that S. aureus was resistant against penicillin, ampicillin, clindamycin, vancomycin, rifampicin, novobiocin, and oxacillin (Figure 2A) whereas S. epidermidis was identified resistant against Streptomycin (0 mm), Oxacillin (0 mm), Tetracyclin (14 mm), and novobiocin (12 mm) (Figure 2B). According to another study, S. epidermidis was present in 46 out of 100 (or 46%) infected hospital samples. The most common types of penicillin resistance were found in S. epidermidis strains (95.6%), followed by tetracycline (91.30%), erythromycin (82.60%), cefazolin (78.26%), and trimethoprim-sulfamethoxazole (73.91%). 90
Qualitative analysis of phytochemicals resulted in the presence of Saponins, fixed oils, flavonoids, alkaloids, starch, and fatty acids were present in both leaves and fruit extracts of Z. armatum. Gums and mucilage, Resins, betacyanins, terpenoids, and coumarins were present in fruits and absent in leaf extracts. Due to the existence of several bioactive substances such as flavonoids, alkaloids, terpenoids, tannins, phenolic compounds, steroids, resins, and other metabolites, plants were thought to have medical potential.91–94 The bioactive components from the plants had been regularly extracted using solvents including methanol, ethanol, acetone, and water. 95 Leaf and fruit extracts of Z. armatum against S. aureus and S. epidermidis resulted in significant antimicrobial activity. The maximum zone of inhibition against S. aureus was produced by methanolic leaf extracts of Z. armatum (ie, 13.33 ± 1.33 mm) and chloroform fruit extracts (ie, 10 ± 5.12 mm). For S. epidermidis (Figure 3B), the best activity was exhibited by benzene leaves extracts of Z. armatum ie, (12.33 ± 1.33 mm) and methanolic fruit extracts by producing a zone of inhibition of 13.67 ± 1.764 mm. According to reports, the type of solvent influences how well the various solvents extract the active components and how they affect the antibacterial activity of these spices.96,97 A study demonstrated that in comparison to aqueous extracts, hybrid organic solvent extracts were more potent against S. aureus. 98
The presence of phytochemicals is known to precipitate cell wall proteins in Gram-positive bacteria, inhibit microtubules, a component of the cytoskeleton, and promote binding to protein receptors that lead to cell death, which may be responsible for the anti-staphylococci effect.99–101 An in silico study was conducted to determine the binding relationship between the bioactive chemicals from flavonoids and alkaloids class of Z. armatum with Penicillin-binding protein 2a (PBP2a) of S. aureus and TcaR-methicillin of S. epidermidis. The flavonoid class of substances appears to target PBP2a most frequently, and several earlier research show flavonoids to have the capacity to modify the bacterial cell walls of MRSA and other types of bacteria.102–104In the present study, flavonoids Nitidine (−8.2 kcal/mol) and Nevadensin (−8.1 kcal/mol) have shown binding affinity with the PBP2a receptor protein higher than selected antibiotics ie, Penicillin (−7.9 kcal/mol), Chloramphenicol (−6.5 kcal/mol), and Oxacillin (−7.9 kcal/mol). Amino acid residue in the allosteric pocket of PBP2a (Lys273) interacted with Nitidine by hydrogen bond and alkyl bond, and Tyr272 formed a carbon-hydrogen bond, pi-alkyl bond, and pi-pi T-shaped bond. Nevadensin amino acids His293, Lys148, and Asp295 have made conventional hydrogen bonds whereas Asp275 and Lys273 made carbon-hydrogen bonds. TcaR of S. epidermidis interacted with Tambuletin (−9.1 kcal/mol) forming seven hydrogen bonds at active TcaR amino acid residues 67 Asn17, Asn20, Gln61, Ala24, Gln31, and Ala38 whereas oxacillin, exhibiting the highest binding affinity (−9.6 kcal/mol) formed only three hydrogen bonds involving Arg110, Asn45, and His42. The evaluation of the binding effectiveness is based on hydrogen bonding, which is one of several forms of interactions, including amide–π interactions, π-π, and H-bond, etc.105–107 SwissADME and toxicity analysis suggest that bioactive compounds from Z. armatum show potential druglikeness and safety profiling as potential lead compounds.
After in vitro testing, in silico analysis advised extracting and purifying the bioactive components from Z. armatum extracts that showed significant interaction with bacterial virulence proteins for use as natural antibiotics against antibiotic-resistant bacteria. However, our study has some limitations such as: the study is focused on two bacterial isolates (S. aureus and S. epidermidis). The potency of Z. armatum extracts against a larger spectrum of bacterial strains would strengthen the generalizability of the findings. Numerous compounds were found in the leaf and fruit extracts by the qualitative analysis of phytochemicals; however, the study could not quantify these compounds. A more thorough understanding of the concentration of bioactive components may be obtained by quantitative analysis.
Conclusions
Based on current investigations, it is concluded that Z. armatum has potential antibacterial activity that may be boosted by enhancing extract concentration. The use of potentially harmful chemically synthesized antibacterial agents can be reduced by using extracts from the plant Z. armatum, which has greater antimicrobial potential than several commercially available antibiotics. These extracts can be used as effective therapeutic agents against strains of S. aureus and S. epidermidis and diseases that arise from them. Additionally, docking investigations revealed that some natural chemicals from Z. armatum that exhibited a significant interaction with bacterial virulence proteins may be extracted and used in the future to develop new medications.
Supplemental Material
sj-docx-2-npx-10.1177_1934578X241237911 - Supplemental material for Antibacterial Profiling of Zanthoxylum armatum Extracts: A Comprehensive Computational and Experimental Study
Supplemental material, sj-docx-2-npx-10.1177_1934578X241237911 for Antibacterial Profiling of Zanthoxylum armatum Extracts: A Comprehensive Computational and Experimental Study by Mamuna Mukhtar, Haris Ahmed Khan and Shumaila Naz in Natural Product Communications
Supplemental Material
sj-docx-3-npx-10.1177_1934578X241237911 - Supplemental material for Antibacterial Profiling of Zanthoxylum armatum Extracts: A Comprehensive Computational and Experimental Study
Supplemental material, sj-docx-3-npx-10.1177_1934578X241237911 for Antibacterial Profiling of Zanthoxylum armatum Extracts: A Comprehensive Computational and Experimental Study by Mamuna Mukhtar, Haris Ahmed Khan and Shumaila Naz in Natural Product Communications
Footnotes
Acknowledgments
We acknowledge the support of the Department of Biosciences, University of Wah for providing the research facilities.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
