Abstract
Cryptotanshinone (CPT) is the main pharmacologically active component of
Introduction
Colorectal cancer (CRC) is the third most common malignant tumor in the world, with an increasing incidence and deaths year-on-year. 1 A combination of chemotherapy and radiotherapy, as well as biological targeting therapies, are commonly used in clinical CRC treatment to improve the quality and duration of patients survival. 2 Moreover, drugs with lower toxicity and better targeting effects for chemotherapy or that assist radiotherapy are being sought and developed to enhance treatment efficacy and safety.
The dried root and rhizome of
Deacetylase sirtuin-3 (SIRT3), a NAD + -dependent class III histone deacetylase, belongs to the Sirtuins family.
14
SIRT3, mainly localized at mitochondria, harvests numerous substrates such as hypoxia-inducible factor 1-alpha (HIF-1a) and tumor suppressor p53 (p53), which are further involved in the development and progression of cancer and neurological diseases, as well as cardiovascular diseases.
15
Tumor tissues often present aberrant SIRT3 and acetylation levels.
16
SIRT3 is highly expressed in CRC, and, therefore, a series of small molecule SIRT3 inhibitors have been investigated for CRC treatment.
17
It is reported that there is a positive correlation between SHMT2 acetylation and SIRT3 expression in 309 CRC tumor samples, and patients with high SIRT3 expression had shorter survival.
18
SIRT3 knockdown was able to significantly inhibit H1299 cell growth and proliferation and induce apoptosis.
19
In our study, we explored the potential targets of CPT-CRC by transcriptome analysis. Further we demonstrated that CPT could suppress CRC cell (SW480 and SW620) proliferation
Results
Transcriptome Sequencing
To elucidate the mechanism of CPT in regulating CRC, we performed transcriptome sequencing. The chemical structure is shown in Figure 1A (PubChem CID: 160254), and with CPT treatment, 1234 differentially expressed genes were identified in SW480 cells. In detail, 679 genes were upregulated and 555 downregulated (fold change cutoff of ≥ 1.3 and adjusted FDR ≤ 0.001) (Figure 1B). Kyoto Encyclopedia of Genes and Genomes (KEGG) (

CPT Inhibits the Proliferation of SW480 and SW620 in vitro and in vivo
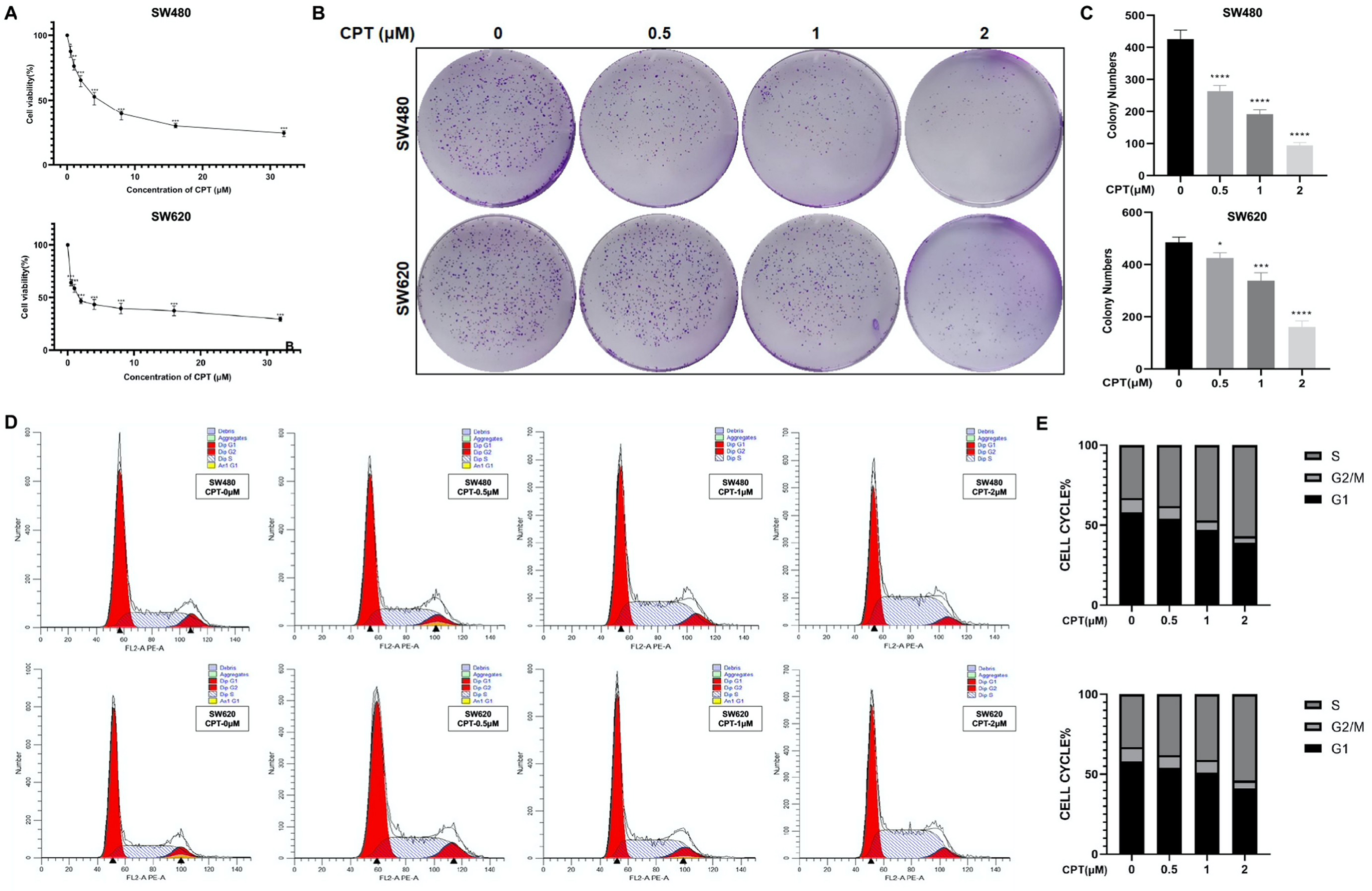
SW480 and SW620 cells were treated with different concentrations of CPT (0, 0.5, 1, 2, 4, 8, 16, and 32 μM) for 48 h, CCK-8 experiment showed that CPT inhibited the viability of SW480 and SW620 in a dose-dependent manner (Figure 2A). Thus, we selected 0.5, 1, and 2 μM concentrations of CPT for subsequent experiments. These CPT concentrations not only significantly inhibited the cell viability, but also suppressed the ability of colony formation of SW480 and SW620 cells (Figure 2B and C). Consistent with this, flow cytometry analysis clarified that CPT arrest of SW480 and SW620 cells was in the S phase, resulting in a decreased percentage of G1 and G2/M phase cells compared to the control group (Figure 2D and E).
To evaluate the pharmacological activity of CPT in the treatment of CRC in

CPT Induces CRC Cell Apoptosis and Triggers Alteration of Mitochondrial Membrane Potential
Flow cytometry analysis showed that CPT induced apoptosis of SW480 and SW620 cells (Figure 4A and B, Supplemental Figure 1A and B). Consistent with this, Hoechst staining showed an increase of densely stained nuclei and apoptotic bodies with increasing CPT concentrations (Figure 4C and D, Supplemental Figure 1C and D), which demonstrated that CPT drives CRC cell (SW480 and SW620) apoptosis. Moreover, JC-1 staining showed that CPT treatment significantly decreased the red fluorescence while inducing the green fluorescence of SW480 and SW620, resulting in a decreased mitochondrial membrane potential, which is a characteristic of apoptosis (Figure 4E, Supplemental Figure 1E). Moreover, apoptosis related proteins were detected. As shown in Figure 4F and G and Supplemental Figure F and G, the expression of Bax and Cytc was increased by CPT in a dose-dependent manner, while Bcl2 and Caspase 3 expression was decreased, which confirmed that CPT can induce apoptosis in CRC cells and trigger the alteration in mitochondrial membrane potential.

SIRT3 Overexpression is Correlated With CRC Patient Survival
Based on the publicly available omics data, we analyzed the relationship between CRC and SIRT3 expression. Analysis revealed that the expression level of SIRT3 was different in various cancer types (Figure 5A). Analysis of clinical cases in the cancer genome atlas (TCGA) database (https://portal.gdc.cancer.gov/; 41 normal samples and 286 CRC samples) indicated that SIRT3 was overexpressed in CRC samples (Figure 5B), which correlated with patient survival (Figure 5C). This suggests that SIRT3 is associated with CRC development, and could be a potential target in CRC treatment.

CPT Targets SIRT3 and Downregulates SIRT3 Protein Expression
To explore further the relationship between CPT and SIRT3, we performed immunofluorescence, Western blot analysis, and molecular docking experiments. Immunofluorescence (Figure 6A) and Western blot analysis (Figure 6B and C) demonstrated that with CPT treatment, the SIRT3 protein level was down-regulated both in SW480 and SW620 cells in a dose-dependent manner. On the other hand, the molecular docking experiment indicated that CPT could bind with Ile154, Pro155, Phe157, Arg158, Phe180, Leu199, Asn229, and Ile230, as well as His248 of SIRT3. The binding energy was −8.6 kcal/mol (<−5 kcal/mol), which indicated that the interaction of CPT and SITR3 is stable (Figure 6D). Thus, these findings indicated that CPT targets SIRT3 and downregulates SIRT3 protein levels.

Discussion
In this study, we explored the potential mechanism of CPT in the treatment of CRC through transcriptome sequencing, which revealed that CPT regulated CRC procession through apoptosis, angiogenesis, and EMT. Further experiments demonstrated that CPT suppresses CRC proliferation
Several signaling pathways are critical in CRC progression, such as the MAPK and PI3K/AKT signaling pathways. It is reported that Mex3a promotes carcinogenesis in CRC by activating the RAP1/MAPK signaling pathway.
20
While inhibition of MAPK signaling was able to inhibit CRC cell proliferation,
21
on the other hand, the PI3K/AKT signaling pathway is critical in CRC cell autophagy and epithelial-mesenchymal transition.
22
Consistent with this, KEGG analysis based on transcriptome sequencing data, identified the PI3K/AKT and MAPK signaling pathways in the CPT-CRC signaling pathways (Figure 1). This relationship was confirmed by Zhang et al, who demonstrated that CPT inhibits CRC cell proliferation by reducing the phosphorylation of PI3K, AKT, and MTOR,
9
indicating that CPT can treat CRC by mediating the indicated signaling pathways. Recently, CPT was identified as a compound with novel anti-CRC pharmacological activity. It is reported that CPT increased the susceptibility of CRC cells to endoplasmic reticulum stress-mediated apoptosis by reducing calpain activity.
23
Our study showed that 0.5, 1, and 2 μM CPT were able to inhibit CRC cell proliferation both
Mitochondria are key metabolic organelles that play a central role in regulating apoptosis. 31 It has been shown that acriflavine impairs mitochondrial function through upregulation of P53 mRNA and protein, which in turn increases chemosensitivity in SW620 cells. 32 In the present study, CPT induced CRC apoptosis and triggered alterations in mitochondrial membrane potential (Figure 4), suggesting that CPT may induce mitochondrial apoptosis by altering mitochondrial metabolism and thereby exerting anti-CRC effects.
The expression level of SIRT3 is different in various cancer tissues, indicating that SIRT3 plays the role of oncogene or tumor suppressor, mediates substrates acetylation, and regulates the indicated signaling pathway. SIRT3 is highly expressed in CRC, oral cancer, lung cancer, melanoma, and renal cancer.33,34 In CRC cells, downregulation of SIRT3 led to a decrease in PTEN expression, which in turn triggered mitochondrial fission to induce apoptosis. 35 In addition, small molecules or external conditions can affect the development and progression of CRC by targeting SIRT3. For instance, δ-valerobetaine induces apoptosis in CRC cells by downregulating the expression of SIRT3. 36 In addition, cisplatin induced mitochondrial methylenetetrahydrofolate dehydrogenase (MTHFD2) acetylation by down-regulating the expression of SIRT3 and inhibiting SIRT3 deacetylation, resulting in decreased NADPH levels and increased ROS content, which in turn reduced CRC viability. 37 These investigations compensate for our flaws. In our study, we identified that CPT down-regulates the protein level, targets Ile154, Pro155, Phe157, Arg158, Phe180, Leu199, Asn229, and Ile230, as well as His248 residues of SIRT3, further functions in inhibiting the proliferation and inducing apoptosis of CRC (Figure 7). On the other hand, SIRT3 has been identified as a tumor suppressor in ovarian, 38 breast, 39 and liver cancers. 40 According to related research, CPT inhibits the proliferation of ovarian cancer cells by down-regulating STAT3, further inducing SIRT3 expression. 41 Whether the regulation of CPT on SIRT3 protein level is STAT3 mediated or direct needs further investigation. However, SIRT3 harvests several substrates, such as KU70. It is reported that SIRT3 mediates the de-acetylation of KU70 and BAX, further inhibiting BAX related apoptosis. We speculate whether CPT could regulate the acetylation of KU70 and interrupt the interaction of KU70 and BAX, further regulating CRC cell apoptosis.42,43 Related experiments will be performed in the future.

Conclusions
In summary, we suggest that CPT inhibits CRC cell proliferation
Materials and Methods
Materials and Reagents
CPT was purchased from Shanghai Yuanye Biotechnology Co., Ltd (99% purity; B21586; Shanghai, China). Anti-B cell lymphoma 2 (Bcl2; 4223S), anti-Bax (2772), anti-cytochrome c (Cytc; 136F3), and anti-Caspase 3 (9662) antibodies were obtained from Cell Signaling Technology (CST; Massachusetts, USA). Anti-GAPDH (10494-1-AP), anti-SIRT3 (10099-1-AP), goat anti-mouse IgG HRP (SA00001-1), and goat anti-Rabbit IgG HRP (SA00001-2) antibodies were obtained from Proteintech (Wuhan, China). The Annexin V-FITC/PI Apoptosis Detection Kit (556547) was purchased from Becton, Dickinson and Company (BD; New Jersey, USA). Hoechst 33342 staining solution (C0031), dimethyl sulfoxide (DMSO; D8370), Triton X-100 (T8200), bovine serum albumin (BSA; A8850), and mitochondrial membrane potential kit (JC-1; M8650) were purchased from Solarbio (Beijing, China). Dulbecco's modified Eagle's medium (DMEM; C11995500BT) and fetal bovine serum (FBS; 16140071) were obtained from Gibco (California, USA). Phosphate buffered saline (PBS; SH30256), trypsin (SH30042), and penicillin–streptomycin dual antibody (SV30010) were from Hyclone (Logan, UT, USA). Cell counting kit-8 kit (CCK-8; AR1199) was purchased from BOSTER Biological Technology Co. Ltd (Wuhan, China). Propidium iodide (PI; CF0031) was obtained from Beijing Dingguo Changsheng Biotechnology (Beijing, China), Paraformaldehyde (BL539A) from Biosharp (Hefei, China), the chemiluminescence reagent (BeyoECL Moon) (P0018FS), and 4’, 6-diamino-2-phenylindole (DAPI) staining solution (C1005) from Biyuntian Biotechnology Co., Ltd (Shanghai, China), Trizol (9108) and TB Green (RR820A) from TaKaRa (Kusatsu, Shiga Japan), and the reverse transcription kit (K1622) from ThermoFisher Scientific (Massachusetts, USA). All other chemicals were obtained from commercial sources.
Animal Experiment
Male BALB/c-nu nude mice (4-6 weeks, weight 20 ± 2 g) were purchased from Changchun Yisi Laboratory Animal Technology Co., Ltd (Changchun, China). The experiment was carried out in the barrier facility of the Laboratory Animal Center of Jilin University and approved by the Laboratory Animal Welfare Ethics Committee of Jilin University (SY20191210). Animal feeding and protocols used were in compliance with the Guide for the Care and Use of Laboratory Animals published by the National Institutes of Health (NIH Publications No. 8023, revised 1978). Nude mice were randomly divided into 4 groups: control-SW480 (n = 11), CPT-SW480 (n = 11), control-SW620 (n = 8), and CPT-SW620 (n = 8). Either SW480 or SW620 cells (5 × 106) were subcutaneously injected into the lateral ribs of nude mice to establish CRC mouse models. After 10 days of culture in the specific pathogen free environment, the control group received normal saline and the CPT group received CPT (20 mg/kg/day) by gavage for 14 days. During treatment, mice were weighed regularly (on days 2, 4, 6, 8, 10, 12, 14), and tumor size was measured with calipers on days 1, 3, 5, 7, 9, 10, 11, and 13.
Tumor volume was calculated using the following formula: tumor volume = 0.5 ×
Cell Culture
Human CRC cells SW480 and SW620 were purchased from Wuhan Punosai Life Technology Co., Ltd (Wuhan, China). Cultures were maintained at 37 °C in a humidified atmosphere using DMEM complete medium (10% FBS, 1% penicillin and streptomycin).
Transcriptome Sequencing
RNA Preparation
SW480 cells were seeded in 10 cm cell culture dishes. SW480 was maintained in DMEM medium containing 10% FBS for 24 h prior to addition of CPT until the cells adhered. CPT (5 μM) was added for 48 h, and cells were collected. Trizol lysis buffer was added to the cell sample tube (1 mL Trizol for 1 × 106 cells). The supernatant was transferred to a centrifuge tube with 300 µL chloroform/isoamyl alcohol (24:1), mixed thoroughly by shaking and centrifuged at 12000 ×
mRNA Library Preparation
Secondary structures were denatured by heating, and oligo dT magnetic beads were used to enrich mRNA. Cleavage reagent was added and reacted until the mRNA was fragmented. Then a synthesis reaction system was added to prepare cDNA and add A bases to the 3’ end to ligate adapters to the cDNA. A PCR reaction system was prepared and the reaction program was set to amplify the ligation product. After the PCR product was denatured into a single strand, a circularization reaction system was prepared to obtain a single-stranded circular product, and the final library was obtained by digesting the uncircularized linear DNA molecules. Fragment size and library concentration were determined using an Agilent 2100 Bioanalyzer. The DNA molecules were sequenced using combined probe-anchored polymerization (cPAS) to obtain sequencing reads of 50 bp/100 bp/150 bp.
RNAdenovo Analysis Method
The raw data were filtered with SOAPnuke (v1.4.0) by (1) Removing reads containing adapters (adapter contamination); (2) Removing reads whose unknown base (“N” base) ratio is more than 5%; (3) Removing reads whose low-quality base ratio (base quality less than or equal to 15) is more than 20%; afterwards clean reads were obtained and stored in FASTQ format. Trinity (v2.0.6) was used to assemble the clean reads and BUSCO to assess the assembly quality. Annotating assembled Unigene with 7 major functional databases (KEGG, GO, NR, NT, SwissProt, Pfam, and KOG), and transcription factors were predicted. PossionDis was performed between-group differential gene analysis with a fold change cutoff of ≥ 1.3 and FDR ≤ 0.001. The pheatmap function was used on the differential gene set to draw a heatmap of differential gene clusters.
Functional Enrichment
According to the GO and KEGG annotation results and official classifications, the differentially expressed genes were functionally classified, the phyper function in R software was used for KEGG enrichment analysis, and the TermFinder package was used for GO enrichment analysis (https://metacpan.org/pod/GO::TermFinder). With a
CCK-8
To determine the effect of CPT on the viability and proliferation of CRC cells, SW480 and SW620 were cultured in 96-well plates at a density of 5000 cells/well for 24 h. SW480 and SW620 cells were treated with CPT dissolved in DMSO at different concentrations (final concentrations of 0.5, 1, 2, 4, 8, 16, and 32 μM) for 48 h. The control group was treated with DMSO. Subsequently, 20 μL CCK-8 solution was added to each well for live cell staining, and incubated at 37 °C for 30 min, protected from light. Then the absorbance of each well at 450 nm was measured.
Colony Formation
SW480 and SW620 cells were seeded in 6-well plates at a density of 500 to 1000 cells/well. CPT (0, 0.5, 1, and 2 μM) was added to the cells after attachment, and fresh complete medium was replaced 48 h later; the medium was changed every 2 days. Two weeks later, they were stained with crystal violet and analyzed using Image J calculation (National Institutes of Health; Bethesda, Maryland, USA).
Cell Cycle
SW480 and SW620 cells in logarithmic growth phase were inoculated into 6-well plates at 2.5 × 105/well for 24 h and treated with CPT for 48 h. Cells were collected with EDTA-free trypsin and fixed overnight in 70% ethanol. Cells were centrifuged for 5 min and stained with PI for 30 min in the dark. Cell cycle changes were detected using a Beckman Coulter flow cytometer (Beckman CytoFLEX; Brea, CA, USA). Cell cycle curves were plotted with GraphPad Prism 7 software (GraphPad Software; San Diego, CA, USA).
Apoptosis
SW480 and SW620 cells were seeded at a density of 2.5 × 105 cells/well and treated with CPT for 48 h. Cells were collected into 1.5-mL Eppendorf tubes with trypsin, washed twice with PBS. Then the Annexin V-FITC/PI Apoptosis Detection Kit (binding buffer, FITC, propidium iodide) was used to complete the relevant operations. The apoptosis rate was detected by flow cytometric (Beckman CytoFLEX; Brea, CA, USA) analysis. SW480 and SW620 cells treated with CPT were then incubated with Hoechst staining solution for 30 min. After washing with PBS, the cell morphology was observed under a fluorescence microscope (Olympus; Tokyo, Japan). SW480 and SW620 cells treated with CPT were also incubated with JC-1 staining working solution for 20 min in the dark and then washed twice with 1× JC-1 buffer. After adding fresh medium, cells were observed under a fluorescence microscope (Olympus; Tokyo, Japan).
Western Blot
SW480 and SW620 cells in logarithmic growth phase were seeded in 6-well plates. Each well contained 1.5 × 105 cells. The first well served as the vehicle control (DMSO) and CPT was added at different concentrations to the remaining wells. After 48 h incubation, cells were collected using trypsin, washed 2 to 3 times with PBS, lysed directly by adding an appropriate amount of 2 × loading buffer, sonicated at 20 Hz for 60 s, and boiled at 95 °C for 15 min. These samples were run on 10% SDS-PAGE gels, and after completion of electrophoresis and transfer, 5% skimmed milk was added and incubated at room temperature for 90 min. Primary antibodies against Bax, Bcl2, Cytc, Caspase 3, and SIRT3 were added and incubated at 4 °C overnight. The membranes were incubated with secondary antibodies for 90 min at room temperature and developed using an ECL kit.
Immunofluorescence
SW480 and SW620 cells were seeded in 6-well plates (1.5 × 105, 2 × 105 cells/well) containing DMEM complete medium for 24 h, and then the cells were exposed to CPT (0.5, 1 μM) for 48 h. Cells were fixed with 4% paraformaldehyde for 15 min at room temperature, permeabilized with 0.3% Triton X-100 for 5 min, and nonspecific protein sites were blocked with 1% BSA for 1 h. Following addition of anti-SIRT3 (1:300) primary antibody, cells were incubated for 1 h at 37 °C, rinsed 3 times with PBS, and incubated with goat anti-mouse IgG/fluorescein isothiocyanate (1:500) for 1 h at 37 °C in the dark. Subsequently, nuclei were labeled using DAPI and images were acquired using confocal fluorescence microscopy (Olympus; Tokyo, Japan).
Molecular Docking
SDF structure files of CPT were obtained from the pubchem website, which further converted into PDB files using OpenBabel 2.3.2 software. The SIRT3 (PDB ID: 4O8Z) structure was obtained from the protein databank. AutoDockVina1.1.2. was used for molecular docking.
Statistical Analysis
All data are presented as the mean ± standard deviation (SD) of 3 independent experiments, and statistical analysis was performed using GraphPad Prism 7 software (GraphPad Software; San Diego, CA, USA). Student's
Supplemental Material
sj-jpg-1-npx-10.1177_1934578X231194808 - Supplemental material for Cryptotanshinone, a Potential SIRT3 Inhibitor, Suppresses Colorectal Cancer Proliferation in vitro and in vivo
Supplemental material, sj-jpg-1-npx-10.1177_1934578X231194808 for Cryptotanshinone, a Potential SIRT3 Inhibitor, Suppresses Colorectal Cancer Proliferation in
Supplemental Material
sj-docx-2-npx-10.1177_1934578X231194808 - Supplemental material for Cryptotanshinone, a Potential SIRT3 Inhibitor, Suppresses Colorectal Cancer Proliferation in vitro and in vivo
Supplemental material, sj-docx-2-npx-10.1177_1934578X231194808 for Cryptotanshinone, a Potential SIRT3 Inhibitor, Suppresses Colorectal Cancer Proliferation in
Footnotes
Acknowledgments
Author Contributions
The conception and design of the study: Donglu Wu, Ye Jin, and Haoming Luo. Data acquisition: Dimeng Song and Zhiyuan Sun. Data analysis and interpretation: Dimeng Song, Xiaoxue Fang, Zhidong Qiu, and Da Liu. Drafting the article or revising it critically for important intellectual content: Dimeng Song, Zhiyuan Sun, and Xiaoxue Fang. Final approval of the version to be submitted: Dimeng Song and Xiaoxue Fang. All authors have read and agreed to the published version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
The animal study was reviewed and approved by the Institutional Animal Care and Use Committee of Jilin University (SY20191210; approved on 10 December 2019).
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the National Natural Science Foundation of China (Grant No. 81903876), Jilin Provincial Department of Education (JJKH20210982KJ), and Jilin Province Youth Science and Technology Talent Support Project (2020025).
Statement of Human Rights
No human participants were included in this study, and informed consent was not required.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not required.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
