Abstract
Introduction
Platycodon grandiflorus (PG) is a perennial herb belonging to the Platycodon genus in the Campanulaceae family. It is primarily found in Northeast China, North Korea, Japan, Russia. PG is commonly used as an auxiliary ingredient in dishes or used directly in pickles. 1 Additionally, it is recognized for its medicinal properties in the treatment of cough, phlegm, sore throat, and other respiratory disorders. 2 PG has a rich culinary heritage and is widely used in various dishes, including tea, kimchi, and side dish, as well as in the production of alcoholic beverages and preserved fruit. 3 It has a diverse composition of active ingredients, including saponins, flavonoids, polysaccharides, phenols, amino acids, and inorganic salts. 4 Among these, triterpenoid saponins, such as Platycodin A, Platycodin B, and Platycodin D are considered to be the key bioactive compounds due to their anti-inflammatory and immune-enhancing pharmacological properties.5–7 Notably, Platycodin D has gained significant attention from researchers due to its therapeutic effects8–10 (Figure 1).

Chemical structure of Platycodin D.
The biosynthetic pathways of triterpenoid saponins in medicinal plants have gradually been elucidated by researchers (Figure 2). Triterpenoid saponins are predominantly synthesized through 2 main pathways: the mevalonate (MVA) pathway in the cytoplasm and the methylerythritol phosphate (MEP) pathway in plastids.11,12 These pathways lead to the production of 3-isopentenyl diphosphate (IPP) and dimethyl allyl pyrophosphate (DMAPP) as the end metabolites, respectively. Key enzymes involved in the MVA and MEP pathways are 3-hydroxy-3-methylglutaryl-CoA reductase (HMGR), deoxyxylulose-5-phosphate synthase (DXS), and 1-deoxy-D-xylulose 5-phosphate reductoisomerase (DXR). 13 Triterpenoid saponins have a skeleton derived from the 30-carbon precursor oxidosqualene, with glycosyl residues are attached. 14 The biosynthesis of the carbocyclic skeleton of saponins involves several steps: isomerization of IPP to DMAPP; biosynthesis of farnesyl pyrophosphate (FPP) from IPP and DMAPP; generation of squalene from FPP by the action of squalene synthase (SS); and the formation of the carbocyclic skeleton through the catalytic action of squalene epoxidase(SE). 15 However, the downstream process of saponin biosynthesis remains unclear (Figure 3).

The MVA pathway and MEP pathway generate IPP and DMAPP, and then IPP and DMAPP synthesize the triterpenoid saponin precursor, 2,3-oxidosqualene.
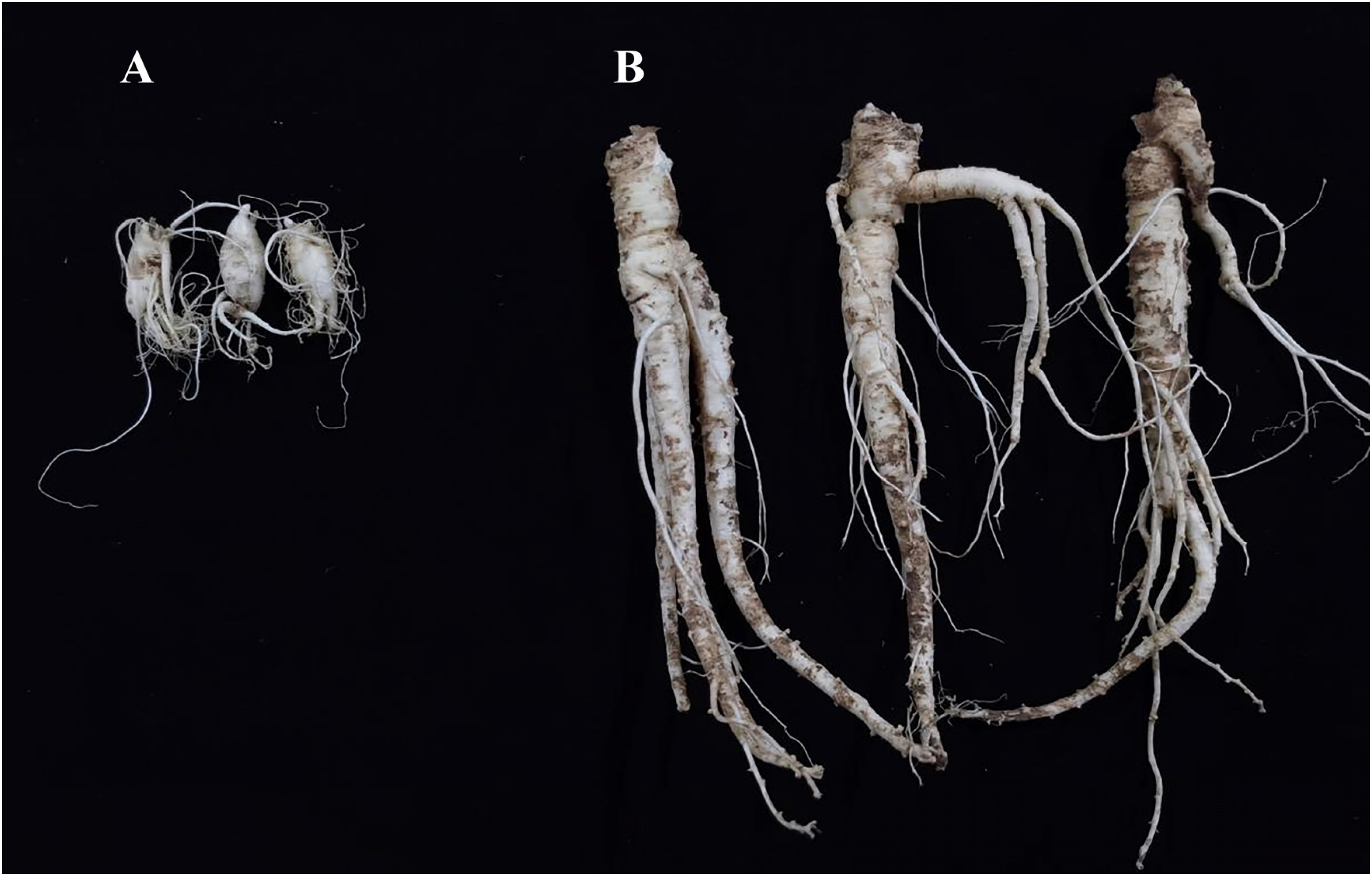
Sample of Platycodon grandiflorus seedlings. A: 6-month-old seedlings; B: 1-year-old seedlings.
Currently, research on PG saponins primarily focuses on optimizing extraction methods, 16 exploring pharmacological activities, toxicology, pharmacokinetics, and other related aspects. 17 However, there has been a lack of systematic investigation into the molecular mechanism underlying the biosynthesis of PG saponins. Therefore, the aim of this study was to examine the key genes and metabolic pathways involved in the biosynthesis of Platycodin D by assessing the expression of key genes associated with triterpenoid saponin synthesis. The findings of this study establish a research foundation for unraveling the molecular mechanism of PG saponin biosynthesis.
Results and Discussion
Total Saponin Content
The total saponin content in 1-year-old roots was 2.98%, while in 6-month-old seedlings, it was 2.25%. The findings revealed a significant difference in the total saponin content between 1-year-old roots and 6-month-old seedlings (P<.05), with the former exhibiting a higher content, as depicted in Figure 4A.

The contents of total saponins and Platycodin D in Platycodon grandiflorus seedlings at different developmental stages. A: total saponins; B: Platycodin D.
Currently, PG is primarily obtained through artificial cultivation in China, and its quality is influenced by factors such as origin, growth stage, and growing environment. Saponins, which are the active components of various medicinal plants, are also among the main active components of PG. Numerous studies have demonstrated their diverse pharmacological activities. For instance, ginsenosides have been found to possess antitumor, antioxidant, anti-fatigue, neuroprotective, and osteoclastogenic inhibitory effects. 18 Steroid saponins isolated from Rhizoma Paridis have shown antitumor, anti-inflammatory, hemostatic, and anti-fibrotic effects. 19 Additionally, Pulsatilla saponins have exhibited the ability to induce apoptosis in cancer cells, inhibit tumor angiogenesis, and provide organ protection through anti-inflammatory and antioxidant measures. 20 In this study, the results revealed a significantly higher total saponin content in 1-year-old PG roots compared to 6-month-old seedlings. These findings suggest that 1-year-old PG roots may possess greater medicinal value than 6-month-old seedlings.
The Content of Platycodin D
The peak area of Platycodin D was detected by ultra-performance liquid chromatography tandem mass spectrometry (UPLC-MS), and the chromatogram in MRM mode is presented in Figure 4. The retention time of Platycodin D was observed to be 8.07 min. The contents of Platycodin D in 1-year-old roots and 6-month-old seedlings quantified as 1700 and 1150 cps, respectively. These results indicated a significantly higher content of Platycodin D in 1-year-old roots compared to 6-month-old seedlings, as depicted in Figures 4B and 5.

MRM mode chromatogram of the Platycodin D. A: Platycodin D control solution; B: Samples of Platycodon grandiflorus.
Platycodin D, one of the major bioactive compounds in the medicinal plant PG, has recently garnered attention from researchers due to its potential therapeutic effects. Previous studies have demonstrated the diverse pharmacological properties of Platycodin D, including antioxidant, anti-inflammatory, anti-cancer, anti-obesity, and hepatoprotective activities. 8 Research on the influence of growth stages on the content of Platycodin D has predominantly focused on comparing 1-year-old seedlings. Studies conducted in Henan province, China have reported that the content of Platycodin D in 2-year-old PG plants is 1.5 times higher than that in 1-year-old plants. 21 Additionally, other studies have observed significantly higher levels of Platycodin D in 2-year-old cultivated PG compared to both annual and 3-year-old PG. 22 In the present study, a significant increase in the content of Platycodin D was observed in 1-year-old roots compared to 6-month-old seedlings. These findings support the notion that growth stages play a significant role in determining the content of Platycodin D. Furthermore, they corroborate the research on total saponins, affirming that the medicinal value of 1-year-old PG roots surpasses that of 6-month-old seedlings. However, the underlying mechanisms responsible for these differences require further investigation.
Expression Analysis of Key Genes Involved in the Biosynthesis of Saponins
Quantitative real-time polymerase chain reaction (PCR) was employed to assess the expression levels of genes involved in saponin biosynthesis pathways. The findings demonstrate differential expression of key genes in the MVA and MEP pathways between 1-year-old and 6-month-old seedlings. Specifically, in the MVA pathway, PGAACT and PGHMGR exhibited higher expression in 6-month-old seedlings, whereas PGMVK and PGMVD displayed higher expression in 1-year-old seedlings. Similarly, in the MEP pathway, PGDXS showed greater expression in 6-month-old seedlings, while PGDXR, PGMCT, PGMDS, PGHDS, and PGHDR were more highly expressed in 1-year-old seedlings. Furthermore, the seedlings exhibited higher expression of genes involved in the synthesis of key enzymes, PGSS and PGSE, compared to 1-year-old roots. Please refer to Figure 6 for further details.

The expression profile of key genes involved in the triterpenoid saponin biosynthesis pathways in Platycodon grandiflorus.
Gene regulation plays a crucial role in the biosynthesis of triterpenoid saponins in plants. 23 Previous studies have shown that the MVA and MEP pathways as 2 essential upstream pathways in saponin biosynthesis.24,25 In this study, we observed high expression of upstream genes, namely PGAACT and PGHMGR, in the MVA pathway of 6-month-old seedlings. It has been demonstrated in previous studies that AACT expression can restrict saponin biosynthesis in medicinal plants. 26 HMGR is a rate-limiting enzyme responsible for regulating the supply of terpene precursors (isopentenyl diphosphate and dimethylallyl diphosphate) and is crucial for preventing uncontrolled accumulation of bioactive saponins while ensures proper plant development and integrity. 27
Additionally, downstream genes such as PGMVK and PGMVD in the MVA pathway showed high expression in 1-year-old seedlings. Previous studies have revealed that saponin content in Panax ginseng significantly decreases under cadmium stress due to MVK suppression. 28 Furthermore, the MVA pathway has been identified as the primary pathway for saponin accumulation in tea tree, with the key gene diphosphopentanoic acid decarboxylase (MVD) playing a critical role in saponin accumulation in tea tree seeds and flowers. 24
The upstream genes of the MEP pathway, such as PGDXS, exhibit high expression levels in 6-month-old seedlings. Previous studies have demonstrated that the DXS expression can reduce the saponin biosynthesis by inhibiting the biotransformation of key metabolites in the saponin biosynthesis pathway. 26 Although the MEP pathway is not considered to be the primary synthetic pathway in ginsenoside synthesis, 29 present study found that downstream genes of the pathway, such as PGDXR, PGMCT, PGMDS, PGHDS, and PGHDR, showed high expression levels in 1-year-old seedlings. These findings suggest the significant involvement of MEP pathway in saponin synthesis in PG seedlings at different growth stages. These results indicate that most of the downstream genes in both the MVA and MEP pathways are highly expressed in 1-year-old seedlings, leading to higher accumulation of IPP and DMAPP, the end products of these pathways, in seedlings of this growth age.
Furthermore, farnesyl diphosphate synthase (FPS), SS, and SE have been implicated in regulating triterpenoid saponin synthesis.30,31 In the present study, FPS and SS were expressed higher in 1-year-old seedlings. Previous research on birch saponins has shown that heterologous expression of FPS in yeast increased squalene expression, resulting in higher production of triterpene saponins, such as bircholic acid, oleanolic acid, and betulinic acid. 32 Additionally, overexpression of FPS in Panax ginseng has been found to significantly increase saponins accumulation. 33 Similarly, overexpression of SS in the hairy roots of Pseudostellaria heterophylla and Panax ginseng led to increased saponins accumulation.34,35 SE is also a critical gene in the saponin biosynthesis pathway of Panax notoginseng, Eleutherococcus Senticosus, and ginseng.36–38 In this study, significantly higher levels of total saponins and Platycodin D in 1-year-old seedlings was observed compared to their 6-month-old counterparts. These results can be explained by the higher expression levels of key genes associated with the triterpenoid saponin synthesis pathway in 1-year-old seedlings, which regulate the essential pathway of saponin synthesis. These findings are consistent with previous investigations involving tea plants and Panax ginseng.24,39
Conclusion
The total saponin and Platycodin D content in 1-year-old PG roots are significantly higher than that in the 6-month seedlings. Moreover, real-time quantitative PCR analysis revealed that most of the genes involved in saponin biosynthesis exhibited higher expression levels in 1-year-old roots. This study provides evidence of the impact of growth stages on the saponin synthesis pathways and the regulation of key genes involved in saponin biosynthesis in PG. Future research can explore the molecular mechanisms underlying the regulation of saponin biosynthesis by developmental stages in PG, including the examination of key metabolites and gene expression levels.
Materials and Methods
Materials
Six-month and 1-year-old seedlings of PG were collected from the medicinal botanical garden of Heilongjiang University of Traditional Chinese Medicine. They were identified by Professor Wei Ma from the School of Pharmacy at the university. The collected seedlings were rinsed with sterile water, dried, frozen with liquid nitrogen, and stored at −80 °C in a refrigerator (Figure 3).
The Plant Total RNA Kit (product batch number: 20210412) was obtained from Hangzhou Xinjing Biochemical Reagent Development Co., Ltd. GeneCopoeia's BlazeTaqtm SYBR Green qPCR Mix 2.0 (product batch number: QP031) and SureScriptTM First-Strand cDNA Synthesis Kit (product batch number: QP056) were purchased from Guangzhou Yijin Biotechnology Co., Ltd. Platycodin D (product batch number: Z8J7B8461) was obtained from Shanghai Yuanye Biotechnology Co., Ltd. Methanol, acetonitrile, acetic acid, and ammonium acetate (chromatographic purity) were sourced from Thermo Fisher Scientific Technology (China) Limited Company.
Methods
The following instruments are used in this study: BioTek Epoch Ultra-microplate spectrophotometer (Berten Instruments Co., Ltd, USA); Agilent AriaMx Real-time PCR instrument (Agilent Technology (China) Co., Ltd); Ultra-High-Performance Liquid Chromatography, 4000 Q Trap LC-MS/MS (Waters Technology (China) Co., Ltd); KQ-500DE numerical control ultrasonic cleaner (Jiangsu Kunshan Ultrasonic Instrument Co., Ltd); SGF-A + 100 electronic analytical balance (Shanghai Gongye Co., Ltd), and an electronic balance (Sartorius Germany).
Extraction of total RNA and preparation of cDNA
The total RNA extraction from the seedlings was performed using the Simgen RNA kit following the manufacturer's instructions. The concentration and purity of the extracted RNA were determined, and the results indicated an A260/280 ratio between 1.8 and 2.0, confirming the purity of the RNA without contamination from DNA or proteins. Additionally, the integrity of the RNA was assessed through agarose gel electrophoresis. Subsequently, the RNA was reverse transcribed into cDNA using the GeneCopoeia SureScriptTM first-strand cDNA synthesis kit. The resulting cDNA was diluted 10-fold and stored in a refrigerator at −20 °C for later use.
Determination of total saponin content
The total content was extracted from 4 g of the coarse powder using 40 mL of ethanol through ultrasonication for 50 min. The crude extract was then subjected to centrifugation at room temperature at a revolution of 5000 r/min for 10 min. This process was repeated 3 times, and the resulting filtrates were combined. Next, the combined filtrate was concentrated to 15 mL in an electrothermal constant-temperature water bath and subsequently cooled. To separate the supernatant, 50 mL of ether was vigorously shaken with the concentrated filtrate, and the resulting supernatant was discarded. The precipitate was then treated with methanol (20, 10, and 5 mL) in 3 steps, heated, cooled, and filtered. Finally, the methanol solutions were combined and concentrated to 15 mL. The concentrated methanol solution was transferred to an evaporating dish and evaporated to dryness in a constant-temperature water bath. The resulting powder was dried in an electric dryer, and its weight was measured to determine the total saponin content. The total saponin content was calculated by dividing the weight of total saponins by the weight of the sample and multiplying it by 100%.
Preparation of the reference standard and test solution
The reference standard of Platycodin D was prepared by accurately weighing and dissolving it in methanol to achieve a concentration of 1 mg/1 mL. Subsequently, the reference standard was diluted to a concentration of 100 μg/L. For the samples, 1 g PG seedlings were ground thoroughly in liquid nitrogen. After adding 5 mL of 70% methanol, the mixture was subjected to ultrasonication for 1 h. The resulting extract was then centrifuged at 12000 rpm/min for 10 min and filtered using a 0.22 μm microporous membrane.
UPLC-MS/MS
The chromatographic analysis was performed using a Ultra-High-Performance Liquid Chromatography, 4000 Q Trap LC-MS/MS (Waters Technology (China) Co., Ltd). Separation was achieved on an Agilent EclipsePlusC18 Column (2.1 mm × 100 mm, 1.8 μm) through gradient elution. The gradient elution conditions were as follows: 0–0.5 min, 15% B; 0.5–10.5min, 15%–90%B; 10.50–11min, 15%B). The mobile phase used was a mixture of 5% formic acid in water-acetonitrile (B), and the flow rate was set at 0.3 mL/min. Full wavelength DAD scanning was employed. During the analysis, the column temperature was maintained at 35 °C, and an injection volume of 3 μL was used.
Mass spectrometry was performed using electrospray ion source, in the MRM (multiple reaction monitoring) detection mode. Nitrogen (N2) gas was used for drying, with a drying gas temperature set at 300 °C and a drying gas flow rate of 5 L/min. The atomizer pressure was set at 310.275 kPa. For sheath gas, the temperature was maintained at 250 °C, and the flow rate was set to 11 L/min. The capillary voltage was set to 3500 V. Platycodin D was detected in the negative ion mode, with the quantitative ion pair of 1247.4/705.5. The voltage used for detection was set at 270 V and energy was set at 100 V.
Real-time PCR primer design and gene expression analysis
The internal reference gene used in this study was glyceraldehyde-3-phosphate dehydrogenase (GAPDH). 40 PCR reactions were performed to amplify the following genes: PGAACT, PGHMGS, PGHMGR, PGMVK, PGPMK, PGMVD, PGDXS, PGDXR, PGMCT, PGCMK, PGMDS, PGHDS, PGHDR, PGGGPS, PGPS, PGSS, and PGSE. The primers were synthesized by Bioengineering (Harbin) Co., Ltd. The primer sequences used in this experiment are listed in Table 1. Real-time PCR was conducted for gene amplification. The total reaction volume of 20 μL, consisted of 10 μl of 2× TransStart green qPCR SuperMix UDG, 1 μL of cDNA, 0.4 μL of PrimerF and PrimerR, and 8.2 μL of nuclease-free Water. The reaction conditions included an initial denaturation at 50 °C for 2 min, followed by 40 cycles of denaturation at 90 °C for 10 s, annealing at 60 °C for 15 s, and extension at 72 °C for 15 s. After the PCR reaction, the temperature was increased from 50 °C to 95 °C, and a dissociation curve was generated to assess the purity of the amplified products. The experiments were conducted in triplicate, and the relative expression of the gene was determined using the 2–ΔΔCT method.
Primer Sequences Used for the Real-Time qPCR.

Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical Approval is not applicable for this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Key Research and development Project, research and demonstration of collection, screening and breeding technology of ginseng and other genuine medicinal materials [grant number 2021YFD1600901]; Heilongjiang Touyan Innovation Team Program [grant number 2019 No. 5].
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
