Abstract
Recently, numerous studies have focused on characterizing the biochemical activities of new natural marine resources as functional foods. Eucheuma spinosum J. Agardh., family Solieraceae, is a potential functional food that is widely used as a natural food source and contains bioactive components. This study aims to identify the bioactivity, drug-likeness, and pharmacokinetics of 3-(3-methoxyphenyl) propanal isolated from E. spinosum, and to discover the molecular interactions between the ligand, 3-(3-methoxyphenyl) propanal, and various antioxidant and antibacterial receptors. The bioactivity of 3-(3-methoxyphenyl) propanal was analyzed using computational Prediction of Activity Spectra for Substance (PASS) analysis. The drug-likeness was examined based on Lipinski's rule and other parameters. The pharmacokinetics of 3-(3-methoxyphenyl) propanal were predicted, and the molecular interactions between the receptors governing the antioxidant and antibacterial activity and 3-(3-methoxyphenyl) propanal were determined using the molecular docking method and molecular dynamics simulation. Based on the PASS analysis, 3-(3-methoxyphenyl) propanal was predicted to inhibit JAK2 expression, which contributes to apoptosis, with a probability of a molecule to be active (Pa) values of 0.733 and 0.508. Additionally, 3-(3-methoxyphenyl) propanal is an oxygen scavenger (Pa 0.408) and antimutagenic (Pa 0.419). Based on the drug-likeness and pharmacokinetic analyses, 3-(3-methoxyphenyl) propanal meets the requirements for an oral drug. Molecular docking revealed that binding affinity of 3-(3-methoxyphenyl) propanal for superoxide dismutase, tyrosinase, glutathione peroxidase, UDP-N-acetylmuramate, and penicillin-binding protein was −5.5, −6.1, −4.2, −6.2, and −4.9 kcal mol−1, respectively. Moreover, according to molecular dynamics simulation, propanal 3-(3-methoxyphenyl) propanal has stronger interaction with tyrosinase than the positive control. Based on PASS analysis and molecular docking results, 3-(3-methoxyphenyl) propanal exhibited promising potential as an antioxidant.
Introduction
Over the past few decades, extensive research has been conducted on functional foods and their bioactive constituents, which provide health benefits. These compounds target the mechanisms that manage, prevent, and/or treat infectious and metabolic diseases. The bioactive constituents in foods comprise powerful active molecules that are naturally present in small quantities, with the ability to regulate biological mechanisms. Functional foods are natural or processed foods that contain biologically active compounds in specific amounts. These foods are effective and nontoxic, and provide clinically proven health benefits for the prevention, management, and treatment of chronic diseases. Generally, there is no legal definition because functional foods are strictly regulated but not recognized by law in most countries.1–3
Marine resources have recently become increasingly intriguing sources of new drugs and healthy foods. 4 Many people consume fresh and dried seaweed, particularly those living in coastal areas. 5 Although, some of the properties of algae have been investigated, many remain unexplored. Compared with other algal species, the red alga Eucheuma spinosum exhibits unique therapeutic properties that require extensive exploration. Red algae are mostly found in tropical, coastal, continental, temperate, and cold water.6,7
E. spinosum is considered the most important source of active secondary metabolites compared to other algae. 8 It contains flavonoids, alkaloids, and derivatives with antibacterial, antifungal, antiviral, 9 and antioxidant properties. 10 Based on the content of secondary metabolites and other health benefits, E. spinosum has been established as a functional food with the potential to decrease the risk of diseases. 11 Several previous studies have reported that the ethyl acetate extract of E. spinosum obtained using the superoxide dismutase (SOD) method has strong antioxidant activity, with an IC50 value of 25 mg/L. 12 In DPPH assay, the antioxidant activity of this extract was relatively weak with an IC50 value of 402.8 mg/L. The antibacterial activity of the methanolic extract of E. spinosum against Staphylococcus aureus was low to medium with an inhibition zone of 4 mm at a maximum concentration of 80 mg/L. 10
Antioxidants are compounds that prevent free radical chain reactions and can consequently prevent cell damage. 13
Antioxidants are usually added to food to slow oxidative degradation and prevent chronic diseases in the body. 14 Free radical scavengers (antioxidants) are important for protecting cells against oxidative stress and maintaining the balance of toxic oxygen species. 15 Several free radical species are known, including superoxide anions (O2−), hydroxyl radicals (OH•), hydrogen peroxide (H2O2), hypochlorite anions (ClO−), and singlet oxygen. 16
Antioxidants are classified into 2 groups based on their mechanisms of action: Enzymatic and nonenzymatic. Enzymatic antioxidants convert dangerous oxidative products into hydrogen peroxide (H2O2) and then into water in a multistep process in the presence of cofactors such as copper, zinc, manganese, and iron. Several enzymatic antioxidants protect the human body from the dangers of reactive oxygen species (ROS), including superoxide dismutase (SOD), catalase (CAT), and glutathione peroxidase (GPx). 17 Tyrosinase is a rate-limiting enzyme involved in melanin synthesis. 18 Nonenzymatic antioxidants, including vitamins C and E, polyphenols, flavonoids, and carotenoids, interrupt free radical chain reactions. 13
In the oral system, saliva serves as a defense mechanism with antioxidant activity. Salivary antioxidant level can be altered by infection, disease, or inflammation. Peroxidase influences oral bacteria to prevent dental caries, 19 which are mainly caused by the bacterium Streptococcus mutans. 20 Five antibacterial pathways have been verified: inhibition of peptidoglycan formation involved in the biosynthesis of bacterial cell walls, inhibition of protein synthesis, inhibition of DNA and RNA synthesis, inhibition of folate synthesis inhibition, and membrane disruption. 21 Cell walls protect bacteria from osmotic pressure and maintain cell shape. 22 Inhibition of bacterial cell wall biosynthesis prevents dental caries. Several enzymes are involved in bacterial cell wall biosynthesis, including UDP-N-acetylmuramate (MurB) and penicillin-binding protein (PBP). 23 MurB catalyzes the conversion of enolpyruvate to D-lactate, resulting in the production of UD-N-acetylmuramate in the first step of cell wall biosynthesis. 24 PBP plays a role in the final step of cell wall biosynthesis by forming cross-links in peptidoglycan and defining bacterial cell walls. 25
Molecular docking is a computational method to visualize atomic interactions between small molecules and protein targets. This method provides information on small molecule activity in the binding site of the target proteins and biochemical processes. 26 Molecular docking includes 2 main stages: predicting the conformation of the ligand, including its location and orientation around the binding site, and aiming for binding affinity. 27 Both steps are correlated by conformational sampling, then all conformations are ranked using a scoring function. The lock and key theory is the most popular ligand–receptor binding mechanism, with the ligand as a key and the receptor as a lock. 28
Based on a general model of the physics driving interatomic interactions, molecular dynamics (MD) simulations forecast how each atom in a protein or other molecular systems will move over time. With femtosecond temporal precision, these simulations may disclose the positions of all the atoms during a wide range of crucial biomolecular processes, including conformational change, ligand binding, and protein folding. Importantly, these simulations can also forecast how biomolecules will react to alterations like mutations, phosphorylation, protonation, or the addition or removal of ligands at the atomic level. 29 This method can be used to identify functionally relevant conformations, as they may be difficult to obtain experimentally, as well as permit transitions between each conformation. 30
This research focuses on examining the molecular interactions between ligands, the isolated compound from E. spinosum (3-(3-methoxyphenyl) propanal, Figure 1), and several receptors related to antioxidant and antibacterial activity using molecular docking and MD simulation (in silico method). 3-(3- Methoxyphenyl) propanal is one of the isolated compounds from E. spinosum. 12 The results were then compared to the specific positive control interactions with receptors. The interactions of the isolated compound with the receptors were considered for further study of E. spinosum as a functional food. The bioactivities of the compound were examined using the Prediction of Activity Spectra of Substances (PASS) online program, and the potential as an oral drug was evaluated based on pharmacokinetics and drug-likeness predictions.

Structure of 3-(3-methoxyphenyl) propanal.
Result and Discussion
Marine natural products are prospective natural sources for the discovery of new functional foods that contain bioactive components, such as flavonoids, alkaloids, saponins, tannins, and corresponding derivatives. Recently, the demand for research and development to discover new potential natural sources of functional foods has increased rapidly. The active compounds responsible for the bioactivity must be properly identified to promote the consumption of new functional foods, in order to provide health advantages for the human body. Owing to the abundance of bioactive components in its extract, including tannins, flavonoids, triterpenoids, and steroids, E.spinosum has potential as a functional food. 31
Molecular Bioactivity Screening of Compound Based on PASS
Based on the PASS analysis (Table 1), 3-(3-methoxyphenyl) propanal possesses several antioxidant and antimicrobial activities. This compound was anticipated to inhibit JAK2 expression, contributing to apoptosis with a probability of a molecule to be active (Pa) values of 0.733 and 0.508. In addition, 3-(3-methoxyphenyl) propanal is an oxygen scavenger (Pa = 0.408) and antimutagenic (Pa = 0.419) agent that plays important roles in regulating oxidative stress. Moreover, this compound was predicted to inhibit tyrosine 3-hydroxylase (a critical agent in melanin formation), with a Pa value of 0.372. However, this compound is thought to be an antiseptic to kill pathogenic microorganisms. In particular, 3-(3-methoxyphenyl) propanal exhibits antitoxoplasma and antipicornavirus potencies, with Pa values of 0.308 and 0.398, respectively.
Prediction of Activity Spectra for Substance (PASS) Screening Data.

Molecular Docking of 3-(3-Methoxyphenyl) to Antioxidant Proteins
Molecular interactions between the antioxidant and antibacterial receptors and the isolated compound were observed using in silico molecular docking. Molecular docking is the process of placing a ligand in an appropriate conformation to a receptor following the lock and key concept. Ligands can be synthesized or isolated from compounds. Molecular docking can be used to determine the binding affinity and interactions between ligands and receptors, including hydrogen bonds and hydrophobic interactions related to the scoring function.28,32,33 To find important positions, the ligand is docked to the entire surface of the protein without any knowledge of the target pocket, which is known as blind docking. 34 During the docking process, the ligand rotates and moves into the fixed three-dimensional (3D) structure of the protein to determine the best-binding modes. 35
A scoring function is required to interpret the docking results, obtain accurate results, and determine the most stable ligand–receptor complex. The scoring function consists of several parameters, including the binding affinity, hydrogen bond, and hydrophobic interaction between ligands and receptors. 36 The binding affinity is related to the free energy change during the binding process, which indicated the strength of attachment of the ligands to the receptors. A more negative binding affinity in kcal/mol indicates stronger binding affinity and a highly stable complex. 33 Hydrogen bonds and hydrophobic interactions are included in the intermolecular bonds. As the number of hydrogen bonds and hydrophobic interactions increases, the stability of the ligand-receptor complex increases.37,38
Molecular Docking of 3-(3-Methoxyphenyl) Propanal to SOD, Tyrosinase, and GPx Enzymes
The docking results obtained using PyRx revealed the binding affinity of [3-(3-methoxyphenyl) propanal (3-(3-methoxyphenyl) propanal) and isobutylamido thiazolyl resorcinol to tyrosinase, 3-(3-methoxyphenyl) propanal and disodium ethylenediaminetetraacetate to SOD, and 3-(3-methoxyphenyl) propanal and thioctic acid to GPx. 3-(3-methoxyphenyl) propanal had a binding affinity score as shown in Table 2.
Binding Affinity Scores of 3-(3-Methoxyphenyl) Propanal and Positive Controls to Tyrosinase, SOD, and GPx Enzymes.

Abbreviations: GPx, glutathione peroxidase; SOD, superoxide dismutase.
Among the 3 receptors related to antioxidant activity, 3-(3-methoxyphenyl) propanal exhibited the strongest binding affinity (lowest value in kcal/mol) 39 when attached to tyrosinase, which inhibits the receptor. The binding affinity of 3-(3-methoxyphenyl) propanal for tyrosinase and SOD was higher (in kcal/mol) indicating weaker binding affinity compared with that of that positive controls, whereas the binding affinity of 3-(3-methoxyphenyl) propanal for thioctic acid had the same value in kcal/mol. 3-(3-Methoxyphenyl) propanal had the same number of interactions with tyrosinase and SOD but showed only 2 interactions with GPx.
Figure 2 illustrates the protein-ligand complexes related to the antioxidant activity, with 3-(3-methoxyphenyl) propanal in red and the positive control in yellow. Figure 3 illustrates ligands and residues involved in the receptor interactions.

Docking pose of 3-(3-methoxyphenyl) propanal and positive control with 3 receptors related to antioxidant activity: Disodium ethylenediaminetetraacetate and 3-(3-methoxyphenyl) propanal with SOD enzyme (

Molecular interactions of 3-(3-methoxyphenyl) propanal with 3 receptors related to antioxidant activity: (
The SOD-3-(3-methoxyphenyl) propanal complex had a binding affinity score of −5.5 kcal/mol. 3-(3-Methoxyphenyl) propanal was bound to 6 residues, VAL7, VAL138, PHE86, LEU211, VAL138, and VAL213, as indicated in Table 3. Disodium ethylenediaminetetraacetate as a positive control had a binding affinity of −6.0 kcal/mol and was bound to 7 different residues: VAL213, ILE82, VAL148, LYS9, CYS146, CYS228, and GLU84. VAL213 contains 2 hydrogens attached to a residue.
Analysis Results of Docked 3-(3-Methoxyphenyl) Propanal and Disodium Ethylenediaminetetraacetate With Superoxide Dismutase (SOD).

The total number of bonds was reduced by 2 compared to the binding of the positive control with SOD, indicating a difference between 3-(3-methoxyphenyl) propanal and the SOD positive control. On average, in the positive control, there was a significantly shorter bond distance between the ligand and the protein. The shorter the bond distance, the more difficult it is to break the bond; therefore, the interaction is more stable. The interaction is considered strong if the bond length is less than 3.00 Å. 40 In addition, the positive control favored hydrogen bond formation because 8 amino acid residues were all hydrogen-bonded. The binding-site positions for both ligands were close to each other. The amino acid residue present in both ligands was VAL213.
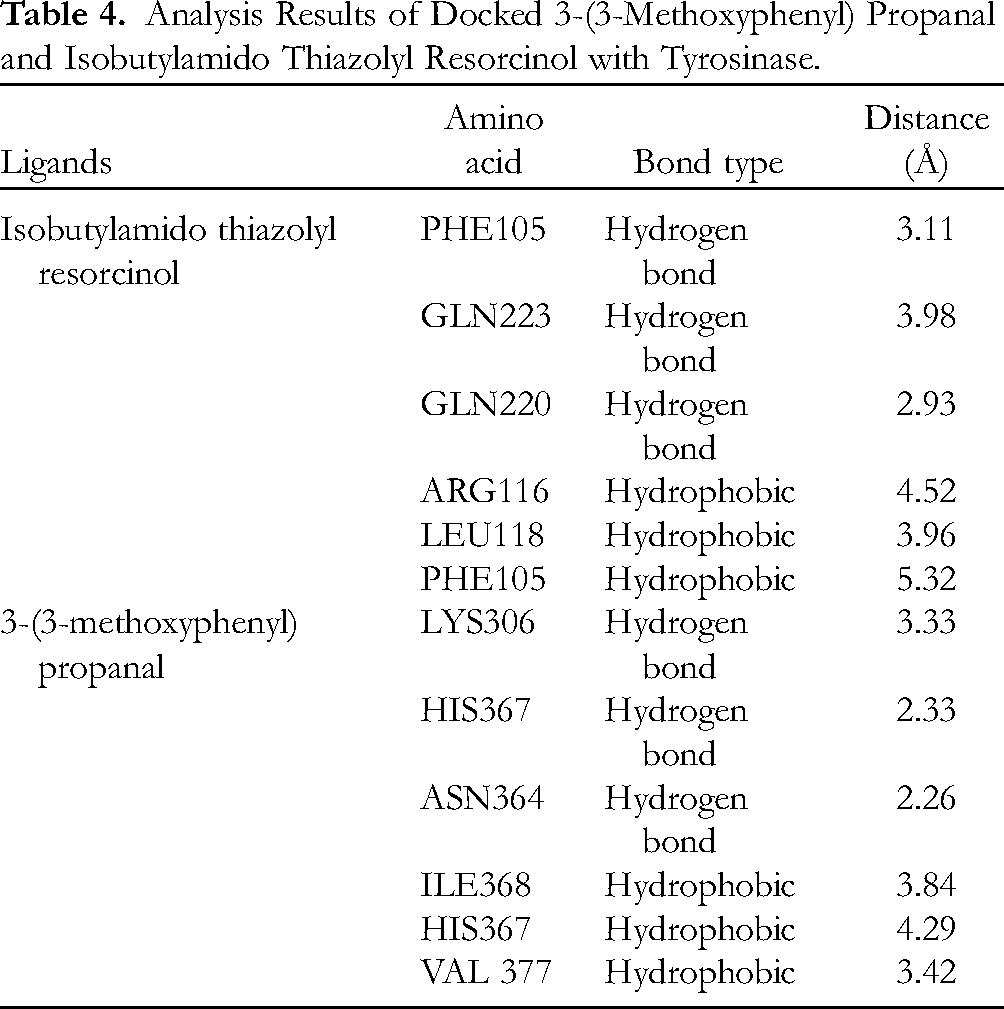
The binding affinity score of the tyrosinase-3-(3-methoxyphenyl) propanal complex was −6.1 kcal/mol. As shown in Table 4, 3-(3-methoxyphenyl) propanal was bonded to 6 residues: LYS306, HIS367, ASN364, ILE368, HIS367, and VAL377. The complexes of 3-(3-methoxyphenyl) propanal-tyrosinase and positive control-tyrosinase had 3 hydrogen bonds and hydrophobic interactions, respectively. Isobutylamido thiazolyl resorcinol, as a positive control, had a binding affinity of −7.3 kcal/mol and was bonded to 6 different residues: PHE105, GLN223, GLN220, ARG116, LEU118, and PHE105. In general, the bond lengths in 3-(3-methoxyphenyl) propanal-tyrosinase complex are shorter than those of the positive control tyrosinase complex. All the residues that bind to both ligands did not show any similarity. There was no similarity in the bonding between the residues and either ligand. The mechanism of phenol-induced tyrosinase activity was determined by the oxidation of phenol to orthoquinones using oxy-tyrosinase. Dopaquinone is formed from
Analysis Results of Docked 3-(3-Methoxyphenyl) Propanal and Isobutylamido Thiazolyl Resorcinol with Tyrosinase.
GPx-3-(3-methoxyphenyl) propanal complex had a binding affinity of −4.2 kcal/mol. As shown in Table 5, 3-(3-methoxyphenyl) propanal was bonded to 2 residues, THR141 and MET140. Thioctic acid, as a positive control, had a binding affinity of −4.2 kcal/mol and was bonded to 6 different residues: MET106, LEU20, ALA21, LEU89, PRO103, and LEU107. Compared to the binding affinity values reported in previous research, reduced glutathione (GSH) was docked to GPx (PDB ID:1GP1) and showed a binding affinity of −5.4 kcal/mol, and was bound to SEC45, PHE145, TRP158, ARG177, ASP126, and ALA129. 42
Docking Data for 3-(3-Methoxyphenyl) Propanal and Thioctic Acid with Glutathione Peroxidase (GPx).

The length of the hydrogen bond formed with 3-(3-methoxyphenyl) propanal was shorter than that of the positive control, whereas the bond length of the hydrophobic 3-(3-methoxyphenyl) propanal was longer than that of the positive control. 3-(3-Methoxyphenyl) propanal and GPx showed fewer interactions, with only 1 hydrogen and 1 hydrophobic bond. Neither 3-(3-methoxyphenyl) propanal nor the positive control was bound to the same residue in GPx. Nevertheless, thioctic acid and GPx formed 3 hydrogen bonds and 3 hydrophobic bonds, respectively.
Molecular Docking of 3-(3-Methoxyphenyl) Propanal to MurB and PBP Enzymes
Natural products are a source of therapeutically viable antibacterial agents. Higher plants synthesize various bioactive compounds that act as antifungal and antibacterial agents. 43 Herein, molecular interactions were also observed between the compound isolated from E.spinosum and receptors related to antibacterial activity (MurB and PBP), which prevent dental caries. These enzymes are involved in the biosynthesis of the bacterial cell wall.
Based on the molecular docking analysis, 3-(3-methoxyphenyl) propanal shows activity against the receptors involved in the early and final steps of cell wall biosynthesis. The docking results of 3-(3-methoxyphenyl) propanal and the specific positive control for each receptor are listed in Table 6. MurB-3-(3-methoxyphenyl) propanal complex had a lower binding affinity score (−6.2 kcal/mol) than the PBP-3-(3-methoxyphenyl) propanal complex (−4.9 kcal/mol), suggesting that MurB-3-(3-methoxyphenyl) propanal exhibits stronger binding affinity than the PBP-3-(3-methoxyphenyl) propanal complex. 3-(3-Methoxyphenyl) propanal had a weaker binding affinity for MurB and PBP enzymes than the positive controls. Compared to the positive controls, more interactions were found in the PBP-3-(3-methoxyphenyl) propanal complex than in MurB-3-(3-methoxyphenyl) propanal.
Binding Affinity Scores of 3-(3-Methoxyphenyl) Propanal and Positive Control Against MurB and PBP.

Abbreviations: MurB, UDP-N-acetylmuramate; PBP, penicillin-binding protein
Ligand positions in MurB and PBP were shown in Figure 4 which were 3-(3-methoxyphenyl) propanal in red and quercetin in yellow. Meanwhile, Figure 5 illustrates 3-interaction of 3-(3-methoxyphenyl) and amino acid residues in the receptor.
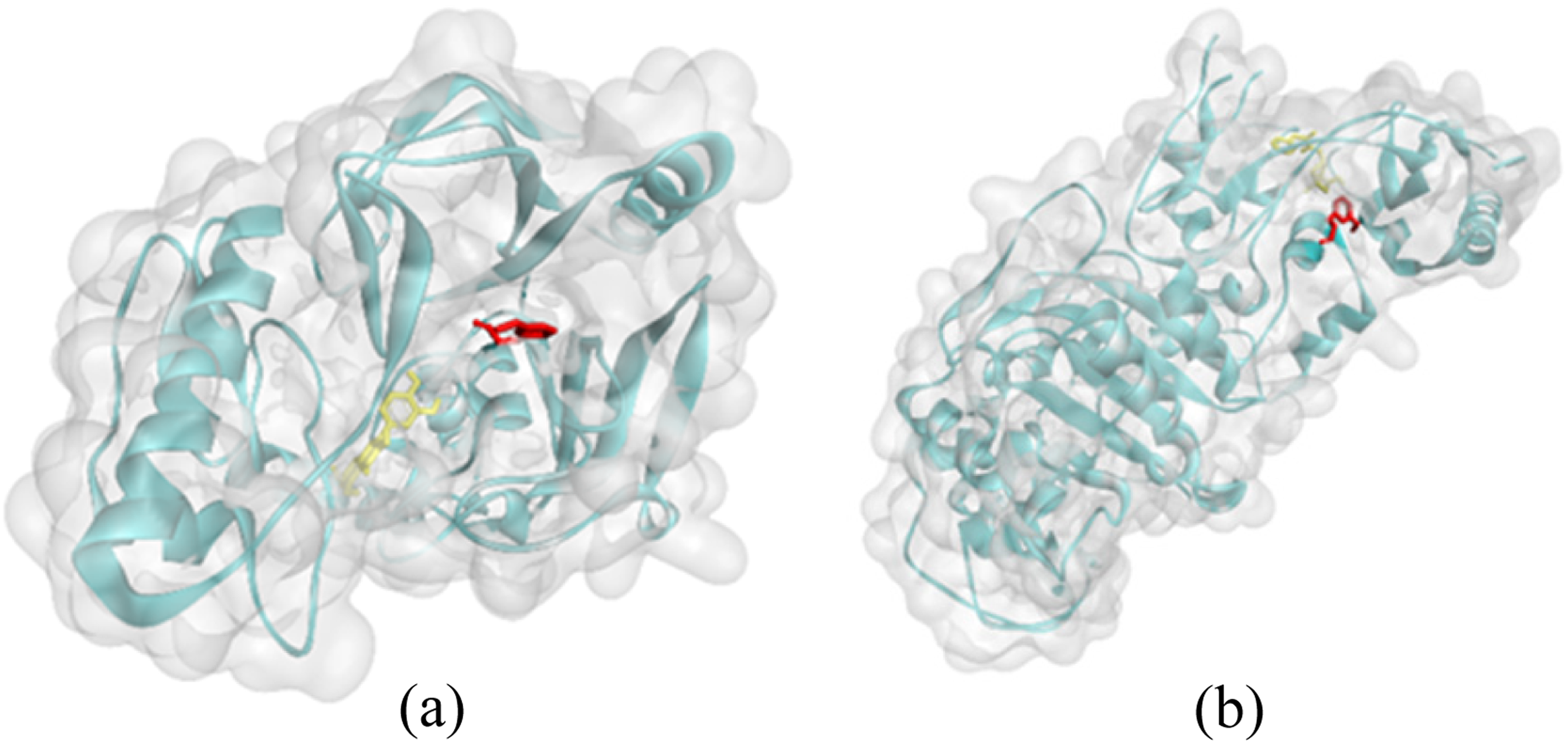
Docking pose of 3-(3-methoxyphenyl) propanal and positive control with 2 receptors related to antibacterial activity, quercetin and 3-(3-methoxyphenyl) propanal with UDP-N-acetylmuramate (MurB) enzyme (
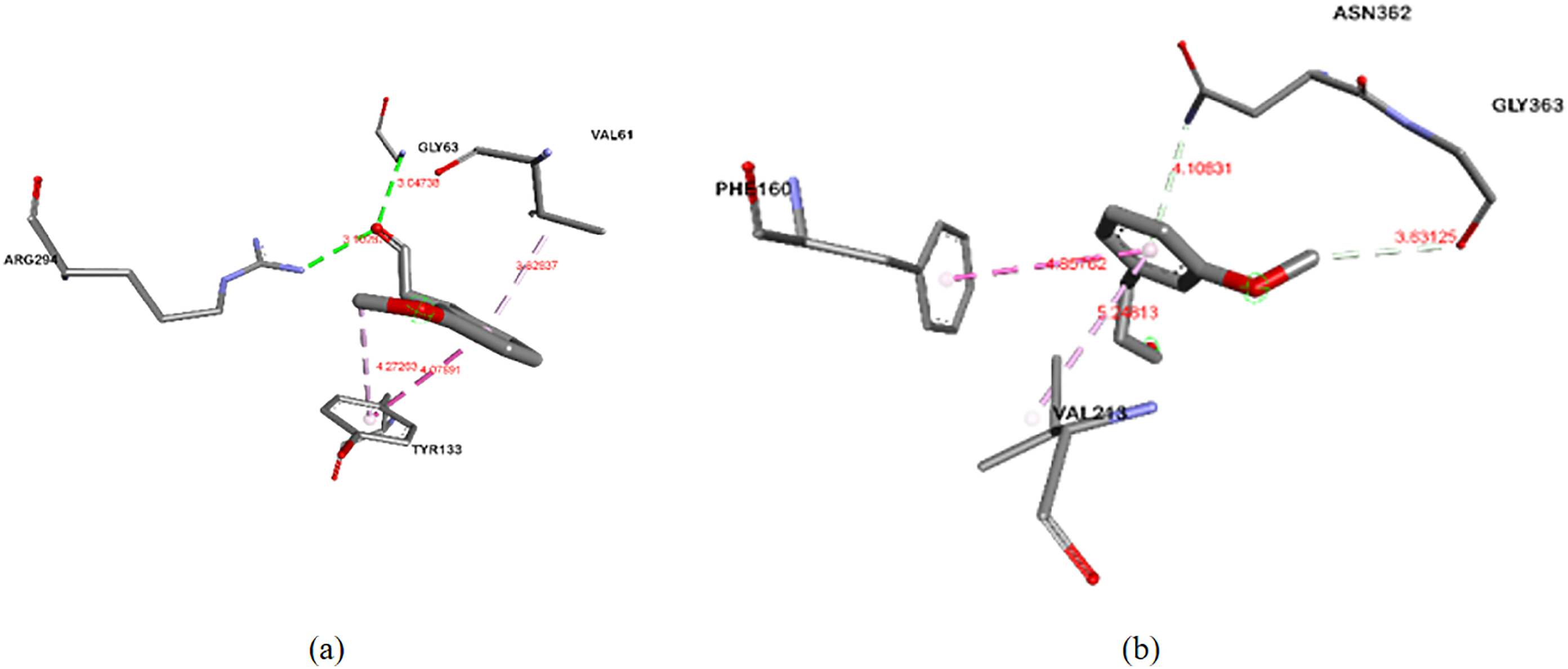
Molecular interactions of 3-(3-methoxyphenyl) propanal with 2 receptors related to antibacterial activity, (
Compared with 3-(3-methoxyphenyl) propanal, quercetin interacts more strongly with the receptor via hydrogen and hydrophobic bonds. Generally, the hydrogen bond length in quercetin is shorter than that in 3-(3-methoxyphenyl) propanal. The bond lengths of hydrophobic bonds in 3-(3-methoxyphenyl) propanal were shorter than those in quercetin. The interactions formed in the ligand-MurB complexes and the ligand-PBP complexes are listed in Tables 7 and 8, respectively.
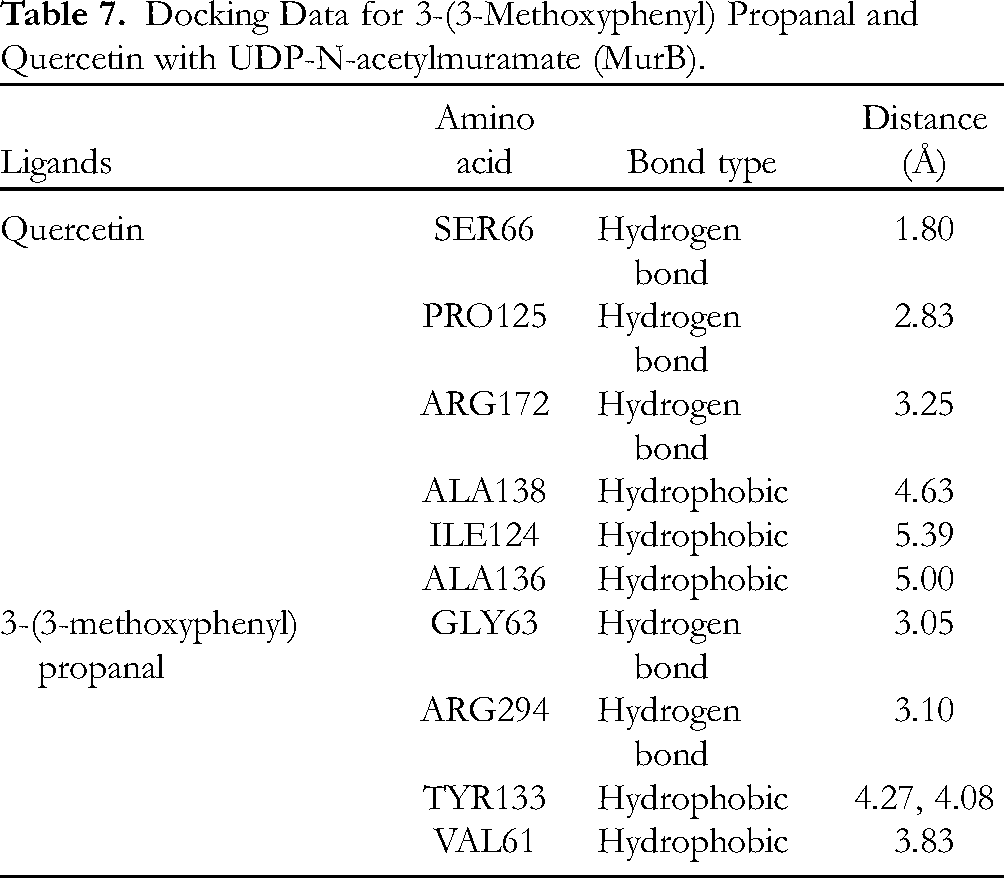
Docking Data for 3-(3-Methoxyphenyl) Propanal and Quercetin with UDP-N-acetylmuramate (MurB).
Docking Data for 3-(3-Methoxyphenyl) Propanal and Penicillin with Penicillin-Binding Protein (PBP).

Table 7 shows that 3-(3-methoxyphenyl) propanal was bound to 4 residues: GLY63, ARG294, TYR133, and VAL61. Quercetin as positive control showed a binding affinity of −8.2 kcal/mol and was bound to 6 different residues: SER66, PRO125, ARG172, ALA138, ILE124, and ALA136. In previous studies, the binding affinities of MurB and 3 different ligands (β-sitosterol, chlorhexidine, and glycopeptides) are reported as −7.6, −8.7, and −7.4 kcal/mol, respectively. β-Sitosterol binds to ARG226, ALA138, ILE124, PRO125, MET134, and ALA136. Chlorhexidine bound to GLU801, ARG683, ASP689, GLU692, THR737, TYR645, VAL573, MET688, and LYS799, while glycopeptides bound to TYR139, ARG209, LYS212, SER222, and PRO125. 32
Table 8 shows that 3-(3-methoxyphenyl) propanal binds to 2 residues: ASN362, GLY363, PHE160, and VAL213. Penicillin, as a positive control, had a binding affinity of −6.9 kcal/mol and was bound to 3 different residues: TRP202, THR265, and VAL267. A previous study of PBP with the same PDB ID docked with β-sitosterol and chlorhexidine provided a binding affinity of −7.8 kcal/mol for β-sitosterol and −9.8 kcal/mol for chlorhexidine. β-Sitosterol binds to GLU231 and PRO210, and chlorhexidine to ILE364, MET360, GLU162, THR230, SER232, ASN362, PHE160, GLN366, ASN358, LYS359, VAL213, TYR234, and TYR369. 32
Determining Drug-Likeness Based on Lipinski's Rule
Based on Lipinski's analysis, the probability of systemic absorption is critical for determining whether a drug can be administered orally. According to Lipinski's evaluation, 3-(3-methoxyphenyl) propanal complies with this rule (Table 9) which means this compound was within a good range for consideration as a drug candidate. Five parameters were evaluated: the solubility (log P), molecular mass, molar reactivity, and hydrogen-bond acceptors and donors. The log P value indicates the coefficient of solubility in lipid/water (range: −0.4 to 5); the greater the log P value, the more hydrophobic the ligand. The log P of 3-(3-methoxyphenyl) propanal is 1.728, indicating its solubility in water. This compound can also diffuse across cell membranes because of its molar mass of 164 kDa, which is less than 500 kDa. Excessively hydrophobic molecules tend to be highly toxic because they are retained longer in the lipid bilayer and cause wider disturbance in the body, reducing the selectivity of binding to the target enzyme. A log P value that is too negative is undesirable because the molecule cannot pass through the lipid bilayer membrane. The number of hydrogen bond donors and acceptors indicates the hydrogen bonding capacity. The higher the hydrogen bonding capacity, the higher the energy required for the absorption process. Generally, 3-(3-methoxyphenyl) propanal meets Lipinski's rule, indicating that it is sufficiently soluble to penetrate the cell membrane via passive diffusion.
Results of Applying Lipinski's Rule.

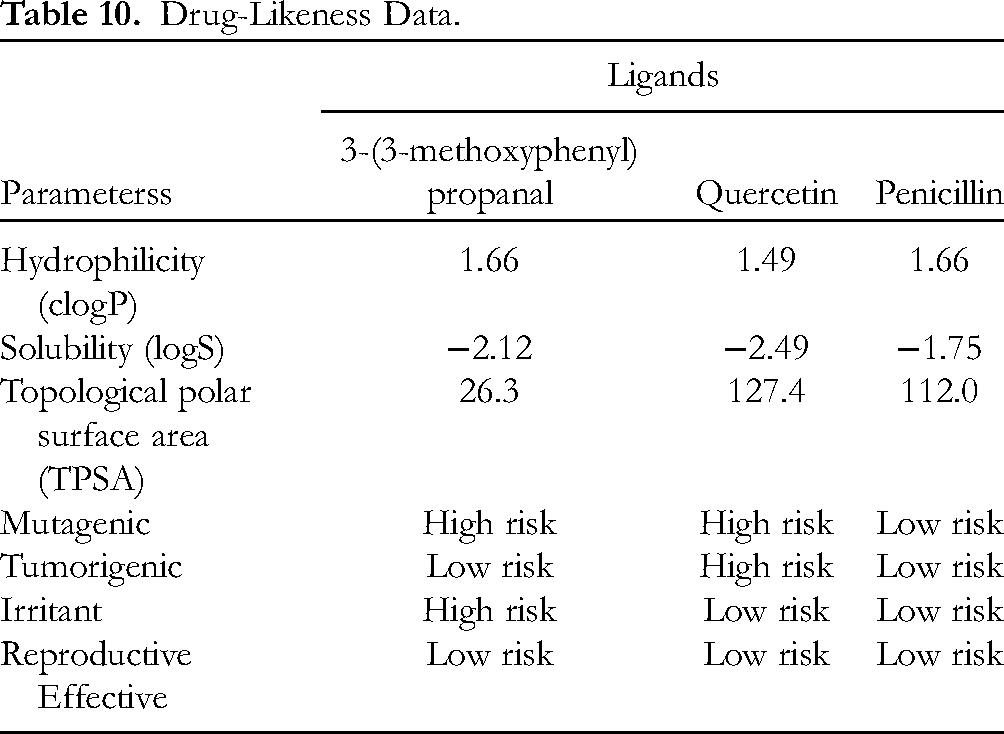
Furthermore, drug-likeness evaluation by ORISIS revealed that 3-(3-methoxyphenyl) propanal exhibits good hydrophilicity and solubility (Table 10). The hydrophilicity is represented by clogP; the lower the hydrophilicity of the compound, the higher the clogP value. 3-(3-Methoxyphenyl) propanal and penicillin had the same clogP value of 1.66, whereas quercetin had a lower value of 1.49. Therefore, it is predicted that quercetin will be easily absorbed and will more readily permeate cells than the other compounds. The value of logS indicates the solubility of 3-(3-methoxyphenyl) propanal in water. A compound is well absorbed if it has good water solubility. The higher the solubility in water, the lower the logS value. Given their hydrophilic properties, the solubility of all ligands is good since their logS values are in the range specified by the rules (greater than −4). The other properties, such as topological polar surface area (TPSA), follow the same trends. All ligands have TPSA values in the range of 0 ≥ TPSA ≤ 132 Å, suggesting that they have good oral bioavailability characteristics. However, 3-(3-methoxyphenyl) propanal is thought to have a high risk of being mutagenic and irritant, while quercetin (
Drug-Likeness Data.
Pharmacokinetic Prediction for 3-(3-Methoxyphenyl) Propanal as an Oral Drug
Table 11 displays the toxicity prediction. All ligands are absorbed in the gastrointestinal (GI) tract and have the same bioavailability score of 0.55. 3-(3-Methoxyphenyl) propanal is almost nontoxic against cytochrome, suggesting that it can be employed as an oral drug.
Pharmacokinetic Prediction of 3-(3-Methoxyphenyl) Propanal.

The absorption, distribution, metabolism, and excretion were further analyzed using Swiss-ADME prediction. Based on the data in Figure 6, all ligands are in the white region, indicating that these ligands can be absorbed by the GI system. However, only 3-(3-methoxyphenyl) propanal was present in the yolk region, suggesting that it can penetrate the brain. Penicillin (

Boiled-egg visualization of pharmacokinetics.
MD Simulation
In this study, a MD simulation was used to highlight the mechanism by which 3-(3-methoxyphenyl) propanal inhibits tyrosinase while employing the positive control isobutylamido thiazolyl resorcinol. Compared to other methodologies, this one can more clearly show how a system interacts at a spatiotemporal resolution.
Based on the graph (Figures 7 and 8), 3-(3-methoxyphenyl) propanal has root mean square deviation (RMSD) values ranging from 0 to 1.8 Å, isobutylamido thiazolyl resorcinol had RMSD values ranging from 0 to 2.05 Å, while the receptor (tyrosinase) has RMSD values ranging from 0 to 2.3 Å. The structure of the receptor and both ligands can be declared stable because they are still in the 2 to 3 Å range; in this range, protein movement is still normal and there is no conformational change.
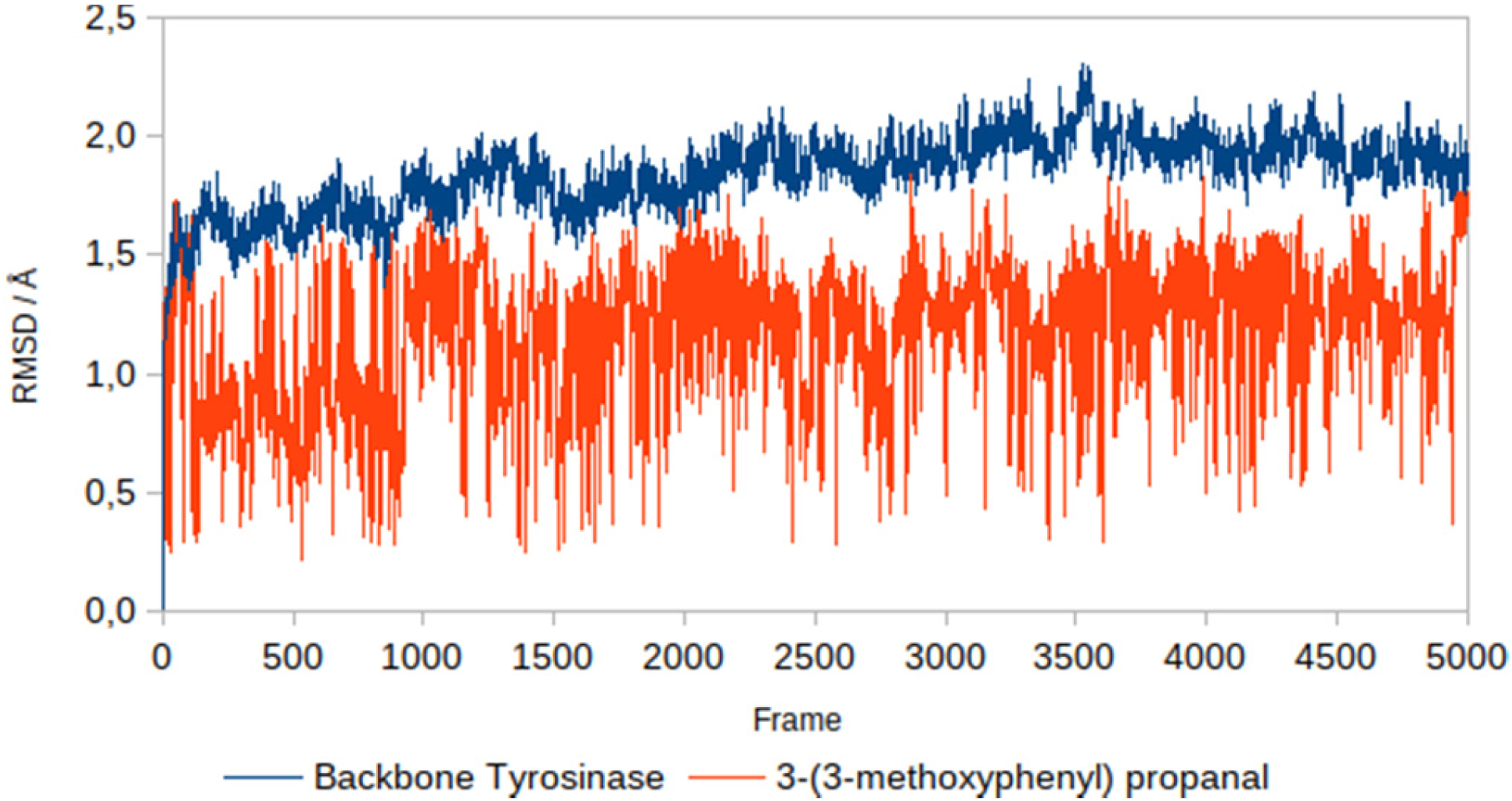
Backbone root mean square deviation (RMSD) of tyrosinase and 3-(3-methoxyphenyl) propanal.
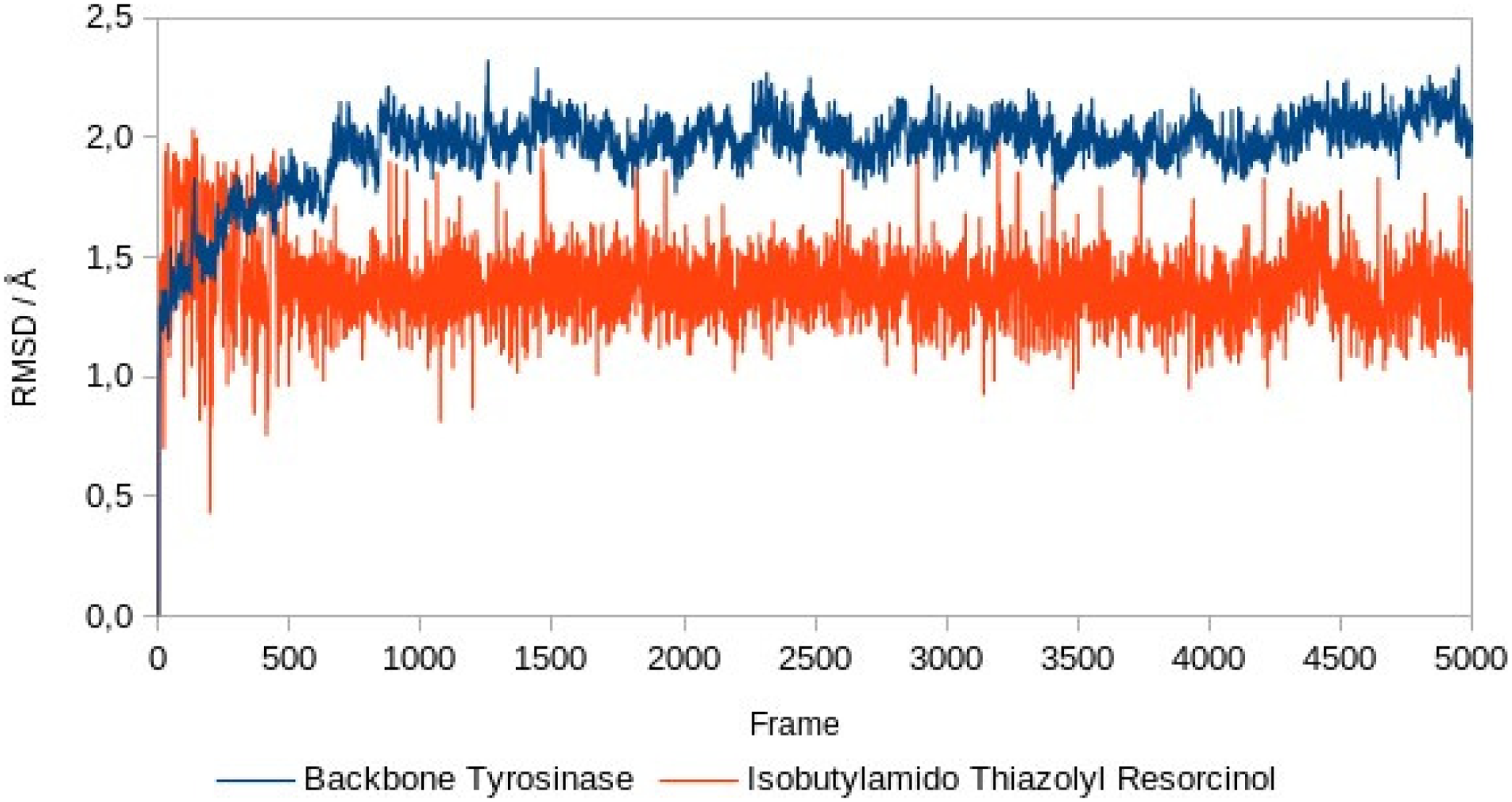
Backbone root mean square deviation (RMSD) of tyrosinase and isobutylamido thiazolyl resorcinol.
The findings demonstrated that as ΔGgas increased, so did ΔGsolv. These 2 energy values add up to form ΔGtotal, which denotes the possibility of the formation of stable bonds. All of these free energy values were negative, indicating that all of the simulated complexes could be completed in the laboratory, based on the free energy values.
The MD simulation results were 3-(3-methoxyphenyl) propanal with a value of −14.2206 kcal/mol compared to the standard, namely isobutylamido thiazolyl resorcinol with a value of −13.1725 kcal/mol, as shown in Table 12. This indicated that 3-(3-methoxyphenyl) propanal is more stable than isobutylamido thiazolyl resorcinol, making it a good candidate as a tyrosinase inhibitor. This outcome contrasts with molecular docking, which takes place in static conditions, as opposed to a MD simulation, which takes place in dynamic situations. The outcomes of the docking and MD simulations are different as a result of this condition.
Interaction Energies of Tyrosinase with Isobutylamido Thiazolyl Resorcinol and Tyrosinase with 3-(3-Methoxyphenyl) Propanal Calculated by MMPBSA.

Isobutylamido thiazolyl resorcinol (Figure 9) and 3-(3-methoxyphenyl) propanal have 1 hydrogen bond interaction with tyrosinase (Figure 10). However, 3-(3-methoxyphenyl) propanal has more interactions than isobutylamido thiazolyl resorcinol. It is suspected that this is why the bond of 3-(3-methoxyphenyl) propanal is stronger than that of isobutylamido thiazolyl resorcinol.

The intermolecular interaction of isobutylamido thiazolyl resorcinol with tyrosinase

The intermolecular interaction of 3-(3-methoxyphenyl) propanal with tyrosinase
Potency 3-(3-Methoxyphenyl) Propanal as Antioxidant and Antibacterial Compound
Based on the binding affinity values obtained from molecular docking, 3-(3-methoxyphenyl) propanal demonstrated weaker potency as an antioxidant and antibacterial agent than the positive control. However, the in vitro DPPH assay of 3-(3-methoxyphenyl) propanal (87.97 ppm) showed good results, with a better IC50 value than that of catechin (91.82 ppm). 3-(3-Methoxyphenyl) propanal showed a weaker binding affinity for the receptors related to the antibacterial activity than for those related to the antioxidant activity. The antibacterial activity of the ethanol extract E. spinosum was examined in vitro using the inhibition zone, MIC, and MBC methods with S aureus as a target. The 5% ethanol extract showed an inhibition zone of 10.1 mm, with a MIC of 100,000 ppm and MBC of 150,000 ppm. E. spinosum is an alga commonly consumed by many people. E. spinosum, which has antioxidant and antibacterial activities, has the potential to be used as an oral drug, and the ROS scavengers make it potentially applicable as a functional food.
From the in silico data, the functional group of 3-(3-methoxyphenyl) propanal, which formed the most bonds with the targeted protein, was a methoxy group. In the SOD and tyrosinase complexes, the methoxy group forms more bonds (3 bonds) than the aldehyde group (1 bond). Even in other complexes (GPx, MurB, and PBP), there is at least 1 methoxy bond, whereas the aldehyde group has no bond with the protein. According to theory, the more bonds that are formed, the greater the probable activity of the compound. Therefore, the methoxy group is thought to have the greatest impact on the activity of E. spinosum. This is in line with a previous study showing that the methoxy group plays a key role in antioxidant activity. The methoxy group enhances the antioxidant properties of phenolic acids by lowering the proton affinity and electron transfer enthalpy, The methoxy group can decrease the bond dissociation enthalpy of the phenolic hydroxyl group and improve the ability of phenolic acids to donate electrons. 44
Furthermore, we compared the antioxidant activity of 3-(3-methoxyphenyl) propanal (

Structures of
Antioxidant Activity of

Conclusions
Based on PASS analysis, molecular docking, and molecular dynamic studies, 3-(3-methoxyphenyl) propanal, isolated from E. spinosum, shows potential as an antioxidant agent. PASS analysis predicted that 3-(3-methoxyphenyl) propanal should have potential activity as an oxygen scavenger, which is critical for reducing oxidative stress in the human body. In addition, molecular dynamic studies show 3-(3-methoxyphenyl) propanal inhibits tyrosinase enzyme. This compound also has the potential to be used as an oral drug. Therefore 3-(3-methoxyphenyl) has the potential to be developed in the future.
Materials and Methods
Materials
The interactions of 3-(3-methoxyphenyl) propanal with 5 receptors having antioxidant and antibacterial activities were evaluated. The receptors, known as macromolecules, were obtained from the UniProt protein database (https://www.uniprot.org) under ID numbers P14679 for tyrosinase, P00441 for SOD, and Q830P3 for MurB. RCSB (https://www.rcsb.org/) is another protein database from which GPx (PDB code: 1GP1) and PBP (PDB code: 6BSQ) were downloaded. The data file format acquired from Uniprot was XML, which was subsequently converted to the protein data bank (PDB) format, whereas the data file format obtained for RCSB was PDB. SOD, tyrosinase, and GPx are associated with antioxidant activity, whereas PBP and MurB are associated with antibacterial activity. Isobutylamido thiazolyl resorcinol (C13H14N2O3S) (CID: 71543007) disodium ethylenediaminetetraacetate (Na2EDTA) (C10H14N2Na2O8) (CID: 8759), thioctic acid (C8H14O2S2) (CID: 864), quercetin (C15H10O7) (CID: 5280343), and penicillin (C16H18N2O4S) (CID: 2349) were used as positive controls. The structures of the isolated compound (CID : 11423769) and positive controls were downloaded from the chemical structures database PubChem (https://pubchem.ncbi.nlm.nih.gov/).
AMBER22 software was used to perform MD on chosen protein-ligand complexes. The protein complex structure was loaded into the Load, Energy, and Minimize Proteins (LEaP) program. The compound was parameterized with the protein FF14SB, the ligand GAFF2, and the partial charge AM1-BCC. The complex was dissolved in a 10 Å box of TIP3P water, crystal water molecules were removed using the docking process, and the addition of Na+ ions neutralized the remaining charges.
Methods
Molecular Docking
Molecular docking was conducted to determine the mechanism of binding between 3-(3-methoxyphenyl) propanal and tyrosinase, superoxide dioxide (SOD), and glutathione peroxidase (GPx), which are enzymes related to antioxidant activity, whereas MurB and PBP enzymes are associated with antibacterial activity. The docking score results predict the activity of the active compound by examining the binding affinity between the target protein and the compound. All proteins are downloaded as PDB files, whereas the 3D structures of 3-(3-methoxyphenyl) propanal and the positive controls were downloaded from PubChem as standard database format files and subsequently converted to PDB files using an online SMILES translator (https://cactus.nci.nih.gov).
The positive controls and 3-(3-methoxyphenyl) propanal (ligands) were docked to a specific protein target or receptor using AutoDock Vina in PyRx 0.8 software.
Lamarkian Genetic Algorithm was applied as hyper-parameters in docking simulations with 10 GA runs since the molecular docking was done using autodock vina in PyRx 0.8 program. Each receptor was loaded as a macromolecule into the program, and either the positive control or 3-(3-methoxyphenyl) propanal was loaded as a ligand. The receptor was free of the ligand; therefore, the docking process was blind to each ligand. Each receptor had a specific grid box as the docking area with X, Y, and Z dimensions. Tyrosinase had a grid box with X, Y, and Z dimensions of 149.6658, 90.7198, and 93.6411, respectively. The SOD scores were 54.7319, 69.7443, and 54.0521 for each dimension, respectively. The GPx grid boxes were 42.1231, 45.8467, and 41.5241 in the X, Y, and Z dimensions, respectively. The MurB grid boxes were 52.2667, 46.2509, and 53.9848 in the X, Y, and Z dimensions, respectively. The PBP grid boxes were 46.6095, 85.4274, and 63.8890 in the X, Y, and Z dimensions, respectively. Aside from grid box, a grid center was also specified for each receptor by using the X, Y, and Z coordinates for each ligand: tyrosinase (X: 5.1705, Y: −7.1671, and Z: −3.7333), SOD (X: 55.3195, Y: 44.0557, and Z: 0.0370), GPx (X: 30.9947, Y: 55.0836, and Z: 41.5241), MurB (X: −21.554, Y: 32.9988, and Z: −4.3529), and PBP (X: 44.6994, Y: 1.1436, and Z: 9.0408).
The conformation with the lowest binding affinity resulting from the docking process was selected for further analysis and visualized using Discovery Studio 2020 Client Software. The analysis process provides information on the interactions between ligands and receptors, including hydrogen bond and hydrophobic interactions, the bond length of each interaction, and the amino acids bound to the ligand. To obtain the best visualization, all interactions between ligand and receptor were illustrated as 3D pictures. The pictures present the interactions by displaying the entire illustration of the receptor and the detailed interactions by showing only the ligand surroundings.
Bioactivity Screening Through PASS Online Program
The bioactivity of 3-(3-methoxyphenyl) propanal was analyzed using the PASS online program at http://www.pharmaexpert.ru/passonline/predict.php. 46
Pharmacokinetic and Drug-Likeness Prediction of Compound
The physicochemical properties of a ligand can be determined using Lipinski's rule analysis, which can be accessed at http://www.scfbio-iitd.res.in/software/drugdesign/lipinski.jsp#anchortag. 47 The PDB file of ligand was loaded into the software to obtain the results automatically. ADMET prediction can provide information on oral bioavailability, cell permeability, metabolism, elimination, and toxicity in terms of pharmacokinetic characteristics and pharmacodynamics. ADMET analysis of 3-(3-methoxyphenyl) propanal was performed using online (http://www.swissadme.ch/) 48 and offline (http://www.organic-chemistry.org/prog/peo/) programs. 49
MD Simulation
During MD simulation, 4 processes take place: minimization, heating, equilibration, and production run. The minimizing step is separated into 4 steps. In the first stage, the system was briefly minimized using 250 steepest descent (SD) and continued with 750 conjugate gradient cycles. For the second and third stages, 5000 SD cycles were utilized with a 10 kcal mol−2 Å−2 restraint force on the protein backbone for the second stage, and, for the third stage, constraint force was applied on all system components except water and Na+ ions. SD with a restraint force of 10 kcal mol−2 Å−2 for all system components was carried out for the final stage of minimization. A 45 ps heating step from 0 to 300 K on a constant volume was followed by a 2 ns equilibration step with restraints on at 300 K constant temperature and constant pressure. In this stage, hydrogen-containing bonds of the complex were constrained using the SHAKE algorithm. At 300 K and constant pressure, the production took place in 100 ns. With a cut-off value of 9.0 Å, the particle mesh Ewald approach was utilized to mimic longer-range interactions.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received the following financial support for the research, authorship, and/or publication of this article: This work was supported by RISTEKDIKTI—the Research Grant of Penelitian Dasar Unggulan Perguruan Tinggi 2017-2018.
