Abstract
Background
Lung adenocarcinoma (LUAD) is a dominant tumor with high morbidity and mortality. In spite of innovations in surgery, chemotherapy, and targeted therapies, the prognosis for patients with LUAD remains unsatisfactory. Therefore, it is particularly important to study the biological markers of the occurrence, development, and prognosis of this disease.
Methods
In this study, 3 datasets were selected from the Gene Expression Omnibus (GEO) database to screen differentially expressed genes (DEGs). Then, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses were performed using Database Annotation, Visualization and Integrated Discovery (DAVID). In addition, a protein-protein interaction network was performed and 10 key genes were analyzed. Gene and protein expression were analyzed using The Cancer Genome Atlas (TCGA) and The Human Protein Atlas (HPA) databases, respectively. Then, overall survival (OS) rate was analyzed using Kaplan-Meier plotter database. The chemical components were downloaded from the HERB database. Finally, the chemical constituents were docked with 10 targets by molecular docking. By using in vitro studies, we evaluated the effects of curcumin and trans-resveratrol on the proliferation of LUAD cells.
Results
A total of 122 upregulated and 434 downregulated genes were identified. Then, 10 key genes (glyceraldehyde-3-phosphate dehydrogenase (GAPDH), interleukin-1 beta (IL1B), von Willebrand factor (VWF), matrix metalloproteinase-9 (MMP9), CD44 antigen (CD44), transcription factor Jun (JUN), platelet endothelial cell adhesion molecule (PECAM1), hematopoietic progenitor cell antigen CD34 (CD34), peroxisome proliferator-activated receptor gamma (PPARG), and collagen alpha-1(I) chain (COL1A1)) were obtained. TCGA results showed that GAPDH, MMP9, and COL1A1 were upregulated, while IL-1B, VWF, CD44, JUN, PECAM1, CD34, and PPARG were downregulated in cancer tissues. Kaplan-Meier plotter analysis also showed that high expression of GAPDH, PPARG, and COL1A1, and low expression of VWF, PECAM1, CD34, PPARG, and COL1A1 were associated with poor prognosis in LUAD. Combined with gene expression, staging, immunohistochemistry, and OS prognostic analysis, the results showed that COL1A1 was a potential target for the occurrence and prognosis of LUAD. In addition, 2 key components (curcumin and trans-resveratrol) had good docking effect with COL1A1. We found that curcumin and trans-resveratrol significantly inhibited the proliferation of LUAD cells and significantly decreased the expression of COL1A1 and MMP9.
Conclusion
In conclusion, our study identified the potential target of LUAD and the main traditional Chinese medicine components (curcumin and trans-resveratrol) for LUAD treatment. This study provides potential therapeutic targets and treatment strategies for the treatment of LUAD.
Introduction
Lung adenocarcinoma (LUAD) is the dominant tumor in all lung cancer pathologic types, with high morbidity and mortality. Despite great advances in the treatment of LUAD due to innovations in surgery, chemotherapy, and targeted therapies, the prognosis for patients with LUAD remains unsatisfactory. 1 In recent years, microarray technology and bioinformatics tools have been used to identify molecular markers related to the occurrence, drug resistance, and prognosis of lung cancer. 2 Therefore, it is particularly important to study the biological markers of the occurrence, development, and prognosis of LUAD.
Curcumin is a diarylheptanoid derived from turmeric, which has tumor prevention and anti-cancer activity. 3 It has been reported that the anti-cancer effect is caused by inhibiting PI3K/AKT/mTOR, nuclear factor kappa B (NF-kB), and STAT3, but the molecular mechanisms involved remain unclear. 4
Trans-resveratrol, a stilbene plant antitoxin, is found in high concentrations in grapes and red wine. Trans-resveratrol has several beneficial functions, including anti-cancer properties. 5 However, the molecular mechanisms remain unclear.
In the present study, we used bioinformatics and molecular docking to explore systematically the diagnosis and traditional Chinese medicine (TCM) treatment of LUAD by analyzing active ingredients, potential target genes, and key pathways.
Materials and Methods
Identification of DEGs in GEO Database
Three genes expression profile microarrays (GSE19188, GSE32863, and GSE116959) were obtained from the Gene Expression Omnibus 6 database (GEO, https://www.ncbi.nlm.nih.gov/geo/). The array data of GSE19188 7 corresponded to 91 LUAD tissues and 65 corresponding para-cancer tissues on the GPL570 platform. The array data of GSE32863 8 corresponded to 58 LUAD tissues and 58 corresponding para-cancer tissues on the GPL6884 platform. The array data of GSE116959 9 corresponded to 57 LUAD tissues and 11 corresponding para-cancer tissues on the GPL17077 platform. Then, the differentially expressed genes (DEGs) were identified by GEO2R (https://www.ncbi.nlm.nih.gov/geo/geo2r/) with criteria of |log FC| > 1.0 and adjust-P < .05. Common genes were obtained using the Venn 2.1.0 tool (https://bioinfogp.cnb.csic.es/tools/venny/index.html).
GO and KEGG Analysis of DEGs
To explore further the characteristics of DEGs, the intersection DEGs were imported by using the Database for Annotation, Visualization and Integrated Discovery (DAVID) 10 (https://david.ncifcrf.gov/). Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed for overlapping DEGs. The bubble map of DEGs was displayed with the online tool 11 (bioinformatics, http://www.bioinformatics.com.cn/). P ≤ .01 and counts ≥ 10 were considered statistically significant.
PPI Analysis
The identified intersection DEGs were imported into STRING (version 11.5, https://cn.string-db.org/) 12 database. Cytoscape software (version 3.8.2) was used to visualize the protein-protein interaction (PPI) network; the high or low expression of genes was marked with red or blue. The degree of each protein node was calculated using Cytohubba, a plug-in called Cytoscape software. The 10 key genes with high degree values were selected.
Verification of GEO Results by TCGA Database
The RNA sequencing fragments per kilobase million data and the corresponding clinical information were downloaded by using The Cancer Genome Atlas 13 (TCGA, https://portal.gdc.cancer.gov/)-LUAD database. A total of 594 samples were collected in LUAD-TCGA, including 59 normal samples and 537 tumor samples. Gene expression and stage were analyzed by using Graph pad prism 8 software.
Protein Expression Level of the key Genes in LUAD
The Human Protein Atlas 14 (HPA, https://www.proteinatlas.org/) is an immunohistochemical (IHC) database based on cancer tissue, normal tissue, and cell protein expression. In this study, the IHC images of 10 key genes of LUAD patients were obtained from this database.
Survival Analysis of the key Genes
From the Kaplan-Meier Plotter 15 (https://kmplot.com/analysis/) database, which provides information on the relationships of more than 54 000 genes (mRNAs, microRNAs, and proteins) with survival rates in 21 cancer types, the top 10 key genes were mapped with the overall survival (OS) of LUAD patients. The criteria were as follows: hazard ratio (HR) within the 95% confidence interval (CI) and log-rank P-value < .05.
Acquisition of TCM Ingredients
HERB 16 (http://herb.ac.cn/) is a database which gathers and reanalyzes 1037 high throughput sequencing experiments. In this study, we reversely enriched TCM components through key targets, and constructed a target-components PPI network.
Molecular Docking
The target genes were downloaded from the PDB 17 database (https://www.rcsb.org/). Then, the 3D structure of curcumin and trans-resveratrol were downloaded from the PubChem 18 database (https://pubchem.ncbi.nlm.nih.gov/). The structure of the protein was dehydrated and hydrogenated by using Auto Dock Tools. Molecular docking verification was performed using Auto Dock Vina software. Finally, PyMoL software was used for visualization.
Cell Line and Drug Treatment
The human LUAD cell line A549 was purchased from Procell (China). The cells were cultured in Ham's F-12K (Procell, China) containing 10% fetal bovine serum (Vica cell, China), 100 U/mL penicillin and 100 μg/mL streptomycin and placed in a humidified incubator at 37 °C in an atmosphere of 100% humidity and 5% CO2.
Curcumin and trans-resveratrol were purchased from TargetMOl (China). Curcumin (Cat. #149135) and trans-resveratrol (Cat. #158951) were dissolved in DMSO (Sigma, USA) and stored at −20 °C. A549 cells were treated with curcumin (75.2 μM) and trans-resveratrol (156.7 μM) for 48 h, respectively.
Cell Viability
A549 cells were seeded in 96-well plates (1 × 104 cells/well) and treated with either curcumin (25, 50, 75, and 100 μM) or trans-resveratrol (50, 100, 150, and 200 μM) at different concentrations for 24 and 48 h, respectively. According to the manufacturer's protocol, a cell counting kit-8 (CCK-8, Cat. #21169949, China) was used to determine cell viability. Then, optical density (OD) was monitored at 450 nm with 650 nm as a reference wavelength using a Spectra Max PLUS 384 (Molecular Devices, USA). Then, the cell viability values were calculated and IC50 values were obtained by using Softmax Pro Software.
Western Blot Analysis
A549 cells were seeded in 6-well plates (1 × 105 cells/well) and treated as described above. In brief, they were washed in PBS, and the cells were directly lysed in a SDS sample buffer. Protein concentration was determined by a BCA method (P0009, Beyotime Biotechnology.). The primary antibodies were rabbit anti-COL1A1 polyclonal antibody (Cat. #CY5120, Abways, diluted to 1:2000), anti-MMP9 polyclonal antibody (Cat. #CY5205, Abways, diluted to 1:2000), and β-actin antibody (Cat. # AC026, abclonal, diluted to 1:50000). The secondary antibodies were goat anti-rabbit (S0001, Affinity, diluted by 1:5000) and goat anti-mouse (AS003, abclonal, diluted by 1:5000). The results were analyzed using SPSS 17.0 software (SPSS, Chicago, IL).
Statistical Analysis
Using GEO2R, genes with an adjusted P-value < .05 were regarded as statistically significant. The GO and KEGG pathways were considered significantly with P < .01. For the Kaplan-Meier plotter analysis, a P-value < .05 was regarded as statistically significant. All statistical tests were performed by SPSS 17.0 statistics software (SPSS, Chicago, IL). All in-vitro experiments were repeated at least 3 times and at least 3 samples were taken at a time.
Results
Identification of DEGs in Lung Adenocarcinoma and Analysis of Key Genes
A total of 828, 514, and 752 upregulated genes were respectively identified in GSE19188, GSE32863, and GSE116959, and 1696, 756, and 1506 downregulated genes were identified in GSE19188, GSE32863, and GSE116959, respectively according to GEO2R. Venn diagrams were used to examine the intersection among the DEGs, including 122 upregulated genes and 434 downregulated genes (Figure 1A and B).

Identification of DEGs. (A) Venn diagram of downregulated DEGs based on the 3 datasets. (B) Venn diagram of upregulated DEGs based on the 3 datasets. (C) Identification of key genes from the DEGs by Cytoscape Red dots represents upregulated genes, and blue dots represent downregulated genes. (D) Identification of 10 key genes by CytoHubba.
Subsequently, to clarify further the interaction between targets, the PPI network, with 514 nodes and 2819 edges, was constructed by STRING database and was visualized by Cytoscape software. To identify better the key targets, 10 key genes with the highest degree were identified by CytoHubba. These were GAPDH, IL1B, VWF, MMP9, CD44, JUN, PECAM1, CD34, PPARG, and COL1A1 (Figure 1C and D).
GO and KEGG Analysis
To understand the possible function and pathway of these DEGs, we explored the influence of DEGs on the function and pathways of genes by GO and KEGG enrichment analysis in DAVID database. The GO function analysis of these genes was conducted into 3 parts, biological process (BP), molecular function (MF), and cellular component (CC). In terms of BP, these DEGs were mostly enriched in “signal transduction,” “cell adhesion,” “cell differentiation,” “apoptotic process,” “positive regulation of gene expression,” “negative regulation of apoptotic process,” “negative regulation of cell proliferation,” “innate immune response,” “inflammatory response,” and “response to drug.” In terms of CC, these DEGs were mostly enriched in “plasma membrane,” “integral component of membrane,” “cytoplasm,” “extracellular region,” “extracellular space,” “extracellular exosome,” “integral component of plasma,” “membrane,” “cell surface,” “Golgi apparatus,” and “external side of plasma membrane.” In terms of MF, these DEGs were mostly enriched in “protein binding,” “identical protein binding,” “calcium ion binding,” “protein homodimerization activity,” “extracellular matrix structural constituent,” “integrin binding,” “RNA polymerase II sequence-specific DNA binding transcription factor binding,” “heparin binding,” “carbohydrate binding,” and “beta-amyloid binding” (Figure 2A).
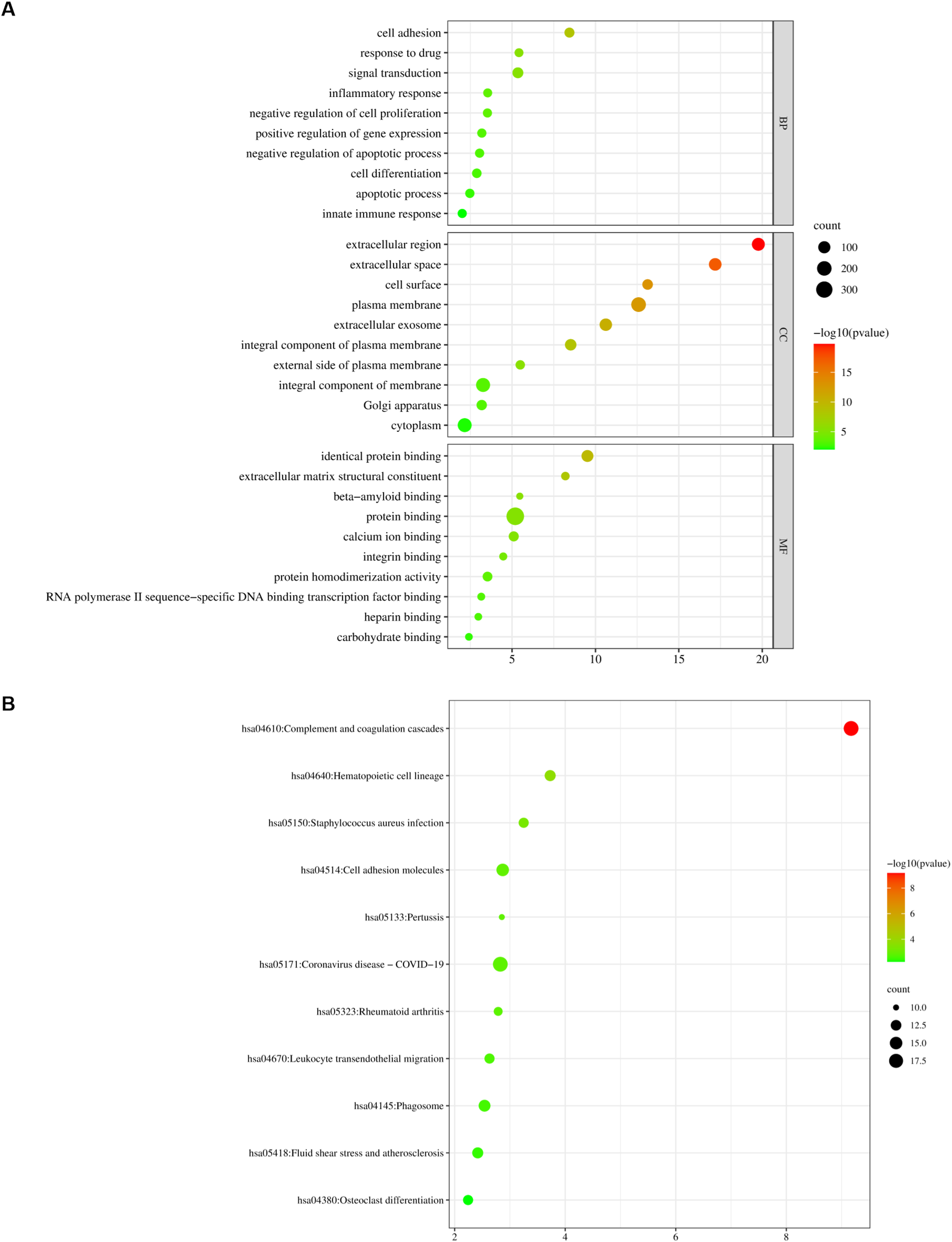
GO and KEGG pathway analysis for key genes. (A) GO analysis for key genes. (B) KEGG pathway analysis for key genes.
The KEGG pathway was changed in “hsa04610: Complement and coagulation cascades,” “hsa05171: Coronavirus disease - COVID-19,” “hsa04514: Cell adhesion molecules,” “hsa04145: Phagosome,” “hsa04640: Hematopoietic cell lineage,” “hsa05418: Fluid shear stress and atherosclerosis,” “hsa05150: Staphylococcus aureus infection,” “hsa04670: Leukocyte trans endothelial migration,” “hsa04380: Osteoclast differentiation,” “hsa05323: Rheumatoid arthritis,” and “hsa05133: Pertussis”. These results showed that these signaling pathway and cell cycle played an important role in LUAD.
Validation of Key Genes in TCGA Database
To verify the reliability of the DEGs in the GEO database, TCGA clinical patient specimens were downloaded. The result indicated that GAPDH, MMP9, and COL1A1 were upregulated, while IL-1B, VWF, CD44, JUN, PECAM1, CD34, and PPARG were downregulated in cancer tissues (Figure 3A). Next, we analyzed the correlation between 10 key genes expression and tumor stage in LUAD. The result showed that the GAPDH, MMP9, and COL1A1 levels gradually increased, but IL-1B, VWF, CD44, JUN, PECAM1, CD34, and PPARG were decreased with stage change. Moreover, the expression of COL1A1 was more significantly affected by stage (Figure 3B).

Transcriptional levels and staging of 10 key genes in LUAD and normal tissues. (A) The transcription levels of 10 key genes in LUAD and normal tissues. (B) The staging of 10 key genes in LUAD and normal tissues.
HPA Database Analysis
Then, to confirm further the importance of 10 key genes in LUAD, we analyzed the levels of protein expressed in the HPA database. As demonstrated in Figure 4, IL-1B was not found, but GAPDH and COL1A1 were highly expressed, while VWF, CD44, JUN, and CD34 were downregulated in LUAD tissues. However, the change in MMP9, PECAM1, and PPARG expression was not obvious (Figure 4). These results showed that these genes played a vital role in LUAD development.

Immunohistochemistry images of (A) GAPDH, (B) VWF, (C) MMP9, (D) CD44, (E) JUN, (F) PECAM1, (G) CD34, (H) PPARG, (I) COL1A1 in LUAD and non-cancerous lung tissues by using HPA database.
Prognostic Analysis
To illustrate the OS time of these genes in liver cancer, the KM plotter was used to draw the survival curves of LUAD patients. The result indicated that high GAPDH, PPARG, and COL1A1 expression and low VWF, PECAM1, CD34, PPARG, and COL1A1 expression were related to poor OS time in LUAD (log-rank P < .05) (Figure 5). These results indicated that GAPDH, PPARG, COL1A1, VWF, PECAM1, CD34, PPARG, and COL1A1 were associated with OS time. COL1A1 gene expression, staging, immunohistochemistry, and OS prognostic analysis suggested that COL1A1 was a potential target for the occurrence and prognosis of LUAD.

OS analysis of 10 key genes in LUAD patients by Kaplan–Meier plotter. (A) GAPDH, (B) IL1B, (C) VWF, (D) MMP9, (E) CD44, (F) JUN, (G) PECAM1, (H) CD34, (I) PPARG, (J) COL1A1.
The Acquisition of Chinese Herb Components
We used HERB database to reverse enrich related TCMs and components. A total of 1969 chemical components were screened (Table S1). Target-component PPI was conducted, including 1979 nodes and 2299 edges (Figure 6). Among these components, the degree of HBIN021985 (curcumin) and HBIN046831 (trans-resveratrol) were high (HBIN021985 = 8, HBIN046831 = 7), indicating that they might be the main anti-cancer components (Table 1).

Target-composition PPI network. Blue rectangles are the TCM ingredient and the red rectangles are the key target.
Top 10 Targets in the PPI Network.

Verification of Molecular Docking
Molecular docking could further verify the relationship between key targets and chemical components. The results showed that the key targets and components had good docking results (Table 2) (Figure S1-S2). Among these, the binding energies of curcumin and COL1A1 (−7.5 kcal/mol), trans-resveratrol and MMP9 (−9.5 kcal/mol), trans-resveratrol and CD44 (−8.5 kcal/mol), and trans-resveratrol and COL1A1 (−8.5 kcal/mol) were all less than −7.5 kcal/mol, having good docking results (Figure 7A). As shown in Figure 7B, the result showed that curcumin formed 3 hydrogen bonds with COL1A1 at amino acid residues ALA-256, PHE-255, and ARG-254. Trans-resveratrol formed 4 hydrogen bonds with COL1A1 at amino acid residues ASN-384, ASN-332, ASP-397, and GLU-290. Trans-resveratrol formed 4 hydrogen bonds with CD44 at amino acid residues SER-112, ASN-94, GLY-40, and HIS-35, and 2 hydrogen bonds with MMP9 at amino acid residues LEU-188 and ALA-189 (Figure 7B). These studies further confirmed the efficacy of these 2 compounds in the treatment of LUAD by targeting these key targets.

Molecular docking of the 2 key chemical composition with the hub targets. (A) The molecular docking of curcumin with COL1A1. (B) The molecular docking of trans-resveratrol with COL1A1. (C) The molecular docking of trans-resveratrol with CD44. (D) The molecular docking of trans-resveratrol with MMP9.
Docking Results of Target Protein and Active Compound (kcal/mol).

Effects of Curcumin and Trans-Resveratrol on Cellular Viability
A549 cells were treated with curcumin and trans-resveratrol for 24 and 48 h to observe the effect of curcumin and trans-resveratrol on cell viability. The results showed that cell viability was correlated with the intervention time and concentration of curcumin and resveratrol (Figure 8A). The IC50 values of curcumin and trans-resveratrol in A549 cells were 75.2 and 156.7 μM at 48 h, respectively (Figure 8B). Therefore, these concentrations were selected as the intervention concentrations of curcumin and resveratrol in subsequent experiments.
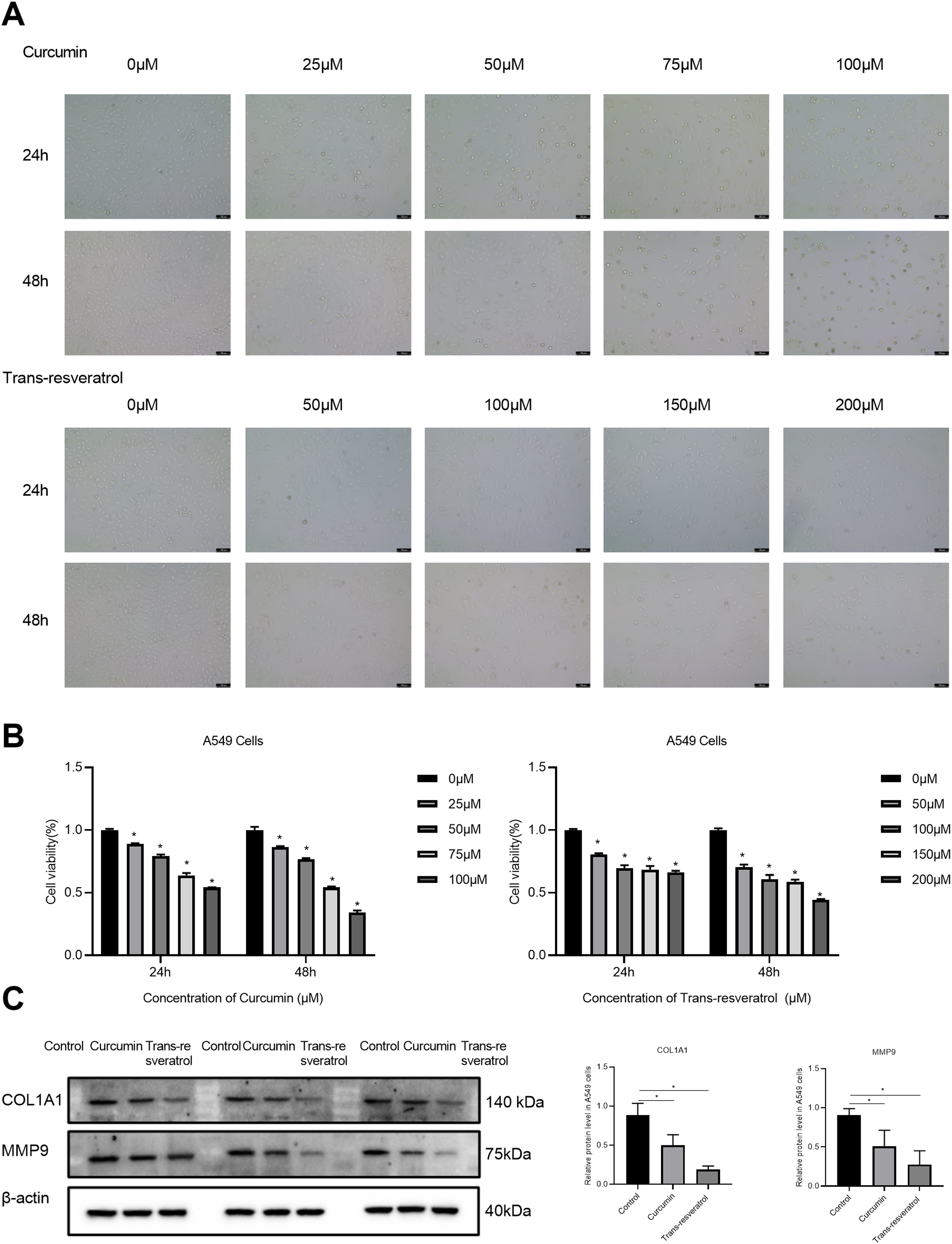
Effects of curcumin and trans-resveratrol on COL1A1 and MMP9 in A549 cells. (A) The decreased cell number was respectively detected at 24 and 48 h after the treatment with different concentrations of curcumin and trans-resveratrol. The infected cells were observed by phase contrast microscope. All the experiments were repeated at least 3 times. (B) CCK8 was used to detect the inhibitory effect of curcumin and trans-resveratrol on the proliferation of A549 cells. (C) Curcumin and trans-resveratrol reduced the protein expression of COL1A1 and MMP9 in A549 cells.
Curcumin and Trans-Resveratrol Decreased the Expression of COL1A1 and MMP9
We selected 2 key targets (COL1A1 and MMP9) for further experimental verification. The results showed that the expressions of COL1A1 and MMP9 in LUAD cells in the different concentration groups were significantly different from those in the control group. The results showed that curcumin and trans-resveratrol could induce down-regulation of COL1A1 and MMP9 expression (Figure 8C).
Discussion
Recently, biomarkers for the diagnosis or treatment of cancer played an important role in the treatment of LUAD, which has attracted more and more attention. 19 Like other cancers, the occurrence, progression, and metastasis of lung cancer is a very complex process involving the aberration of multiple genes and pathways. 20
In this study, we identified 556 DGEs, including 122 upregulated and 434 downregulated genes. Then, it was found through GO and KEGG enrichment analysis that these genes were mainly involved in cell cycle function and pathways. We also screened out 10 key genes, including glyceraldehyde-3-phosphate dehydrogenase (GAPDH), interleukin-1 beta (IL1B), von Willebrand factor (VWF), matrix metalloproteinase-9 (MMP9), CD44 antigen (CD44), transcription factor Jun (JUN), platelet endothelial cell adhesion molecule (PECAM1), hematopoietic progenitor cell antigen CD34 (CD34), peroxisome proliferator-activated receptor gamma (PPARG), and collagen alpha-1(I) chain (COL1A1). Besides, a PPI network was analyzed for the interaction between the DGEs. The TCGA and Kaplan-Meier plotter databases were used to analyze the expression, staging, and survival of 10 key genes. In addition, Chinese medicine ingredients were screened using HERB database. Finally, the key target was verified by molecular docking with TCM components.
COL1A1 is one of the most abundant proteins in the human body. In addition, COL1A1 promotes embryonic stem cell proliferation, which is related to the response to hypoxia and oxidative stress, promoting tumor cell growth and metastasis. 21 Similarly, a study also indicated that in non-SCLC cell lines, COL1A1 was involved in the induction of epithelial-mesenchymal transition (EMT) through the transforming growth factor (TGF)-β-dependent pathway. 22 MMP9 is a type IV collagenase and high expression of MMP9 promotes the progression of lung cancer. 23 Studies showed that the expression of MMP-9 was higher in lung cancer patients. The serum level of MMP-9 in the experimental group was closely related to the TNM stage, differentiation degree, lymph node metastasis, smoking history, and tissue form. 24 Moreover, external carcinogens induced MMP9 production and EMT progression by activating the Shp2/ERK1/2/JNK/Smad2/3 signaling pathways. 25 In addition, with certain diseases or age, MMPs could be a characteristic of cancer. 26 CD44 was a non-kinase transmembrane glycoprotein that promoted cancer cell proliferation and invasion. 27 The expression level of CD44 was down-regulated in LUAD tissues, and the low CD44 expression group had a poor prognosis. 28 Similarly, we found that GAPDH, MMP9, and COL1A1 were upregulated, while IL-1B, VWF, CD44, JUN, PECAM1, CD34, and PPARG were downregulated in cancer tissues, and high expression of GAPDH, PPARG, COL1A1, and low expression of VWF, PECAM1, CD34, PPARG, COL1A1 had a poor prognosis in LUAD.
Our molecular analysis results showed that the binding energies of the 2 active ingredients and 10 target proteins were all ≤ −5 kcal/mol, indicating that the above active ingredients had a good binding effect on the target. Curcumin is a main bioactive component of the Indian spice turmeric. Curcumin displays a vast range of pharmacological properties, including antitumor, anti-inflammatory, and antioxidant activity. 29 Studies showed that curcumin reduced the expression of COL1A1, MMP-9, and CD44, inhibiting proliferation, migration, and invasion of lung cancer cells.30–32 In addition, curcumin inhibited the increase of GAPDH expression, played an anti-bacterial role, 33 and counteracted the high glucose-induced drug resistance of hepatocellular cancer cells. 34 Additionally, curcumin inhibited hepatocyte growth factor (HGF) promoted CD34, EMT expression 35 and inhibited tobacco smoke-induced MAPK/AP-1 activation, including JUN proteins, inhibiting lung cancer progression. 36 Curcumin also played an anti-inflammatory role by either activating PPARG 37 or inhibiting the expression of IL1B, 38 alleviating lipopolysaccharide-induced acute lung injury. Curcumin improved hepatic fibrosis induced by carbon tetrachloride in rats by inhibiting various angiogenic factors, including VWF and VEGFR. 39 In addition, curcumin reduced the expression of PECAM-1 regulated by NF-κB, inhibiting the growth of breast cancer. 40
Trans-resveratrol is a naturally occurring polyphenol, found mainly in red wine and peanuts. In recent years, trans-resveratrol has demonstrated anti-tumor, anti-oxidation, anti-inflammatory, and other effects. 41 Studies indicated that trans-resveratrol attenuated inflammation and oxidative stress by reducing the expression of IL1B, 42 GAPDH, 43 and VWF. 44 Trans-resveratrol also inhibited the metastasis of LUAD by inhibiting NF-κB pathway mediated by heme oxygenase-1 (HO-1) and down-regulating MMP9 expression. 45 In addition, trans-resveratrol derivatives inhibited the expression of JUN and reduced the invasion of lung cancer. 46 Studies also showed that trans-resveratrol inhibited the expression of PECAM-1 and reduced the adhesion of inflammatory cells to endothelial cells, which reduced the severity of acute lung injury complicated with severe acute pancreatitis. 47 Furthermore, trans-resveratrol effectively improved liver fibrosis by decreasing COL1A1 and PPARG expression. 48 Moreover, trans-resveratrol reduced the expression of CD34 49 and CD44, 50 inhibiting tumor formation. In this study, we found that curcumin and trans-resveratrol inhibited the proliferation of A549 cells and reduced COL1A1 and MMP9 expression.
Unfortunately, the study may have some limitations. There were no clinical lung cancer patients available. The specific mechanism of key gene involvement in lung cancer remains unclear. Therefore, further clinical studies are needed to verify the reliability of the results in order to identify the genes most closely related to the pathogenesis of lung cancer and provide new therapies for lung cancer.
Conclusion
In summary, the key targets and TCM components of LUAD were studied based on bioinformatics combined with network pharmacology. In addition, this study demonstrated that 10 key genes, including GAPDH, IL1B, VWF, MMP9, CD44, JUN, PECAM1, CD34, PPARG, and COL1A1, and 2 key components, curcumin and trans-resveratrol, were screened. We revealed that curcumin and trans-resveratrol played a critical role in LUAD by affecting GAPDH, IL1B, VWF, MMP9, CD44, JUN, PECAM1, CD34, PPARG, and COL1A1. Besides, our molecular docking studies also showed that the main active components (curcumin and trans-resveratrol) could dock well with GAPDH, IL1B, VWF, MMP9, CD44, JUN, PECAM1, CD34, PPARG, and COL1A1, providing an important basis for further investigation. However, this study also has some limitations, and further pharmacological and clinical studies are needed to verify our results.
Supplemental Material
sj-tif-1-npx-10.1177_1934578X231169370 - Supplemental material for A Systematic Study of Traditional Chinese Medicine for the Treatment of Lung Adenocarcinoma Using a Reverse Network of Key Targets Based on Bioinformatics and Molecular Docking: Curcumin and Trans-Resveratrol as Potential Drug Candidates for Lung Adenocarcinoma
Supplemental material, sj-tif-1-npx-10.1177_1934578X231169370 for A Systematic Study of Traditional Chinese Medicine for the Treatment of Lung Adenocarcinoma Using a Reverse Network of Key Targets Based on Bioinformatics and Molecular Docking: Curcumin and Trans-Resveratrol as Potential Drug Candidates for Lung Adenocarcinoma by Wujun Wang, Zhu Zeng, Junli Zhang and Zengtao Sun in Natural Product Communications
Supplemental Material
sj-tif-2-npx-10.1177_1934578X231169370 - Supplemental material for A Systematic Study of Traditional Chinese Medicine for the Treatment of Lung Adenocarcinoma Using a Reverse Network of Key Targets Based on Bioinformatics and Molecular Docking: Curcumin and Trans-Resveratrol as Potential Drug Candidates for Lung Adenocarcinoma
Supplemental material, sj-tif-2-npx-10.1177_1934578X231169370 for A Systematic Study of Traditional Chinese Medicine for the Treatment of Lung Adenocarcinoma Using a Reverse Network of Key Targets Based on Bioinformatics and Molecular Docking: Curcumin and Trans-Resveratrol as Potential Drug Candidates for Lung Adenocarcinoma by Wujun Wang, Zhu Zeng, Junli Zhang and Zengtao Sun in Natural Product Communications
Supplemental Material
sj-docx-3-npx-10.1177_1934578X231169370 - Supplemental material for A Systematic Study of Traditional Chinese Medicine for the Treatment of Lung Adenocarcinoma Using a Reverse Network of Key Targets Based on Bioinformatics and Molecular Docking: Curcumin and Trans-Resveratrol as Potential Drug Candidates for Lung Adenocarcinoma
Supplemental material, sj-docx-3-npx-10.1177_1934578X231169370 for A Systematic Study of Traditional Chinese Medicine for the Treatment of Lung Adenocarcinoma Using a Reverse Network of Key Targets Based on Bioinformatics and Molecular Docking: Curcumin and Trans-Resveratrol as Potential Drug Candidates for Lung Adenocarcinoma by Wujun Wang, Zhu Zeng, Junli Zhang and Zengtao Sun in Natural Product Communications
Footnotes
Availability of Data and Materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Author Contributions
Conceptualization: Zengtao Sun. Data curation: Wujun Wang, Zhu Zeng. Methodology: Wujun Wang, Zhu Zeng. Visualization: Junli Zhang. Writing-original draft: Wujun Wang, Zhu Zeng, Junli Zhang, Zengtao Sun. All authors read and approved the initial manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Not applicable.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Statement of Human and Animal Rights
Not applicable.
Statement of Informed Consent
Not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
