Abstract
Introduction
Sargassum is a genus of brown algae, commonly distributed worldwide.
1
Sargassum micracanthum (Kutzing) Endlicher is a member of the family Sargassaceae, which comprises about 193 species.
2
The species has a coriaceous thallus up to 50 cm in length and grows on rocks in the low intertidal to upper subtidal zones of open coasts.
3
Previous studies have reported that Sargassum species contain compounds with various biological activities, such as anti-oxidative, cholinesterase inhibitory, neuroprotective, anti-cancer, anti-inflammatory, and anti-viral activities.4-7 The compounds include chromanes, chromenes, meroterpenoids, farnesylactones, and plastoquinones.8-11 Chemical investigation of a methanol extract of S. micracanthum led to the isolation of one novel butenolide (

Structures of compounds
Results and Discussion
Compound
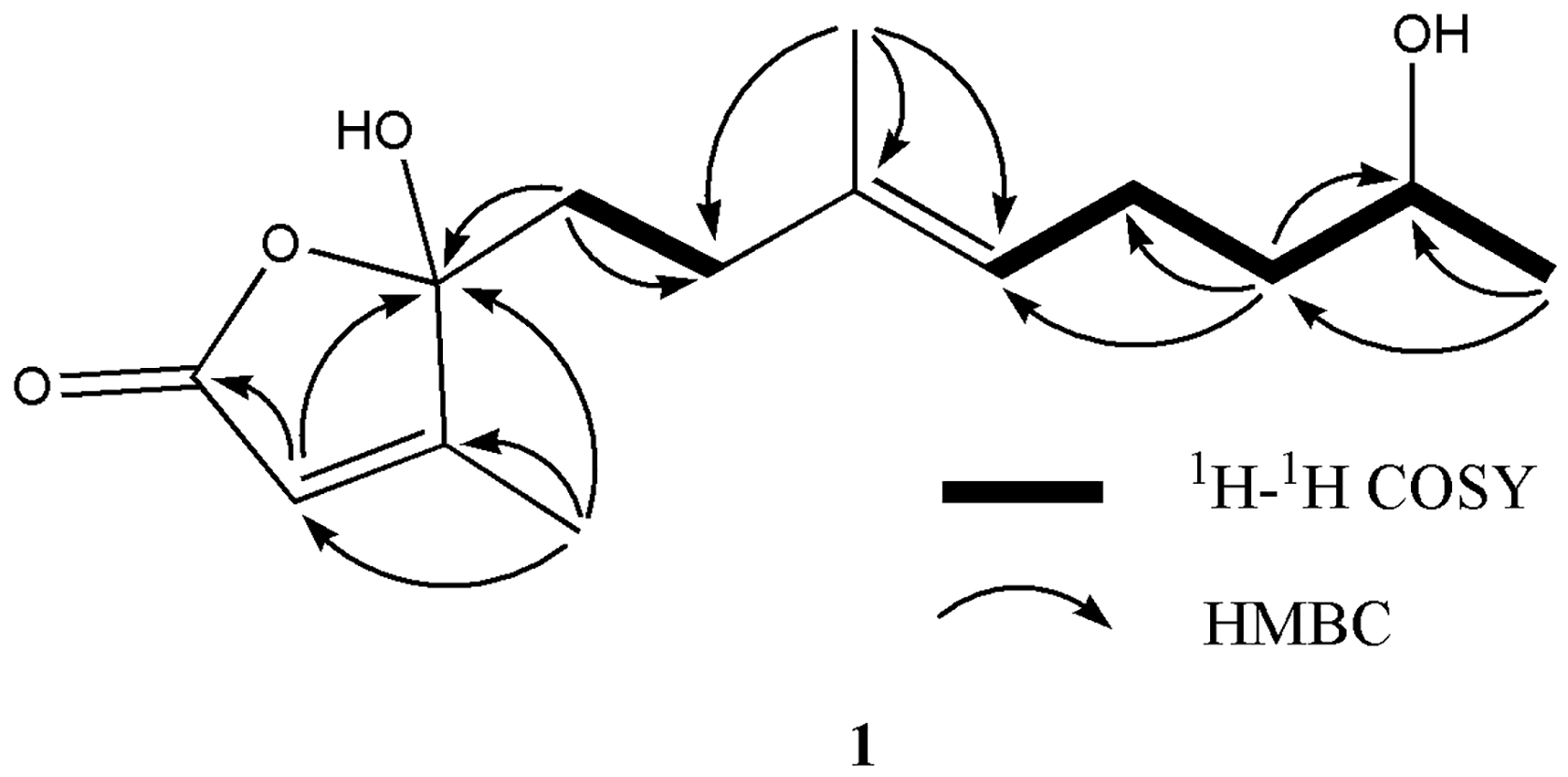
HMBC and 1H-1H COSY correlations of compound
1H- and 13C-NMR Spectroscopic Data of

Compound
To investigate the anti-inflammatory activities of compounds
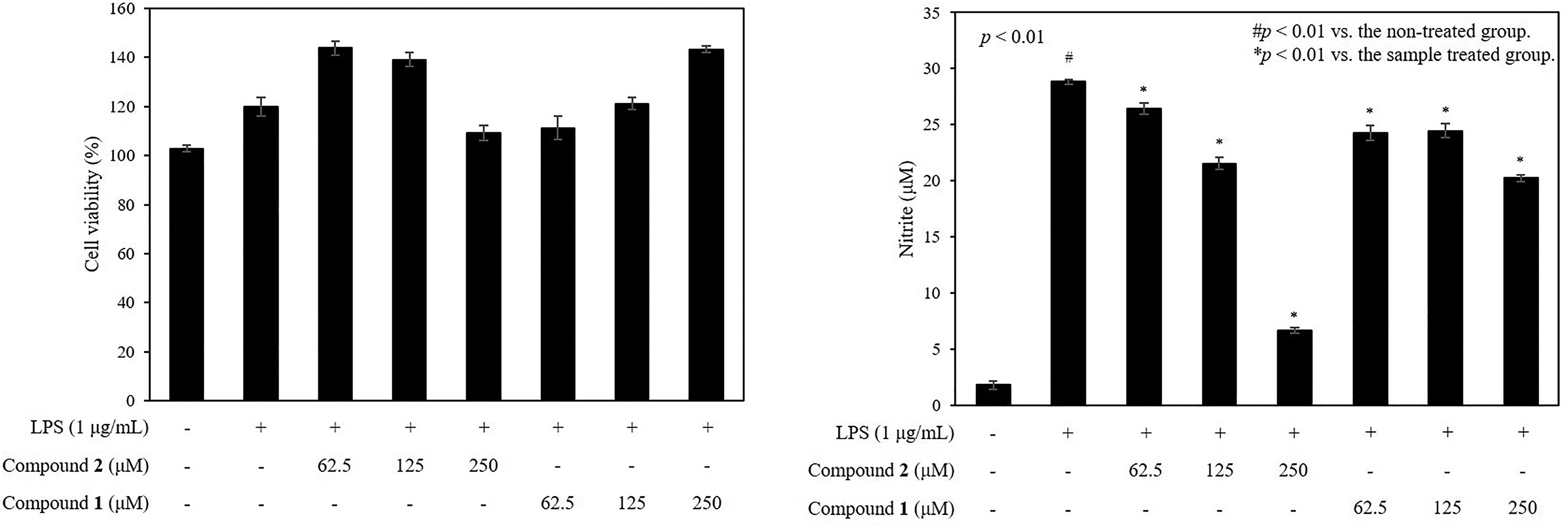
The inhibitory effects of compounds

The effect of compounds
Conclusions
A new butenolide, sargassumin C (
Sargassumin C (1)
Colorless oil
IR (film) νmax : 3353, 2920, 2849, 1738, 1644, 1450, 1114, 1015 cm−1.
UV (MeOH) : λmax (log ε) 213 (3.5) nm.
HRESIMS
m/z 253.1448 [M-H]− (Calcd for C14H21O4 [M-H]−, 253.1440, Δ = + 0.8 mmu)
1H (500 MHz) and 13C NMR (125 MHz) spectra are shown in Table 1.
2-hydroxy-(5E,9E)-6,10,14-trimethylpentadeca-5,9-dien-12-one (2)
Colorless oil
HRESIMS
m/z 303.2294 [M+H]+ (Calcd for C18H33O2 [M+H]+, 303.2300, Δ = - 0.6 mmu)
1H NMR (500 MHz, CDCl3) : δ 5.20 (1H, t, J = 6.4 Hz, H-9), 5.13 (1H, t, J = 6.4 Hz, H-5), 3.78 (1H, m, H-2), 2.98 (2H, s, H-11), 2.26 (1H, d, J = 6.8 Hz, H-13), 2.11 (2H, m, H-4), 2.09 (1H, m, H-14), 2.06 (2H, m, H-8), 2.00 (2H, m, H-7), 1.60 (3H, s, 6-CH3), 1.58 (3H, s, 10-CH3), 1.48 (2H, m, H-3), 1.17 (3H, d, J = 6.1 Hz, H-1), and 0.87 (6H, d, J = 6.4 Hz, 14-CH3 and 15)
13C NMR (125 MHz, CDCl3) : δ 209.8 (C-12), 135.4 (C-6), 129.6 (C-9), 129.2 (C-10), 124.4 (C-5), 68.1 (C-2), 54.6 (C-11), 50.7 (C-13), 39.3 (C-3 and C-7), 26.8 (C-8), 24.6 (C-14), 24.5 (C-4), 23.6 (C-1), 22.7 (C-15 and 14-CH3), 16.6 (10-CH3), and 16.1 (6-CH3).
Experimental
General Experimental Procedures
NMR spectra were recorded in CDCl3 using a JNM-ECZ500R FT-NMR spectrometer (JEOL, Tokyo, Japan). Optical rotation was measured on a JASCO P-2000 Digital Polarimeter (Jasco, Tokyo, Japan). The CD spectrum was recorded on a JASCO J-715 spectropolarimeter. The UV spectrum was measured on a Thermo Scientific multiskan GO spectrophotometer (Thermo Fisher Scientific, Massachusetts, US), and the IR spectrum on a JASCO FT-IR 4100 spectrometer (Jasco, Tokyo, Japan). For medium-pressure liquid chromatography (Buchi, Flawil, Switzerland), a C18 column was used (40 g, Biotage, Uppsala, Sweden), and for high performance liquid chromatography (Waters Acquity Arc, Waters, Milford, US) a C18 column (4.6 μm × 250 mm and 10.0 μm × 250 mm, Cosmosil, Kyoto, Japan) was used for the isolation process. HRESIMS was performed with a SCIEX X500R Q-TOF LC-MS/MS spectrometer (SCIEX, Massachusetts, US).
Plant Material
Sargassum micracanthum was collected from Udo-myeon, Jeju Island, Korea, in June 2018 and identified by Dr Jeong Chan Kang (Research institute for Basic Sciences, Jeju National University). A voucher specimen (MABIK AL00079431) was deposited at the National Marine Biodiversity Institute of Korea (MABIK).
Extraction and Isolation
Dried whole plants (450 g) of Sargassum micracanthum were ground and extracted with MeOH (2 L × 3) at room temperature for 1 day. The MeOH extract was partitioned with n-hexane, chloroform (CHCl3), ethyl acetate (EtOAc), and n-butanol, respectively. The CHCl3-soluble portion was mixed with the EtOAc-soluble portion due to their similar HPLC profiles. The combined extract was subjected to medium-pressure liquid chromatography (MPLC) eluting with ACN/H2O (10:90 to 100:0) to obtain sub-fractions (1-6). Sub-fraction 1 was further separated by MPLC, followed by semi-HPLC eluting with 30% ACN in water to give compound
NO and ELISA Assay
The production of NO and pro-inflammatory cytokines such as IL-6, IL-1β, TNF-α, and IL-10 was assessed using Griess reagent (Thermo Fisher, Massachusetts, US) and an ELISA kit (Abcam, Cambridge, UK), according to the manufacturer's instructions. Briefly, RAW264.7 cells were seeded onto either 24-well plates (1.5 × 105 cells/well) or 6-well plates (6 × 105 cells/well), and incubated for 18 h at 37 °C. After incubation, the cells were treated with various concentration of compounds
Preparation of (S)- and (R)-MTPA Esters (2a and 2b) of 2
Compound
(S)-MTPA ester (
Colorless oil
HRESIMS
m/z 519.2690 [M+Na]+ (Calcd for C28H39F3O4Na [M+Na]+, 519.2698, Δ = −0.8 mmu)
1H NMR (500 MHz, CDCl3)
δ 5.19 (1H, t, J = 6.8 Hz, H-9), 5.12 (1H, m, H-5), 5.02 (1H, m, H-2), 2.99 (2H, s, H-11), 2.26 (2 h, d, J = 6.8 Hz, H-13), 2.10 (2H, m, H-4), 2.09 (2H, m, H-14), 1.97 (2H, m, H-7), 1.91 (2H, m, H-8), 1.67 (2H, m H-3), 1.58 (3H, s, 10-CH3), 1.43 (3H, s, 6-CH3), 1.32 (3H, s, H-1), 0.87 (6H, d, J = 6.8 Hz, 14-CH3 and 15).
(R)-MTPA ester (
Colorless oil
HRESIMS
m/z 519.2688 [M+Na]+ (Calcd for C28H39F3O4Na [M+Na]+, 519.2698, Δ = −1.0 mmu)
1H NMR (500 MHz, CDCl3)
δ 5.20 (1H, t, J = 6.8 Hz, H-9), 5.11 (1H, m, H-5), 5.07 (1H, m, H-2), 2.99 (2H, s, H-11), 2.26 (2 h, d, J = 6.8 Hz, H-13), 2.11 (2H, m, H-4), 2.10 (2H, m, H-14), 2.04 (2H, m, H-8), 2.00 (2H, m, H-7), 1.73 (2H, m, H-3), 1.58 (3H, s, 10-CH3), 1.55 (3H, s, 6-CH3, overlap), 1.24 (3H, s, H-1), 0.87 (6H, d, J = 6.8 Hz, 14-CH3 and 15).
Supplemental Material
sj-docx-1-npx-10.1177_1934578X221137411 - Supplemental material for Sargassumin C, a Novel Butenolide from Sargassum micracanthum
Supplemental material, sj-docx-1-npx-10.1177_1934578X221137411 for Sargassumin C, a Novel Butenolide from Sargassum micracanthum by Ji-Yul Kim, Gun-Woo Oh, Jeong Min Lee, Hyun-Soo Kim, Dae-Won Ki, Seok-Chun Ko, Mi-Jin Yim, Kyung Woo Kim, Dae-Sung Lee and Kyunghwa Baek in Natural Product Communications
Footnotes
Acknowledgements
This research was supported by the National Marine Biodiversity Institute of Korea Research Program 2022M00500.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the the National Marine Biodiversity Institute of Korea Research Program, (grant number 2022M00500).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
